Approaches to the Structure-Based Design of Antivirulence Drugs: Therapeutics for the Post-Antibiotic Era
Abstract
1. Introduction
2. Principles of Structure-Based Drug Design
2.1. Rationale and Target Selection
2.2. Methods of SBDD
3. Substrate- and Known Inhibitor-Based Design
4. Virtual Screening of Chemical Libraries
4.1. Background
4.2. AutoDock
4.3. Glide
4.4. GOLD
5. De Novo Ligand Design Based on Protein Structure
5.1. Background
5.2. LUDI
5.3. De Novo Binding Protein Design
6. Conclusions and Outlook
Funding
Acknowledgments
Conflicts of Interest
References
- Chopra, I.; Hesse, L.; O’Neill, A.J. Exploiting current understanding of antibiotic action for discovery of new drugs. J. Appl. Microbiol. 2002, 92, 4S–15S. [Google Scholar] [CrossRef] [PubMed]
- Knowles, D.J.C. New strategies for antibacterial drug design. Trends Microbiol. 1997, 5, 379–383. [Google Scholar] [CrossRef]
- White, A.R. BSAC Working Party on The Urgent Need: Regenerating Antibacterial Drug Discovery and Development Effective antibacterials: At what cost? The economics of antibacterial resistance and its control. J. Antimicrob. Chemother. 2011, 66, 1948–1953. [Google Scholar] [CrossRef] [PubMed]
- Kmietowicz, Z. Few novel antibiotics in the pipeline, WHO warns. BMJ 2017, 358, j4339. [Google Scholar] [CrossRef] [PubMed]
- Spellberg, B.; Guidos, R.; Gilbert, D.; Bradley, J.; Boucher, H.W.; Scheld, W.M.; Bartlett, J.G.; Edwards, J.; Infectious Diseases Society of America. The epidemic of antibiotic-resistant infections: A call to action for the medical community from the Infectious Diseases Society of America. Clin. Infect. Dis. 2008, 46, 155–164. [Google Scholar] [CrossRef] [PubMed]
- Talbot, G.H.; Bradley, J.; Edwards, J.E.; Gilbert, D.; Scheld, M.; Bartlett, J.G. Bad bugs need drugs: An update on the development pipeline from the Antimicrobial Availability Task Force of the Infectious Diseases Society of America. Clin. Infect. Dis. 2006, 42, 657–668. [Google Scholar] [CrossRef]
- Chopra, I.; Schofield, C.; Everett, M.; O’Neill, A.; Miller, K.; Wilcox, M.; Frère, J.-M.; Dawson, M.; Czaplewski, L.; Urleb, U.; et al. Treatment of health-care-associated infections caused by Gram-negative bacteria: A consensus statement. Lancet Infect Dis 2008, 8, 133–139. [Google Scholar] [CrossRef]
- Ventola, C.L. The Antibiotic Resistance Crisis. P T 2015, 40, 277–283. [Google Scholar]
- Dickey, S.W.; Cheung, G.Y.C.; Otto, M. Different drugs for bad bugs: Antivirulence strategies in the age of antibiotic resistance. Nat. Rev. Drug Discov. 2017, 16, 457–471. [Google Scholar] [CrossRef]
- Spees, A.M.; Wangdi, T.; Lopez, C.A.; Kingsbury, D.D.; Xavier, M.N.; Winter, S.E.; Tsolis, R.M.; Bäumler, A.J. Streptomycin-Induced Inflammation Enhances Escherichia coli Gut Colonization Through Nitrate Respiration. mBio 2013, 4, e00430-13. [Google Scholar] [CrossRef]
- Maura, D.; Ballok, A.E.; Rahme, L.G. Considerations and caveats in anti-virulence drug development. Curr. Opin. Microbiol. 2016, 33, 41–46. [Google Scholar] [CrossRef] [PubMed]
- Falkow, S. What is a pathogen? Am. Soc. Microbiol. News 1991, 63, 356–359. [Google Scholar]
- Keller, M.A.; Stiehm, E.R. Passive immunity in prevention and treatment of infectious diseases. Clin. Microbiol. Rev. 2000, 13, 602–614. [Google Scholar] [CrossRef] [PubMed]
- Schmitt, C.K.; Meysick, K.C.; O’Brien, A.D. Bacterial toxins: Friends or foes? Emerg. Infect. Dis. 1999, 5, 224–234. [Google Scholar] [CrossRef] [PubMed]
- Cegelski, L.; Marshall, G.R.; Eldridge, G.R.; Hultgren, S.J. The biology and future prospects of antivirulence therapies. Nat. Rev. Microbiol. 2008, 6, 17–27. [Google Scholar] [CrossRef] [PubMed]
- Rasko, D.A.; Sperandio, V. Anti-virulence strategies to combat bacteria-mediated disease. Nat. Rev. Drug Discov. 2010, 9, 117–128. [Google Scholar] [CrossRef]
- Escaich, S. Antivirulence as a new antibacterial approach for chemotherapy. Curr. Opin. Chem. Biol. 2008, 12, 400–408. [Google Scholar] [CrossRef]
- Totsika, M. Benefits and Challenges of Antivirulence Antimicrobials at the Dawn of the Post-Antibiotic Era. Curr. Med. Chem. 2016, 6, 30–37. [Google Scholar] [CrossRef]
- Kalia, V.C. Quorum sensing inhibitors: An overview. Biotechnol. Adv. 2013, 31, 224–245. [Google Scholar] [CrossRef]
- Baron, C. Antivirulence drugs to target bacterial secretion systems. Curr. Opin. Microbiol. 2010, 13, 100–105. [Google Scholar] [CrossRef]
- Cusumano, C.K.; Hultgren, S.J. Bacterial adhesion—A source of alternate antibiotic targets. IDrugs 2009, 12, 699–705. [Google Scholar] [PubMed]
- Totsika, M. Disarming pathogens: Benefits and challenges of antimicrobials that target bacterial virulence instead of growth and viability. Future Med. Chem. 2017, 9, 267–269. [Google Scholar] [CrossRef] [PubMed]
- Zambelloni, R.; Marquez, R.; Roe, A.J. Development of Antivirulence Compounds: A Biochemical Review. Chem. Biol. Drug Des. 2015, 85, 43–55. [Google Scholar] [CrossRef] [PubMed]
- Overbye, K.M.; Barrett, J.F. Antibiotics: Where did we go wrong? Drug Discov. Today 2005, 10, 45–52. [Google Scholar] [CrossRef]
- Erickson, J.; Neidhart, D.J.; VanDrie, J.; Kempf, D.J.; Wang, X.C.; Norbeck, D.W.; Plattner, J.J.; Rittenhouse, J.W.; Turon, M.; Wideburg, N. Design, activity, and 2.8 A crystal structure of a C2 symmetric inhibitor complexed to HIV-1 protease. Science 1990, 249, 527–533. [Google Scholar] [CrossRef] [PubMed]
- Roberts, N.A.; Martin, J.A.; Kinchington, D.; Broadhurst, A.V.; Craig, J.C.; Duncan, I.B.; Galpin, S.A.; Handa, B.K.; Kay, J.; Kröhn, A. Rational design of peptide-based HIV proteinase inhibitors. Science 1990, 248, 358–361. [Google Scholar] [CrossRef] [PubMed]
- Dorsey, B.D.; Levin, R.B.; McDaniel, S.L.; Vacca, J.P.; Guare, J.P.; Darke, P.L.; Zugay, J.A.; Emini, E.A.; Schleif, W.A. L-735,524: The Design of a Potent and Orally Bioavailable HIV Protease Inhibitor. J. Med. Chem. 1994, 37, 3443–3451. [Google Scholar] [CrossRef]
- McCauley, J. Relenza. Curr. Biol. 1999, 9, R796. [Google Scholar] [CrossRef]
- Stratton, M.S.; Alberts, D.S. Current application of selective COX-2 inhibitors in cancer prevention and treatment. Oncology 2002, 16, 37–51. [Google Scholar]
- Deininger, M.; Buchdunger, E.; Druker, B.J. The development of imatinib as a therapeutic agent for chronic myeloid leukemia. Blood 2005, 105, 2640–2653. [Google Scholar] [CrossRef]
- Hong, W.; Zeng, J.; Xie, J. Antibiotic drugs targeting bacterial RNAs. Acta Pharm. Sin. B 2014, 4, 258–265. [Google Scholar] [CrossRef]
- Nissen, P.; Hansen, J.; Ban, N.; Moore, P.B.; Steitz, T.A. The Structural Basis of Ribosome Activity in Peptide Bond Synthesis. Science 2000, 289, 920–930. [Google Scholar] [CrossRef] [PubMed]
- Yu, H. Extending the size limit of protein nuclear magnetic resonance. Proc. Natl. Acad. Sci. USA 1999, 96, 332–334. [Google Scholar] [CrossRef] [PubMed]
- Bartesaghi, A.; Merk, A.; Banerjee, S.; Matthies, D.; Wu, X.; Milne, J.L.S.; Subramaniam, S. 2.2 Å resolution cryo-EM structure of β-galactosidase in complex with a cell-permeant inhibitor. Science 2015, 348, 1147–1151. [Google Scholar] [CrossRef]
- Vyas, V.K.; Ukawala, R.D.; Ghate, M.; Chintha, C. Homology Modeling a Fast Tool for Drug Discovery: Current Perspectives. Indian J. Pharm. Sci. 2012, 74, 1–17. [Google Scholar] [CrossRef] [PubMed]
- Enyedy, I.J.; Lee, S.L.; Kuo, A.H.; Dickson, R.B.; Lin, C.Y.; Wang, S. Structure-based approach for the discovery of bis-benzamidines as novel inhibitors of matriptase. J. Med. Chem. 2001, 44, 1349–1355. [Google Scholar] [CrossRef] [PubMed]
- Xiang, Z. Advances in Homology Protein Structure Modeling. Curr. Protein Pept. Sci. 2006, 7, 217–227. [Google Scholar] [CrossRef] [PubMed]
- Arnold, K.; Bordoli, L.; Kopp, J.; Schwede, T. The SWISS-MODEL workspace: A web-based environment for protein structure homology modelling. Bioinformatics 2006, 22, 195–201. [Google Scholar] [CrossRef]
- Kelley, L.A.; Mezulis, S.; Yates, C.M.; Wass, M.N.; Sternberg, M.J. The Phyre2 web portal for protein modelling, prediction and analysis. Nat. Protoc. 2015, 10, 845–858. [Google Scholar] [CrossRef]
- Webb, B.; Sali, A. Comparative Protein Structure Modeling Using MODELLER. Curr. Protoc. Bioinformatics 2014, 47, 5.6.1–5.6.32. [Google Scholar] [CrossRef]
- Kemmish, H.; Fasnacht, M.; Yan, L. Fully automated antibody structure prediction using BIOVIA tools: Validation study. PLoS ONE 2017, 12, e0177923. [Google Scholar] [CrossRef] [PubMed]
- Halgren, T.A. Identifying and Characterizing Binding Sites and Assessing Druggability. J. Chem. Inf. Model. 2009, 49, 377–389. [Google Scholar] [CrossRef] [PubMed]
- Villoutreix, B.O.; Renault, N.; Lagorce, D.; Sperandio, O.; Montes, M.; Miteva, M.A. Free resources to assist structure-based virtual ligand screening experiments. Curr. Protein Pept. Sci. 2007, 8, 381–411. [Google Scholar] [CrossRef] [PubMed]
- Scott, D.E.; Bayly, A.R.; Abell, C.; Skidmore, J. Small molecules, big targets: Drug discovery faces the protein–protein interaction challenge. Nat. Rev. Drug Discov. 2016, 15, 533–550. [Google Scholar] [CrossRef] [PubMed]
- Jones, G.; Willett, P.; Glen, R.C.; Leach, A.R.; Taylor, R. Development and validation of a genetic algorithm for flexible docking. J. Mol. Biol. 1997, 267, 727–748. [Google Scholar] [CrossRef] [PubMed]
- Schnecke, V.; Swanson, C.A.; Getzoff, E.D.; Tainer, J.A.; Kuhn, L.A. Screening a peptidyl database for potential ligands to proteins with side-chain flexibility. Proteins 1998, 33, 74–87. [Google Scholar] [CrossRef]
- Claussen, H.; Buning, C.; Rarey, M.; Lengauer, T. FlexE: Efficient molecular docking considering protein structure variations. J. Mol. Biol. 2001, 308, 377–395. [Google Scholar] [CrossRef] [PubMed]
- Simmons, K.J.; Chopra, I.; Fishwick, C.W.G. Structure-based discovery of antibacterial drugs. Nat. Rev. Microbiol. 2010, 8, 501–510. [Google Scholar] [CrossRef] [PubMed]
- Pagadala, N.S.; Syed, K.; Tuszynski, J. Software for molecular docking: A review. Biophys. Rev. 2017, 9, 91–102. [Google Scholar] [CrossRef]
- Chan, D.C.M.; Laughton, C.A.; Queener, S.F.; Stevens, M.F.G. Structural Studies on Bioactive Compounds. 34. Design, Synthesis, and Biological Evaluation of Triazenyl-Substituted Pyrimethamine Inhibitors of Pneumocystis carinii Dihydrofolate Reductase. J. Med. Chem. 2001, 44, 2555–2564. [Google Scholar] [CrossRef] [PubMed]
- Varney, M.D.; Marzoni, G.P.; Palmer, C.L.; Deal, J.G.; Webber, S.; Welsh, K.M.; Bacquet, R.J.; Bartlett, C.A.; Morse, C.A. Crystal-structure-based design and synthesis of benz[cd]indole-containing inhibitors of thymidylate synthase. J. Med. Chem. 1992, 35, 663–676. [Google Scholar] [CrossRef] [PubMed]
- Schmid, M.B. Seeing is believing: The impact of structural genomics on antimicrobial drug discovery. Nat. Rev. Microbiol. 2004, 2, 739–746. [Google Scholar] [CrossRef]
- Smith, K.M.; Bu, Y.; Suga, H. Library Screening for Synthetic Agonists and Antagonists of a Pseudomonas aeruginosa Autoinducer. Chem. Biol. 2003, 10, 563–571. [Google Scholar] [CrossRef]
- Persson, T.; Hansen, T.H.; Rasmussen, T.B.; Skindersø, M.E.; Givskov, M.; Nielsen, J. Rational design and synthesis of new quorum-sensing inhibitors derived from acylated homoserine lactones and natural products from garlic. Org. Biomol. Chem. 2005, 3, 253–262. [Google Scholar] [CrossRef]
- Akram, O.N.; DeGraff, D.J.; Sheehan, J.H.; Tilley, W.D.; Matusik, R.J.; Ahn, J.-M.; Raj, G.V. Tailoring Peptidomimetics for Targeting Protein–Protein Interactions. Mol. Cancer Res. 2014, 12, 967–978. [Google Scholar] [CrossRef] [PubMed]
- Larzábal, M.; Mercado, E.C.; Vilte, D.A.; Salazar-González, H.; Cataldi, A.; Navarro-Garcia, F. Designed Coiled-Coil Peptides Inhibit the Type Three Secretion System of Enteropathogenic Escherichia coli. PLoS ONE 2010, 5, e9046. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Faucher, F.; Zhang, W.; Wang, S.; Neville, N.; Poole, K.; Zheng, J.; Jia, Z. Structure-guided disruption of the pseudopilus tip complex inhibits the Type II secretion in Pseudomonas aeruginosa. PLoS Pathog. 2018, 14, e1007343. [Google Scholar] [CrossRef]
- Qvit, N.; Rubin, S.J.S.; Urban, T.J.; Mochly-Rosen, D.; Gross, E.R. Peptidomimetic therapeutics: Scientific approaches and opportunities. Drug Discov. Today 2017, 22, 454–462. [Google Scholar] [CrossRef]
- Pinkner, J.S.; Remaut, H.; Buelens, F.; Miller, E.; Aberg, V.; Pemberton, N.; Hedenström, M.; Larsson, A.; Seed, P.; Waksman, G.; et al. Rationally designed small compounds inhibit pilus biogenesis in uropathogenic bacteria. Proc. Natl. Acad. Sci. USA 2006, 103, 17897–17902. [Google Scholar] [CrossRef]
- Lee, Y.M.; Almqvist, F.; Hultgren, S.J. Targeting virulence for antimicrobial chemotherapy. Curr. Opin. Pharmacol. 2003, 3, 513–519. [Google Scholar] [CrossRef]
- Cegelski, L.; Pinkner, J.S.; Hammer, N.D.; Cusumano, C.K.; Hung, C.S.; Chorell, E.; Åberg, V.; Walker, J.N.; Seed, P.C.; Almqvist, F.; et al. Small-molecule inhibitors target Escherichia coli amyloid biogenesis and biofilm formation. Nat. Chem. Biol. 2009, 5, 913–919. [Google Scholar] [CrossRef] [PubMed]
- Schnecke, V.; Kuhn, L.A. Virtual screening with solvation and ligand-induced complementarity. Perspect. Drug Discov. Des. 2000, 20, 171–190. [Google Scholar] [CrossRef]
- Alberg, D.G.; Schreiber, S.L. Structure-based design of a cyclophilin-calcineurin bridging ligand. Science 1993, 262, 248–250. [Google Scholar] [CrossRef] [PubMed]
- Taylor, R.D.; Jewsbury, P.J.; Essex, J.W. A review of protein-small molecule docking methods. J. Comput. Aided Mol Des 2002, 16, 151–166. [Google Scholar] [CrossRef] [PubMed]
- Shoichet, B.K.; McGovern, S.L.; Wei, B.; Irwin, J.J. Lead discovery using molecular docking. Curr. Opin. Chem. Biol. 2002, 6, 439–446. [Google Scholar] [CrossRef]
- Ramírez, D. Computational Methods Applied to Rational Drug Design. Open Med. Chem. J. 2016, 10, 7–20. [Google Scholar] [CrossRef]
- Goodsell, D.S.; Morris, G.M.; Olson, A.J. Automated docking of flexible ligands: Applications of AutoDock. J. Mol. Recognit. 1996, 9, 1–5. [Google Scholar] [CrossRef]
- Prasad, J.C.; Goldstone, J.V.; Camacho, C.J.; Vajda, S.; Stegeman, J.J. Ensemble modeling of substrate binding to cytochromes P450: Analysis of catalytic differences between CYP1A orthologs. Biochemistry 2007, 46, 2640–2654. [Google Scholar] [CrossRef]
- Källblad, P.; Mancera, R.L.; Todorov, N.P. Assessment of Multiple Binding Modes in Ligand−Protein Docking. J. Med. Chem. 2004, 47, 3334–3337. [Google Scholar] [CrossRef]
- Limongelli, V.; Marinelli, L.; Cosconati, S.; Braun, H.A.; Schmidt, B.; Novellino, E. Ensemble-docking approach on BACE-1: Pharmacophore perception and guidelines for drug design. ChemMedChem 2007, 2, 667–678. [Google Scholar] [CrossRef]
- Case, D.A.; Cheatham, T.E.; Darden, T.; Gohlke, H.; Luo, R.; Merz, K.M.; Onufriev, A.; Simmerling, C.; Wang, B.; Woods, R.J. The Amber Biomolecular Simulation Programs. J. Comput. Chem. 2005, 26, 1668–1688. [Google Scholar] [CrossRef] [PubMed]
- Trott, O.; Olson, A.J. AutoDock Vina: Improving the speed and accuracy of docking with a new scoring function, efficient optimization and multithreading. J. Comput. Chem. 2010, 31, 455–461. [Google Scholar] [CrossRef] [PubMed]
- Ng, M.C.K.; Fong, S.; Siu, S.W.I. PSOVina: The hybrid particle swarm optimization algorithm for protein-ligand docking. J. Bioinform. Comput. Biol. 2015, 13, 1541007. [Google Scholar] [CrossRef] [PubMed]
- Swietnicki, W.; Carmany, D.; Retford, M.; Guelta, M.; Dorsey, R.; Bozue, J.; Lee, M.S.; Olson, M.A. Identification of Small-Molecule Inhibitors of Yersinia pestis Type III Secretion System YscN ATPase. PLoS ONE 2011, 6, e19716. [Google Scholar] [CrossRef] [PubMed]
- Zsoldos, Z.; Reid, D.; Simon, A.; Sadjad, S.B.; Johnson, A.P. eHiTS: A new fast, exhaustive flexible ligand docking system. J. Mol. Graph. Model. 2007, 26, 198–212. [Google Scholar] [CrossRef] [PubMed]
- Friesner, R.A.; Banks, J.L.; Murphy, R.B.; Halgren, T.A.; Klicic, J.J.; Mainz, D.T.; Repasky, M.P.; Knoll, E.H.; Shelley, M.; Perry, J.K.; et al. Glide: A New Approach for Rapid, Accurate Docking and Scoring. 1. Method and Assessment of Docking Accuracy. J. Med. Chem. 2004, 47, 1739–1749. [Google Scholar] [CrossRef] [PubMed]
- Abagyan, R.; Totrov, M.; Kuznetsov, D. ICM—A New Method for Protein Modeling and Design: Applications to Docking and Structure Prediction from the Distorted Native Conformation. J. Comput. Chem. 1994, 15, 488–506. [Google Scholar] [CrossRef]
- Böhm, H.J. The computer program LUDI: A new method for the de novo design of enzyme inhibitors. J. Comput. Aided Mol. Des. 1992, 6, 61–78. [Google Scholar] [CrossRef] [PubMed]
- Gillet, V.; Johnson, A.P.; Mata, P.; Sike, S.; Williams, P. SPROUT: A program for structure generation. J. Comput. Aided Mol. Des. 1993, 7, 127–153. [Google Scholar] [CrossRef] [PubMed]
- Caflisch, A.; Miranker, A.; Karplus, M. Multiple copy simultaneous search and construction of ligands in binding sites: Application to inhibitors of HIV-1 aspartic proteinase. J. Med. Chem. 1993, 36, 2142–2167. [Google Scholar] [CrossRef]
- Khodaverdian, V.; Pesho, M.; Truitt, B.; Bollinger, L.; Patel, P.; Nithianantham, S.; Yu, G.; Delaney, E.; Jankowsky, E.; Shoham, M. Discovery of Antivirulence Agents against Methicillin-Resistant Staphylococcus aureus. Antimicrob. Agents Chemother. 2013, 57, 3645–3652. [Google Scholar] [CrossRef] [PubMed]
- Annapoorani, A.; Umamageswaran, V.; Parameswari, R.; Pandian, S.K.; Ravi, A.V. Computational discovery of putative quorum sensing inhibitors against LasR and RhlR receptor proteins of Pseudomonas aeruginosa. J. Comput. Aided Mol. Des. 2012, 26, 1067–1077. [Google Scholar] [CrossRef] [PubMed]
- Otto, M. Staphylococcus aureus toxins. Curr. Opin. Microbiol. 2014, 17, 32–37. [Google Scholar] [CrossRef] [PubMed]
- Rasmussen, T.B.; Givskov, M. Quorum-sensing inhibitors as anti-pathogenic drugs. Int. J. Med. Microbiol. 2006, 296, 149–161. [Google Scholar] [CrossRef] [PubMed]
- Duprez, W.; Bachu, P.; Stoermer, M.J.; Tay, S.; McMahon, R.M.; Fairlie, D.P.; Martin, J.L. Virtual Screening of Peptide and Peptidomimetic Fragments Targeted to Inhibit Bacterial Dithiol Oxidase DsbA. PLoS ONE 2015, 10, e0133805. [Google Scholar] [CrossRef] [PubMed]
- Totsika, M.; Heras, B.; Wurpel, D.J.; Schembri, M.A. Characterization of Two Homologous Disulfide Bond Systems Involved in Virulence Factor Biogenesis in Uropathogenic Escherichia coli CFT073. J. Bacteriol. 2009, 191, 3901–3908. [Google Scholar] [CrossRef] [PubMed]
- Kurth, F.; Duprez, W.; Premkumar, L.; Schembri, M.A.; Fairlie, D.P.; Martin, J.L. Crystal structure of the dithiol oxidase DsbA enzyme from proteus mirabilis bound non-covalently to an active site peptide ligand. J. Biol. Chem. 2014, 289, 19810–19822. [Google Scholar] [CrossRef] [PubMed]
- Duprez, W.; Premkumar, L.; Halili, M.A.; Lindahl, F.; Reid, R.C.; Fairlie, D.P.; Martin, J.L. Peptide inhibitors of the Escherichia coli DsbA oxidative machinery essential for bacterial virulence. J. Med. Chem. 2015, 58, 577–587. [Google Scholar] [CrossRef]
- Chang, C.-Y.; Krishnan, T.; Wang, H.; Chen, Y.; Yin, W.-F.; Chong, Y.-M.; Tan, L.Y.; Chong, T.M.; Chan, K.-G. Non-antibiotic quorum sensing inhibitors acting against N-acyl homoserine lactone synthase as druggable target. Sci. Rep. 2014, 4, 7245. [Google Scholar] [CrossRef]
- Gould, T.A.; Schweizer, H.P.; Churchill, M.E.A. Structure of the Pseudomonas aeruginosa acyl-homoserinelactone synthase LasI. Mol. Microbiol. 2004, 53, 1135–1146. [Google Scholar] [CrossRef]
- Kimyon, Ö.; Ulutürk, Z.İ.; Nizalapur, S.; Lee, M.; Kutty, S.K.; Beckmann, S.; Kumar, N.; Manefield, M. N-Acetylglucosamine Inhibits LuxR, LasR and CviR Based Quorum Sensing Regulated Gene Expression Levels. Front. Microbiol. 2016, 7, 1313. [Google Scholar] [CrossRef] [PubMed]
- Boehm, H.J.; Boehringer, M.; Bur, D.; Gmuender, H.; Huber, W.; Klaus, W.; Kostrewa, D.; Kuehne, H.; Luebbers, T.; Meunier-Keller, N.; et al. Novel inhibitors of DNA gyrase: 3D structure based biased needle screening, hit validation by biophysical methods, and 3D guided optimization. A promising alternative to random screening. J. Med. Chem. 2000, 43, 2664–2674. [Google Scholar] [CrossRef]
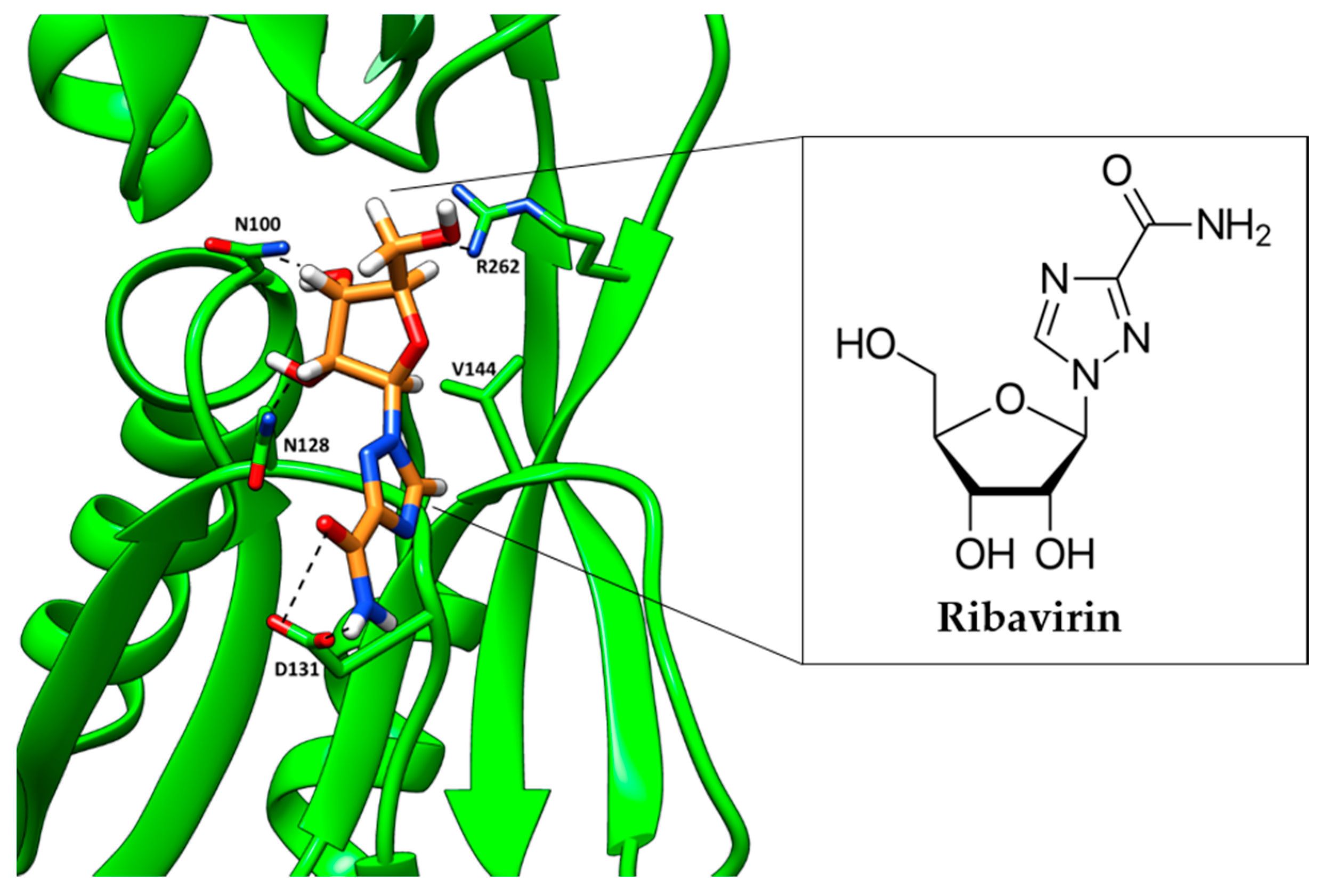
- Mandal, R.S.; Ta, A.; Sinha, R.; Theeya, N.; Ghosh, A.; Tasneem, M.; Bhunia, A.; Koley, H.; Das, S. Ribavirin suppresses bacterial virulence by targeting LysR-type transcriptional regulators. Sci. Rep. 2016, 6, 39454. [Google Scholar] [CrossRef] [PubMed]
- Arya, R.; Ravikumar, R.; Santhosh, R.S.; Princy, S.A. SarA based novel therapeutic candidate against Staphylococcus aureus associated with vascular graft infections. Front. Microbiol. 2015, 6, 416. [Google Scholar] [CrossRef] [PubMed]
- Arya, R.; Princy, S.A. Computational approach to design small molecule inhibitors and identify SarA as a potential therapeutic candidate. Med. Chem. Res. 2013, 22, 1856–1865. [Google Scholar] [CrossRef]
- Chevalier, A.; Silva, D.-A.; Rocklin, G.J.; Hicks, D.R.; Vergara, R.; Murapa, P.; Bernard, S.M.; Zhang, L.; Lam, K.-H.; Yao, G.; et al. Massively parallel de novo protein design for targeted therapeutics. Nature 2017, 550, 74–79. [Google Scholar] [CrossRef]
- Rossi, F.; Girardello, R.; Cury, A.P.; Di Gioia, T.S.R.; de Almeida, J.N.; da Silva Duarte, A.J. Emergence of colistin resistance in the largest university hospital complex of São Paulo, Brazil, over five years. Braz. J. Infect. Dis. 2017, 21, 98–101. [Google Scholar] [CrossRef]
- Payne, D.J.; Gwynn, M.N.; Holmes, D.J.; Pompliano, D.L. Drugs for bad bugs: Confronting the challenges of antibacterial discovery. Nat. Rev. Drug Discov. 2007, 6, 29–40. [Google Scholar] [CrossRef]


| Program | Search strategy | Flexible ligand? | Flexible protein side chains? | Description | Free for academia? | |
|---|---|---|---|---|---|---|
| Virtual screening | AutoDock [67,74] | GA/MC | yes | yes | Exhaustive search of interaction energy grid followed by simulated annealing energy scoring | yes |
| GOLD [45] | GA | yes | yes | Positions ligand and minimizes energy via an evolutionary algorithm | no | |
| eHITS [75] | IC | yes | no | Ligands are divided into rigid fragments, which are docked individually, then reconstructed | no | |
| GLIDE [76] | Hybrid | yes | no | Protein-ligand coulomb-vdW energy minimization and an empirical GlideScore | no | |
| FlexE [47] | IC | yes | yes | Incremental construction by sampling ligand conformations and target ensembles | no | |
| ICM [77] | MC | yes | yes | Monte-Carlo energy minimization of protein and ligand | no | |
| De novo ligand design | LUDI [78] | EG | yes | no | Empirically-derived energy scoring of fragments | no |
| SPROUT [79] | AI | yes | no | Template atoms or fragments are linked to make skeletons | no | |
| MCSS [80] | Hybrid | yes | no | CHARMm-based exhaustive search for target site functional group minima | no |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Neville, N.; Jia, Z. Approaches to the Structure-Based Design of Antivirulence Drugs: Therapeutics for the Post-Antibiotic Era. Molecules 2019, 24, 378. https://doi.org/10.3390/molecules24030378
Neville N, Jia Z. Approaches to the Structure-Based Design of Antivirulence Drugs: Therapeutics for the Post-Antibiotic Era. Molecules. 2019; 24(3):378. https://doi.org/10.3390/molecules24030378
Chicago/Turabian StyleNeville, Nolan, and Zongchao Jia. 2019. "Approaches to the Structure-Based Design of Antivirulence Drugs: Therapeutics for the Post-Antibiotic Era" Molecules 24, no. 3: 378. https://doi.org/10.3390/molecules24030378
APA StyleNeville, N., & Jia, Z. (2019). Approaches to the Structure-Based Design of Antivirulence Drugs: Therapeutics for the Post-Antibiotic Era. Molecules, 24(3), 378. https://doi.org/10.3390/molecules24030378






