Comparison of the Interactions of Different Growth Factors and Glycosaminoglycans
Abstract
1. Introduction
2. Results
2.1. Kinetics Measurements of Growth Factor–Heparin Interactions
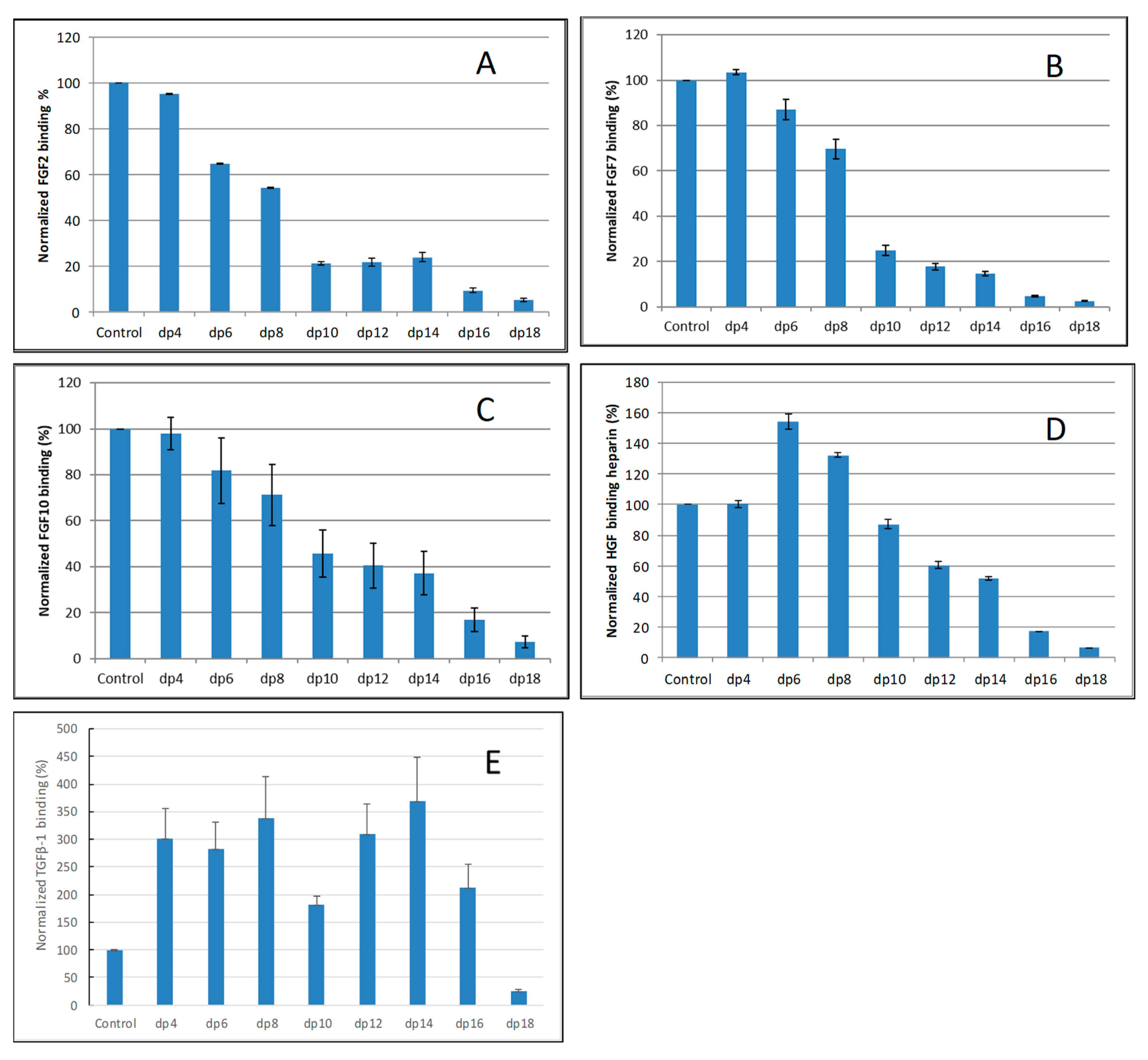
2.2. Solution Competition Study on the Interaction between Heparin (Surface) and Growth Factor to Heparin-Derived Oligosaccharides (in Solution) Using SPR
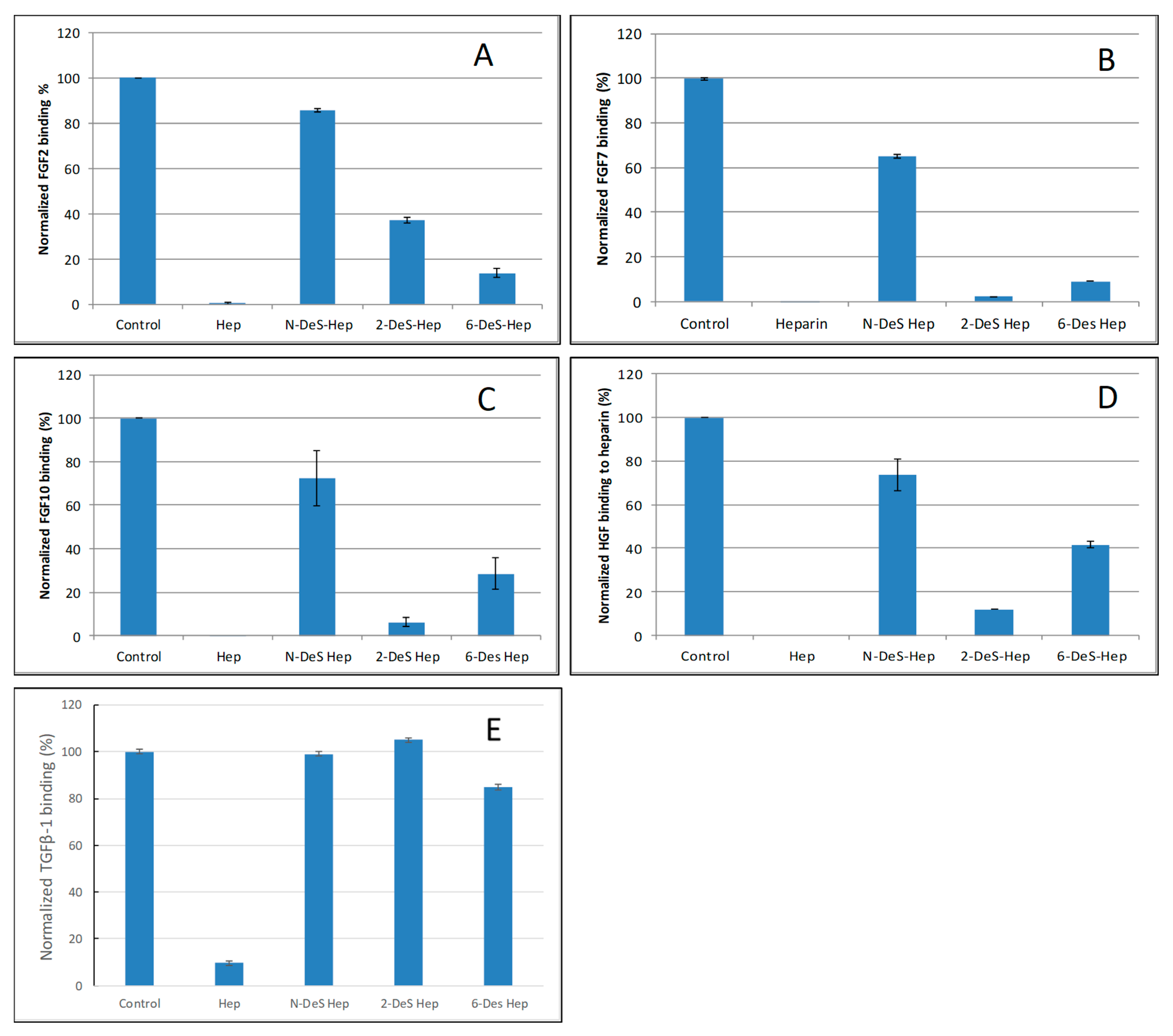
2.3. SPR Solution Competition Study of Different Chemically Modified Heparins
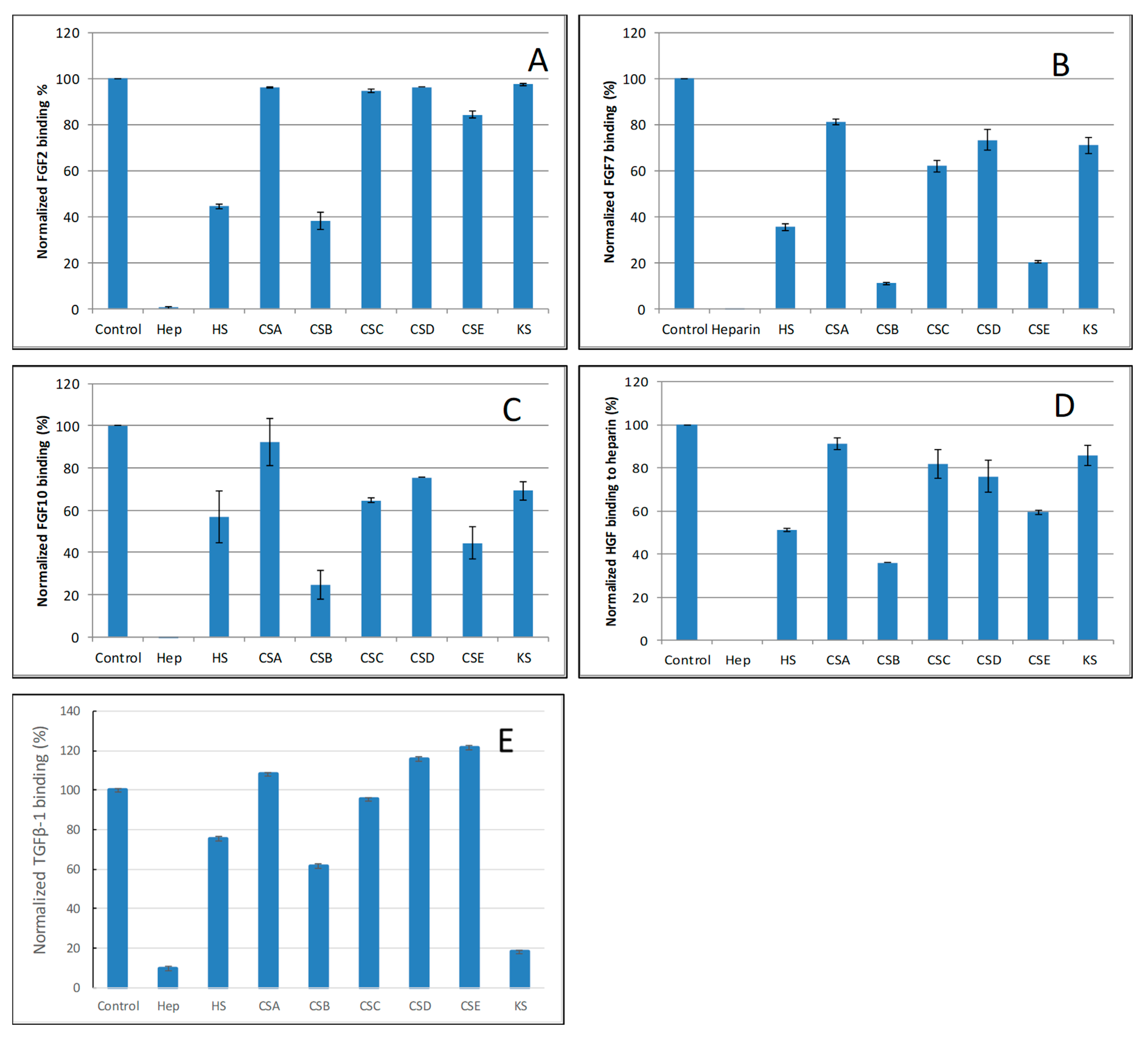
2.4. SPR Solution Competition Study of Different GAGs
3. Discussion
4. Materials and Methods
4.1. Materials
4.2. Preparation of Heparin Biochip
4.3. Kinetic Measurement of Interaction between Growth Factor and Heparin Using BIAcore
4.4. Solution Competition Study between Heparin on the Chip Surface and Heparin-Derived Oligosaccharides in Solution Using SPR
4.5. Solution Competition Study between Heparin on Chip Surface and GAGs, Chemical Modified Heparin in Solution Using SPR
5. Conclusions
Author Contributions
Funding
Conflicts of Interest
Abbreviations
| GAG | glycosaminoglycan |
| SPR | surface plasmon resonance |
| HS | heparan sulfate |
| CSA | chondroitin sulfate A |
| CSB | chondroitin sulfate B |
| CSC | chondroitin sulfate C |
| CSD | chondroitin sulfate D |
| CSE | chondroitin sulfate E |
| KS | keratan sulfate |
| SA | streptavidin |
| dp | degree of polymerization |
| FGF | fibroblast growth factors |
| HGF | hepatocyte growth factor |
| TGF | transforming growth factor |
References
- Cross, M.; Dexter, T.M. Growth factors in development, transformation, and tumorigenesis. Cell 1991, 64, 271–280. [Google Scholar] [CrossRef]
- Aaronson, S.A. Growth factors and cancer. Science 1991, 254, 1146–1153. [Google Scholar] [CrossRef] [PubMed]
- Milani, S.; Herbst, H.; Schuppan, D.; Stein, H.; Surrenti, C. Transforming growth factors beta 1 and beta 2 are differentially expressed in fibrotic liver disease. Am. J. Pathol. 1991, 139, 1221–1229. [Google Scholar] [PubMed]
- Ludwicka, A.; Ohba, T.; Trojanowska, M.; Yamakage, A.; Strange, C.; Smith, E.A.; Leroy, E.C.; Sutherland, S.; Silver, R.M. Elevated levels of platelet derived growth factor and transforming growth factor-beta 1 in bronchoalveolar lavage fluid from patients with scle-roderma. J. Rheumatol. 1995, 22, 1876–1883. [Google Scholar] [PubMed]
- Holgate, S.T.; Davies, D.E.; Lackie, P.M.; Wilson, S.J.; Puddicombe, S.M.; Lordan, J.L. Epithelial-mesenchymal interactions in the pathogenesis of asthma. J. Allergy Clin. Immunol. 2000, 105, 193–204. [Google Scholar] [CrossRef]
- Zhang, X.; Ibrahimi, O.A.; Olsen, S.K.; Umemori, H.; Mohammadi, M.; Ornitz, D.M. Receptor specificity of the fibroblast growth factor family. The complete mammalian FGF family. J. Biol. Chem. 2006, 281, 15694–15700. [Google Scholar] [CrossRef]
- Dong, M.; Blobe, G.C. Role of transforming growth factor-β in hematologic malignancies. Blood 2006, 107, 4589–4596. [Google Scholar] [CrossRef] [PubMed]
- Witsch, E.; Sela, M.; Yarden, Y. Roles for growth factors in cancer progression. Physiology 2010, 25, 85–101. [Google Scholar] [CrossRef]
- Domouzoglou, E.M.; Naka, K.K.; Vlahos, A.P.; Papafaklis, M.I.; Michalis, L.K.; Tsatsoulis, A.; Maratos-Flier, E. Fibroblast growth factors in cardiovascular disease: The emerging role of FGF21. Am. J. Physiol. Heart Circ. Physiol. 2015, 309, H1029–H1038. [Google Scholar] [CrossRef]
- Itoh, N.; Ohta, H.; Nakayama, Y.; Konishi, M. Roles of FGF Signals in Heart Development, Health, and Disease. Front. Cell Dev. Biol. 2016, 4, 110. [Google Scholar] [CrossRef]
- Lee, K.; Silva, E.A.; Mooney, D.J. Growth factor delivery-based tissue engineering: General approaches and a review of recent developments. J. R. Soc. Interface 2010, 8, 153–170. [Google Scholar] [CrossRef] [PubMed]
- Linhardt, R.J.; Kerns, R.J.; Vlahov, I.R. Heparin and heparin oligosaccharides: Preparation, analysis, application and biological activities. In Biochemical Functions and Biotechnology of Natural and Artificial polymers; Yalpani, M., Ed.; ATL Press, Science Publishers: Mt. Prospect, IL, USA, 1996; pp. 46–62. [Google Scholar]
- Linhardt, R.J.; Toida, T. Role of glycosaminoglycans in cellular communication. Acc. Chem. Res. 2004, 37, 431–438. [Google Scholar] [CrossRef] [PubMed]
- Iozzo, R.V.; Zoeller, J.J.; Nyström, A. Basement membrane proteoglycans: Modulators Par Excellence of cancer growth and angiogenesis. Mol. Cells. 2009, 27, 503–513. [Google Scholar] [CrossRef] [PubMed]
- Capila, I.; Linhardt, R.J. Heparin–protein interactions. Angew. Chem. Int. Ed. 2002, 41, 390–412. [Google Scholar] [CrossRef]
- Kjellén, L.; Lindahl, U. Specificity of glycosaminoglycan-protein interactions. Curr. Opin. Struct. Biol. 2018, 50, 101–108. [Google Scholar] [CrossRef] [PubMed]
- Xu, D.; Esko, J.D. Demystifying heparan sulfate-protein interactions. Annu. Rev. Biochem. 2014, 83, 129–157. [Google Scholar] [CrossRef] [PubMed]
- Powell, A.K.; Yates, E.A.; Fernig, D.G.; Turnbull, J.E. Interactions of heparin/heparan sulfate with proteins: Appraisal of structural factors and experimental approaches. Glycobiology 2004, 14, 17R–30R. [Google Scholar] [CrossRef]
- Wang, L.; Fuster, M.; Sriramarao, P.; Esko, J.D. Endothelial heparan sulfate deficiency impairs L-selectin-and chemokine-mediated neutrophil trafficking during inflammatory responses. Nat. Immunol. 2005, 6, 902–910. [Google Scholar] [CrossRef]
- Häcker, U.; Nybakken, K.; Perrimon, N. Heparan sulphate proteoglycans: The sweet side of development. Nat. Rev. Mol. Cell Biol. 2005, 6, 530–541. [Google Scholar] [CrossRef]
- Faham, S.; Hileman, R.E.; Fromm, J.R.; Linhardt, R.J.; Rees, D.C. Heparin structure and interactions with basic fibroblast growth factor. Science 1996, 271, 1116–1120. [Google Scholar] [CrossRef]
- DiGabriele, A.D.; Lax, I.; Chen, D.I.; Svahn, C.M.; Jaye, M.; Schlessinger, J.; Hendrickson, W.A. Structure of a heparin-linked biologically active dimer of fibroblast growth factor. Nature 1988, 393, 812–817. [Google Scholar] [CrossRef] [PubMed]
- Pellegrini, L.; Burke, D.F.; von Delft, F.; Mulloy, B.; Blundell, T.L. Crystal structure of fibroblast growth factor receptor ectodomain bound to ligand and heparin. Nature 2000, 407, 1029–1034. [Google Scholar] [CrossRef] [PubMed]
- Ricard-Blum, S. Protein–glycosaminoglycan interaction networks: Focus on heparan sulfate. Perspect. Sci. 2017, 11, 62–69. [Google Scholar] [CrossRef]
- Li, Y.; Sun, C.; Yates, E.A.; Jiang, C.; Wilkinson, M.C.; Fernig, D.G. Heparin binding preference and structures in the fibroblast growth factor family parallel their evolutionary diversification. Open Biol. 2016, 6, 150275. [Google Scholar] [CrossRef] [PubMed]
- Asada, M.; Shinomiya, M.; Suzuki, M.; Honda, E.; Sugimoto, R.; Ikekita, M.; Imamura, T. Glycosaminoglycan affinity of the complete fibroblast growth factor family. Biochim Biophys. Acta. 2009, 1790, 40–48. [Google Scholar] [CrossRef]
- Ashikari-Hada, S.; Habuchi, H.; Kariya, Y.; Itoh, N.; Reddi, A.H.; Kimata, K. Characterization of growth factor-binding structures in heparin/heparan sulfate using an octasaccharide library. J. Biol. Chem. 2004, 279, 2346–2354. [Google Scholar] [CrossRef]
- Rahmoune, H.; Rudland, P.S.; Gallagher, J.T.; Fernig, D.G. Hepatocyte growth factor/scatter factor has distinct classes of binding site in heparan sulfate from mammary cells. Biochemistry 1988, 37, 6003–6008. [Google Scholar] [CrossRef]
- McCaffrey, T.A.; Falcone, D.J.; Du, B. Transforming growth factor-beta 1 is a heparin-binding protein: Identification of putative heparin-binding regions and isolation of heparins with varying affinity for TGF-beta 1. J. Cell Physiol. 1992, 152, 430–440. [Google Scholar] [CrossRef]
- Itoh, N. FGF10: A multifunctional mesenchymal-epithelial signaling growth factor in development, health, and disease. Cytokine Growth Factor Rev. 2016, 28, 63–69. [Google Scholar] [CrossRef]
- Tong, L.; Zhou, J.; Rong, L.; Seeley, E.J.; Pan, J.; Zhu, X.; Liu, J.; Wang, Q.; Tang, X.; Qu, J.; et al. Fibroblast Growth Factor-10 (FGF-10) Mobilizes Lung-resident Mesenchymal Stem Cells and Protects Against Acute Lung Injury. Sci. Rep. 2016, 6, 21642. [Google Scholar] [CrossRef]
- Patel, V.N.; Likar, K.M.; Zisman-Rozen, S.; Cowherd, S.N.; Lassiter, K.S.; Sher, I.; Yates, E.A.; Turnbull, J.E.; Ron, D.; Hoffman, M.P. Specific heparan sulfate structures modulate FGF10-mediated submandibular gland epithelial morphogenesis and differentiation. J. Biol. Chem. 2008, 283, 9308–9317. [Google Scholar] [CrossRef] [PubMed]
- Ye, S.; Luo, Y.; Lu, W.; Jones, R.B.; Linhardt, R.J.; Capila, I.; Toida, T.; Kan, M.; Pelletier, H.; McKeehan, W.L. Structural basis for interaction of FGF-1, FGF-2, and FGF-7 with different heparan sulfate motifs. Biochemistry 2001, 40, 14429–14439. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Casas-Finet, J.R.; Heath Coats, R.; Kaufman, J.D.; Stahl, S.J.; Wingfield, P.T.; Rubin, J.S.; Bottaro, D.P.; Byrd, R.A. Identification and dynamics of a heparin-binding site in hepatocyte growth factor. Biochemistry 1999, 38, 14793–14802. [Google Scholar] [CrossRef] [PubMed]
- Ozen, E.; Gozukizil, A.; Erdal, E.; Uren, A.; Bottaro, D.P.; Atabey, N. Heparin inhibits Hepatocyte Growth Factor induced motility and invasion of hepatocellular carcinoma cells through early growth response protein 1. PLoS ONE 2012, 7, e42717. [Google Scholar] [CrossRef] [PubMed]
- Lee, J.; Wee, S.; Gunaratne, J.; Chua, R.J.; Smith, R.A.; Ling, L.; Fernig, D.G.; Swaminathan, K.; Nurcombe, V.; Cool, S.M. Structural determinants of heparin-transforming growth factor-β1 interactions and their effects on signaling. Glycobiology 2015, 25, 1491–1504. [Google Scholar] [CrossRef] [PubMed]
- Lyon, M.; Rushton, G.; Gallagher, J.T. The interaction of the transforming growth factor-betas with heparin/heparan sulfate is isoform-specific. J. Biol. Chem. 1997, 272, 18000–18006. [Google Scholar] [CrossRef] [PubMed]
- Catlow, K.R.; Deakin, J.A.; Wei, Z.; Delehedde, M.; Fernig, D.G.; Gherardi, E.; Gallagher, J.T.; Pavão, M.S.; Lyon, M. Interactions of hepatocyte growth factor/scatter factor with various glycosaminoglycans reveal an important interplay between the presence of iduronate and sulfate density. J. Biol. Chem. 2008, 283, 5235–5248. [Google Scholar] [CrossRef] [PubMed]
- Yates, E.A.; Santini, F.; Guerrini, M.; Naggi, A.; Torri, G.; Casu, B. 1H and 13C NMR spectral assignments of the major sequences of twelve systematically modified heparin derivatives. Carbohydr. Res. 1996, 294, 15–27. [Google Scholar] [CrossRef]
- Linhardt, R.J.; Turnbull, J.E.; Wang, H.M.; Loganathan, D.; Gallagher, J.T. Examination of the substrate specificity of heparin and heparan sulfate lyases. Biochemistry 1990, 29, 2611–2617. [Google Scholar] [CrossRef]
- Kim, S.Y.; Zhang, F.; Gong, W.; Chen, K.; Xia, K.; Liu, F.; Gross, R.; Wang, J.M.; Linhardt, R.J.; Cotton, M.L. Copper regulates the interactions of antimicrobial piscidin peptides from fish mast cells with formyl peptide receptors and heparin. J. Biol. Chem. 2018, 293, 15381–15396. [Google Scholar] [CrossRef]
Sample Availability: Samples of heparin/heparin derivatives and GAGs are available from the authors. |

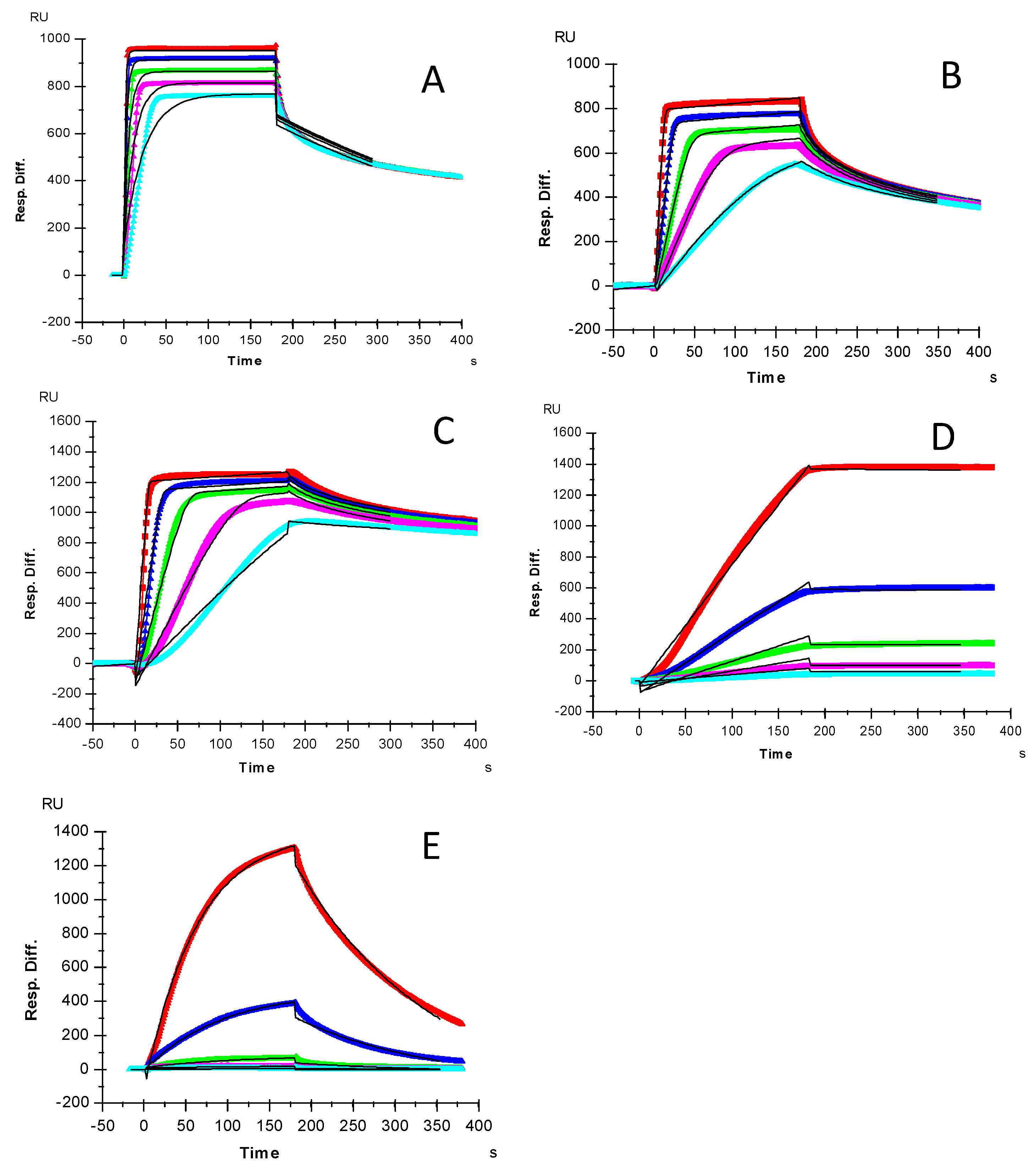
| Interactions | ka (1/MS) | kd (1/S) | KD (M) |
|---|---|---|---|
| FGF2/Heparin | 2.4 × 106 (±5.9 × 104) | 2.9 × 10−3 (±4.7 × 10−5) | 1.2 × 10−9 |
| FGF7/Heparin | 5.2 × 106 (±7.2 × 104) | 0.025 (±3.4 × 10−4) | 4.9 × 10−9 |
| FGF10/Heparin | 5.7 × 106 (±1.7 × 104) | 7.1 × 10−3 (±2.1 × 10−5) | 1.3 × 10−9 |
| HGF/Heparin | 4.2 × 103 (±104) | 5.7 ×10−7 (±3.6 × 10−6) | 1.4 × 10−10 |
| TGFβ-1/heparin | 1.0 × 105 (±342) | 9.2 × 10−3 (±4.0 × 10−5) | 5.9 × 10−8 |
| Interactions | Size | Sulfation | GAGs |
|---|---|---|---|
| FGF2/Heparin | >dp4 | NS > 2S > 6S | Hep >> HS ≈ CSB >> CSE > CSA, C, D, KS |
| FGF7/Heparin | >dp6 | NS > 2S ≈ 6S | Hep >> CSB > HS > CSE > CSA, C, D, KS |
| FGF10/Heparin | >dp6 | NS > 6S > 2S | Hep >> CSB > HS > CSE > CSA, C, D, KS |
| HGF/Heparin | >dp10 | NS > 6S > 2S | Hep >> CSB > HS > CSE > CSA, C, D, KS |
| TGFβ-1/heparin | >dp18 | NS ≈ 2S > 6S | Hep > KS > CSB > HS > CSA, C, D, E |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zhang, F.; Zheng, L.; Cheng, S.; Peng, Y.; Fu, L.; Zhang, X.; Linhardt, R.J. Comparison of the Interactions of Different Growth Factors and Glycosaminoglycans. Molecules 2019, 24, 3360. https://doi.org/10.3390/molecules24183360
Zhang F, Zheng L, Cheng S, Peng Y, Fu L, Zhang X, Linhardt RJ. Comparison of the Interactions of Different Growth Factors and Glycosaminoglycans. Molecules. 2019; 24(18):3360. https://doi.org/10.3390/molecules24183360
Chicago/Turabian StyleZhang, Fuming, Lanhong Zheng, Shuihong Cheng, Yanfei Peng, Li Fu, Xing Zhang, and Robert J. Linhardt. 2019. "Comparison of the Interactions of Different Growth Factors and Glycosaminoglycans" Molecules 24, no. 18: 3360. https://doi.org/10.3390/molecules24183360
APA StyleZhang, F., Zheng, L., Cheng, S., Peng, Y., Fu, L., Zhang, X., & Linhardt, R. J. (2019). Comparison of the Interactions of Different Growth Factors and Glycosaminoglycans. Molecules, 24(18), 3360. https://doi.org/10.3390/molecules24183360






