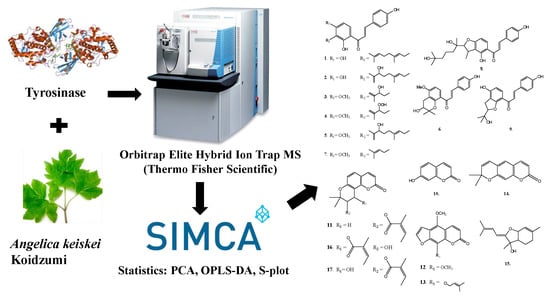
Characterizing Tyrosinase Modulators from the Roots of Angelica keiskei Using Tyrosinase Inhibition Assay and UPLC-MS/MS as the Combinatorial Novel Approach
Abstract
1. Introduction
2. Results
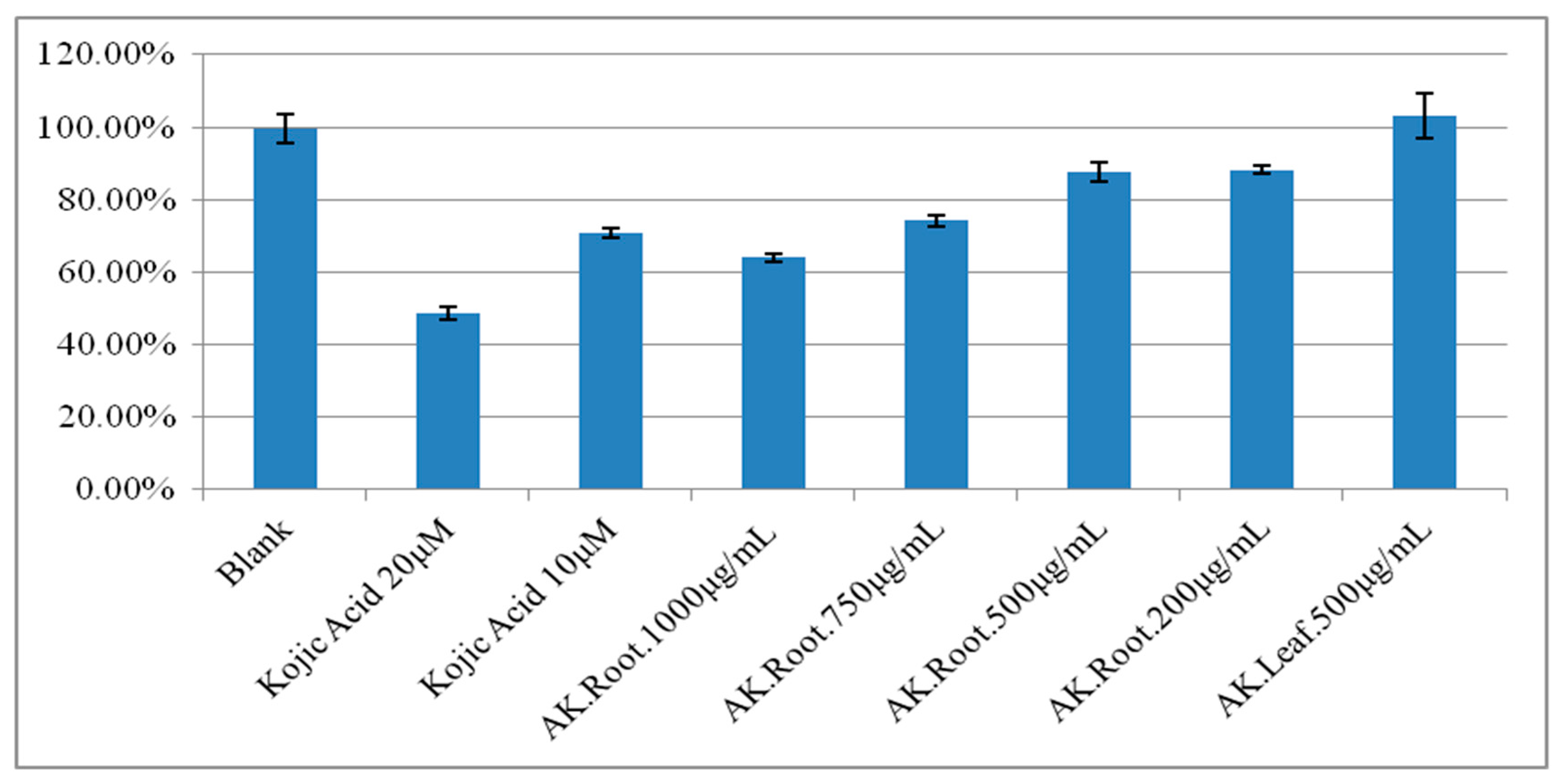
2.1. The Tyrosinase Modulatory Effect of Angelica keiskei
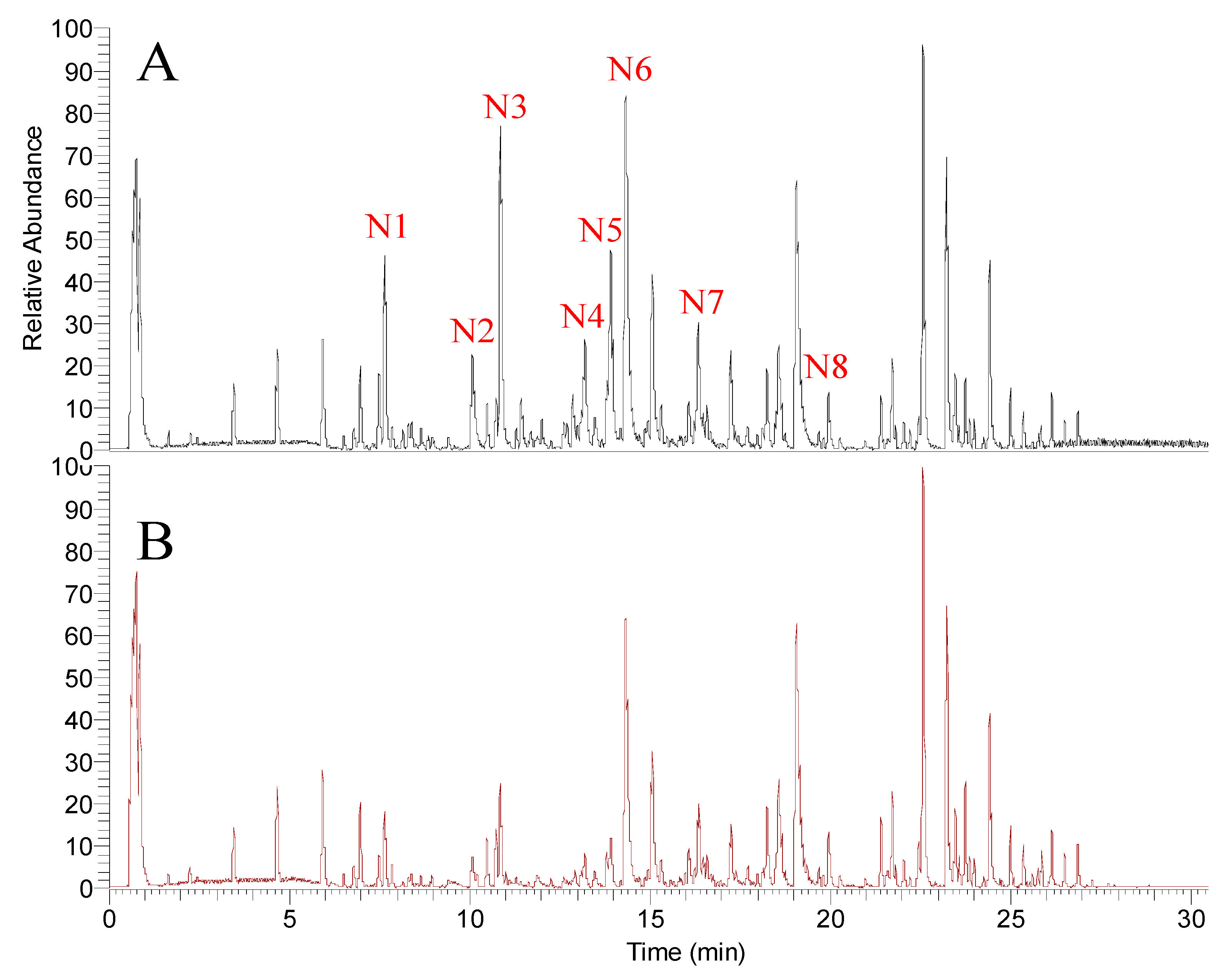
2.2. Screening Tyrosinase Modulator by UPLC-MS/MS
2.3. Multivariate Analysis to Differentiate Processed A. keiskei
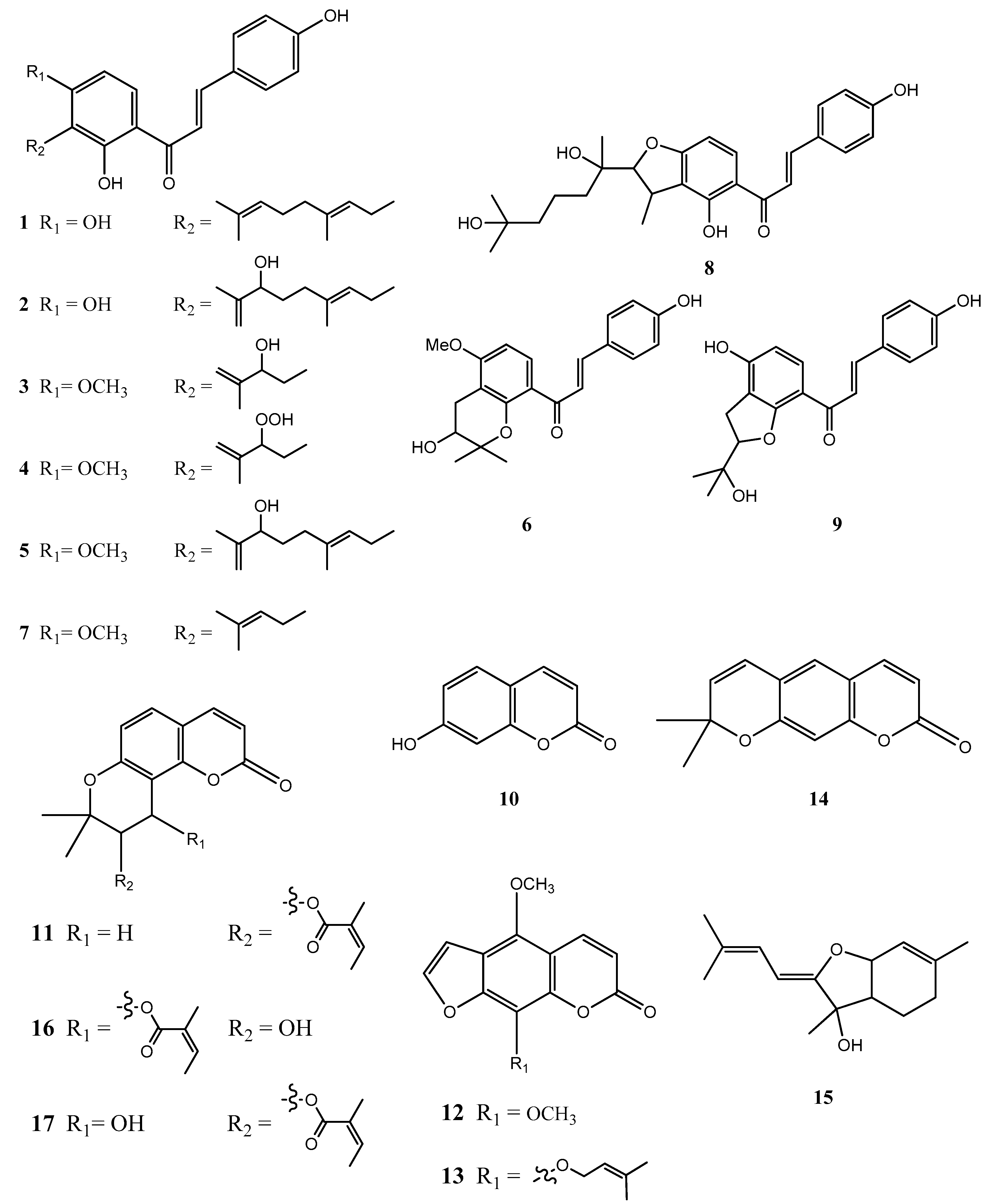
2.4. Structural Characterization of Chalcones and Coumarines
3. Materials and Methods
3.1. Reagents and Materials
3.2. Sample Preparation
3.2.1. Crude Extract Preparation and Effective Compound Isolation
3.2.2. Tyrosinase Assay
- SE = absorbance of sample experiment (with tyrosine)
- SC = absorbance of sample control (without tyrosine)
- EC = absorbance of experimental control (with tyrosine)
- BC = absorbance of blank control (without tyrosine)
3.3. UPLC-MS/MS Analysis
3.4. In Vitro Tyrosinase Inhibition Assay by UPLC-MS/MS Analysis
3.5. Mass Fragmentation Analyses and Compound Identifications
3.6. Statistics
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Korać, R.; Khambholja, K. Potential of herbs in skin protection from ultraviolet radiation. Pharmacogn. Rev. 2011, 5, 164–173. [Google Scholar] [CrossRef] [PubMed]
- D’Mello, S.; Finlay, G.; Baguley, B.; Askarian-Amiri, M. Signaling Pathways in Melanogenesis. Int. J. Mol. Sci. 2016, 17, 1144. [Google Scholar] [CrossRef]
- Pillaiyar, T.; Manickam, M.; Namasivayam, V. Skin whitening agents: Medicinal chemistry perspective of tyrosinase inhibitors. J. Enzym. Inhib. Med. Chem. 2017, 32, 403–425. [Google Scholar]
- Picardo, M.; Maresca, V.; Eibenschutz, L.; De Bernardo, C.; Rinaldi, R.; Grammatico, P. Correlation Between Antioxidants and Phototypes in Melanocytes Cultures. A Possible Link of Physiologic and Pathologic Relevance. J. Investig. Dermatol. 1999, 113, 424–425. [Google Scholar] [PubMed]
- Pillaiyar, T.; Namasivayam, V.; Manickam, M.; Jung, S.-H. Inhibitors of melanogenesis: An updated review. J. Med. Chem. 2018, 61, 7395–7418. [Google Scholar] [PubMed]
- Akihisa, T.; Tokuda, H.; Hasegawa, D.; Ukiya, M.; Kimura, Y.; Enjo, F.; Suzuki, T.; Nishino, H. Chalcones and Other Compounds from the Exudates of Angelica keiskei and Their Cancer Chemopreventive Effects. J. Nat. Prod. 2006, 69, 38–42. [Google Scholar] [CrossRef]
- Kil, Y.-S.; Choi, S.-K.; Lee, Y.-S.; Jafari, M.; Seo, E.-K. Chalcones from Angelica keiskei: Evaluation of Their Heat Shock Protein Inducing Activities. J. Nat. Prod. 2015, 78, 2481–2487. [Google Scholar] [CrossRef]
- Akihisa, T.; Tokuda, H.; Ukiya, M.; Iizuka, M.; Schneider, S.; Ogasawara, K.; Mukainaka, T.; Iwatsuki, K.; Suzuki, T.; Nishino, H. Chalcones, coumarins, and flavanones from the exudate of Angelica keiskei and their chemopreventive effects. Cancer Lett. 2003, 201, 133–137. [Google Scholar] [CrossRef]
- Li, L.; Aldini, G.; Carini, M.; Chen, C.Y.O.; Chun, H.-K.; Cho, S.-M.; Park, K.-M.; Correa, C.R.; Russell, R.M.; Blumberg, J.B.; et al. Characterisation, extraction efficiency, stability and antioxidant activity of phytonutrients in Angelica keiskei. Food Chem. 2009, 115, 227–232. [Google Scholar] [CrossRef]
- Aoki, N.; Muko, M.; Ohta, E.; Ohta, S. C-geranylated chalcones from the stems of Angelica keiskei with superoxide-scavenging activity. J. Nat. Prod. 2008, 71, 1308–1310. [Google Scholar] [CrossRef]
- Kimura, Y.; Baba, K. Antitumor and antimetastatic activities of Angelica keiskei roots, part 1: Isolation of an active substance, xanthoangelol. Int. J. Cancer 2003, 106, 429–437. [Google Scholar] [CrossRef] [PubMed]
- Shin, J.E.; Choi, E.J.; Jin, Q.; Jin, H.G.; Woo, E.R. Chalcones isolated from Angelica keiskei and their inhibition of IL-6 production in TNF-alpha-stimulated MG-63 cell. Arch. Pharmacal Res. 2011, 34, 437–442. [Google Scholar] [CrossRef] [PubMed]
- Luo, L.; Wang, R.; Wang, X.; Ma, Z.; Li, N. Compounds from Angelica keiskei with NQO1 induction, DPPH scavenging and α-glucosidase inhibitory activities. Food Chem. 2012, 131, 992–998. [Google Scholar]
- Caesar, L.; Cech, N. A Review of the Medicinal Uses and Pharmacology of Ashitaba. Planta Med. 2016, 82, 1236–1245. [Google Scholar] [PubMed]
- Chen, Y.; Kong, L.-D.; Xia, X.; Kung, H.-F.; Zhang, L. Behavioral and biochemical studies of total furocoumarins from seeds of Psoralea corylifolia in the forced swimming test in mice. J. Ethnopharmacol. 2005, 96, 451–459. [Google Scholar] [CrossRef]
- Akihisa, T.; Kikuchi, T.; Nagai, H.; Ishii, K.; Tabata, K.; Suzuki, T. 4-Hydroxyderricin from Angelica keiskei roots induces caspase-dependent apoptotic cell death in HL60 human leukemia cells. J. Oleo Sci. 2011, 60, 71–77. [Google Scholar]
- Sumiyoshi, M.; Taniguchi, M.; Baba, K.; Kimura, Y. Antitumor and antimetastatic actions of xanthoangelol and 4-hydroxyderricin isolated from Angelica keiskei roots through the inhibited activation and differentiation of M2 macrophages. Phytomedicine 2015, 22, 759–767. [Google Scholar] [CrossRef]
- Gupta, D.K.; Verma, M.K.; Anand, R.; Khajuria, R.K. Development of a validated UPLC–qTOF-MS/MS method for determination of bioactive constituent from Glycyrrhiza glabra. J. Pharm. Anal. 2013, 3, 205–210. [Google Scholar] [CrossRef]
- Aldini, G.; Regazzoni, L.; Pedretti, A.; Carini, M.; Cho, S.M.; Park, K.M.; Yeum, K.J. An integrated high resolution mass spectrometric and informatics approach for the rapid identification of phenolics in plant extract. J. Chromatogr. A 2011, 1218, 2856–2864. [Google Scholar]
- Arung, E.T.; Furuta, S.; Sugamoto, K.; Shimizu, K.; Ishikawa, H.; Matsushita, Y.-I.; Kondo, R. The inhibitory effects of representative chalcones contained in Angelica keiskei on melanin biosynthesis in B16 melanoma cells. Nat. Prod. Commun. 2012, 7, 1007–1010. [Google Scholar] [CrossRef]
- Asthana, S.; Zucca, P.; Vargiu, A.V.; Sanjust, E.; Ruggerone, P.; Rescigno, A. Structure–Activity Relationship Study of Hydroxycoumarins and Mushroom Tyrosinase. J. Agric. Food Chem. 2015, 63, 7236–7244. [Google Scholar] [CrossRef] [PubMed]
- Chen, C.H.; Chen, P.Y.; Wang, K.C.; Lee, C.K. Rapid identification of the antioxidant constituent of Koelreuteria henryi. J. Chin. Chem. Soc. 2010, 57, 404–410. [Google Scholar] [CrossRef]
- Jayasinghe, L.; Balasooriya, B.A.I.S.; Padmini, W.C.; Hara, N.; Fujimoto, Y. Geranyl chalcone derivatives with antifungal and radical scavenging properties from the leaves of Artocarpus nobilis. Phytochemistry 2004, 65, 1287–1290. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.W.; Curtis-Long, M.J.; Yuk, H.J.; Wang, Y.; Song, Y.H.; Jeong, S.H.; Park, K.H. Quantitative analysis of phenolic metabolites from different parts of Angelica keiskei by HPLC-ESI MS/MS and their xanthine oxidase inhibition. Food Chem. 2014, 153, 20–27. [Google Scholar] [CrossRef] [PubMed]
- Baba, K.; Nakata, K.; Taniguchi, M.; Kido, T.; Kozawa, M. Chalcones from Angelica keiskei. Phytochemistry 1990, 29, 3907–3910. [Google Scholar] [CrossRef]
- Sugamoto, K.; Matsusita, Y.-I.; Matsui, K.; Kurogi, C.; Matsui, T. Synthesis and antibacterial activity of chalcones bearing prenyl or geranyl groups from Angelica keiskei. Tetrahedron 2011, 67, 5346–5359. [Google Scholar] [CrossRef]
- Ashraf, Z.; Rafiq, M.; Seo, S.-Y.; Babar, M.M.; Zaidi, N.-U.-S.S. Design, synthesis and bioevaluation of novel umbelliferone analogues as potential mushroom tyrosinase inhibitors. J. Enzym. Inhib. Med. Chem. 2015, 30, 874–883. [Google Scholar] [CrossRef] [PubMed]
- Orhan, I.; Tosun, F.; Skalicka-Woźniak, K. Cholinesterase, tyrosinase inhibitory and antioxidant potential of randomly selected Umbelliferous plant species and chromatographic profile of Heracleum platytaenium Boiss. and Angelica sylvestris L. var. sylvestris. J. Serb. Chem. Soc. 2016, 81, 357–368. [Google Scholar] [CrossRef]
Sample Availability: Samples of the compounds are not available from the authors. |
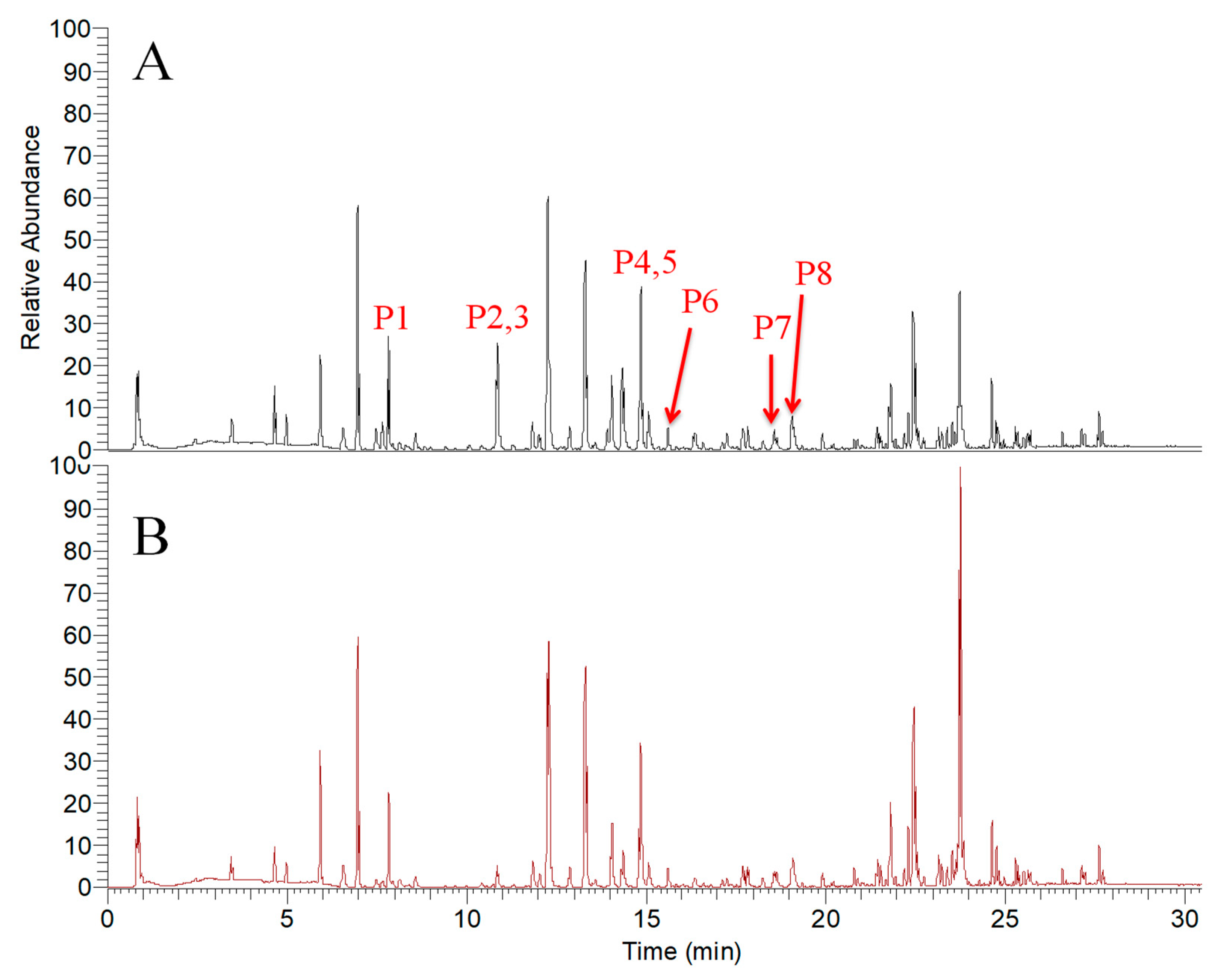
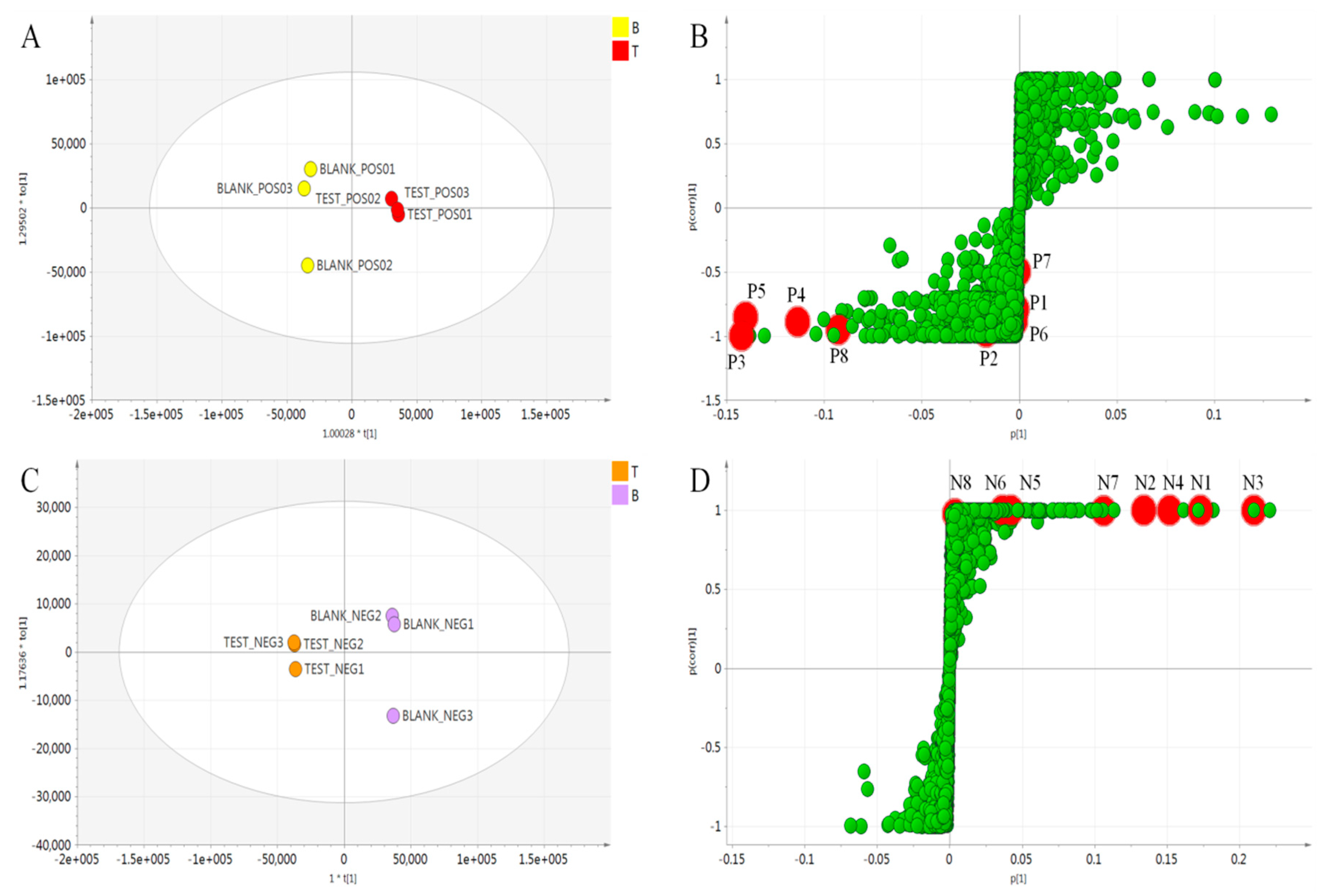
| No. | tR (min) | Measured m/z [M + H]+ | Theoretical Formula [M + H]+ | Error (ppm) | HCD (eV) | Fragment Ions (m/z) Abundance Rate (%) | Identification |
|---|---|---|---|---|---|---|---|
| P1 | 7.93 | 247.0605 | C13H11O5 | 1.8 | 50 | 247.0608 (15) 232.0373 (40) 217.0137 (100) | Isopimpinellin |
| P2 | 10.8 | 163.0387 | C9H7O3 | −1.7 | 50 | 163.0389 (51) 133.0282 (72) 105.0332 (100) | Umbelliferone |
| P3 | 10.84 | 355.1542 | C21H23O5 | 0.5 | 50 | 235.0959 (46) 217.0855 (45) 181.0493 (83) 163.0384 (100) | Xanthoangelol H |
| P4 | 14.86 | 229.0861 | C14H13O3 | 0.9 | 50 | 229.0856 (100) 187.0386 (23) 175.0387 (50) 159.0438 (22) | Xanthyletin |
| P5 | 14.88 | 329.1384 | C19H21O5 | 0.2 | 30 | 245.0807 (54) 227.0686 (100) | Selinidin |
| P6 | 15.52 | 235.1697 | C15H23O2 | 1.7 | 50 | 235.1683 (100) 179.106 (82) | Ashitabaol A |
| P7 | 18.56 | 423.2170 | C26H31O5 | 0.9 | 30 | 283.0945 (100) 181.0499 (23) 163.0375 (42) 131.5666 (18) | Xanthoangelol G |
| P8 | 19.04 | 339.1592 | C21H23O4 | 0.2 | 40 | 283.0945 (32) 181.0484 (99) 163.0379 (100) | 4-Hydroxyderricin |
| No. | tR (min) | Measured m/z [M + H]+ | Theoretical Formula [M + H]+ | Error (ppm) | HCD (eV) | Fragment Ions (m/z) Abundance Rate (%) | Identification |
|---|---|---|---|---|---|---|---|
| N1 | 7.65 | 369.1320 | C21H21O6 | −3.4 | 50 | 369.1389 (100) 297.0811 (1.3) 119.0517 (0.9) | Xanthoangelol E |
| N2 | 10.08 | 299.0905 | C17H15O5 | −2.9 | 40 | 299.0959 (100) 119.0516 (10) | Phellopterin |
| N3 | 10.87 | 353.1370 | C21H21O5 | −3.7 | 50 | 353.1436 (100) 239.1103 (2.4) 119.0518 (32) | Xanthoangelol H |
| N4 | 13.2 | 339.1216 | C20H19O5 | −3.1 | 50 | 339.1274 (100) 119.0517 (17) | (2E)-1-[4-hydroxy-2-(2-hydroxy-2-propanyl)-2,3-dihydro-1-benzofuran-7-yl]-3-(4-hydroxyphenyl)-2-propen-1-one |
| N5 | 13.82 | 439.1737 | C25H27O7 | −3.2 | 50 | 439.1779 (100) 319.1223 (3.4) 119.0525 (2.2) | Xanthokeismin B |
| N6 | 14.32 | 353.1373 | C21H21O5 | −2.9 | 50 | 353.1435 (100) 283.1014 (12) 119.0517 (32) | Xanthoangelol D |
| N7 | 16.35 | 407.1837 | C25H27O5 | −3.8 | 50 | 407.1913 (100) 287.1324 (5.5) 119.0517 (2.2) 243.142 (0.8) | Xanthoangelol B |
| N8 | 19.95 | 391.1892 | C25H27O4 | −3.0 | 50 | 391.1956 (100) 271.137 (11) 119.0518 (4) | Xanthoangelol |
| Compounds | l-tyrosine IC50 (μM) (%) 1 |
|---|---|
| Kojic acid 2 | 3.8 ± 0.2 |
| Xathoangelol | 15.87 ± 1.21 |
| 4-hydroyderricin | 60.14 ± 2.29 |
| Lasepitin | >100 μM (No inhibition at 100 μM) |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, J.-H.; Mei, H.-C.; Kuo, I.-C.; Lee, T.-H.; Chen, Y.-H.; Lee, C.-K. Characterizing Tyrosinase Modulators from the Roots of Angelica keiskei Using Tyrosinase Inhibition Assay and UPLC-MS/MS as the Combinatorial Novel Approach. Molecules 2019, 24, 3297. https://doi.org/10.3390/molecules24183297
Lee J-H, Mei H-C, Kuo I-C, Lee T-H, Chen Y-H, Lee C-K. Characterizing Tyrosinase Modulators from the Roots of Angelica keiskei Using Tyrosinase Inhibition Assay and UPLC-MS/MS as the Combinatorial Novel Approach. Molecules. 2019; 24(18):3297. https://doi.org/10.3390/molecules24183297
Chicago/Turabian StyleLee, Jia-Hao, Hui-Ching Mei, I-Chih Kuo, Tzong-Huei Lee, Yu-Hsin Chen, and Ching-Kuo Lee. 2019. "Characterizing Tyrosinase Modulators from the Roots of Angelica keiskei Using Tyrosinase Inhibition Assay and UPLC-MS/MS as the Combinatorial Novel Approach" Molecules 24, no. 18: 3297. https://doi.org/10.3390/molecules24183297
APA StyleLee, J.-H., Mei, H.-C., Kuo, I.-C., Lee, T.-H., Chen, Y.-H., & Lee, C.-K. (2019). Characterizing Tyrosinase Modulators from the Roots of Angelica keiskei Using Tyrosinase Inhibition Assay and UPLC-MS/MS as the Combinatorial Novel Approach. Molecules, 24(18), 3297. https://doi.org/10.3390/molecules24183297