G-Quadruplex-Based Fluorescent Turn-On Ligands and Aptamers: From Development to Applications
Abstract
1. Introduction
2. Fluorescent Turn-on G-Quadruplex Ligand
2.1. Porphyrins
2.1.1. Application of NMM and its Derivatives (TMPyP4 and TMPipEOPP)
2.1.2. Mechanism of NMM and its Derivatives (TMPyP4 and TMPipEOPP)
2.2. Benzothiozole
2.2.1. Application of ThT, its Derivatives (ThT-DB, ThT-HE, & ThT-NE), and IMT
2.2.2. Mechanism of ThT, ThT-NE, and IMT
2.3. Triphenylmethane (TPM)
2.3.1. Application of CV and MG
2.3.2. Mechanism of CV and MG
2.4. Other Ligands Reported in the Literature as G4 Fluorescence Turn-On Ligands
2.5. Future Perspectives of the Development and Applications of Fluorescent Turn-On Ligands
3. G-Quadruplex-Containing Nucleic Acid Aptamers
3.1. Biosensing Applications of G-Quadruplex-Containing Aptamers
3.2. Bioimaging Applications of G-Quadruplex-Containing Aptamers
3.3. Therapeutic Applications of G-Quadruplex-Containing Aptamers
3.4. Current Challenges and Future Perspectives of the Development and Applications of G4-Containing Aptamers
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Davis, J.T. G-quartets 40 years later: From 5’-GMP to molecular biology and supramolecular chemistry. Angew. Chem. 2004, 43, 668–698. [Google Scholar] [CrossRef] [PubMed]
- The Role of Cations in Determining Quadruplex Structure and Stability. In Quadruplex Nucleic Acids; Neidle, S., Balasubramanian, S., Eds.; The Royal Society of Chemistry: London, UK, 2006; pp. 100–130. [Google Scholar]
- Huppert, J.L.; Balasubramanian, S. Prevalence of quadruplexes in the human genome. Nucleic Acids Res. 2005, 33, 2908–2916. [Google Scholar] [CrossRef] [PubMed]
- Hazel, P.; Huppert, J.; Balasubramanian, S.; Neidle, S. Loop-Length-Dependent Folding of G-Quadruplexes. J. Am. Chem. Soc. 2004, 126, 16405–16415. [Google Scholar] [CrossRef] [PubMed]
- Guedin, A.; Gros, J.; Alberti, P.; Mergny, J.L. How long is too long? Effects of loop size on G-quadruplex stability. Nucleic Acids Res. 2010, 38, 7858–7868. [Google Scholar] [CrossRef] [PubMed]
- Varizhuk, A.; Ischenko, D.; Tsvetkov, V.; Novikov, R.; Kulemin, N.; Kaluzhny, D.; Vlasenok, M.; Naumov, V.; Smirnov, I.; Pozmogova, G. The expanding repertoire of G4 DNA structures. Biochimie 2017, 135, 54–62. [Google Scholar] [CrossRef] [PubMed]
- Mukundan, V.T.; Phan, A.T. Bulges in G-quadruplexes: Broadening the definition of G-quadruplex-forming sequences. J. Am. Chem. Soc. 2013, 135, 5017–5028. [Google Scholar] [CrossRef] [PubMed]
- Das, K.; Srivastava, M.; Raghavan, S.C. GNG Motifs Can Replace a GGG Stretch during G-Quadruplex Formation in a Context Dependent Manner. PLoS ONE 2016, 11, e0158794. [Google Scholar] [CrossRef] [PubMed]
- Qin, M.; Chen, Z.; Luo, Q.; Wen, Y.; Zhang, N.; Jiang, H.; Yang, H. Two-quartet G-quadruplexes formed by DNA sequences containing four contiguous GG runs. J. Phys. Chem. B 2015, 119, 3706–3713. [Google Scholar] [CrossRef]
- Li, X.M.; Zheng, K.W.; Zhang, J.Y.; Liu, H.H.; He, Y.D.; Yuan, B.F.; Hao, Y.H.; Tan, Z. Guanine-vacancy-bearing G-quadruplexes responsive to guanine derivatives. Proc. Natl. Acad. Sci. USA 2015, 112, 14581–14586. [Google Scholar] [CrossRef]
- Heddi, B.; Martin-Pintado, N.; Serimbetov, Z.; Kari, T.M.; Phan, A.T. G-quadruplexes with (4n-1) guanines in the G-tetrad core: Formation of a G-triad.water complex and implication for small-molecule binding. Nucleic Acids Res. 2016, 44, 910–916. [Google Scholar] [CrossRef]
- Lim, K.W.; Phan, A.T. Structural basis of DNA quadruplex-duplex junction formation. Angew. Chem. 2013, 52, 8566–8569. [Google Scholar] [CrossRef] [PubMed]
- Bing, T.; Zheng, W.; Zhang, X.; Shen, L.; Liu, X.; Wang, F.; Cui, J.; Cao, Z.; Shangguan, D. Triplex-quadruplex structural scaffold: A new binding structure of aptamer. Sci. Rep. 2017, 7, 15467. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Wu, Y.; Zhang, W. G-quadruplex structures and their interaction diversity with ligands. ChemMedChem 2014, 9, 899–911. [Google Scholar] [CrossRef] [PubMed]
- Dvorkin, S.A.; Karsisiotis, A.I.; Webba da Silva, M. Encoding canonical DNA quadruplex structure. Sci. Adv. 2018, 4, eaat3007. [Google Scholar] [CrossRef] [PubMed]
- Rhodes, D.; Lipps, H.J. G-quadruplexes and their regulatory roles in biology. Nucleic Acids Res. 2015, 43, 8627–8637. [Google Scholar] [CrossRef] [PubMed]
- Fay, M.M.; Lyons, S.M.; Ivanov, P. RNA G-Quadruplexes in Biology: Principles and Molecular Mechanisms. J. Mol. Biol. 2017, 429, 2127–2147. [Google Scholar] [CrossRef] [PubMed]
- Hansel-Hertsch, R.; Di Antonio, M.; Balasubramanian, S. DNA G-quadruplexes in the human genome: Detection, functions and therapeutic potential. Nat. Rev. Mol. Cell Biol. 2017, 18, 279–284. [Google Scholar] [CrossRef]
- Harris, L.M.; Merrick, C.J. G-quadruplexes in pathogens: A common route to virulence control? PLoS Pathog. 2015, 11, e1004562. [Google Scholar] [CrossRef]
- Kikin, O.; D’Antonio, L.; Bagga, P.S. QGRS Mapper: A web-based server for predicting G-quadruplexes in nucleotide sequences. Nucleic Acids Res. 2006, 34, W676–W682. [Google Scholar] [CrossRef]
- Bedrat, A.; Lacroix, L.; Mergny, J.-L. Re-evaluation of G-quadruplex propensity with G4Hunter. Nucleic Acids Res. 2016, 44, 1746–1759. [Google Scholar] [CrossRef]
- Garant, J.M.; Perreault, J.P.; Scott, M.S. Motif independent identification of potential RNA G-quadruplexes by G4RNA screener. Bioinformatics (Oxf. Engl.) 2017, 33, 3532–3537. [Google Scholar] [CrossRef] [PubMed]
- Sahakyan, A.B.; Chambers, V.S.; Marsico, G.; Santner, T.; Di Antonio, M.; Balasubramanian, S. Machine learning model for sequence-driven DNA G-quadruplex formation. Sci. Rep. 2017, 7, 14535. [Google Scholar] [CrossRef] [PubMed]
- del Villar-Guerra, R.; Trent, J.O.; Chaires, J.B. G-Quadruplex Secondary Structure Obtained from Circular Dichroism Spectroscopy. Angew. Chem. Int. Ed. Engl. 2018, 57, 7171–7175. [Google Scholar] [CrossRef] [PubMed]
- Mergny, J.-L.; Phan, A.-T.; Lacroix, L. Following G-quartet formation by UV-spectroscopy. FEBS Lett. 1998, 435, 74–78. [Google Scholar] [CrossRef]
- Scalabrin, M.; Palumbo, M.; Richter, S.N. Highly Improved Electrospray Ionization-Mass Spectrometry Detection of G-Quadruplex-Folded Oligonucleotides and Their Complexes with Small Molecules. Anal. Chem. 2017, 89, 8632–8637. [Google Scholar] [CrossRef]
- Webba da Silva, M. NMR methods for studying quadruplex nucleic acids. Methods (San Diego Calif.) 2007, 43, 264–277. [Google Scholar] [CrossRef] [PubMed]
- Mendez, M.A.; Szalai, V.A. Fluorescence of unmodified oligonucleotides: A tool to probe G-quadruplex DNA structure. Biopolymers 2009, 91, 841–850. [Google Scholar] [CrossRef]
- Han, H.; Hurley, L.H.; Salazar, M. A DNA polymerase stop assay for G-quadruplex-interactive compounds. Nucleic Acids Res. 1999, 27, 537–542. [Google Scholar] [CrossRef]
- Williamson, J.R.; Raghuraman, M.K.; Cech, T.R. Monovalent cation-induced structure of telomeric DNA: The G-quartet model. Cell 1989, 59, 871–880. [Google Scholar] [CrossRef]
- Kwok, C.K.; Balasubramanian, S. Targeted Detection of G-Quadruplexes in Cellular RNAs. Angew. Chem. Int. Ed. Engl. 2015, 54, 6751–6754. [Google Scholar] [CrossRef]
- Kwok, C.K.; Sahakyan, A.B.; Balasubramanian, S. Structural Analysis using SHALiPE to Reveal RNA G-Quadruplex Formation in Human Precursor MicroRNA. Angew. Chem. Int. Ed. 2016, 55, 8958–8961. [Google Scholar] [CrossRef] [PubMed]
- Chambers, V.S.; Marsico, G.; Boutell, J.M.; Di Antonio, M.; Smith, G.P.; Balasubramanian, S. High-throughput sequencing of DNA G-quadruplex structures in the human genome. Nat. Biotechnol. 2015, 33, 877–881. [Google Scholar] [CrossRef] [PubMed]
- Hänsel-Hertsch, R.; Beraldi, D.; Lensing, S.V.; Marsico, G.; Zyner, K.; Parry, A.; Di Antonio, M.; Pike, J.; Kimura, H.; Narita, M.; et al. G-quadruplex structures mark human regulatory chromatin. Nat. Genet. 2016, 48, 1267. [Google Scholar] [CrossRef] [PubMed]
- Kwok, C.K.; Marsico, G.; Sahakyan, A.B. rG4-seq reveals widespread formation of G-quadruplex structures in the human transcriptome. Nat. Methods 2016, 13, 841–844. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.U.; Bartel, D.P. RNA G-quadruplexes are globally unfolded in eukaryotic cells and depleted in bacteria. Science ( N.Y.) 2016, 353, aaf5371. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.Y.; Lejault, P.; Chevrier, S.; Boidot, R.; Robertson, A.G.; Wong, J.M.Y.; Monchaud, D. Transcriptome-wide identification of transient RNA G-quadruplexes in human cells. Nat. Commun. 2018, 9, 4730. [Google Scholar] [CrossRef] [PubMed]
- Collie, G.W.; Parkinson, G.N. The application of DNA and RNA G-quadruplexes to therapeutic medicines. Chem. Soc. Rev. 2011, 40, 5867–5892. [Google Scholar] [CrossRef] [PubMed]
- Tian, T.; Xiao, H.; Zhou, X. A Review: G-Quadruplex’s Applications in Biological Target Detection and Drug Delivery. Curr. Top. Med. Chem. 2015, 15, 1988–2001. [Google Scholar] [CrossRef]
- Platella, C.; Riccardi, C.; Montesarchio, D.; Roviello, G.N.; Musumeci, D. G-quadruplex-based aptamers against protein targets in therapy and diagnostics. Biochim. Biophys. Acta Gener. Subj. 2017, 1861, 1429–1447. [Google Scholar] [CrossRef]
- Michalowski, D.; Chitima-Matsiga, R.; Held, D.M.; Burke, D.H. Novel bimodular DNA aptamers with guanosine quadruplexes inhibit phylogenetically diverse HIV-1 reverse transcriptases. Nucleic Acids Res. 2008, 36, 7124–7135. [Google Scholar] [CrossRef]
- Andreola, M.-L.; Pileur, F.; Calmels, C.; Ventura, M.; Tarrago-Litvak, L.; Toulmé, J.-J.; Litvak, S. DNA Aptamers Selected against the HIV-1 RNase H Display in Vitro Antiviral Activity. Biochemistry 2001, 40, 10087–10094. [Google Scholar] [CrossRef] [PubMed]
- Mashima, T.; Matsugami, A.; Nishikawa, F.; Nishikawa, S.; Katahira, M. Unique quadruplex structure and interaction of an RNA aptamer against bovine prion protein. Nucleic Acids Res. 2009, 37, 6249–6258. [Google Scholar] [CrossRef] [PubMed]
- Rosenberg, J.E.; Bambury, R.M.; Van Allen, E.M.; Drabkin, H.A.; Lara, P.N., Jr.; Harzstark, A.L.; Wagle, N.; Figlin, R.A.; Smith, G.W.; Garraway, L.A.; et al. A phase II trial of AS1411 (a novel nucleolin-targeted DNA aptamer) in metastatic renal cell carcinoma. Investig. New Drugs 2014, 32, 178–187. [Google Scholar] [CrossRef] [PubMed]
- Weerasinghe, P.; Li, Y.; Guan, Y.; Zhang, R.; Tweardy, D.J.; Jing, N. T40214/PEI complex: A potent therapeutics for prostate cancer that targets STAT3 signaling. Prostate 2008, 68, 1430–1442. [Google Scholar] [CrossRef] [PubMed]
- Kwok, C.K.; Tang, Y.; Assmann, S.M.; Bevilacqua, P.C. The RNA structurome: Transcriptome-wide structure probing with next-generation sequencing. Trends Biochem. Sci. 2015, 40, 221–232. [Google Scholar] [CrossRef] [PubMed]
- Kwok, C.K.; Marsico, G.; Balasubramanian, S. Detecting RNA G-Quadruplexes (rG4s) in the Transcriptome. Cold Spring Harb. Perspect. Biol. 2018, 10. [Google Scholar] [CrossRef]
- Kwok, C.K.; Merrick, C.J. G-Quadruplexes: Prediction, Characterization, and Biological Application. Trends Biotechnol. 2017, 35, 997–1013. [Google Scholar] [CrossRef]
- Chan, K.L.; Peng, B.; Umar, M.I.; Chan, C.Y.; Sahakyan, A.B.; Le, M.T.N.; Kwok, C.K. Structural analysis reveals the formation and role of RNA G-quadruplex structures in human mature microRNAs. Chem. Commun. 2018, 54, 10878–10881. [Google Scholar] [CrossRef]
- Neidle, S.; Balasubramanian, S. Quadruplex Nucleic Acids; The Royal Society of Chemistry: London, UK, 2006. [Google Scholar]
- Zhang, S.; Sun, H.; Wang, L.; Liu, Y.; Chen, H.; Li, Q.; Guan, A.; Liu, M.; Tang, Y. Real-time monitoring of DNA G-quadruplexes in living cells with a small-molecule fluorescent probe. Nucleic Acids Res. 2018, 46, 7522–7532. [Google Scholar] [CrossRef]
- Shivalingam, A.; Izquierdo, M.A.; Le Marois, A.; Vysniauskas, A.; Suhling, K.; Kuimova, M.K.; Vilar, R. The interactions between a small molecule and G-quadruplexes are visualized by fluorescence lifetime imaging microscopy. Nat. Commun. 2015, 6, 8178. [Google Scholar] [CrossRef]
- Ma, D.L.; Dong, Z.Z.; Vellaisamy, K.; Cheung, K.M.; Yang, G.; Leung, C.H. Luminescent Strategies for Label-Free G-Quadruplex-Based Enzyme Activity Sensing. Chem. Rec. 2017, 17, 1135–1145. [Google Scholar] [CrossRef] [PubMed]
- Ma, D.L.; Zhang, Z.; Wang, M.; Lu, L.; Zhong, H.J.; Leung, C.H. Recent Developments in G-Quadruplex Probes. Chem. Biol. 2015, 22, 812–828. [Google Scholar] [CrossRef] [PubMed]
- Ma, D.L.; Wang, M.; Lin, S.; Han, Q.B.; Leung, C.H. Recent Development of G-Quadruplex Probes for Cellular Imaging. Curr. Top. Med. Chem. 2015, 15, 1957–1963. [Google Scholar] [PubMed]
- Vummidi, B.R.; Alzeer, J.; Luedtke, N.W. Fluorescent probes for G-quadruplex structures. ChemBioChem Eur. J. Chem. Biol. 2013, 14, 540–558. [Google Scholar] [CrossRef] [PubMed]
- Leung, K.H.; He, H.Z.; Ma, V.P.; Chan, D.S.; Leung, C.H.; Ma, D.L. A luminescent G-quadruplex switch-on probe for the highly selective and tunable detection of cysteine and glutathione. Chem. Commun. 2013, 49, 771–773. [Google Scholar] [CrossRef] [PubMed]
- Dong, Z.-Z.; Lu, L.; Wang, W.; Li, G.; Kang, T.-S.; Han, Q.; Leung, C.-H.; Ma, D.-L. Luminescent detection of nicking endonuclease Nb.BsmI activity by using a G-quadruplex-selective iridium(III) complex in aqueous solution. Sens. Actuators B Chem. 2017, 246, 826–832. [Google Scholar] [CrossRef]
- Lin, S.; Lu, L.; Liu, J.B.; Liu, C.; Kang, T.S.; Yang, C.; Leung, C.H.; Ma, D.L. A G-quadruplex-selective luminescent iridium(III) complex and its application by long lifetime. Biochim. Biophys. Acta. Gener. Subj. 2017, 1861, 1448–1454. [Google Scholar] [CrossRef]
- Lin, S.; Kang, T.S.; Lu, L.; Wang, W.; Ma, D.L.; Leung, C.H. A G-quadruplex-selective luminescent probe with an anchor tail for the switch-on detection of thymine DNA glycosylase activity. Biosens. Bioelectron. 2016, 86, 849–857. [Google Scholar] [CrossRef]
- Lin, S.; Lu, L.; Kang, T.S.; Mergny, J.L.; Leung, C.H.; Ma, D.L. Interaction of an Iridium(III) Complex with G-Quadruplex DNA and Its Application in Luminescent Switch-On Detection of Siglec-5. Anal. Chem. 2016, 88, 10290–10295. [Google Scholar] [CrossRef]
- Lu, L.; Wang, W.; Yang, C.; Kang, T.-S.; Leung, C.-H.; Ma, D.-L. Iridium(iii) complexes with 1,10-phenanthroline-based N^N ligands as highly selective luminescent G-quadruplex probes and application for switch-on ribonuclease H detection. J. Mater. Chem. B 2016, 4, 6791–6796. [Google Scholar] [CrossRef]
- Ma, D.-L.; Che, C.-M.; Yan, S.-C. Platinum(II) Complexes with Dipyridophenazine Ligands as Human Telomerase Inhibitors and Luminescent Probes for G-Quadruplex DNA. J. Am. Chem. Soc. 2009, 131, 12. [Google Scholar] [CrossRef] [PubMed]
- Yat-Wah Man, B.; Shiu-Hin Chan, D.; Yang, H.; Ang, S.-W.; Yang, F.; Yan, S.-C.; Ho, C.-M.; Wu, P.; Che, C.-M.; Leungy, C.-H.; et al. A selective G-quadruplex-based luminescent switch-on probe for the detection of nanomolar silver(I) ions in aqueous solutionw. Chem. Commun. 2010, 46, 3. [Google Scholar] [CrossRef]
- Wang, P.; Leung, C.H.; Ma, D.L.; Yan, S.C.; Che, C.M. Structure-based design of platinum(II) complexes as c-myc oncogene down-regulators and luminescent probes for G-quadruplex DNA. Chemistry 2010, 16, 6900–6911. [Google Scholar] [CrossRef] [PubMed]
- Suntharalingam, K.; White, A.J.; Vilar, R. Synthesis, structural characterization, and quadruplex DNA binding studies of platinum(II)-terpyridine complexes. Inorg. Chem. 2009, 48, 9427–9435. [Google Scholar] [CrossRef] [PubMed]
- Naud-Martin, D.; Landras-Guetta, C.; Verga, D.; Ghosh, D.; Achelle, S.; Mahuteau-Betzer, F.; Bombard, S.; Teulade-Fichou, M.P. Selectivity of Terpyridine Platinum Anticancer Drugs for G-quadruplex DNA. Molecules 2019, 24, 404. [Google Scholar]
- Piraux, G.; Bar, L.; Abraham, M.; Lavergne, T.; Jamet, H.; Dejeu, J.; Marcelis, L.; Defrancq, E.; Elias, B. New Ruthenium-Based Probes for Selective G-Quadruplex Targeting. Chemistry 2017, 23, 11872–11880. [Google Scholar] [CrossRef] [PubMed]
- Wachter, E.; Howerton, B.S.; Hall, E.C.; Parkin, S.; Glazer, E.C. A new type of DNA “light-switch”: A dual photochemical sensor and metalating agent for duplex and G-quadruplex DNA. Chem. Commun. 2014, 50, 311–313. [Google Scholar] [CrossRef]
- Wachter, E.; Moya, D.; Parkin, S.; Glazer, E.C. Ruthenium Complex “Light Switches” that are Selective for Different G-Quadruplex Structures. Chemistry 2016, 22, 550–559. [Google Scholar] [CrossRef]
- Gama, S.; Rodrigues, I.; Mendes, F.; Santos, I.C.; Gabano, E.; Klejevskaja, B.; Gonzalez-Garcia, J.; Ravera, M.; Vilar, R.; Paulo, A. Anthracene-terpyridine metal complexes as new G-quadruplex DNA binders. J. Inorg. Biochem. 2016, 160, 275–286. [Google Scholar] [CrossRef]
- Tong, L.L.; Li, L.; Chen, Z.; Wang, Q.; Tang, B. Stable label-free fluorescent sensing of biothiols based on ThT direct inducing conformation-specific G-quadruplex. Biosens. Bioelectron. 2013, 49, 420–425. [Google Scholar] [CrossRef]
- Huang, H.; Zhang, J.; Harvey, S.E.; Hu, X.; Cheng, C. RNA G-quadruplex secondary structure promotes alternative splicing via the RNA-binding protein hnRNPF. Genes Dev. 2017, 31, 2296–2309. [Google Scholar] [CrossRef] [PubMed]
- Tang, X.; Wu, K.; Zhao, H.; Chen, M.; Ma, C. A Label-Free Fluorescent Assay for the Rapid and Sensitive Detection of Adenosine Deaminase Activity and Inhibition. Sensors 2018, 18, 2441. [Google Scholar] [CrossRef] [PubMed]
- Huber, M.D.; Lee, D.C.; Maizels, N. G4 DNA unwinding by BLM and Sgs1p: Substrate specificity and substrate-specific inhibition. Nucleic Acids Res. 2002, 30, 3954–3961. [Google Scholar] [CrossRef] [PubMed]
- Hu, D.; Pu, F.; Huang, Z.; Ren, J.; Qu, X. A quadruplex-based, label-free, and real-time fluorescence assay for RNase H activity and inhibition. Chemistry 2010, 16, 2605–2610. [Google Scholar] [CrossRef] [PubMed]
- Yingfu Li, C.R.G.; Sen, D. Recognition of Anionic Porphyrins by DNA Aptamers. Biochemistry 1996, 35, 12. [Google Scholar] [CrossRef]
- Haribabu Arthanari, S.B.; Kawano, T.L.; Bolton, P.H. Fluorescent dyes specific for quadruplex DNA. Nucleic Acids Res. 1998, 26, 5. [Google Scholar] [CrossRef]
- Nicoludis, J.M.; Barrett, S.P.; Mergny, J.L.; Yatsunyk, L.A. Interaction of human telomeric DNA with N-methyl mesoporphyrin IX. Nucleic Acids Res. 2012, 40, 5432–5447. [Google Scholar] [CrossRef]
- Tippana, R.; Xiao, W.; Myong, S. G-quadruplex conformation and dynamics are determined by loop length and sequence. Nucleic Acids Res. 2014, 42, 8106–8114. [Google Scholar] [CrossRef]
- Sabharwal, N.C.; Savikhin, V.; Turek-Herman, J.R.; Nicoludis, J.M.; Szalai, V.A.; Yatsunyk, L.A. N-methylmesoporphyrin IX fluorescence as a reporter of strand orientation in guanine quadruplexes. FEBS J. 2014, 281, 1726–1737. [Google Scholar] [CrossRef]
- Ren, J.; Qin, H.; Wang, J.; Luedtke, N.W.; Wang, E.; Wang, J. Label-free detection of nucleic acids by turn-on and turn-off G-quadruplex-mediated fluorescence. Ana. Bioanal. Chem. 2011, 399, 2763–2770. [Google Scholar] [CrossRef]
- Li, M.; Zhao, A.; Ren, J.; Qu, X. N-Methyl Mesoporphyrin IX as an Effective Probe for Monitoring Alzheimer’s Disease beta-Amyloid Aggregation in Living Cells. ACS Chem. Neurosci. 2017, 8, 1299–1304. [Google Scholar] [CrossRef] [PubMed]
- Sun, Y.; Zhao, C.; Yan, Z.; Ren, J.; Qu, X. Simple and sensitive microbial pathogen detection using a label-free DNA amplification assay. Chem. Commun. 2016, 52, 5. [Google Scholar] [CrossRef] [PubMed]
- Waller, Z.A.; Pinchbeck, B.J.; Buguth, B.S.; Meadows, T.G.; Richardson, D.J.; Gates, A.J. Control of bacterial nitrate assimilation by stabilization of G-quadruplex DNA. Chem. Commun. 2016, 52, 13511–13514. [Google Scholar] [CrossRef] [PubMed]
- Zhu, L.N.; Zhao, S.J.; Wu, B.; Li, X.Z.; Kong, D.M. A new cationic porphyrin derivative (TMPipEOPP) with large side arm substituents: A highly selective G-quadruplex optical probe. PLoS ONE 2012, 7, e35586. [Google Scholar] [CrossRef] [PubMed]
- Gabelica, V.; Maeda, R.; Fujimoto, T.; Yaku, H.; Murashima, T.; Sugimoto, N.; Miyoshi, D. Multiple and cooperative binding of fluorescence light-up probe thioflavin T with human telomere DNA G-quadruplex. Biochemistry 2013, 52, 5620–5628. [Google Scholar] [CrossRef] [PubMed]
- Paramasivan, S.; Bolton, P.H. Mix and measure fluorescence screening for selective quadruplex binders. Nucleic Acids Res. 2008, 36, e106. [Google Scholar] [CrossRef] [PubMed]
- Hu, D.; Huang, Z.; Pu, F.; Ren, J.; Qu, X. A label-free, quadruplex-based functional molecular beacon (LFG4-MB) for fluorescence turn-on detection of DNA and nuclease. Chemistry 2011, 17, 1635–1641. [Google Scholar] [CrossRef]
- Zhao, C.; Wu, L.; Ren, J.; Qu, X. A label-free fluorescent turn-on enzymatic amplification assay for DNA detection using ligand-responsive G-quadruplex formation. Chem. Commun. 2011, 47, 5461–5463. [Google Scholar] [CrossRef]
- Yao, X.; Song, D.; Qin, T.; Yang, C.; Yu, Z.; Li, X.; Liu, K.; Su, H. Interaction between G-Quadruplex and Zinc Cationic Porphyrin: The Role of the Axial Water. Sci. Rep. 2017, 7, 10951. [Google Scholar] [CrossRef]
- Wheelhouse, R.T.; Sun, D.; Han, H.; Han, F.X.; Hurley, L.H. Cationic Porphyrins as Telomerase Inhibitors: The Interaction of Tetra-(N-methyl-4-pyridyl)porphine with Quadruplex DNA. J. Am. Chem. Soc. 1998, 120, 2. [Google Scholar] [CrossRef]
- Biancalana, M.; Koide, S. Molecular mechanism of Thioflavin-T binding to amyloid fibrils. Biochim. Biophysi. Acta 2010, 1804, 1405–1412. [Google Scholar] [CrossRef] [PubMed]
- Inbar, P.; Li, C.Q.; Takayama, S.A.; Bautista, M.R.; Yang, J. Oligo(ethylene glycol) derivatives of thioflavin T as inhibitors of protein-amyloid interactions. ChemBioChem Eur. J. Chem. Biol. 2006, 7, 1563–1566. [Google Scholar] [CrossRef]
- Mohanty, J.; Barooah, N.; Dhamodharan, V.; Harikrishna, S.; Pradeepkumar, P.I.; Bhasikuttan, A.C. Thioflavin T as an efficient inducer and selective fluorescent sensor for the human telomeric G-quadruplex DNA. J. Am. Chem. Soc. 2013, 135, 367–376. [Google Scholar] [CrossRef] [PubMed]
- Liu, S.; Peng, P.; Wang, H.; Shi, L.; Li, T. Thioflavin T binds dimeric parallel-stranded GA-containing non-G-quadruplex DNAs: A general approach to lighting up double-stranded scaffolds. Nucleic Acids Res. 2017, 45, 12080–12089. [Google Scholar] [CrossRef]
- Renaud de la Faverie, A.; Guedin, A.; Bedrat, A.; Yatsunyk, L.A.; Mergny, J.L. Thioflavin T as a fluorescence light-up probe for G4 formation. Nucleic Acids Res. 2014, 42, e65. [Google Scholar] [CrossRef] [PubMed]
- Lee, I.J.; Patil, S.P.; Fhayli, K.; Alsaiari, S.; Khashab, N.M. Probing structural changes of self assembled i-motif DNA. Chem. Commun. 2015, 51, 3747–3749. [Google Scholar] [CrossRef] [PubMed]
- Liu, X.; Hua, X.; Fan, Q.; Chao, J.; Su, S.; Huang, Y.Q.; Wang, L.; Huang, W. Thioflavin T as an Efficient G-Quadruplex Inducer for the Highly Sensitive Detection of Thrombin Using a New Foster Resonance Energy Transfer System. ACS Appl. Mater. Interfaces 2015, 7, 16458–16465. [Google Scholar] [CrossRef]
- Zeng, H.; Zhu, Y.; Ma, L.; Xia, X.; Li, Y.; Ren, Y.; Zhao, W.; Yang, H.; Deng, R. G-quadruplex specific dye-based ratiometric FRET aptasensor for robust and ultrafast detection of toxin. Dyes Pigments 2019, 164, 35–42. [Google Scholar] [CrossRef]
- Yuka Kataoka, H.F.; Kasahara, Y.; Yoshihara, T.; Tobita, S.; Kuwahara, M. Minimal Thioflavin T Modifications Improve Visual Discrimination of Guanine-Quadruplex Topologies and Alter Compound-induced Topological Structures. Anal. Chem. 2014, 86, 12078–12084. [Google Scholar] [CrossRef] [PubMed]
- Zhang, S.; Sun, H.; Chen, H.; Li, Q.; Guan, A.; Wang, L.; Shi, Y.; Xu, S.; Liu, M.; Tang, Y. Direct visualization of nucleolar G-quadruplexes in live cells by using a fluorescent light-up probe. Biochim. Biophys. Acta Gener. Subj. 2018, 1862, 1101–1106. [Google Scholar] [CrossRef]
- Guan, A.J.; Zhang, X.F.; Sun, X.; Li, Q.; Xiang, J.F.; Wang, L.X.; Lan, L.; Yang, F.M.; Xu, S.J.; Guo, X.M.; et al. Ethyl-substitutive Thioflavin T as a highly-specific fluorescence probe for detecting G-quadruplex structure. Sci. Rep. 2018, 8, 2666. [Google Scholar] [CrossRef] [PubMed]
- Fleming, A.M.; Ding, Y.; Alenko, A.; Burrows, C.J. Zika Virus Genomic RNA Possesses Conserved G-Quadruplexes Characteristic of the Flaviviridae Family. ACS Infect. Dis. 2016, 2, 674–681. [Google Scholar] [CrossRef] [PubMed]
- Vinyard, W.A.; Fleming, A.M.; Ma, J.; Burrows, C.J. Characterization of G-Quadruplexes in Chlamydomonas reinhardtii and the Effects of Polyamine and Magnesium Cations on Structure and Stability. Biochemistry 2018, 57, 6551–6561. [Google Scholar] [CrossRef]
- Zahin, M.; Dean, W.L.; Ghim, S.J.; Joh, J.; Gray, R.D.; Khanal, S.; Bossart, G.D.; Mignucci-Giannoni, A.A.; Rouchka, E.C.; Jenson, A.B.; et al. Identification of G-quadruplex forming sequences in three manatee papillomaviruses. PLoS ONE 2018, 13, 23. [Google Scholar] [CrossRef] [PubMed]
- Luo, X.; Xue, B.; Feng, G.; Zhang, J.; Lin, B.; Zeng, P.; Li, H.; Yi, H.; Zhang, X.L.; Zhu, H.; et al. Lighting up the Native Viral RNA Genome with a Fluorogenic Probe for the Live-Cell Visualization of Virus Infection. J. Am. Chem. Soc. 2019, 141, 5182–5191. [Google Scholar] [CrossRef]
- Laguerre, A.; Hukezalie, K.; Winckler, P.; Katranji, F.; Chanteloup, G.; Pirrotta, M.; Perrier-Cornet, J.M.; Wong, J.M.; Monchaud, D. Visualization of RNA-Quadruplexes in Live Cells. J. Am. Chem. Soc. 2015, 137, 8521–8525. [Google Scholar] [CrossRef]
- Jeng, S.C.; Chan, H.H.; Booy, E.P.; McKenna, S.A.; Unrau, P.J. Fluorophore ligand binding and complex stabilization of the RNA Mango and RNA Spinach aptamers. RNA 2016, 22, 1884–1892. [Google Scholar] [CrossRef]
- Xu, S.; Li, Q.; Xiang, J.; Yang, Q.; Sun, H.; Guan, A.; Wang, L.; Liu, Y.; Yu, L.; Shi, Y.; et al. Thioflavin T as an efficient fluorescence sensor for selective recognition of RNA G-quadruplexes. Sci. Rep. 2016, 6, 24793. [Google Scholar] [CrossRef] [PubMed]
- Guo, J.H.; Zhu, L.N.; Kong, D.M.; Shen, H.X. Triphenylmethane dyes as fluorescent probes for G-quadruplex recognition. Talanta 2009, 80, 607–613. [Google Scholar] [CrossRef] [PubMed]
- Stanoeva, T.; Neshchadin, D.; Gescheidt, G.; Ludvik, J.; Lajoie, B.; Batchelor, S.N. An Investigation into the Initial Degradation Steps of Four Major Dye Chromophores: Study of Their One-Electron Oxidation and Reduction by EPR, ENDOR, Cyclic Voltammetry, and Theoretical Calculations. J. Phys. Chem. A 2005, 109, 7. [Google Scholar] [CrossRef]
- Kong, D.M.; Ma, Y.E.; Wu, J.; Shen, H.X. Discrimination of G-quadruplexes from duplex and single-stranded DNAs with fluorescence and energy-transfer fluorescence spectra of crystal violet. Chemistry 2009, 15, 901–909. [Google Scholar] [CrossRef] [PubMed]
- Li, F.; Feng, Y.; Zhao, C.; Tang, B. Crystal violet as a G-quadruplex-selective probe for sensitive amperometric sensing of lead. Chem. Commun. 2011, 47, 11909–11911. [Google Scholar] [CrossRef]
- Song, J.; Wu, F.-Y.; Wan, Y.-Q.; Ma, L.-H. Ultrasensitive turn-on fluorescent detection of trace thiocyanate based on fluorescence resonance energy transfer. Talanta 2015, 132, 619–624. [Google Scholar] [CrossRef] [PubMed]
- Chen, Y.; Wang, J.; Zhang, Y.; Xu, L.; Gao, T.; Wang, B.; Pei, R. Selection and characterization of a DNA aptamer to crystal violet. Photochem. Photobiol. Sci. 2018, 17, 800–806. [Google Scholar] [CrossRef] [PubMed]
- He, H.Z.; Ma, V.P.; Leung, K.H.; Chan, D.S.; Yang, H.; Cheng, Z.; Leung, C.H.; Ma, D.L. A label-free G-quadruplex-based switch-on fluorescence assay for the selective detection of ATP. Analyst 2012, 137, 1538–1540. [Google Scholar] [CrossRef]
- De-Ming, K.; Ma, Y.-E.; Guo, J.-H.; Yang, W.; Shen, H.-X. Fluorescent Sensor for Monitoring Structural Changes of G-Quadruplexes and Detection of Potassium Ion. Anal. Chem. 2009, 81, 7. [Google Scholar] [CrossRef]
- Jin, Y.; Bai, J.; Li, H. Label-free protein recognition using aptamer-based fluorescence assay. Analyst 2010, 135, 1731–1735. [Google Scholar] [CrossRef] [PubMed]
- Kong, D.M.; Guo, J.H.; Yang, W.; Ma, Y.E.; Shen, H.X. Crystal violet-G-quadruplex complexes as fluorescent sensors for homogeneous detection of potassium ion. Biosens. Bioelectron. 2009, 25, 88–93. [Google Scholar] [CrossRef] [PubMed]
- Zheng, B.; Cheng, S.; Dong, H.; Liang, H.; Liu, J.; Lam, M.H.-W. Label Free Determination of Potassium Ions Using Crystal Violet and Thrombin-Binding Aptamer. Anal. Lett. 2014, 47, 1726–1736. [Google Scholar] [CrossRef]
- Ivancic, V.A.; Ekanayake, O.; Lazo, N.D. Binding Modes of Thioflavin T on the Surface of Amyloid Fibrils Studied by NMR. ChemPhysChem 2016, 17, 2461–2464. [Google Scholar] [CrossRef] [PubMed]
- Zhang, X.Y.; Luo, H.Q.; Li, N.B. Crystal violet as an i-motif structure probe for reversible and label-free pH-driven electrochemical switch. Anal. Biochem. 2014, 455, 55–59. [Google Scholar] [CrossRef] [PubMed]
- Introduction: Quadruplexes and their Biology. In Therapeutic Applications of Quadruplex Nucleic Acids; Neidle, S., Ed.; Science Elsevier: London, UK, 2012; pp. 1–20. [Google Scholar]
- Gunaratnam, M.; Greciano, O.; Martins, C.; Reszka, A.P.; Schultes, C.M.; Morjani, H.; Riou, J.F.; Neidle, S. Mechanism of acridine-based telomerase inhibition and telomere shortening. Biochem. Pharmacol. 2007, 74, 679–689. [Google Scholar] [CrossRef] [PubMed]
- Perrone, R.; Nadai, M.; Frasson, I.; Poe, J.A.; Butovskaya, E.; Smithgall, T.E.; Palumbo, M.; Palu, G.; Richter, S.N. A dynamic G-quadruplex region regulates the HIV-1 long terminal repeat promoter. J. Med. Chem. 2013, 56, 6521–6530. [Google Scholar] [CrossRef] [PubMed]
- Le, D.D.; di Antonio, M.; Chan, L.K.M.; Balasubramanian, S. G-quadruplex ligands exhibit differential G-tetrad selectivity. Chem. Commun. 2015, 51. [Google Scholar] [CrossRef] [PubMed]
- Koirala, D.; Dhakal, S.; Ashbridge, B.; Sannohe, Y.; Rodriguez, R.; Sugiyama, H.; Balasubramanian, S.; Mao, H. A single-molecule platform for investigation of interactions between G-quadruplexes and small-molecule ligands. Nat. Chem. 2011, 3, 782–787. [Google Scholar] [CrossRef] [PubMed]
- Muller, S.; Kumari, S.; Rodriguez, R.; Balasubramanian, S. Small-molecule-mediated G-quadruplex isolation from human cells. Nat. Chem. 2010, 2, 1095–1098. [Google Scholar] [CrossRef] [PubMed]
- Muller, S.; Sanders, D.A.; Di Antonio, M.; Matsis, S.; Riou, J.F.; Rodriguez, R.; Balasubramanian, S. Pyridostatin analogues promote telomere dysfunction and long-term growth inhibition in human cancer cells. Org. Biomol. Chem. 2012, 10, 6537–6546. [Google Scholar] [CrossRef] [PubMed]
- Rodriguez, R.; Miller, K.M.; Forment, J.V.; Bradshaw, C.R.; Nikan, M.; Britton, S.; Oelschlaegel, T.; Xhemalce, B.; Balasubramanian, S.; Jackson, S.P. Small-molecule-induced DNA damage identifies alternative DNA structures in human genes. Nat. Chem. Biol. 2012, 8, 301–310. [Google Scholar] [CrossRef]
- Mahmood, T.; Paul, A.; Ladame, S. Synthesis and spectroscopic and DNA-binding properties of fluorogenic acridine-containing cyanine dyes. J. Org. Chem. 2010, 75, 204–207. [Google Scholar] [CrossRef]
- Percivalle, C.; Mahmood, T.; Ladame, S. Two-in-one: A pH-sensitive, acridine-based, fluorescent probe binds G-quadruplexes in oncogene promoters. Med. Chem. Commun. 2013, 4, 211–215. [Google Scholar] [CrossRef]
- Machireddy, B.; Kalra, G.; Jonnalagadda, S.; Ramanujachary, K.; Wu, C. Probing the Binding Pathway of BRACO19 to a Parallel-Stranded Human Telomeric G-Quadruplex Using Molecular Dynamics Binding Simulation with AMBER DNA OL15 and Ligand GAFF2 Force Fields. J. Chem. Inf. Model. 2017, 57, 2846–2864. [Google Scholar] [CrossRef] [PubMed]
- Jin, B.; Zhang, X.; Zheng, W.; Liu, X.; Qi, C.; Wang, F.; Shangguan, D. Fluorescence light-up probe for parallel G-quadruplexes. Anal. Chem. 2014, 86, 943–952. [Google Scholar] [CrossRef] [PubMed]
- Yang, C.; Hu, R.; Li, Q.; Li, S.; Xiang, J.; Guo, X.; Wang, S.; Zeng, Y.; Li, Y.; Yang, G. Visualization of Parallel G-Quadruplexes in Cells with a Series of New Developed Bis(4-aminobenzylidene)acetone Derivatives. ACS Omega 2018, 3, 10487–10492. [Google Scholar] [CrossRef]
- Chen, S.-B.; Wu, W.-B.; Hu, M.-H.; Ou, T.-M.; Gu, L.-Q.; Tan, J.-H.; Huang, Z.-S. Discovery of a New Fluorescent Light-Up Probe Specific to Parallel G-Quadruplexes. Chem. Commun. 2014, 50. [Google Scholar] [CrossRef] [PubMed]
- Hu, M.H.; Chen, S.B.; Wang, B.; Ou, T.M.; Gu, L.Q.; Tan, J.H.; Huang, Z.S. Specific targeting of telomeric multimeric G-quadruplexes by a new triaryl-substituted imidazole. Nucleic Acids Res. 2017, 45, 1606–1618. [Google Scholar] [CrossRef]
- Hu, M.H.; Zhou, J.; Luo, W.H.; Chen, S.B.; Huang, Z.S.; Wu, R.; Tan, J.H. Development of a Smart Fluorescent Sensor that Specifically Recognizes the c-MYC G-Quadruplex. Anal. Chem. 2019. [Google Scholar] [CrossRef] [PubMed]
- Kotar, A.; Wang, B.; Shivalingam, A.; Gonzalez-Garcia, J.; Vilar, R.; Plavec, J. NMR Structure of a Triangulenium-Based Long-Lived Fluorescence Probe Bound to a G-Quadruplex. Angew. Chem. 2016, 55, 12508–12511. [Google Scholar] [CrossRef]
- Yan, J.W.; Chen, S.B.; Liu, H.Y.; Ye, W.J.; Ou, T.M.; Tan, J.H.; Li, D.; Gu, L.Q.; Huang, Z.S. Development of a new colorimetric and red-emitting fluorescent dual probe for G-quadruplex nucleic acids. Chem. Commun. 2014, 50, 6927–6930. [Google Scholar] [CrossRef]
- Chen, S.B.; Hu, M.H.; Liu, G.C.; Wang, J.; Ou, T.M.; Gu, L.Q.; Huang, Z.S.; Tan, J.H. Visualization of NRAS RNA G-Quadruplex Structures in Cells with an Engineered Fluorogenic Hybridization Probe. J. Am. Chem. Soc. 2016, 138, 10382–10385. [Google Scholar] [CrossRef]
- Chen, X.C.; Chen, S.B.; Dai, J.; Yuan, J.H.; Ou, T.M.; Huang, Z.S.; Tan, J.H. Tracking the Dynamic Folding and Unfolding of RNA G-Quadruplexes in Live Cells. Angew. Chem. 2018. [Google Scholar] [CrossRef]
- Laguerre, A.; Stefan, L.; Larrouy, M.; Genest, D.; Novotna, J.; Pirrotta, M.; Monchaud, D. A twice-as-smart synthetic G-quartet: PyroTASQ is both a smart quadruplex ligand and a smart fluorescent probe. J. Am. Chem. Soc. 2014, 136, 12406–12414. [Google Scholar] [CrossRef] [PubMed]
- Laguerre, A.; Wong, J.M.; Monchaud, D. Direct visualization of both DNA and RNA quadruplexes in human cells via an uncommon spectroscopic method. Sci. Rep. 2016, 6, 32141. [Google Scholar] [CrossRef]
- Flinders, J.; DeFina, S.C.; Brackett, D.M.; Baugh, C.; Wilson, C.; Dieckmann, T. Recognition of planar and nonplanar ligands in the malachite green-RNA aptamer complex. ChemBioChem Eur. J. Chem. Biol. 2004, 5, 62–72. [Google Scholar] [CrossRef] [PubMed]
- Dai, R.; Wang, X.; Wang, Z.; Mu, S.; Liao, J.; Wen, Y.; Lv, J.; Huang, K.; Xiong, X. A sensitive and label-free sensor for melamine and iodide by target-regulating the formation of G-quadruplex. Microchem. J. 2019, 146, 592–599. [Google Scholar] [CrossRef]
- Hsu, J.C.; Chen, E.H.; Snoeberger, R.C., 3rd.; Luh, F.Y.; Lim, T.S.; Hsu, C.P.; Chen, R.P. Thioflavin T and its photoirradiative derivatives: Exploring their spectroscopic properties in the absence and presence of amyloid fibrils. J. Phys. Chem. B 2013, 117, 3459–3468. [Google Scholar] [CrossRef]
- Iliuk, A.B.; Hu, L.; Tao, W.A. Aptamer in bioanalytical applications. Anal. Chem. 2011, 83, 4440–4452. [Google Scholar] [CrossRef]
- Tuerk, C.; Gold, L. Systematic Evolution of Ligands by Exponential Enrichment: RNA Ligands to Bacteriophage T4 DNA Polymerase. Science 1990, 249, 505–510. [Google Scholar] [CrossRef]
- Song, S.; Wang, L.; Li, J.; Fan, C.; Zhao, J. Aptamer-based biosensors. Trends Anal. Chem. 2008, 27, 108–117. [Google Scholar] [CrossRef]
- Li, F.; Zhang, H.; Wang, Z.; Newbigging, A.M.; Reid, M.S.; Li, X.-F.; Le, X.C. Aptamers Facilitating Amplified Detection of Biomolecules. Anal. Chem. 2014, 87, 274–292. [Google Scholar] [CrossRef]
- Gatto, B.; Palumbo, M.; Sissi, C. Nucleic Acid Aptamers Based on the G-Quadruplex Structure: Therapeutic and Diagnostic Potential. Curr. Med. Chem. 2009, 16, 1248–1265. [Google Scholar] [CrossRef] [PubMed]
- Phan, A.T. Human telomeric G-quadruplex: Structures of DNA and RNA sequences. FEBS J. 2010, 277, 1107–1117. [Google Scholar] [CrossRef] [PubMed]
- Bock, L.C.; Griffin, L.C.; Latham, J.A.; Vermaas, E.H.; Toole, J.J. Selection of single-stranded DNA molecules that bind and inhibit human thrombin. Nature 1992, 355, 564–566. [Google Scholar] [CrossRef]
- Macaya, R.F.; Schultiz, P.; Smith, F.W.; Roe, J.A.; Feigon, J. Thrombin-binding DNA aptamer forms a unimolecular quadruplex structure in solution. Proc. Natl. Acad. Sci. USA 1993, 90, 3745–3749. [Google Scholar] [CrossRef] [PubMed]
- Jasinski, D.; Haque, F.; Binzel, D.W.; Guo, P. Advancement of the Emerging Field of RNA Nanotechnology. ACS Nano 2017, 11, 1142–1164. [Google Scholar] [CrossRef] [PubMed]
- Xia, Y.; Zhang, R.; Wang, Z.; Tian, J.; Chen, X. Recent advances in high-performance fluorescent and bioluminescent RNA imaging probes. Chem. Soc. Rev. 2017, 46, 2824–2843. [Google Scholar] [CrossRef]
- Paige, J.S.; Nguyen-Duc, T.; Song, W.; Jaffrey, S.R. Fluorescence imaging of cellular metabolites with RNA. Science 2012, 335, 1194. [Google Scholar] [CrossRef]
- Shieh, Y.A.; Yang, S.J.; Wei, M.F.; Shieh, M.J. Aptamer-based tumor-targeted drug delivery for photodynamic therapy. ACS Nano 2010, 4, 1433–1442. [Google Scholar] [CrossRef] [PubMed]
- Meng, H.; Liu, H.; Kuai, H.; Peng, R.; Mo, L.; Zhang, X. Aptamer-integrated DNA nanostructures for biosensing, bioimaging and cancer therapy. Chem. Soc. Rev. 2016, 45, 2583–2602. [Google Scholar] [CrossRef] [PubMed]
- Dhiman, A.; Kalra, P.; Bansal, V.; Bruno, J.G.; Sharma, T.K. Aptamer-based point-of-care diagnostic platforms. Sens. Actuators B Chem. 2017, 246, 535–553. [Google Scholar] [CrossRef]
- Chang, H.; Tang, L.; Wang, Y.; Jiang, J.; Li, J. Graphene fluorescence resonance energy transfer aptasensor for the thrombin detection. Anal. Chem. 2010, 82, 2341–2346. [Google Scholar] [CrossRef]
- Ge, J.; Ou, E.-C.; Yu, R.-Q.; Chu, X. A novel aptameric nanobiosensor based on the self-assembled DNA-MoS2 nanosheet architecture for biomolecule detection. J. Mater. Chem. B 2014, 2, 625–628. [Google Scholar] [CrossRef]
- Liu, X.; Freeman, R.; Willner, I. Amplified fluorescence aptamer-based sensors using exonuclease III for the regeneration of the analyte. Chemistry 2012, 18, 2207–2211. [Google Scholar] [CrossRef] [PubMed]
- Yan, S.; Huang, R.; Zhou, Y.; Zhang, M.; Deng, M.; Wang, X.; Weng, X.; Zhou, X. Aptamer-based turn-on fluorescent four-branched quaternary ammonium pyrazine probe for selective thrombin detection. Chem. Commun. 2011, 47, 1273–1275. [Google Scholar] [CrossRef] [PubMed]
- Ji, D.; Wang, H.; Ge, J.; Zhang, L.; Li, J.; Bai, D.; Chen, J.; Li, Z. Label-free and rapid detection of ATP based on structure switching of aptamers. Anal. Biochem. 2017, 526, 22–28. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z.; Sharon, E.; Freeman, R.; Liu, X.; Willner, I. Fluorescence detection of DNA, adenosine-5’-triphosphate (ATP), and telomerase activity by zinc(II)-protoporphyrin IX/G-quadruplex labels. Anal. Chem. 2012, 84, 4789–4797. [Google Scholar] [CrossRef] [PubMed]
- Wei, Y.; Chen, Y.; Li, H.; Shuang, S.; Dong, C.; Wang, G. An exonuclease I-based label-free fluorometric aptasensor for adenosine triphosphate (ATP) detection with a wide concentration range. Biosens. Bioelectron. 2015, 63, 311–316. [Google Scholar] [CrossRef]
- Xu, L.; Shen, X.; Hong, S.; Wang, J.; Zhang, Y.; Wang, H.; Zhang, J.; Pei, R. Turn-on and label-free fluorescence detection of lead ions based on target-induced G-quadruplex formation. Chem. Commun. 2015, 51, 8165–8168. [Google Scholar] [CrossRef] [PubMed]
- Cheng, H.; Qiu, X.; Zhao, X.; Meng, W.; Huo, D.; Wei, H. Functional Nucleic Acid Probe for Parallel Monitoring K(+) and Protoporphyrin IX in Living Organisms. Anal. Chem. 2016, 88, 2937–2943. [Google Scholar] [CrossRef]
- Yang, L.; Qing, Z.; Liu, C.; Tang, Q.; Li, J.; Yang, S.; Zheng, J.; Yang, R.; Tan, W. Direct Fluorescent Detection of Blood Potassium by Ion-selective Formation of Intermolecular G-quadruplex and Ligand Binding. Anal. Chem. 2016. [Google Scholar] [CrossRef]
- Lord, S.J.; Lee, H.L.; Moerner, W.E. Single-molecule spectroscopy and imaging of biomolecules in living cells. Anal. Chem. 2010, 82, 2192–2203. [Google Scholar] [CrossRef]
- Johnsson, N.; Johnsson, K. Chemical tools for biomolecular imaging. ACS Chem. Biol. 2007, 2, 31–38. [Google Scholar] [CrossRef] [PubMed]
- Paige, J.S.; Wu, K.Y.; Jaffrey, S.R. RNA mimics of green fluorescent protein. Science 2011, 333, 642–646. [Google Scholar] [CrossRef] [PubMed]
- Han, K.Y.; Leslie, B.J.; Fei, J.; Zhang, J.; Ha, T. Understanding the photophysics of the Spinach-DFHBI RNA aptamer-fluorogen complex to improve live-cell RNA imaging. J. Am. Chem. Soc. 2013, 135, 19033–19038. [Google Scholar] [CrossRef] [PubMed]
- Song, W.; Strack, R.L.; Jaffrey, S.R. Imaging bacterial protein expression using genetically encoded RNA sensors. Nat. Methods 2013, 10, 873–875. [Google Scholar] [CrossRef] [PubMed]
- Strack, R.L.; Disney, M.D.; Jaffrey, S.R. A superfolding Spinach2 reveals the dynamic nature of trinucleotide repeat-containing RNA. Nat. Methods 2013, 10, 1219–1224. [Google Scholar] [CrossRef] [PubMed]
- Okuda, M.; Fourmy, D.; Yoshizawa, S. Use of Baby Spinach and Broccoli for imaging of structured cellular RNAs. Nucleic Acids Res. 2017, 45, 1404–1415. [Google Scholar] [CrossRef] [PubMed]
- Warner, K.D.; Chen, M.C.; Song, W.; Strack, R.L.; Thorn, A.; Jaffrey, S.R.; Ferre-D’Amare, A.R. Structural basis for activity of highly efficient RNA mimics of green fluorescent protein. Nat. Struct. Mol. Biol. 2014, 21, 658–663. [Google Scholar] [CrossRef]
- Jepsen, M.D.E.; Sparvath, S.M.; Nielsen, T.B.; Langvad, A.H.; Grossi, G.; Gothelf, K.V.; Andersen, E.S. Development of a genetically encodable FRET system using fluorescent RNA aptamers. Nat. Commun. 2018, 9, 18. [Google Scholar] [CrossRef]
- Chou, S.H.; Chin, K.H.; Wang, A.H. DNA aptamers as potential anti-HIV agents. Trends Biochem. Sci. 2005, 30, 231–234. [Google Scholar] [CrossRef]
- de Soultrait, V.R.; Lozach, P.-Y.; Altmeyer, R.; Tarrago-Litvak, L.; Litvak, S.; Andréola, M.L. DNA Aptamers Derived from HIV-1 RNase H Inhibitors are Strong Anti-integrase Agents. J. Mol. Biol. 2002, 324, 195–203. [Google Scholar] [CrossRef]
- Jing, N.; Xiong, W.; Guan, Y.; Pallansch, L.; Wang, S. Potassium-dependent folding: A key to intracellular delivery of G-quartet oligonucleotides as HIV inhibitors. Biochemistry 2002, 41, 5397–5403. [Google Scholar] [CrossRef] [PubMed]
- Li, T.; Shi, L.; Wang, E.; Dong, S. Multifunctional G-quadruplex aptamers and their application to protein detection. Chemistry 2009, 15, 1036–1042. [Google Scholar] [CrossRef] [PubMed]
- Chen, W.H.; Yang Sung, S.; Fadeev, M.; Cecconello, A.; Nechushtai, R.; Willner, I. Targeted VEGF-triggered release of an anti-cancer drug from aptamer-functionalized metal–organic framework nanoparticles. Nanoscale 2018, 10, 4650–4657. [Google Scholar] [CrossRef] [PubMed]
- Huang, H.; Suslov, N.B.; Li, N.S.; Shelke, S.A.; Evans, M.E.; Koldobskaya, Y.; Rice, P.A.; Piccirilli, J.A. A G-quadruplex-containing RNA activates fluorescence in a GFP-like fluorophore. Nat. Chem. Biol. 2014, 10, 686–691. [Google Scholar] [CrossRef] [PubMed]
- Trachman, R.J., 3rd.; Demeshkina, N.A.; Lau, M.W.L.; Panchapakesan, S.S.S.; Jeng, S.C.Y.; Unrau, P.J.; Ferre-D’Amare, A.R. Structural basis for high-affinity fluorophore binding and activation by RNA Mango. Nat. Chem. Biol. 2017, 13, 807–813. [Google Scholar] [CrossRef] [PubMed]
- Singh, R.; Lillard, J.W., Jr. Nanoparticle-based targeted drug delivery. Exp. Mol. Pathol. 2009, 86, 215–223. [Google Scholar] [CrossRef]
- Weerasinghe, P.; Garcia, G.E.; Zhu, Q.; Yuan, P.; Feng, L.; Mao, L.; Jing, N. Inhibition of Stat3 activation and tumor growth suppression of non-small cell lung cancer by G-quartet oligonucleotides. Int. J. Oncol. 2007, 31, 129–136. [Google Scholar] [CrossRef]
- Paeschke, K.; Simonsson, T.; Postberg, J.; Rhodes, D.; Lipps, H.J. Telomere end-binding proteins control the formation of G-quadruplex DNA structures in vivo. Nat. Struct. Mol. Biol. 2005, 12, 847–854. [Google Scholar] [CrossRef]
- Stoddart, C.A.; Rabin, L.; Hincenbergs, M.; Moreno, M.E.; Steppes, V.L.; Leeds, J.M.; Truong, L.A.; Wyatt, J.R.; Ecker, D.J.; Mccune, J.M. Inhibition of human immunodeficiency virus type 1 infection in SCID-hu Thy/Liv mice by the G-Quartet-forming oligonucleotide, ISIS 5320. Antimicrob. Agents Chemother. 1998, 42, 2113–2115. [Google Scholar] [CrossRef]
- Pileur, F.; Andreola, M.-L.; Dausse, E.; Michel, J.; Moreau, S.; Yamada, H.; Gaidamakov, S.A.; Crouch, R.J.; Toulmé, J.-J.; Cazenave, C. Selective inhibitory DNA aptamers of the human RNase H1. Nucleic Acids Res. 2003, 31, 5776–5788. [Google Scholar] [CrossRef]
- Jones, L.A.; Clancy, L.E.; Rawlinson, W.D.; White, P.A. High-affinity aptamers to subtype 3a hepatitis C virus polymerase display genotypic specificity. Antimicrob. Agents Chemother. 2006, 50, 3019–3027. [Google Scholar] [CrossRef] [PubMed]
- Shum, K.T.; Tanner, J.A. Differential inhibitory activities and stabilisation of DNA aptamers against the SARS coronavirus helicase. ChemBioChem Eur. J. Chem. Biol. 2008, 9, 3037–3045. [Google Scholar] [CrossRef] [PubMed]
- Yoshida, W.; Mochizuki, E.; Takase, M.; Hasegawa, H.; Morita, Y.; Yamazaki, H.; Sode, K.; Ikebukuro, K. Selection of DNA aptamers against insulin and construction of an aptameric enzyme subunit for insulin sensing. Biosens. Bioelectron. 2009, 24, 1116–1120. [Google Scholar] [CrossRef] [PubMed]
- Okazawa, A.; Maeda, H.; Fukusaki, E.; Katakura, Y.; Kobayashi, A. In vitro selection of Hematoporphyrin binding DNA aptamers. Bioorg. Med. Chem. Lett. 2000, 10, 2653–2656. [Google Scholar] [CrossRef]
- Li, T.; Dong, S.; Wang, E. G-quadruplex aptamers with peroxidase-like DNAzyme functions: Which is the best and how does it work? Chem. Asian J. 2009, 4, 918–922. [Google Scholar] [CrossRef] [PubMed]
- Filonov, G.S.; Moon, J.D.; Svensen, N.; Jaffrey, S.R. Broccoli: Rapid selection of an RNA mimic of green fluorescent protein by fluorescence-based selection and directed evolution. J. Am. Chem. Soc. 2014, 136, 16299–16308. [Google Scholar] [CrossRef]
- Warner, K.D.; Sjekloca, L.; Song, W.; Filonov, G.S.; Jaffrey, S.R.; Ferre-D’Amare, A.R. A homodimer interface without base pairs in an RNA mimic of red fluorescent protein. Nat. Chem. Biol. 2017, 13, 1195–1201. [Google Scholar] [CrossRef]
- Trachman, R.J., 3rd.; Abdolahzadeh, A.; Andreoni, A.; Cojocaru, R.; Knutson, J.R.; Ryckelynck, M.; Unrau, P.J.; Ferre-D’Amare, A.R. Crystal Structures of the Mango-II RNA Aptamer Reveal Heterogeneous Fluorophore Binding and Guide Engineering of Variants with Improved Selectivity and Brightness. Biochemistry 2018, 57, 3544–3548. [Google Scholar] [CrossRef]
- Trachman, R.J., 3rd.; Autour, A.; Jeng, S.C.Y.; Abdolahzadeh, A.; Andreoni, A.; Cojocaru, R.; Garipov, R.; Dolgosheina, E.V.; Knutson, J.R.; Ryckelynck, M.; et al. Structure and functional reselection of the Mango-III fluorogenic RNA aptamer. Nat. Chem. Biol. 2019, 15, 472–479. [Google Scholar] [CrossRef]
- Levesque, D.; Beaudoin, J.D.; Roy, S.; Perreault, J.P. In vitro selection and characterization of RNA aptamers binding thyroxine hormone. Biochem. J. 2007, 403, 129–138. [Google Scholar] [CrossRef]
- Mori, T.; Oguro, A.; Ohtsu, T.; Nakamura, Y. RNA aptamers selected against the receptor activator of NF-kappaB acquire general affinity to proteins of the tumor necrosis factor receptor family. Nucleic Acids Res. 2004, 32, 6120–6128. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.; Zhang, Y.; Ye, G.; Yang, Z.; Zhang, L.; Zhang, L. In vitro selection of G-rich RNA aptamers that target HIV-1 integrase. Sci. China Series B Chem. 2008, 51, 401–413. [Google Scholar] [CrossRef]
- Weiss, S.; Proske, D.; Neumann, M.; Groschup, M.H.; Kretzschmar, H.A.; Famulok, M.; Winnacker, E.-L. RNA aptamers specifically interact with the prion protein PrP. J. Virol. 1997, 71, 8790–8797. [Google Scholar] [PubMed]
- Homann, M.; Lorger, M.; Engstler, M.; Zacharias, M.; Göringer, H.U. Serum-stable RNA aptamers to an invariant surface domain of live african trypanosomes. Comb. Chem. High Throughput Screen. 2006, 9, 491–499. [Google Scholar] [PubMed]
| Class | Ligand and Commercial Availability (CAS no.) | Structure and Fluorescence Properties | Representative Applications | Advantages and Limitations | Ref. |
|---|---|---|---|---|---|
| Porphyrin | N-methyl mesoporphyrin IX (NMM), Yes (142234-85-3) | 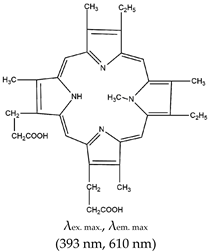 | — Highly specific parallel telomeric G4s binding, stabilization, and structural rearrangement. — Specific inhibitor of G4 unwinding by helicase (of Saccharomyces cerevisiae and human BML). In the presence of NMM the helicase gets trapped on the NMM-G4 complex. — Real time specific G4 based fluorescence assay for RNase sensing and inhibition. — A label free sensor for sensing iodide and melamine. Based on thymine-melamine-thymine (T-M-T)/thymine-Hg2+-thymine (T-Hg2+-T) and G4-NMM complex. — Highly sensitive microbial pathogen sensor based on quaternized magnetic nanoparticle exonuclease III assisted DNA amplification assays. Based on conformational transition from hairpin to G4 (assisted by Exo III nuclease) and subsequent specific interaction (of the G4) with NMM. | — Asymmetric anionic porphyrin — Inhibitors of different enzymes — Easy to develop label free sensors — Selective and sensitive G4 ligand — Allow real time study of G4 — Specific binding to parallel telomeric G4 — Microbial pathogen sensing — In live cell imaging, a Stokes shift and a red-shift emission were observed when applied, both of which were higher than the emissions seen with a different class of ligand, thioflavin T (ThT) — Not shown to provide visual discrimination of various G4s | [75,76,79,84,146,147] |
| 5,10,15,20-tetra-{4-[2-(1-methyl-1-piperidinyl)ethoxy]phenyl porphyrin (TMPipEOPP), No | 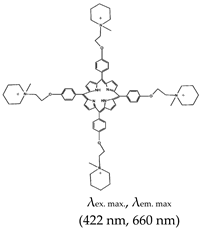 | — G4s specific probe that allow visual discrimination between G4s, duplexes, and single stranded DNAs. | — Cationic porphyrin — Aid visual differentiation of various G4s — Not shown to have enzymatic potentials | [86] | |
| Pyridinium, 4,4′,4′′,4′′′-(21H,23H-porphine-5,10,15,20-tetrayl)tetrakis[1-methyl-, 4-methylbenzenesulfonate (1:4) (TMPyP4), Yes (36951-72-1) | 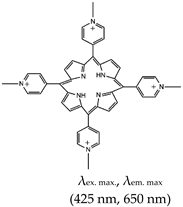 | — A ligand specific metabolic regulation of nitrate assimilation in Paracoccus denitrificans (a Gram-negative soil bacterium), based on stabilization of G4. | — Cationic porphyrin — Metabolic regulator nitrate assimilation — Not shown to provide visual discrimination of various G4s. — Not shown to have inhibitory potentials | [85] | |
| Benzothiozole | 3,6-dimethyl-2-(4-dimethylaminophenyl) benzothiazolium cation Thioflavin T (ThT), Yes (2390-54-7) | 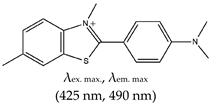 | — Sensitive and efficient G4 fluorescence sensor for human telomeric DNA. — Fluorescence sensor for the determination of cysteine and glutathione. Also, shown to detects biothiols in blood samples. Based on ThT ability to induced conformational specific G4. — Highly sensitive sensor for thrombin detection using Förster resonance energy transfer (FRET). Based on ThT ability to induce G4 and used as an acceptor with a conjugated polymer as the donor. — Specific fluorescence probe for G4 formation. Based on direct interaction between ThT and the target. — Aptasensor for Adenosine deaminase activity and inhibition. — Fluorescence probe for the detection of G4s in Chlamydomonas reinhardtii. — Fluorescence probe for the detection of G4s in zika virus. — Fluorescence probe for the detection of G4s in papillomaviruses. | — Has low background fluorescence intensity, which translates to a high signal-to-noise ratio — Highly sensitive probe for the detection of G4s in different species and in different samples — Induced conformational specific G4 — Shown to have enzymatic activity and inhibition — An ethyl substituted ThT allows naked-eye visualization of G4 in solution under ultraviolet light — Highly specific to parallel G4s — In live cells imaging, it produces less emissions compared to what was observed with a different class of ligand, NMM — In live cell imaging, its green fluorescence can easily coincide with the intrinsic fluorescence of the cell’s other components — ThT induced G4s can potentially cause topological changes — Produces false positive and false negative results — It binds tightly to non-G4 G–A-rich containing sequences and dimerizes them into a parallel double-stranded mode — Difficult to use for effective monitoring of G4s in the chromatin of live cells because of its inability to stain the nuclei | [72,74,97,99,104,105,106,110,148] |
| N-Isopropyl-2-(4-N,N-dimethylanilino)-6-methylbenzothiazole (IMT), No |  | — Real time fluorescence probe for monitoring the formation of G4 in live cells and its response to chemical treatment demonstrated. | — Live cell monitoring of G4 formation in real time — Selectively bind G4s in a cell’s chromatin (with negligible cytotoxicity) — Toxicity analysis only performed using single method instead of using two different methods in parallel | [51] | |
| ThT-NE, No | 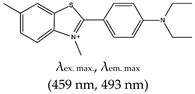 | — Cell permeable and highly specific G4 based fluorescence turn-on probe for real time imaging of native viral RNA genome in hepatitis C virus (HCV). This method was shown to allow subcellular monitoring and continuous live-cell monitoring of infected cells. | — Allows real time subcellular and continuous live-cell monitoring of native viral RNA genome — Toxicity effect to cells not shown/reported | [107] | |
| Triphenylmethane (TPM) | Crystal Violet (CV), Yes (548-62-9) |  | — Label free fluorescence aptasensor for specific detection of CV based on G4 interaction with CV. — A label-free G4 based fluorescence turn-on probe for the selective detection of ATP in aqueous medium. This is based on the ability of CV to specifically binds to G4. — Live cells visualization of G4 role in alternative splicing via RNA-binding protein hnRNPF. — G4-based fluorescence aptasensor for the selective detection of thrombin protein. Based on CV-G4 fluorescence. — Fluorescence probe for monitoring G4 structural differences (as a function of cation) and sensing of K+. — Fluorescence probe for homogenous detection of K+ based on the fluorescence intensity changes of CV-G4 complex. | — Distinguishes intramolecular from intermolecular G4s — Distinguishes single DNA strands from duplex DNAs — Widely employed in biosensing — Distinguishes between parallel and anti-parallel G4 topologies (preferentially binds to anti-parallel G4s) | [73,116,117,118,119,120] |
| Malachite Green (MG), Yes (569-64-2) | 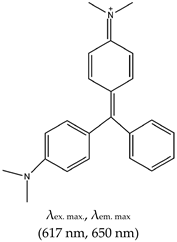 | — Fluorescence G4 based aptasensor for binding recognition to MG ligand. | — Widely employed in (bio)sending — Not shown to allow naked eye visualization of G4s in solution | [146] | |
| Triangulenium | Morpholino containing bis-substituted triangulenium (DAOTA-M2), No | 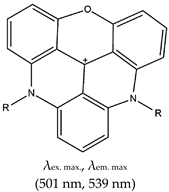 | — Fluorescence probe for G4 visualization in live cells, based fluorescence lifetime imaging microscopy. This probe was demonstrated to be cell permeable, have low toxicity, and be localized in the nucleus. | — Allows live cell visualization of G4 — High cell permeability — Low cytotoxicity — Can localize in the nucleus — One-to-one G4-specific sensor — Allows the visualization of interactions between ligands and G4s by fluorescence lifetime microscopy | [52] |
| Imidazole | Ethyl 2-(6-(4-(4-((4-(4,5-bis(4-(4-methylpiperazin-1-yl)phenyl)-1H-imidazol-2-yl)phenoxy)methyl)-1H-1,2,3-triazol-1-yl)butoxy)-3-oxo-3H-xanthen-9-yl)benzoate (IZFL-2), No | 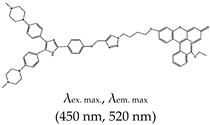 | — A tunable fluorescence activation probe for the specific detection of c-Myc G4. This was demonstrated to differentiate between wild-type c-Myc G4 and other G4s. | — Its fluorescence can be tuned — Distinctive smart sensor specific only for c-Myc G4s — Not yet demonstrated in live cells | [139] |
| 2,4,5-triaryl-substituted imidazole (IZCM-1), No | 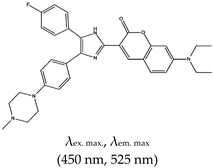 | — Fluorescence turn-on probe for the specific detection of parallel G4 without affecting their topology and thermal stability. | — Effectively and specifically binds to parallel G4s | [137] | |
| [2-(4-(4,5-bis(4-(4-methylpiperazin-1-yl)phenyl)-1H-imidazol-2-yl)phenyl)-6-(4-methylpiperazin-1-yl)-1H-benzo[de]isoquinoline-1,3(2H)-dione] (IZNP-1), No | 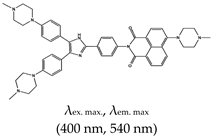 | — Fluorescence turn-on probe for the specific targeting of telomeric multimeric G4 structures, shown to occur via intercalation into the pocket between two G-quartet units. | — Can discriminate between telomeric multimeric G4s and monomeric G4s— Induce apoptosis and senescence in cancer cells | [138] | |
| Acridine | 3,6,9-trisubstitutedAcridine; cyanine dye 1, No | 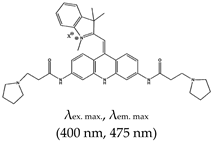 | — Water soluble dual function probe for G4 specific binding; pH sensitive and fluorescence probe for G4 stabilization and detection that operate by a push–pull mechanism. | — Highly water soluble — pH sensitive — Application not demonstrated in vivo | [133] |
| Alkaloid | (E)-3-((7-(diethylamino)-2-oxo-2H-chromen-3-yl)methylene)-6,7-difluoro-4-methyl-9-oxo-1,2,3,9-tetrahydropyrrolo[2,1-b]quinazolin-4-ium iodide (ISCH-1), No | 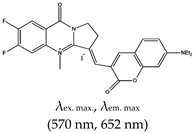 | — Multifunctional (colorimetric and red-emitting fluorescence) turn-on probe for specific G4 detection. This method is ideal for reliability and diverse applications. | — Multifunctional (colorimetric and fluorescence) — Reliable and potential for numerous applications — Not shown to allow specific targeting of G4s in a given region (such as the 5′-UTR) | [141] |
| (E)-3-((7-(diethylamino)-2-oxo-2H-chromen-3-yl)methylene)-7-fluoro-4-methyl-9-oxo-6-(prop-2-yn-1-yloxy)-1,2,3,9-tetrahydropyrrolo[2,1-b]quinazolin-4-ium (ISCH-oa1), No |  | — G-quadruplex-triggered fluorogenic hybridization (GTFH) probe, that selectively allows the visualization of the G-quadruplexes that form in a particular region interest (NRAS mRNA 5′-UTR region was demonstrated) both in vitro and in cells. The ligand consists of two segments, which are a fluorescent light-up fluorophore and oligonucleotide sequence that can hybridize with the sequence adjacent to the guanine rich sequence in the NRAS mRNA 5′-UTR or other regions of interest. | — Allows specific targeting of G4s in a particular region such as 5′-UTR — Can be use both in vivo and in vitro — Cannot detect the in situ spots of a given RNA in single cell — Requires RNAs to be transfected into cells to increase their concentration | [142] | |
| (E)-2-(2-(7-(diethylamino)-2-oxo-2H-chromen-3-yl)vinyl)-6-fluoro-1-methyl-7-(4-methylpiperazin-1-yl)quinolin-1-ium iodide (QUMA-1), No | 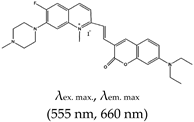 | — Highly selective fluorescence turn-on probe for real time and continuous tracking and monitoring of rG4 structural dynamics in live cells, this application has been demonstrated in through live cell imaging. Also, applied in visualization of rG4s unwinding by helicase. | — Allows live cell monitoring and tracking of rG4s — Allows the imaging of rG4 unwinding — Fluorescence intensity decreases in the presence of competing G4s ligands | [143] | |
| Acetone | Bis(4-aminobenzylidene)acetone derivative referred to as GD3, No | 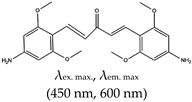 | — An effective red emitting fluorescence turn-on ligand for parallel G4s structures. Its biological application was demonstrated in fixed cells and shown to allow the visualization and monitoring of G4s structures. It was also shown to occur based on dipole moment created in the microenvironment of the ligand and restriction of the fluorophore resulting in altered charge transfer in the system, hence an enhanced light-up observed | — Red emitting ligand — Specific for parallel G4s — Allows monitoring of G4s in fixed cells | [136] |
| Pyrene | Pyrene template-assembled synthetic G-quartet (PyroTASQ), No |  | — Multitasking G4s smart probe (stabilizing ligand and fluorescence turn-on probe). This ligand and probe were demonstrated to recognize and bind to both DNA and RNA G4s, and shown to occur through an interesting approach; in which the ligand causes a ‘quadruplex-promoted conformational switch’ that leads to assembling of four guanines into a G-quartet, and subsequently the pyrene’s fluorescence is release | — Multifunctional (stabilization and fluorescence turn-on) — Can bind to both DNA and RNA G4s — Failed for in vivo studies as it forms aggregates in cells | [144] |
| Naphthalene | Naptho-template-assembled synthetic G-quartet (N-TASQ), No | 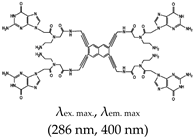 | — Multitasking G4s smart probe stabilizing ligand and fluorescence turn-on probe for live cell visualization of RNA G4s using multi-photon microscopy technique, while both RNA and DNA G4s were visualized using confocal microscopy. The interaction occurs through similar approach with Pyro-TASQ. | — Multifunctional (stabilization and fluorescence turn-on) — Can bind to both DNA and RNA G4s — G4 visualization in live cells using the multi-photon microscopy — Allows both RNA and DNA G4 imaging using confocal microscopy — No binding competition with other G4 ligands shown | [108,145] |
| Aptamer | Target | Sequence (5′-3′) | Length | Ref. |
|---|---|---|---|---|
| DNA G-quadruplex-containing aptamers | ||||
| TBA | Thrombin | d(GGTTGGTGTGGTTGG) | 15 | [156] |
| AS1411 | Nucleolin | d(GGTGGTGGTGGTTGTGGTGGTGGTGG) | 26 | [185] |
| T40214 | Stat3 a | d(GGGCGGGCGGGCGGGC) | 16 | [190] |
| T40231 | Stat3 | d(GGTGGGTGGGTGGG) | 14 | [190] |
| 22AG | Human TEBPs b | d(AGGGTTAGGGTTAGGGTTAGGG) | 22 | [95] |
| N.A. | Ciliate TEBPs | d(TTTTGGGGTTTTGGGG) | 16 | [191] |
| ISIS5320 | HIV-1 gp120 | d(TTGGGGTT) | 8 | [192] |
| 93del | HIV-1 integrase | d(GGGGTGGGAGGAGGGT) | 16 | [182] |
| 112del | HIV-1 integrase | d(CGGGTGGGTGGGTGGT) | 16 | [183] |
| T30695 | HIV-1 integrase | d(GGGTGGGTGGGTGGGT) | 16 | [182] |
| RT5 | HIV-1 reverse transcriptase | d(CAGGCGCCGGGGGGGTGGGAATACAGTGATCAGCG) | 35 | [41] |
| RT6 | HIV-1 reverse transcriptase | d(CAGGCGTTAGGGAAGGGCGTCGAAAGCAGGGTGGG) | 35 | [41] |
| RT47 | HIV-1 reverse transcriptase | d(CAGGCCTTGGGCGGGCCGGGACAATGGAGAGATTT) | 35 | [41] |
| ODN93 | HIV-1 reverse transcriptase | d(GGGGGTGGGAGGAGGGTAGGCCTTAGGTTTCTGA) | 34 | [193] |
| r10/43. | HCV RdRp c | d(GGGCGTGGTGGGTGGGGTACTAATAATGTGCGTTTG) | 36 | [194] |
| G5 | SARS Coronavirus Helicase | d(AGCGGGCATATGGTGGTGGGTGGTATGGTC) | 30 | [195] |
| N.A. | Insulin | d(GGTGGTGGGGGGGGTTGGTAGGGTGTCTTC) | 30 | [196] |
| N.A. | Hematoporphyrin IX | d(ATGGGGTCGGGCGGGCCGGGTGTC) | 24 | [197] |
| PS2M | Hemin | d(GTGGGTAGGGCGGGTTGG) | 18 | [198] |
| ABA | ATP | d(ACCTGGGGGAGTATTGCGGAGGAAGGT) | 27 | [167] |
| RNA G-quadruplex-containing aptamers | ||||
| Spinach | DFHBI d | r(GACGCAACUGAAUGAAAUGGUGAAGGACGGGUCCAGGUGUGGCUGCUUCGGCAGUGCAGCUUGUUGAGUAGAGUGUGAGCUCCGUAACUAGUCGCGUC) | 98 | [175] |
| Spinach mini | DFHBI | r(GACGCGACCGAAAUGGUGAAGGACGGGUCCAGUGCUUCGGCACUGUUGAGUAGAGUGUGAGCUCCGUAACUGGUCGCGUC) | 80 | [175] |
| Spinach1.2 | DFHBI | r(GACGCGACCGAAUGAAAUGGUGAAGGACGGGUCCAGCCGGCUGCUUCGGCAGCCGGCUUGUUGAGUAGAGUGUGAGCUCCGUAACUGGUCGCGUC) | 95 | [178] |
| Spinach2 | DFHBI | r(GAUGUAACUGAAUGAAAUGGUGAAGGACGGGUCCAGUAGGCUGCUUCGGCAGCCUACUUGUUGAGUAGAGUGUGAGCUCCGUAACUAGUUACAUC) | 95 | [178] |
| Spinach2 mini | DFHBI | r(GAUGUAACUGAAAUGGUGAAGGACGGGUCCAGUGCUUCGGCACUGUUGAGUAGAGUGUGAGCUCCGUAACUAGUUACAUC) | 80 | [179] |
| Baby Spinach | DFHBI | r(GGUGAAGGACGGGUCCAGUAGUUCGCUACUGUUGAGUAGAGUGUGAGCUCC) | 51 | [180] |
| Broccoli | DFHBI | r(GAGACGGUCGGGUCCAGAUAUUCGUAUCUGUCGAGUAGAGUGUGGGCUC) | 49 | [199] |
| Corn | DFHO e | r(CGAGGAAGGAGGUCUGAGGAGGUCACUG) | 28 | [200] |
| Mango | Thiazole orange-biotin | r(GGCACGUACGAAGGGACGGUGCGGAGAGGAGAGUACGUG) | 39 | [109] |
| Mango-II | Thiazole orange-biotin | r(GCGUACGAAGGAGAGGAGAGGAAGAGGAGAGUACGC) | 36 | [201] |
| Mango-III | Thiazole orange-biotin | r(GCUACGAAGGAAGGAUUGGUAUGUGGUAUAUUCGUAGC) | 38 | [202] |
| ApT4-A | Thyroxine hormone | r(GGUGGAGGGGGACGUGCUGCAUCCGCAGUGCGUCUUGGGUUGUG) | 44 | [203] |
| N.A. | Human receptor activator of NF-kB | r(ACGGAUUCGUAUGGGUGGGAUCGGGAAGGGCUACGAACACCGU) | 43 | [204] |
| N.A. | HIV-1 integrase | r(GGAGGGAGGGGAU) or r(GGAGUUAGGGGCU) | 13 | [205] |
| N.A. | Prion protein rPrP23-231 | r(CACUGCUACCUUAGAGUAGGAGCGGGACGAGGGGUUGUUGGGACGUGGGUAUGAUCCAUACAUUAGGAAGCUGGUGAGCUGGCACC) | 86 | [206] |
| N2 | Trypanosome | r(AAGAAGCGCGCGAGGCAGGACGAGGCAGGUGAGCGCUGUCCGA) | 43 | [207] |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Umar, M.I.; Ji, D.; Chan, C.-Y.; Kwok, C.K. G-Quadruplex-Based Fluorescent Turn-On Ligands and Aptamers: From Development to Applications. Molecules 2019, 24, 2416. https://doi.org/10.3390/molecules24132416
Umar MI, Ji D, Chan C-Y, Kwok CK. G-Quadruplex-Based Fluorescent Turn-On Ligands and Aptamers: From Development to Applications. Molecules. 2019; 24(13):2416. https://doi.org/10.3390/molecules24132416
Chicago/Turabian StyleUmar, Mubarak I., Danyang Ji, Chun-Yin Chan, and Chun Kit Kwok. 2019. "G-Quadruplex-Based Fluorescent Turn-On Ligands and Aptamers: From Development to Applications" Molecules 24, no. 13: 2416. https://doi.org/10.3390/molecules24132416
APA StyleUmar, M. I., Ji, D., Chan, C.-Y., & Kwok, C. K. (2019). G-Quadruplex-Based Fluorescent Turn-On Ligands and Aptamers: From Development to Applications. Molecules, 24(13), 2416. https://doi.org/10.3390/molecules24132416






