In Vitro Ecological Response of the Human Gut Microbiome to Bioactive Extracts from Edible Wild Mushrooms
Abstract
1. Introduction
2. Results
2.1. In Vitro Effects of a Microbiome Environment on the Fingerprints of Bioactive Compounds
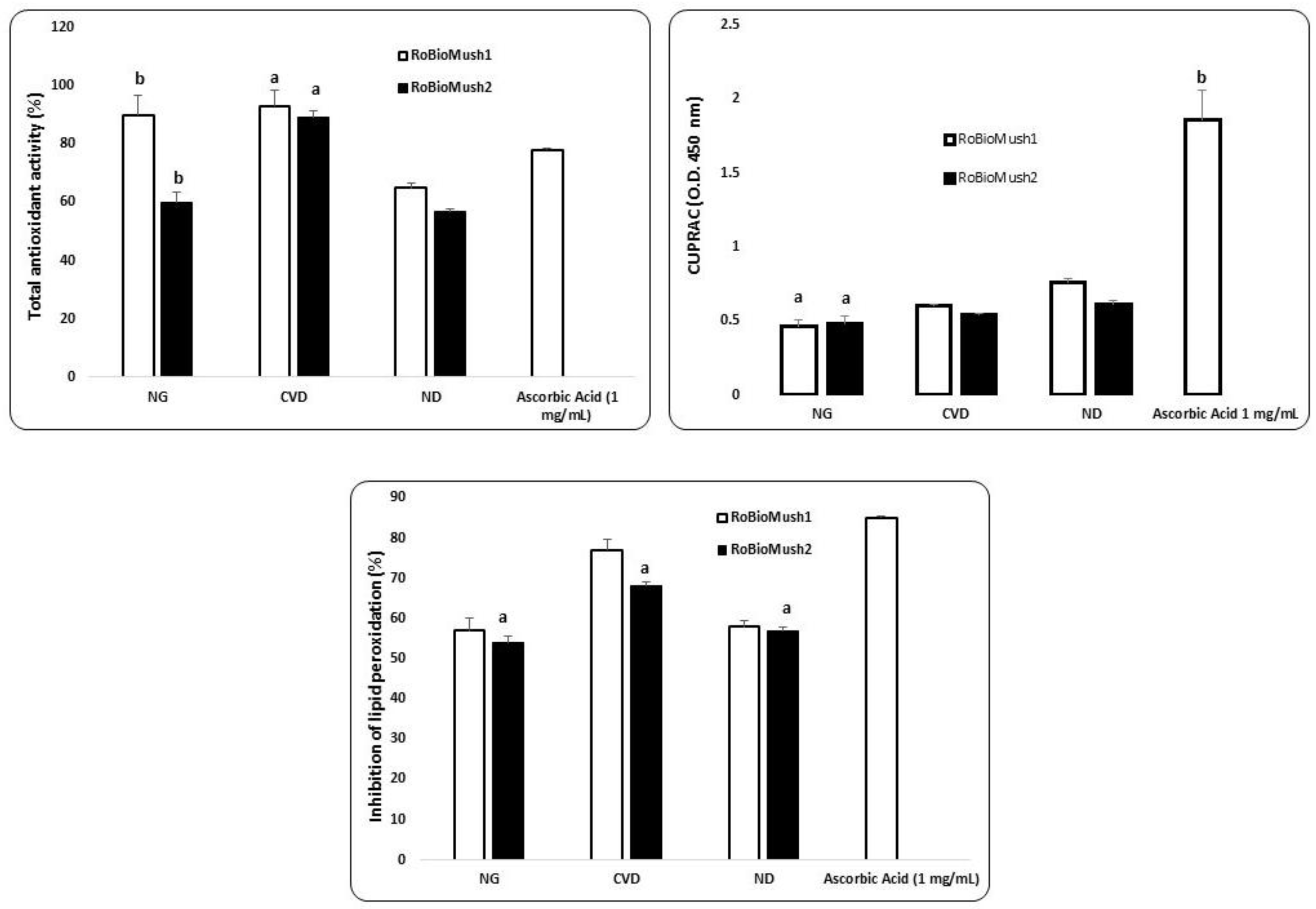
2.2. In Vitro Effects of the Microbiome Environment on Antioxidant Capacity
2.3. In Vitro Effects of the Microbiome Environment on Metabolic Activity
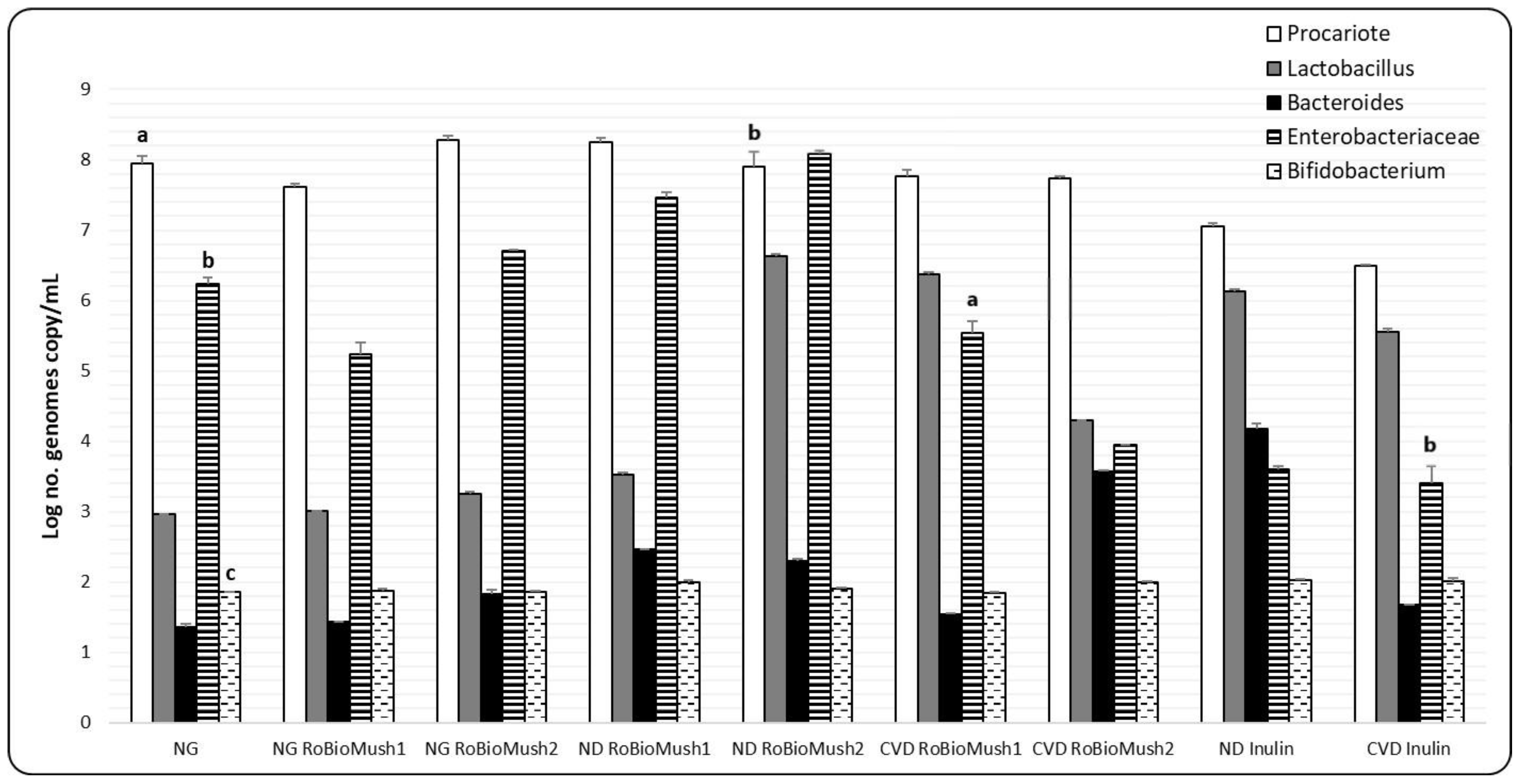
2.4. Quantifying the Number of Microbial Cells from the Microbiota
3. Discussion
4. Materials and Methods
4.1. Standards and Reagents
4.2. Obtaining Atomized Extracts
4.3. In Vitro Microbiome Modulation
4.4. Quantification of Short-Chain Organic Acids by Capillary Zone Electrophoresis (CZE)
4.5. Quantification of Polyphenols and Carbohydrates
4.5.1. Polyphenol Analysis by CZE
4.5.2. Carbohydrate Analysis by CZE
4.6. Determination of In Vitro Antioxidant Activity
4.7. Microbial Fingerprint Analysis
4.8. Statistical Analysis
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Shimizu, T.; Mori, K.; Ouchi, K.; Kushida, M.; Tsuduki, T. Effects of Dietary Intake of Japanese Mushrooms on Visceral Fat Accumulation and Gut Microbiota in Mice. Nutrients 2018, 10, 610. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.-J.; Li, Y.; Zhou, T.; Xu, D.-P.; Zhang, P.; Li, S.; Li, H.-B. Bioactivities and Health Benefits of Mushrooms Mainly from China. Molecules 2016, 21, 938. [Google Scholar] [CrossRef] [PubMed]
- Cordona, F.; Andres-Lacueva, C.; Tulipani, S.; Tinahones, F.J.; Queipo-Ortuno, M.I. Benefits of polyphenols on gut microbiota and implications in human health. J. Nutr. Biochem. 2013, 24, 1415–1422. [Google Scholar] [CrossRef] [PubMed]
- The Guardian. Available online: https://www.theguardian.com/news/2018/mar/26/the-human-microbiome-why-our-microbes-could-be-key-to-our-health (accessed on 12 August 2018).
- Valverde, M.E.; Hernández-Pérez, T.; Paredes-López, O. Edible mushrooms: Improving human health and promoting quality life. Int. J. Microbiol. 2015, 2015, 376–387. [Google Scholar] [CrossRef] [PubMed]
- Vaz, J.A.; Barros, L.; Martins, A.; Santos-Buelga, C.; Vasconcelos, M.H.; Ferreira, I.C.F.R. Chemical composition of wild edible mushrooms and antioxidant properties of their water soluble polysaccharidic and ethanolic fractions. Food Chem. 2011, 126, 610–616. [Google Scholar] [CrossRef]
- Vamanu, E.; Nita, S. Antioxidant capacity and the correlation with major phenolic compounds, anthocyanin, and tocopherol content in various extracts from the wild edible Boletus edulis mushroom. Biomed Res. Int. 2013, 2013. [Google Scholar] [CrossRef] [PubMed]
- Kozarski, M.; Klaus, A.; Jakovljevic, D.; Todorovic, N.; Vunduk, J.; Petrović, P.; Niksic, M.; Vrvic, M.M.; van Griensven, L. Antioxidants of Edible Mushrooms. Molecules 2015, 20, 19489–19525. [Google Scholar] [CrossRef] [PubMed]
- Vamanu, E.; Pelinescu, D. Effects of mushroom consumption on the microbiota of different target groups—Impact of polyphenolic composition and mitigation on the microbiome fingerprint. LWT Food Sci. Technol. 2017, 85, 262–268. [Google Scholar] [CrossRef]
- Correa, V.G.; Gonçalves, G.A.; de Sá-Nakanishi, A.B.; Ferreira, I.C.F.R.; Barros, L.; Dias, M.I.; Koehnlein, E.A.; de Souza, C.G.M.; Peralta, R.M.; Bracht, A. Effects of in vitro digestion and in vitro colonic fermentation on stability and functional properties of yerba mate (Ilex paraguariensis A. St. Hil.) beverages. Food Chem. 2017, 237, 453–460. [Google Scholar] [CrossRef] [PubMed]
- Preidis, G.A.; Versalovic, J. Targeting the human microbiome with antibiotics, probiotics, and prebiotics: Gastroenterology enters the metagenomics era. Gastroenterology 2009, 136, 2015–2031. [Google Scholar] [CrossRef] [PubMed]
- Neville, B.A.; d’Enfert, C.; Bougnoux, M.E. Candida albicans commensalism in the gastrointestinal tract. FEMS Yeast Res. 2015, 15. [Google Scholar] [CrossRef] [PubMed]
- Correa, R.C.G.; Haminiuk, C.W.I.; Barros, L.; Dias, M.I.; Calhelha, R.C.; Kato, C.G.; Correa, V.G.; Peralta, R.M.; Ferreira, I.C.F.R. Stability and biological activity of Merlot (Vitis vinifera) grape pomace phytochemicals after simulated in vitro gastrointestinal digestion and colonic fermentation. J. Funct. Foods 2017, 36, 410–417. [Google Scholar] [CrossRef]
- Gatea, F.; Teodor, E.D.; Paun, G.; Matei, A.O.; Radu, G.L. Capillary electrophoresis method validation for organic acids assessment in probiotics. Food Anal. Meth. 2015, 8, 1335–1340. [Google Scholar] [CrossRef]
- Vamanu, E. Bioactive capacity of some Romanian wild edible mushrooms consumed mainly by local communities. Nat. Prod. Res. 2018, 32, 440–444. [Google Scholar] [CrossRef] [PubMed]
- Cardoso, R.V.C.; Calhelha, R.C.; Fernandes, A.; Barros, L.; Oliveira, M.B.P.P.; Martins, A.; Ferreira, I.C.F.R. Development of nutraceutical formulations based on the mycelium of Pleurotus ostreatus and Agaricus bisporus. Food Funct. 2017, 21, 2155–2164. [Google Scholar] [CrossRef] [PubMed]
- Vamanu, E.; Nita, S. Bioactive compounds, antioxidant and anti-inflammatory activities of extracts from Cantharellus cibarius. Rev. Chim. 2014, 65, 372–379. [Google Scholar]
- Rocha, A.L.; Duque, F.; Monteiro, M.; Tallarico Adorno, M.A.; Sakamoto, I.K.; Sivieri, K. An exploratory study on the influence of orange juice on gut microbiota using a dynamic colonic model. Food Res. Int. 2016, 84, 160–169. [Google Scholar] [CrossRef]
- Rodríguez Carrio, J.; Salazar, N.; Margolles, A.; González, S.; Gueimonde, M.; de los Reyes-Gavilán, C.G.; Suárez, A. Free fatty acids profiles are related to gut microbiota signatures and short-chain fatty acids. Front. Immunol. 2017, 8, 823. [Google Scholar] [CrossRef] [PubMed]
- Gutiérrez-Díaz, I.; Fernández-Navarro, T.; Salazar, N.; Bartolomé, B.; Moreno-Arribas, M.V.; de Andres-Galiana, E.J.; Fernández-Martínez, J.L.; de los Reyes-Gavilán, C.G.; Gueimonde, M.; González, S. Adherence to a mediterranean diet influences the fecal metabolic profile of microbial-derived phenolics in a Spanish cohort of middle age and older people. J. Agric. Food Chem. 2017, 25, 586–595. [Google Scholar] [CrossRef] [PubMed]
- Clemente, J.C.; Ursell, L.K.; Parfrey, L.W.; Knight, R. The Impact of the gut microbiota on human health: An integrative view. Cell 2012, 148, 1258–1270. [Google Scholar] [CrossRef] [PubMed]
- Vamanu, E.; Sârbu, I.; Pop, O.; Pop Erdelyi, A.; Ene, M. Bioactive Product from Edible Wild Mushrooms for Improving Colon Microbial Fingerprint. Patent Application A 2017 00459, 07 July 2017. [Google Scholar]
- Erdelyi Pop, A.; Vamanu, A.; Pop, O.V.; Vamanu, E. Preliminary in vitro activity and correlation with chemical composition of Momordica charantia, Vaccinium myrtillus and Vaccinium vitis-idaea after enzymatic extraction process. Rev. Chim. 2015, 66, 1687–1691. [Google Scholar]
- Vamanu, E.; Sarbu, I.; Nedelcu, I.; Pelinescu, D. Study of PROEXO product influence on infant microbiota in an in vitro colonic fermentation system. Annal. Microbiol. 2015, 65, 1189–1193. [Google Scholar] [CrossRef]
- Vamanu, E.; Pelinescu, D.; Sarbu, I. Comparative fingerprinting of the human microbiota in diabetes and cardiovascular disease. J. Med. Food 2016, 19, 1188–1195. [Google Scholar] [CrossRef] [PubMed]
- Galli, V.; Barbas, C. Capillary electrophoresis for the analysis of short-chain organic acids in coffee. J. Chromatogr. 2004, A 1032, 299–304. [Google Scholar] [CrossRef]
- Gatea, F.; Teodor, E.D.; Matei, A.O.; Badea, G.I.; Radu, G.L. Capillary electrophoresis method for 20 polyphenols separation in propolis and plant extracts. Food Anal. Meth. 2015, 8, 1197–1206. [Google Scholar] [CrossRef]
- Chen, J.; Yang, F.; Guo, H.; Wu, F.; Wang, X. Optimized hydrolysis and analysis of Radix asparagi polysaccharide monosaccharide composition by capillary zone electrophoresis. J. Sep. Sci. 2015, 38, 2327–2331. [Google Scholar] [CrossRef] [PubMed]
- Gatea, F.; Teodor, E.D.; Seciu, A.M.; Radu, L.E. Monosaccharides composition and cytostatic activity of polysaccharide fraction of Phemeranthus confertiflorus L. Farmacia 2017, 65, 796–801. [Google Scholar]
- Pallag, A.M.; Jurca, T.; Pasca, B.; Sirbu, V.; Honiges, A.; Costuleanu, M. Analysis of phenolic compounds composition by HPLC and assessment of antioxidant capacity in Equisetum arvense L. extracts. Rev. Chim. 2016, 67, 1623–1627. [Google Scholar]
- Daker, M.; Abdullah, N.; Vikineswary, S.; Goh, P.C.; Kuppusamy, U.R. Antioxidant from maize and maize fermented by Marasmiellus sp. as stabiliser of lipid-rich foods. Food Chem. 2008, 107, 1092–1098. [Google Scholar] [CrossRef]
- Abdullah, N.; Ismail, S.M.; Aminudin, N.; Shuib, A.S.; Lau, B.F. Evaluation of selected culinary-medicinal mushrooms for antioxidant and ACE inhibitory activities. Evid. Based Complement. Altern. Med. 2012, 2012. [Google Scholar] [CrossRef] [PubMed]
- Ketabi, A.; Dieleman, L.A.; Ganzle, M.G. Influence of isomalto-oligosaccharides on intestinal microbiota in rats. J. Appl. Microbiol. 2011, 110, 1297–1306. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Samples of the RoBioMush 1 are available from the authors. |
| Product (Atomized Formula) | Glucides (g/g Atomized Samples) | Phenolic Acids (µg/g Atomized Samples) | Flavonoids (µg/g Atomized Samples) |
|---|---|---|---|
| RoBioMush1 | Xylose 0.033 ± 0.001 a Arabinose 0.041 ± 0.008 b Glucose 0.192 ± 0.018 c Rhamnose 0.115 ± 0.017 c Mannose 0.007 ± 0.000 Galactose 0.016 ± 0.001 a | Cinnamic acid 1.84 ± 0.03 a Chlorogenic acid 2.41 ± 0.02 a Sinapic acid 0.89 ± 0.00 Ferulic acid 0.2 ± 0.00 Coumaric acid 0.08 ± 0.00 | Rutin 11.7 ± 0.62 b Naringenin 0.08 ± 0.00 Luteolin 0.58 ± 0.00 Isoquercitrin 3.6 ± 0.01 a Umbelliferone 0.65 ± 0.03 a |
| RoBioMush2 | Xylose 0.04 ± 0.002 a Arabinose 0.033 ± 0.002 a Glucose 0.239 ± 0.008 b Rhamnose 0.138 ± 0.001 a Mannose 0.018 ± 0.000 Galactose 0.026 ± 0.001 a | Cinnamic acid 1.45 ± 0.03 a Chlorogenic acid 1.36 ± 0.04 a | Rutin 17.6 ± 0.45 a Naringenin 0.41 ± 0.00 Luteolin 0.55 ± 0.00 Isoquercitrin 2.11 ± 0.02 a Umbelliferone 2.04 ± 0.01 a |
| Bioactive Compounds (µg/mL) | NG * RoBioMush1 | NG RoBioMush2 | ND * RoBioMush1 | ND RoBioMush2 | CVD * RoBioMush1 | CVD RoBioMush2 |
|---|---|---|---|---|---|---|
| (A) Monosaccharides after in vitro simulation | ||||||
| Xylose | 250.70 ± 6.46 d | - | 174.99 ± 2.76 b | 172.38 ± 0.92 | 246.79 ± 4.62 b | - |
| Arabinose | - | - | 127.93 ± 4.73 b | 105.04 ± 3.94 a | - | - |
| Glucose | 34.76 ± 1.07 c | - | 133.12 ± 3.21 | 70.32 ± 2.14 b | - | 168.13 ± 3.16 |
| Rhamnose | - | - | 237.76 ± 2.40 a | - | - | - |
| Mannose | - | - | 224.33 ± 6.50 | 83.14 ± 0.93 | - | 13.53 ± 3.70 c |
| Galactose | 81.66 ± 1.94 c | 89 ± 1.05 c | 230.69 ± 3.89 c | 396.68 ± 5.19 c | 2.33 ± 0.00 | 126.14 ± 2.59 |
| Galacturonic acid | - | 233.83 ± 5.90 d | - | 75.74 ± 2.21 | - | 76.26 ± 4.42 b |
| Glucuronic acid | - | - | - | - | - | 122.06 ± 2.91 a |
| (B) The phenolic fraction after in vitro simulation | ||||||
| Rutin | - | - | 12.88 ± 0.49 a | - | - | - |
| Naringenin | - | 5.46 ± 0.11 a | - | - | - | - |
| Isoquercitrin | - | 83.44 ± 1.36 b | 2.29 ± 0.37 | - | - | - |
| Umbelliferone | - | - | 4.90 ± 0.61 b | 32.23 ± 1.06 a | - | - |
| Cinnamic acid | - | 3.0 ± 0.11 a | - | 5.3 ± 0.37 b | - | - |
| Chlorogenic acid | - | - | - | 2.99 ± 1.07 a | - | |
| Sinapic acid | - | - | - | - | - | 86.67 ± 1.64 a |
| Kaempferol | - | - | - | - | - | - |
| Luteolin | - | - | - | - | - | 1.11 ± 0.00 |
| Coumaric acid | - | - | - | 1.57 ± 0.20 | - | - |
| (C) Organic acids | ||||||
| Acetic acid | 0.48 ± 0.01 | 1.09 ± 0.00 | 1.56 ± 0.01 | 2.67 ± 0.11 a | 1.57 ± 0.01 | 0.55 ± 0.03 |
| Propionic acid | - | 0.17 ± 0.01 | 1,07 ± 0.00 | 0.73 ± 0.01 | 0.15 ± 0.00 | 0.07 ± 0.00 |
| Lactic acid | 2.78 ± 0.01 | 1.39 ± 0.12 a | 16.86 ± 2.11 b | 23.10 ± 2.69 a | 5.35 ± 0.02 | 7.14 ± 0.45 a |
| Butyric acid | 2.09 ± 0.18 a | 0.55 ± 0.03 | 2.22 ± 0.07 | 2.56 ± 0.08 | 1.46 ± 0.03 | 1.03 ± 0.10 b |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Vamanu, E.; Gatea, F.; Sârbu, I. In Vitro Ecological Response of the Human Gut Microbiome to Bioactive Extracts from Edible Wild Mushrooms. Molecules 2018, 23, 2128. https://doi.org/10.3390/molecules23092128
Vamanu E, Gatea F, Sârbu I. In Vitro Ecological Response of the Human Gut Microbiome to Bioactive Extracts from Edible Wild Mushrooms. Molecules. 2018; 23(9):2128. https://doi.org/10.3390/molecules23092128
Chicago/Turabian StyleVamanu, Emanuel, Florentina Gatea, and Ionela Sârbu. 2018. "In Vitro Ecological Response of the Human Gut Microbiome to Bioactive Extracts from Edible Wild Mushrooms" Molecules 23, no. 9: 2128. https://doi.org/10.3390/molecules23092128
APA StyleVamanu, E., Gatea, F., & Sârbu, I. (2018). In Vitro Ecological Response of the Human Gut Microbiome to Bioactive Extracts from Edible Wild Mushrooms. Molecules, 23(9), 2128. https://doi.org/10.3390/molecules23092128