Abstract
Exocarpium Citri grandis (ECG) is an important Traditional Chinese Medicine (TCM) for the treatment of cough and phlegm, and the flavonoids contained were considered the main effective components. To date, the systematic chemical profiling of these flavonoids and derived in vivo metabolites in human have not been well investigated. ECG was extracted using boiling water and then provided to volunteers for oral administration. Following the ingestion, urine samples were collected from volunteers over 48 h. The extract and urine samples were analyzed using ultra-fast liquid chromatography/quadrupole-time-of-flight tandem mass spectrometry (UFLC-Q-TOF-MS/MS) system to screen and identify flavonoids and derived in vivo metabolites. A total of 18 flavonoids were identified in the ECG extract, and 20 metabolites, mainly glucuronide and sulfate conjugates, were screened in urine samples collected post consumption. The overall excretion of naringenin metabolites corresponded to 5.45% of intake and occurred mainly within 4–12 h after the ingestion. Meanwhile, another 29 phenolic catabolites were detected in urine. Obtained data revealed that flavonoids were abundant in the ECG extract, and these components underwent extensive phase II metabolism in humans. These results provided valuable information for further study of the pharmacology and mechanism of action of ECG.
1. Introduction
Flavonoids, which are widely distributed in plants, have been proven to exert a wide range of biological and pharmacological activities [1,2,3]. The rapid screening and identification of flavonoids and derived in vivo metabolites have attracted significant attention and played a vital role in the pharmaceutical discovery process [4,5,6].
Exocarpium Citri grandis (ECG) (Huajuhong in Chinese), the epicarp of C. grandis ‘Tomentosa’, was officially listed in the first section of the 2015 edition of “Pharmacopoeia of the People’s Republic of China” and used as a primary ingredient in many famous Traditional Chinese Medicine (TCM) prescriptions [7,8]. ECG has been used as an effective antitussive and expectorant for many years [9,10]. Modern studies have demonstrated that ECG exhibited its antitussive effect via peripheral nerves [11,12,13]. In addition, contained flavonoids have been proven to be the main effective components in ECG [14,15,16]. Until now, the determination of flavonoids in ECG was mainly performed in HPLC, and only a few compounds were detected [17,18,19]. Existing results on in vivo metabolism of flavonoids in ECG are restricted in rats [20,21]. Furthermore, investigation on the systematic chemical profiling of these flavonoids and derived in vivo metabolites in humans is non-existent. Therefore, it is essential to determine the number of flavonoids in ECG and how these components are metabolized and then excreted.
The main objective of this work was to identify flavonoids in the ECG extract and derived metabolites in human urine after oral administration. With the ultra-fast liquid chromatography/quadrupole-time-of-flight tandem mass spectrometry (UFLC-Q-TOF-MS/MS) system, flavonoids in the ECG extract were systematically identified and quantified. After the consumption of 250 mL of the extract above, urine samples were collected from volunteers and then analyzed to identify metabolites. Furthermore, urinary metabolites were quantified to clarify the excretive profiles of flavonoids. The results obtained in this study would be valuable for further study of the pharmacology and mechanism of action of ECG.
2. Results and Discussion
2.1. Identification and Quantification of Flavonoids in ECG Extract
With the high resolution UFLC-Q-TOF-MS/MS system, a total of 18 flavonoids were identified in the ECG extract. These flavonoids were derived from eight aglycones, including naringenin, apigenin, diosmetin, eriodictyol, luteolin, hesperitin, isosakuranetin, and kaempferol. The detailed information, including compound description, formula, retention time (RT), and characteristic fragment ions were showed in Table 1 (Structures and product ion spectra were provided in Part I of Supplementary Materials). Among them, the main components (naringenin and hesperetin derivatives) were quantified. These derivatives were naringin (331.51 μmol/250 mL), naringin-4′-O-glucoside (2.89 μmol), naringenin (8.36 μmol), narirutin (1.40 μmol), narirutin-4′-O-glucoside (1.97 μmol), and hesperidin (0.08 μmol). In total, there are 346 μmol glycosides of naringenin and 0.08 μmol hesperidin in the 250 mL ECG extract.

Table 1.
Identification of flavonoids in the ECG extract by UFLC-Q-TOF-MS/MS.
Abundant naringin, which was considered the main effective component in ECG [14], was detected in the aqueous extract. These results reveal that a traditional boiling water extraction is a simple and feasible method to take advantage of ECG.
As shown in Table 1, three flavonoid aglycones (naringenin, kaempferol, and apigenin), and twelve flavonoid-O-glycosides (mainly flavonoid-7-O-glycosides, including naringin, hesperidin, neoeriocitrin, et al.) were identified in the ECG extract. Retro Diels-Alder (RDA) reactions were common in the MS fragmentation of flavonoids [22,23]. In the case of naringin, the signal at m/z 459 and 119 could be explained by cleavage of bonds 1 and 3 in the C ring. The fragment ion at m/z 313 and 151 were yielded by the neutral loss of a rhamnose moiety and subsequent neutral loss of a glucose moiety from the ion at m/z 459. The characteristic ion at m/z 271, which was proposed as deprotonated naringenin, was formed by the successive neutral loss of a rhamnose and a glucose moiety from naringin. An RDA reaction related the cleavage of bonds 1 and 3, and deprotonated naringenin also gave the fragment ion at m/z 151 and 119. Meanwhile, as a result of the cleavage of bond 5, the product ion at m/z 177 and 93 was generated from deprotonated naringenin. The product ion spectra and proposed fragmentations of deprotonated naringin were shown in Figure 1. Besides the twelve flavonoid-O-glycosides above, three flavonoid-C-glycosides (vicenin-2, lucenin-2,4′-methyl ether, and luteolin-6-C-glucoside) were also detected. A loss of 120 Da (C4H8O4), which was proposed as a fragment of glucose moiety, was detected in these three flavonoid-C-glycosides. However, the neutral loss of glycoside moiety was only identified in the fragmentation of luteolin-6-C-glucoside. Different from flavonoid-O-glycosides, RDA reactions were not observed in these flavonoid-C-glycosides. These results indicated that the glycosides bonding with carbon atoms exert a greater impact on structural characteristics of flavonoid-glycosides than that when bonding with oxygen atoms.
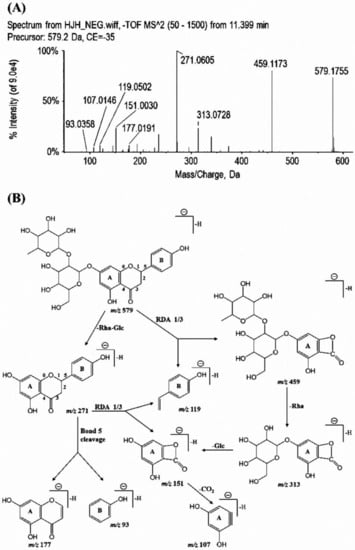
Figure 1.
Product ion spectra (A) and proposed fragmentation pattern (B) of deprotonated naringin.
2.2. Identification and Quantification of Metabolites in Urine
After acute intake of 250 mL ECG extract, a total of 20 metabolites were detected and identified in urine samples. Table 2 showed the compound description, elemental composition, elution time, and characteristic fragment ions of these metabolites (structures and product ion spectra were provided in Part III of Supplementary Materials). In addition, typical extracted ion chromatograms of identified metabolites were illustrated in Figure 2.

Table 2.
Identification of flavonoid metabolites in human urine after the consumption of 250 mL ECG extract by UFLC-Q-TOF-MS/MS.
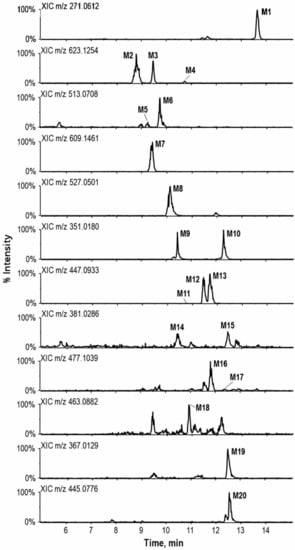
Figure 2.
Extracted ion chromatograms of metabolites in urine samples. For peak identification, see Table 2.
Three metabolites (M11, M12, and M13), which yielded deprotonated molecular ion of m/z 447 and characteristic fragments of m/z 271, were detected in urine (shown in Table 2). With authentic standards, M12 and M13 were unambiguously identified as naringenin-7-O-glucuronide and naringenin-4′-O-glucuronide, respectively. Due to its low acidity, the 5-OH was generally considered to be the least reactive among the three possible conjunct sites for naringenin (7-, 4′-, and 5-OH) [24]. Furthermore, combined with an early study [25], M11 was tentatively characterized as naringenin-5-O-glucuronide. After the loss of a moiety weighted 176 Da (glucuronyl moiety), M2, M3, and M4 all gave characteristic fragments at m/z 271, 175, 151, which were similar to that of naringenin-O-glucuronides. With reported results [26,27], M2, M3, and M4 were identified as naringenin-4′,7-O-diglucuronide, naringenin-5,7-O-diglucuronide, and naringenin-4′,5-O-diglucuronide, correspondingly. M9 and M10, which gave the deprotonated ion at m/z 351 (80 Da (SO3) more than that of naringenin) and exhibited a similar fragmentation pattern as naringenin, were supposed as naringenin-O-sulfates. Furthermore, given the least reactivity of 5-OH for naringenin [24] and the relative retention times of naringenin sulfates, M9 and M10 were further identified as naringenin-4′-O-sulfate and naringenin-7-O-sulfate, respectively. In addition to the above metabolites, naringenin-O-glucoside-O-sulfates, naringenin-O-glucoside-O-glucuronide, and naringenin-O-glucuronide-O-sulfate were also detected in some urine samples. However, due to the unavailability of reference standards for these metabolites, other methodologies will be needed to elucidate their final structure.
As for hesperetin, eriodictyol, and apigenin, probably due to little ingestion, several metabolites were identified. Co-chromatography with an authentic standard established that M14 was hesperetin-3′-O-sulfate. According to a study carried out by Brand and co-researchers [28], hesperetin was mainly conjugated at positions 7- and 3′-OH, and hesperetin-3′-O-sulfate was eluted earlier than hesperetin-7-O-sulfate. Thus, M15 could be tentatively identified as hesperetin-7-O-sulfate. Compared with reference standards, M16 and M17 were definitely identified as hesperetin-7-O-glucuronide and hesperetin-3′-O-glucuronide, respectively. Similar to the fragmentation pattern of naringenin glucuronides, the hesperetin glucuronides produced two major characteristic ions via the loss of the glucuronyl moiety (giving an ion of m/z 301) or the aglycon hesperetin (giving an ion of m/z 175), respectively. Following the loss of respective ligand (glucuronide or sulfate), M18 and M19 both gave characteristic fragments at m/z 151 and 135, which were proposed as the products of Retro Diels-Alder (RDA) reactions. With reported results [29], M18 and M19 were identified as eriodictyol-O-glucuronide and eriodictyol-O-sulfate, respectively. However, it is difficult to assign the specified conjunct sites of ligands for these two metabolites due to the four binding sites of eriodictyol (5-, 7-, 3′-, and 4′-OH). Based on an early report [30], M20 was tentatively identified as apigenin glucuronide. Nevertheless, given three hydroxyls on the apigenin skeleton, M20 could be tentatively identified as the 5-, 7-, or 4′-glucuronide.
Mediated by lactase-phlorizin hydrolase and intestinal microbiota, the hydrolysis of flavonoid-O-glycosides was generally considered the first and determinant step in the absorption of flavonoids [31]. Consistent with reported results [32], free flavonoid-O-glycosides (naringin, narirutin, hesperidin, neoeriocitrin, et al.) were not detected in urine samples. Following the hydrolysis of flavonoid-O-glycosides, corresponding aglycones were generated and subsequently engaged in glucuronidation and sulfation, giving rise to a series of conjunct metabolites. As shown in Table 3, flavonoid aglycone glucuronides and sulfates (mainly naringenin glucuronides and sulfates) were the major metabolites in urine after the intake of a 250 mL ECG extract, aligned with reported studies [33,34,35]. Based on identified metabolites shown in Table 2, it was rational to speculate that flavonoids derived from ECG underwent extensive phase II metabolism in the human body. Different with the metabolism profile in rats [20,21], naringin glucuronides, naringin sulfates, and other naringin derivatives were not detected in human urine, showing species variations.

Table 3.
Quantification of metabolites of naringenin and hesperetin in the urine after the ingestion of 250 mL ECG extract containing glycosides of naringenin (346 μmol), and hesperetin (0.08 μmol).
In a study conducted by Zhang and Brodbelt [24], naringenin glucuronides, naringenin sulfates, naringenin glucuronide sulfates, and a naringenin diglucuronide were screened as the metabolites of naringin and narirutin in urine after consumption of grapefruit juice. However, the specific conjunct sites of ligands for naringenin and the urinary excretion were undefined. As to the current work, with more authentic standards and updated findings, a total of 19 conjugative metabolites that were derived from naringenin, hesperetin, eriodictyol, and apigenin were identified in urine collected after the consumption of ECG extracts. Meanwhile, the urinary excretions of major metabolites were quantified along with the metabolite identification.
Naringenin and hesperetin metabolites, whose content was much higher than other metabolites, were quantified in urine of 0–48 h after the supplementation (shown in Table 3). There no metabolites detected in the urine collected prior to the ingestion. As shown in Table 3, naringenin-7-O-glucuronide, and naringenin-4′-O-glucuronide were the predominated metabolites of naringenin in urine after the intake of ECG extract, followed by naringenin-O-glucoside-O-sulfate (M6, RT = 9.9 min), free naringenin, naringenin-O-glucuronide-O-sulfate, and naringenin-O-glucoside-O-glucuronide. The overall excretion of naringenin metabolites corresponded to 5.45% of intake and occurred mainly within 4–12 h after the ingestion (Table 3). Several hesperetin metabolites, including hesperetin-O-glucuronides and hesperetin-O-sulfates, were excreted in little amounts, probably due to little ingestion of hesperidin. However, the overall recovery of hesperetin was higher than that of naringenin, which was equivalent to 58.7% of intake (Table 3).
Catalyzed by intestinal microflora, unabsorbed flavonoids were further metabolized into phenolic catabolites [36,37]. In this work, a total of 29 phenolic catabolites were screened and identified in urine samples collected after the oral administration of ECG extract. (Detailed information, structures, and product ion spectra were shown in part III of Supplementary Materials). However, most of these catabolites were also detected in urine samples collected before the oral administration. Generally, phenolic catabolites maintain a high concentration level in humans [38]. Abundant polyphenols in diets inevitably yield a number of catabolites in human intestines [39]. Meanwhile, several phenolic acids are generated in the metabolic processes of the human body [27]. For example, 3-(4′-hydroxy)-phenylpropionic acid is a major metabolite of tyrosine in humans [40]. Therefore, although a low-flavonoid diet was followed by the volunteers, phenolic catabolites were also detected in urine samples collected before the oral administration. Hence, without other methodologies, it is difficult to speculate about the exact amounts of phenolic catabolites derived from specific flavonoids in ECG extract.
3. Experimental
3.1. Chemicals and Reagents
The reference standards naringin, hesperidin, and apigenin were obtained from the National Institute for the Control of Pharmaceutical and Biological Products (Beijing, China). Naringenin, hesperetin, and MS grade formic acid were purchased from Sigma-Aldrich (St. Louis, MO, USA). Hesperetin-7-O-glucuronide, hesperetin-3′-O-glucuronide, and hesperetin-7-O-sulfate were acquired from Toronto Research Chemicals (Toronto, ON, Canada). Naringenin-7-O-glucuronide was purchased from Cayman Chemical Company (Ann Arbor, MI, USA). Naringenin-4′-O-glucuronide was obtained from Shanghai ZZBIO Co., Ltd. (Shanghai, China). Naringenin-7-O-glucoside was acquired from ChromaDex Inc. (Irvine, CA, USA). Rhoifolin was purchased from Shanghai Tauto Biotech Co., Ltd. (Shanghai, China). And kaempferol was obtained from Shanghai Yuanye Biological Co., Ltd. (Shanghai, China). The stable isotope labeled internal standard, naringin-d4, was supplied from Artis-Chem Co. Ltd. (Shanghai, China).
MS grade methanol was purchased from Fisher Scientific Inc. (Fair Lawn, NJ, USA). Acetonitrile of HPLC grade was obtained from Honeywell B&J Chemicals Inc. (Morristown, NJ, USA). Milli-Q grade water was purified by reverse-osmosis and filtered through a 0.22 μm membrane filter before use.
ECG was obtained from Huazhou Huajuhong Medicinal Materials Development Co., Ltd. (with GMP certificate). These samples were authenticated by Prof. Wenbo Liao from Sun Yat-sen University. In addition, the voucher specimens were deposited in our laboratory.
3.2. Preparation of ECG Extract
The ECG extract used in the feeding study was prepared with common processing methods in China. Briefly, ECG was cut into small pieces. In addition, 60-g fragmented samples were weighed and extracted with 1500 mL boiling water for 30 min. After filtering, the aqueous extract without residue was used in the study and a partial aqueous extract was stored at −80 °C for chemical profile analysis.
3.3. Study Design
Five healthy volunteers (three men and two women), aged 23–28 years and with BMI (Body Mass Index, in kg/m2) from 19.4 to 22.8 were recruited. These volunteers were nonsmokers, non-pregnant female, and not taking medication. All subjects were informed about the objectives, method, and risks of this study. All of them were asked to sign informed consent before their inclusion in the trial. Volunteers were required to follow a diet low in flavonoids, which excluded citrus fruit derived food, cruciferous vegetables, tomato, soybean, fresh ginger, chamomile, apple, grape, and beverages such as coffee, tea, coke, soda water, fruit juice, and wine, for 48 h before the ingestion.
On the day of supplementation, after an overnight fast, each volunteer drank 250 mL ECG extract. Volunteers were provided with a light breakfast (bread, eggs, and water) 2 h after the supplementation, and remained on a low-flavonoid diet for a further 48 h until the final urine samples were collected. Urine samples were collected once prior to drinking and over six time periods (0–4, 4–8, 8–12, 12–24, 24–36, 36–48 h) after consumption. Urine excreted in each time period was mixed, measured for its volume, and then stored at −80 °C before analysis.
3.4. Sample Preparation
All reference standards were accurately weighed, dissolved in methanol, stored at 4 °C and brought to room temperature before use. The stable isotope labeled internal standard naringin-d4 was dissolved in acetonitrile and prepared at 15 μg/mL used for protein precipitation. A sample preparation processes for ECG extract and urine were the same. An aliquot of 100 μL of the liquid sample above was transferred into a 1.5 mL polypropylene tube. After adding 200 μL volume acetonitrile for dissolving naringin-d4, the sample was vortex-mixed for 3 min and centrifuged at 15,000× g for 30 min at 25 °C. Finally, an aliquot of 10 μL supernatant was injected into the UFLC-Q-TOF-MS/MS for analysis.
3.5. UFLC-Q-TOF-MS/MS Analysis
Sample analysis was carried out on a Shimadzu UFLC XR instrument (Shimadzu Corp., Kyoto, Japan) equipped with an on-line degasser, a binary pump, and an autosampler. Chromatographic separation was performed on a Phenomenex Kinetex C18 column (3.0 × 150 mm, 2.6 μm, 100 Å; Phenomenex, Torrance, CA, USA) at 40 °C and eluted at a flow rate of 0.3 mL/min. The mobile phase was composed of 0.1% aqueous formic acid (v/v) (A) and methanol with 0.1% formic acid (v/v) (B). The following gradient elution program was used: linear gradient from 5% to 65% B (0–10 min), 65–100% B (10–30 min), and isocratic 100% B for 10 min. A 5-min post-run time was set to equilibrate the column.
Without splitting, the UFLC effluent was introduced directly to a hybrid triple quadrupole time-of-flight mass spectrometer (Triple TOFTM 5600 plus; AB Sciex, Foster City, CA, USA) equipped with an electrospray ionization source. The main instrumental conditions were as follows: ion source gas 1 and gas 2 were both 55 psi, curtain gas was 35 psi, ion source temperature was 550 °C, ion spray voltage floating was 5500 V in positive mode while 4500 V in negative mode, collision energy was 35 V, collision energy spread was 25 V, and declustering potential was 80 V. Nitrogen was used as nebulizer and auxiliary gas. Aqueous extract of ECG was analyzed in both positive and negative ionization modes, and the TOF-MS scan range was from m/z 100 to 1500. While urine samples were determined only in negative ionization mode for stronger signal response. Data acquisition was carried out using Analyst® TF 1.6 software (AB Sciex, Foster City, CA, USA) in IDA (information-dependent acquisition) mode. Identifications were based on chromatographic elution time, chemical composition, MS fragmentation pattern, and comparisons with available standards and references, as well as the mass spectral library (LibraryView, Version 1.0; AB Sciex, Foster City, CA, USA).
Naringenin and hesperetin derivatives, including partial compounds in ECG extract, as well as metabolites in urine, were quantified with chromatographic peak areas acquired in TOF-MS full scan and expressed relative to available authentic standards. Meanwhile, subjected to the unavailability of authentic standards, partial derivatives were prudently quantified as corresponding structural analogues, aligned with another study [32]. Naringin-4′-O-glucoside, narirutin, and narirutin-4′-O-glucoside were quantified as naringin. Naringenin-5-O-glucuronide, naringenin-O-diglucuronide, and naringenin-O-glucuronide-O-sulfate were quantified as naringenin-7-O-glucuronide. Naringenin-O-glucoside-O-glucuronide, and naringenin-O-glucoside-O-sulfate were quantified as naringenin-7-O-glucoside. Hesperetin-5-O-glucuronide was quantified as hesperetin-7-O-glucuronide. Naringenin-4′-O-sulfate, naringenin-7-O-sulfate, and hesperetin-3′-O-sulfate were quantified as hesperetin-7-O-sulfate.
Calibration curves that were used in the quantification of constituents in ECG extract were prepared from reference compounds dissolved in 50% methanol (v/v), while that used in the quantification of urinary metabolites was prepared from stock solutions by diluting with urine collected before the administration. All of these calibration curves covered the concentration range from 5 to 500 ng/mL. Calibration curves were constructed and fitted by linear regression analysis (R2 > 0.99) to plot the peak area ratio of analyte relative to the internal standard against the analyte concentrations. The intra-run precision, which ranges from 1.4% to 10.7%, was considered acceptable. Partial samples, whose concentrations exceeded the range of calibration curve, were diluted according to the actual situation.
4. Conclusions
In summary, using the UFLC-Q-TOF-MS/MS system, a total of 18 flavonoids was detected in the ECG extract, and naringenin derivatives as well as hesperidin contained within were quantified. Five volunteers were recruited and required to drink 250 mL ECG extract. Urine samples were collected after the supplementation and then analyzed with UFLC-Q-TOF-MS/MS. Finally, a total of 20 metabolites were identified and partly quantified in urine. Based on detected metabolites, flavonoids derived from ECG were considered to undergo extensive in vivo phase II metabolism (mainly hydrolysis, glucuronidation, and sulfation). Although 29 phenolic catabolites were detected in urine, it is hard to determine the exact amounts of phenolic catabolites derived from specific flavonoids due to multiple interferences. These results could be helpful for further study of the pharmacology and mechanism of action of ECG.
Supplementary Materials
The following are available online at http://www.mdpi.com/1420-3049/23/4/895/s1.
Acknowledgments
This work was supported by the National Major Scientific and Technical Special Project of China (No. 2015ZX09101014), and the Applied Science and Technology R&D Special Fund Project of Guangdong Province (No. 2015B020234004).
Author Contributions
X.Z. and H.Y. conceived and designed the study. X.Z., Y.Z., H.L., P.L., W.Z., Y.L. and Y.B. performed the experiments. X.Z. and W.P. analyzed the data, and W.S. played an important role in interpreting the results. X.Z. and H.Y. wrote the paper. X.Z., W.S. and H.Y. reviewed and edited the manuscript. All authors read and approved the manuscript.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Cushnie, T.P.; Lamb, A.J. Recent advances in understanding the antibacterial properties of flavonoids. Int. J. Antimicrob. Agents 2011, 38, 99–107. [Google Scholar] [CrossRef] [PubMed]
- Friedman, M. Overview of antibacterial, antitoxin, antiviral, and antifungal activities of tea flavonoids and teas. Mol. Nutr. Food Res. 2007, 51, 116–134. [Google Scholar] [CrossRef] [PubMed]
- Witkowska-Banaszczak, E. Flavonoids from Trollius europaeus flowers and evaluation of their biological activity. J. Pharm. Pharmacol. 2018, 70, 550–558. [Google Scholar] [CrossRef] [PubMed]
- Chalet, C.; Hollebrands, B.; Janssen, H.G.; Augustijns, P.; Duchateau, G. Identification of phase-II metabolites of flavonoids by liquid chromatography-ion-mobility spectrometry-mass spectrometry. Anal. Bioanal. Chem. 2017, 410, 471–482. [Google Scholar] [CrossRef] [PubMed]
- Du, L.Y.; Qian, D.W.; Shang, E.X.; Liu, P.; Jiang, S.; Guo, J.M.; Su, S.L.; Duan, J.A.; Xu, J.; Zhao, M. UPLC-Q-TOF/MS-based screening and identification of the main flavonoids and their metabolites in rat bile, urine and faeces after oral administration of Scutellaria baicalensis extract. J. Ethnopharmacol. 2015, 169, 156–162. [Google Scholar] [CrossRef] [PubMed]
- Gad, M.H.; Tuenter, E.; El-Sawi, N.; Younes, S.; El-Ghadban, E.M.; Demeyer, K.; Pieters, L.; Heyden, Y.V.; Mangelings, D. Identification of some Bioactive Metabolites in a Fractionated Methanol Extract from Ipomoea aquatica (Aerial Parts) through TLC, HPLC, UPLC-ESI-QTOF-MS and LC-SPE-NMR Fingerprints Analyses. Phytochem. Anal. 2018, 29, 5–15. [Google Scholar]
- Yuan, Y.; Long, P.; Jiang, C.; Li, M.; Huang, L. Development and characterization of simple sequence repeat (SSR) markers based on a full-length cDNA library of Scutellaria baicalensis. Genomics 2015, 105, 61–67. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Liu, M.; Hu, J.; Su, W. Systematic chemical profiling of Citrus grandis ‘Tomentosa’ by ultra-fast liquid chromatography/diode-array detector/quadrupole time-of-flight tandem mass spectrometry. J. Pharm. Biomed. Anal. 2014, 90, 167–179. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Zou, W.; Yang, C.; Peng, W.; Su, W. Metabolism and excretion studies of oral administered naringin, a putative antitussive, in rats and dogs. Biopharm. Drug Dispos. 2012, 33, 123–134. [Google Scholar] [CrossRef] [PubMed]
- Jiang, K.; Song, Q.; Wang, L.; Xie, T.; Wu, X.; Wang, P.; Yin, G.; Ye, W.; Wang, T. Antitussive, expectorant and anti-inflammatory activities of different extracts from Exocarpium Citri grandis. J. Ethnopharmacol. 2014, 156, 97–101. [Google Scholar] [CrossRef] [PubMed]
- Lin, B.; Li, P.; Wang, Y.; Peng, W.; Wu, Z.; Su, W.; Ji, H. The expectorant activity of naringenin. Pulm. Pharmacol. Ther. 2008, 21, 259–263. [Google Scholar] [CrossRef] [PubMed]
- Luo, Y.; Zhang, C.; Li, P.; Nie, Y.; Wu, H.; Shen, J.; Su, W. Naringin attenuates enhanced cough, airway hyperresponsiveness and airway inflammation in a guinea pig model of chronic bronchitis induced by cigarette smoke. Int. Immunopharmacol. 2012, 13, 301–307. [Google Scholar] [CrossRef] [PubMed]
- Gao, S.; Li, P.; Yang, H.; Fang, S.; Su, W. Antitussive effect of naringin on experimentally induced cough in Guinea pigs. Planta Med. 2011, 77, 16–21. [Google Scholar] [CrossRef] [PubMed]
- Li, P.; Wang, Y.; Wu, Z.; Peng, W.; Yang, C.; Nie, Y.; Liu, M.; Luo, Y.; Zou, W.; Liu, Y. The pre-clinical studies of naringin, an innovative drug, derived from Citri Grandis Exocarpium (Huajuhong). Acta Sci. Nat. Univ. Sunyatseni 2015, 54, 1–5. [Google Scholar]
- Liang, Y.; Huang, Z.; Chen, H.; Zhang, T.; Ito, Y. Preparative Isolation and Purification of Two Closely Related Glycosidic Flavonoids from Exocarpium Citri Grandis by High-Speed Countercurrent Chromatography. J. Liq. Chromatogr. Relat. Technol. 2007, 30, 419–430. [Google Scholar] [CrossRef]
- Wang, Y. Research Progress of Citri Grandis Exocarpium. Mod. Traditional Chin. Med. Mater. Med. World Sci. Technol. 2017, 19, 1076–1082. [Google Scholar]
- Liu, X.; Chen, Y.; Lin, L.; Chen, D.; Zhuang, M. Simultaneous Determination of Naringin and Rhoifolin in Exocarpium Citri Grandis by RP-HPLC. Traditional Chin. Drug Res. Clin. Pharmacol. 2010, 21, 640–642. [Google Scholar]
- Yu, X.; Liu, Q.; Xie, Z.; Lam, S.; Xu, X. Chromatographic Fingerprint Analysis of Exocarpium Citri Grandis by High-Performance Liquid Chromatography Coupled with Diode-Array Detector. Food Anal. Meth. 2015, 8, 1868–1875. [Google Scholar] [CrossRef]
- Yu, X.X.; Liu, Q.D.; Wu, J.W.; Liang, Z.K.; Zhao, M.Q.; Xu, X.J. Simultaneous determination of four major constituents in Citri Grandis Exocarpium by HPLC-DAD. Acta Chromatogr. 2015, 28, 1–15. [Google Scholar] [CrossRef]
- Sun, G.; Qian, D.; Duan, J.; Li, X.; Wan, J.; Gou, J. UPLC-Q-TOF/MS analysis of naringin and naringenin and its metabolites in rat plasma after intragastric administration of alcohol extract of Exocarpium Citri Grandis. China J. Chin. Mater. Med. 2010, 35, 1580–1586. [Google Scholar]
- Sun, G.; Qian, D.; Duan, J.; Li, X.; Wan, J. UPLC-Q-TOF/MS analysis of naringin and naringenin and its metabolites in rat urine and feces after intragastric administration of alcohol extract of Exocarpium Citri Grandis. Acta Pharm. Sin. 2010, 45, 761–766. [Google Scholar]
- Ma, Y.; Li, Q.; Van den Heuvel, H.; Claeys, M. Characterization of flavone and flavonol aglycones by collision-induced dissociation tandem mass spectrometry. Rapid Commun. Mass Spectrom. 1997, 11, 1357–1364. [Google Scholar] [CrossRef]
- Fabre, N.; Rustan, I.; De, H.E.; Quetin-Leclercq, J. Determination of flavone, flavonol, and flavanone aglycones by negative ion liquid chromatography electrospray ion trap mass spectrometry. J. Am. Soc. Mass Spectrom. 2001, 12, 707–715. [Google Scholar] [CrossRef]
- Zhang, J.M.; Brodbelt, J.S. Screening flavonoid metabolites of naringin and narirutin in urine after human consumption of grapefruit juice by LC-MS and LC-MS/MS. Analyst 2004, 129, 1227–1233. [Google Scholar] [CrossRef] [PubMed]
- Zeng, X.; Bai, Y.; Peng, W.; Su, W. Identification of naringin metabolites in human urine and feces. Eur. J. Drug Metab. Pharm. 2017, 42, 647–656. [Google Scholar] [CrossRef] [PubMed]
- Mullen, W.; Archeveque, M.A.; Edwards, C.A.; Matsumoto, H.; Crozier, A. Bioavailability and metabolism of orange juice flavanones in humans: Impact of a full-fat yogurt. J. Agric. Food Chem. 2008, 56, 11157–11164. [Google Scholar] [CrossRef] [PubMed]
- Pereiracaro, G.; Ludwig, I.A.; Polyviou, T.; Malkova, D.; García, A.; Morenorojas, J.M.; Crozier, A. Identification of plasma and urinary metabolites and catabolites derived from orange juice (poly)phenols: Analysis by high-performance liquid chromatography–high-resolution mass spectrometry. J. Agric. Food Chem. 2016, 64, 5724–5735. [Google Scholar] [CrossRef] [PubMed]
- Brand, W.; Boersma, M.G.; Bik, H.; Hoek-van den Hil, E.F.; Vervoort, J.; Barron, D.; Meinl, W.; Glatt, H.; Williamson, G.; van Bladeren, P.J. Phase II metabolism of hesperetin by individual UDP-glucuronosyltransferases and sulfotransferases and rat and human tissue samples. Drug Metab. Dispos. 2010, 38, 617–625. [Google Scholar] [CrossRef] [PubMed]
- Kim, U.; Han, S.B.; Kwon, O.S.; Yoo, H.H. Identfication of phase I and phase II metabolites of hesperetin in rat liver microsomes by liquid chromatography-electrospray ionization-tandem mass spectrometry. Mass Spectrom. Lett. 2011, 2, 53–56. [Google Scholar] [CrossRef]
- Qu, J.; Wang, Y.; Luo, G.; Wu, Z. Identification and determination of glucuronides and their aglycones in Erigeron breviscapus by liquid chromatography-tandem mass spectrometry. J. Chromatogr. A 2001, 928, 155–162. [Google Scholar] [CrossRef]
- Chen, Z.; Zheng, S.; Li, L.; Jiang, H. Metabolism of flavonoids in human: A comprehensive review. Curr. Drug Metab. 2014, 15, 48–61. [Google Scholar] [CrossRef] [PubMed]
- Pereira-Caro, G.; Oliver, C.M.; Weerakkody, R.; Singh, T.; Conlon, M.; Borges, G.; Sanguansri, L.; Lockett, T.; Roberts, S.A.; Crozier, A. Chronic administration of a microencapsulated probiotic enhances the bioavailability of orange juice flavanones in humans. Free Radic. Biol. Med. 2015, 84, 206–214. [Google Scholar] [CrossRef] [PubMed]
- Borges, G.; Mullen, W.; Mullan, A.; Lean, M.E.; Roberts, S.A.; Crozier, A. Bioavailability of multiple components following acute ingestion of a polyphenol-rich juice drink. Mol. Nutr. Food Res. 2010, 54 (Suppl. 2), S268–S277. [Google Scholar] [CrossRef] [PubMed]
- Mullen, W.; Borges, G.; Lean, M.E.J.; Roberts, S.A.; Crozier, A. Identification of metabolites in human plasma and urine after consumption of a polyphenol-rich juice drink. J. Agric. Food Chem. 2010, 58, 2586–2595. [Google Scholar] [CrossRef] [PubMed]
- Zeng, X.; Su, W.; Bai, Y.; Chen, T.; Yan, Z.; Wang, J.; Su, M.; Zheng, Y.; Peng, W.; Yao, H. Urinary metabolite profiling of flavonoids in Chinese volunteers after consumption of orange juice by UFLC-Q-TOF-MS/MS. J. Chromatogr. B 2017, 1061, 79–88. [Google Scholar] [CrossRef] [PubMed]
- Pereiracaro, G.; Borges, G.; van der Hooft, J.; Clifford, M.N.; Del, R.D.; Lean, M.E.; Roberts, S.A.; Kellerhals, M.B.; Crozier, A. Orange juice (poly)phenols are highly bioavailable in humans. Am. J. Clin. Nutr. 2014, 100, 1378–1384. [Google Scholar] [CrossRef] [PubMed]
- Zou, W.; Luo, Y.; Liu, M.; Chen, S.; Wang, S.; Nie, Y.; Cheng, G.; Su, W.; Zhang, K. Human intestinal microbial metabolism of naringin. Eur. J. Drug Metab. Pharmacokinet. 2015, 40, 363–367. [Google Scholar] [CrossRef] [PubMed]
- Jenner, A.M.; Rafter, J.; Halliwell, B. Human fecal water content of phenolics: The extent of colonic exposure to aromatic compounds. Free Radic. Biol. Med. 2005, 38, 763–772. [Google Scholar] [CrossRef] [PubMed]
- Saura-Calixto, F.; Serrano, J.; Goñi, I. Intake and bioaccessibility of total polyphenols in a whole diet. Food Chem. 2007, 101, 492–501. [Google Scholar] [CrossRef]
- Feng, L.; He, Y.-Q.; Xu, G.-H.; Hu, H.; Guo, L.; Wan, Y.-Q. Determination of Tyrosine and Its Metabolites in Human Serum with Application to Cancer Diagnosis. Anal. Lett. 2014, 47, 1275–1289. [Google Scholar] [CrossRef]
Sample Availability: Not Available. |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).