Spiroketones and a Biphenyl Analog from Stems and Leaves of Larrea nitida and Their Inhibitory Activity against IL-6 Production
Abstract
:1. Introduction
2. Results and Discussion
3. Materials and Methods
3.1. General Experimental Procedures
3.2. Plant Material
3.3. Extraction and Isolation
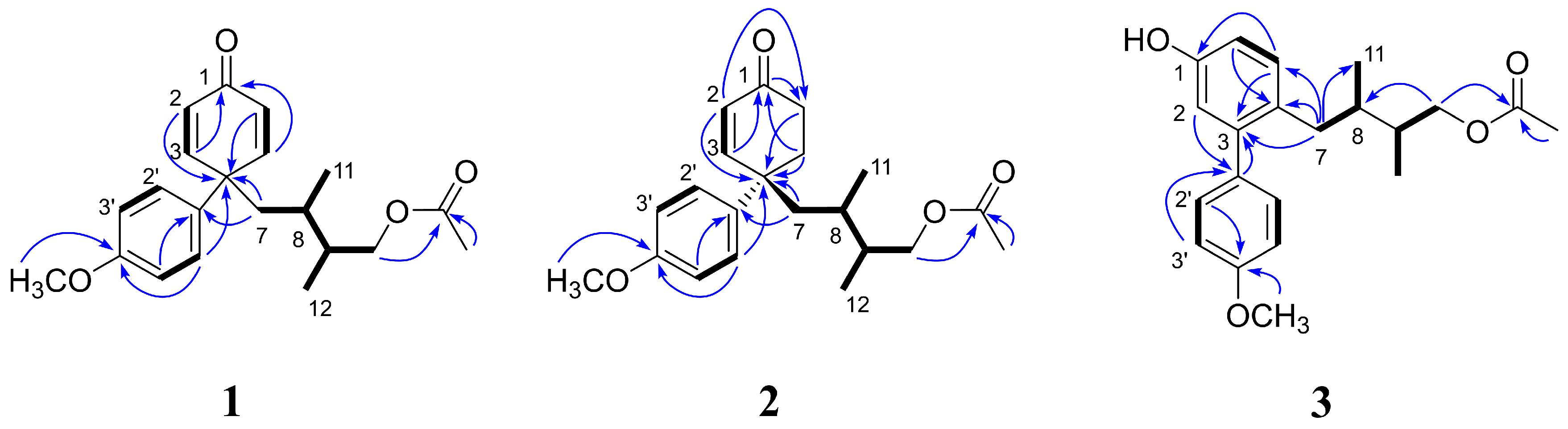
- Nitidaone A (1): white amorphous solid; −42.1° (c 0.5, MeOH); UV (MeOH) λmax (log ε) 256 (3.60), 272 (3.41) nm; 1H and 13C NMR data, see Table 1; HRESIMS m/z 343.1918 [M + H]+ (calcd. for C21H26O4, 343.1909).
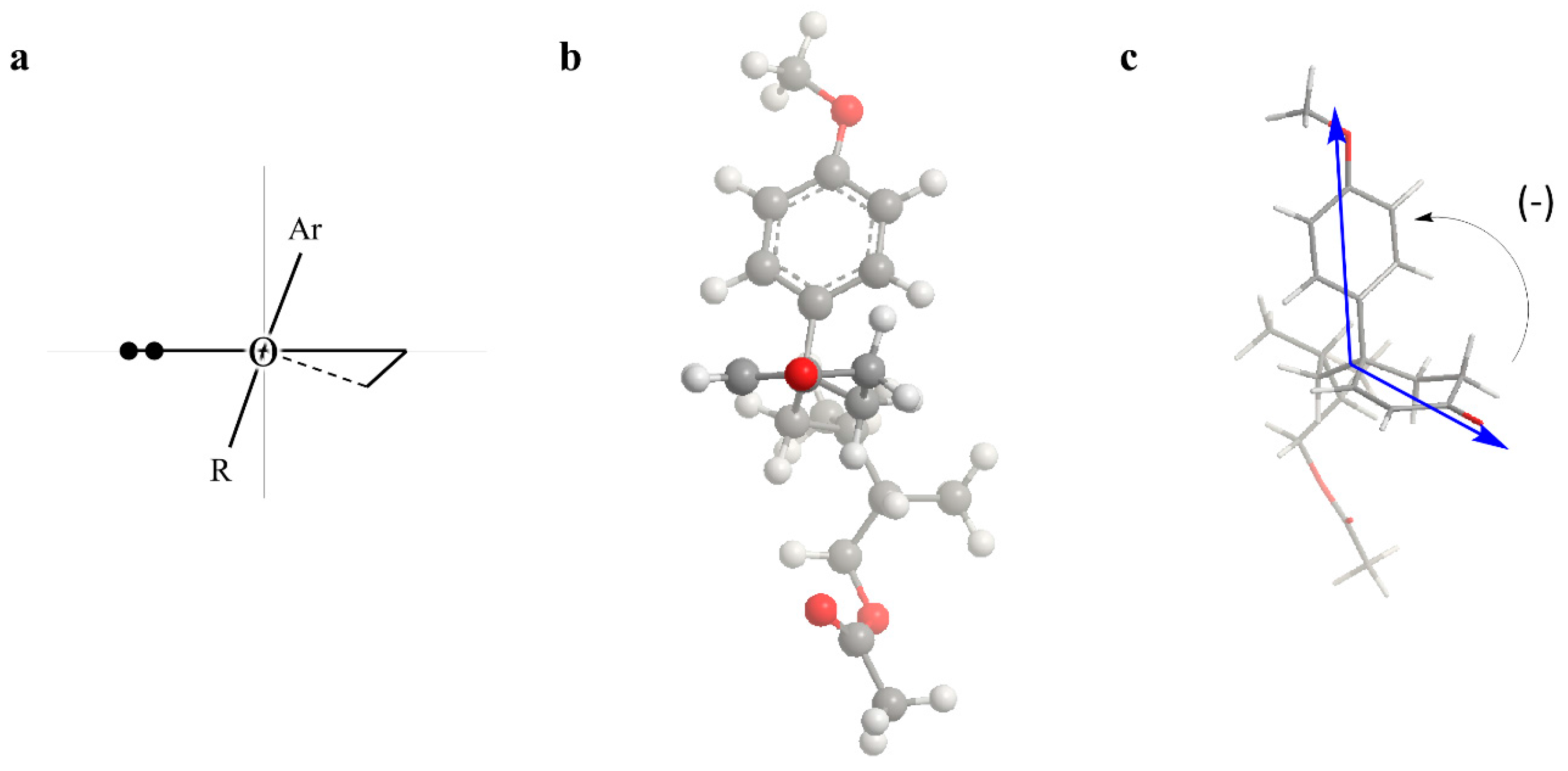
- Nitidaone B (2): white amorphous solid; −53.9° (c 0.1, MeOH); UV (MeOH) λmax (log ε) 208 (3.79), 212 (3.82) nm; CD (MeOH) λmax (Δε) 208 (−0.05), 216 (0.08), 233 (−0.30), 339 (−0.04); 1H and 13C NMR data, see Table 1; HRESIMS m/z 345.2069 [M + H]+ (calcd. for C21H28O4, 345.2066).
- Nitidaol (3): white amorphous solid; −4.94° (c 0.2, MeOH); UV (MeOH) λmax (log ε) 257 (3.63), 281 (3.60) nm; 1H and 13C NMR data, see Table 2; HRESIMS m/z 343.1913 [M + H]+ (calcd. for C21H26O4, 343.1909).
3.4. Interleukin-6 Determination
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Lia, V.; Comas, C.I.; Cortés, M.C.; Hunziker, J.H. Isozyme variation in Larrea ameghinoi and Larrea nitida (Zygophyllaceae): Genetic diversity and its bearing on their relationship. Genetica 1999, 106, 197–207. [Google Scholar] [CrossRef] [PubMed]
- Lia, V.V.; Confalonieri, V.A.; Comas, C.I.; Hunziker, J.H. Molecular Phylogeny of Larrea and Its Allies (Zygophyllaceae): Reticulate Evolution and the Probable Time of Creosote Bush Arrival to North America. Mol. Phylogenet. Evol. 2001, 21, 309–320. [Google Scholar] [CrossRef] [PubMed]
- Hunziker, J.H.; Palacios, R.A.; De Valesi, A.G.; Poggio, L. Species disjunctions in Larrea: Evidence from morphology, cytogenetics, phenolic compounds, and seed albumins. Ann. Mo. Bot. Gard. 1972, 59, 224–233. [Google Scholar] [CrossRef]
- Sakakibara, M.; Difeo, D.; Nakatani, N.; Timmermann, B.; Mabry, T.J. Flavonoid methyl ethers on the external leaf surface of Larrea tridentata and L. Divaricata. Phytochemistry 1976, 15, 727–731. [Google Scholar] [CrossRef]
- Abou-Gazar, H.; Bedir, E.; Takamatsu, S.; Ferreira, D.; Khan, I.A. Antioxidant lignans from Larrea tridentata. Phytochemistry 2004, 65, 2499–2505. [Google Scholar] [CrossRef] [PubMed]
- Leonforte, J.F. Contact dermatitis from Larrea (creosote bush). J. Am. Acad. Dermatol. 1986, 14, 202–207. [Google Scholar] [CrossRef]
- Agüero, M.B.; Svetaz, L.; Baroni, V.; Lima, B.; Luna, L.; Zacchino, S.; Saavedra, P.; Wunderlin, D.; Feresin, G.E.; Tapia, A. Urban propolis from San Juan province (Argentina): Ethnopharmacological uses and antifungal activity against Candida and dermatophytes. Ind. Crops Prod. 2014, 57, 166–173. [Google Scholar] [CrossRef]
- Agüero, M.B.; Svetaz, L.; Sánchez, M.; Luna, L.; Lima, B.; López, M.L.; Zacchino, S.; Palermo, J.; Wunderlin, D.; Feresin, G.E.; et al. Argentinean Andean propolis associated with the medicinal plant Larrea nitida Cav. (Zygophyllaceae). HPLC-MS and GC-MS characterization and antifungal activity. Food Chem. Toxicol. 2011, 49, 1970–1978. [Google Scholar] [CrossRef] [PubMed]
- Torres, R.; Urbina, F.; Morales, C.; Modak, B.; Monache, F.D. Antioxidant properties of lignans and rerulic acid from the resinous exudate of Larrea nitida. J. Chil. Chem. Soc. 2003, 48. [Google Scholar] [CrossRef]
- Ahn, H.-N.; Jeong, S.-Y.; Bae, G.-U.; Chang, M.; Zhang, D.; Liu, X.; Pei, Y.; Chin, Y.-W.; Lee, J.; Oh, S.-R.; et al. Selective estrogen receptor modulation by Larrea nitida on MCF-7 cell proliferation and immature rat uterus. Biomol. Ther. 2014, 22, 347–354. [Google Scholar] [CrossRef] [PubMed]
- Pel, P.; Kim, Y.-M.; Chin, Y.-W. Chemical Constituents with Anti-allergic Activity from the Barks of Cinnamomum cambodianum Collected in Cambodia. Bull. Korean Chem. Soc. 2015, 36, 384–387. [Google Scholar] [CrossRef]
- Quan, G.-H.; Chae, H.-S.; Song, H.H.; Ahn, K.-S.; Lee, H.-K.; Kim, Y.-H.; Oh, S.-R.; Chin, Y.-W. Anti-allergic flavones from Arthraxon hispidus. Chem. Pharm. Bull. (Tokyo) 2013, 61, 920–926. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.-M.; Chae, H.-S.; Lee, E.J.; Yang, M.H.; Park, J.H.; Yoon, K.D.; Kim, J.; Ahn, H.C.; Choi, Y.H.; Chin, Y.-W. A Citrus flavonoid, 6-demethoxytangeretin, suppresses production and gene expression of interleukin-6 in Human Mast Cell-1 via anaplastic lymphoma kinase and mitogen-activated protein kinase pathways. Biol. Pharm. Bull. 2014, 37, 871–876. [Google Scholar] [CrossRef] [PubMed]
- Yokosuka, A.; Matsuo, Y.; Jitsuno, M.; Adachi, K.; Mimaki, Y. Larrealignans A and B, novel lignan glycosides from the aerial parts of Larrea tridentata. Chem. Pharm. Bull. (Tokyo) 2011, 59, 1467–1470. [Google Scholar] [CrossRef] [PubMed]
- Hu, H.; Zhang, X.; Wu, Y. Antitumor activity of stilbenoids and flavonoids isolated from Acanthopanax brachypus. Russ. J. Gen. Chem. 2014, 84, 1434–1441. [Google Scholar] [CrossRef]
- Matsuda, H.; Morikawa, T.; Toguchida, I.; Yoshikawa, M. Structural requirements of flavonoids and related compounds for aldose reductase inhibitory activity. Chem. Pharm. Bull. (Tokyo) 2002, 50, 788–795. [Google Scholar] [CrossRef] [PubMed]
- Nieuwenhuis, J.J.; Strelow, F.; Strauss, H.F.; Wiechers, A. (4R)-(−)-O-Methyljoubertiamine and O-methyldihydrojoubertiamine, two minor alkaloids from Sceletium subvelutium L. Bolus. J. Chem. Soc. Perkin Trans. 1 1981, 284–286. [Google Scholar] [CrossRef]
- Dauben, W.G.; Shaffer, G.W.; Vietmeyer, N.D. Alkyl-substitution effects in the photochemistry of 2-cyclohexenones. J. Org. Chem. 1968, 33, 4060–4069. [Google Scholar] [CrossRef]
- Pavia, D.L.; Lampman, G.M.; Kriz, G.S.; Vyvyan, J.R. Introduction to Spectroscopy, 4th ed.; Brooks Cole: Belmont, CA, USA, 2009; pp. 387–408, ISBN-13 978-0-495-11478-9. [Google Scholar]
Sample Availability: Not available. |
| Nitidaone A (1) a | Nitidaone B (2) a | |||||
|---|---|---|---|---|---|---|
| Position | δC | Type | δH | δC | Type | δH (J in Hz) |
| 1 | 188.6 | C | 202.0 | C | ||
| 2 | 128.5 | CH | 6.29, dd (10.0, 1.9) | 129.3 | CH | 6.06, d (10.3) |
| 3 | 158.4 | CH | 7.19, dd (10.0, 3.0) | 159.8 | CH | 7.38, dd (10.3, 1.5) |
| 4 | 50.5 | C | 44.7 | C | ||
| 5 | 157.9 | CH | 7.10, dd (10.0, 3.0) | 36.3 | CH | 2.29–2.40, overlap; 2.14–2.24, overlap |
| 6 | 129.2 | CH | 6.32, dd (10.0,1.9) | 35.6 | CH | 2.29-2.40, overlap; 2.14–2.24, overlap |
| 7 | 43.2 | CH2 | 2.29, dd (13.9, 2.4); 1.95, dd (13.9, 7.6) | 46.7 | CH2 | 1.94, d (11.7); 1.61–1.67, overlap |
| 8 | 32.7 | CH | 1.45, m | 31.9 | CH | 1.62–1.68, overlap |
| 9 | 39.8 | CH | 1.77, m | 40.1 | CH | 1.63–1.69, overlap |
| 10 | 68.1 | CH2 | 4.03, dd (11.1, 6.9); 3.85, dd (11.1, 6.8) | 68.2 | CH2 | 3.90, dd (11.0, 6.6); 3.78, dd (11.0, 6.7) |
| 11 | 19.8 | CH3 | 0.93, d (7.0) | 19.2 | CH3 | 0.71, d (6.4) |
| 12 | 13.9 | CH3 | 0.89, d (7.0) | 13.7 | CH3 | 0.83, d (6.9) |
| 1′ | 133.4 | C | 137.0 | C | ||
| 2′, 6′ | 129.0 | CH | 7.28, d (9.0) | 129.3 | CH | 7.29, d (8.9) |
| 3′, 5′ | 115.5 | CH | 6.89, d (9.0) | 115.1 | CH | 6.90, d (8.9) |
| 4′ | 160.6 | C | 160.0 | C | ||
| 4′-OCH3 | 55.9 | CH3 | 3.76, s | 55.8 | CH3 | 3.78, s |
| COCH3 | 21.0 | CH3 | 2.00, s | 21.0 | CH3 | 1.98, s |
| OCO | 173.1 | C | 173.0 | C | ||
| Nitidaol (3) a | |||
|---|---|---|---|
| Position | δC | Type | δH (J in Hz) |
| 1 | 156.4 | C | |
| 2 | 118.2 | CH | 6.58, d (2.7) |
| 3 | 144.5 | C | |
| 4 | 131.1 | C | |
| 5 | 132.6 | CH | 7.01, d (8.3) |
| 6 | 114.9 | CH | 6.67, dd (8.3, 2.7) |
| 7 | 36.8 | CH2 | 2.78 dd (13.6, 4.9) |
| 2.19, dd (13.6, 9.7) | |||
| 8 | 37.8 | CH | 1.47, m |
| 9 | 38.2 | CH | 1.55, m |
| 10 | 68.7 | CH2 | 3.66, m |
| 11 | 16.8 | CH3 | 0.65, d (6.9) |
| 12 | 14.1 | CH3 | 0.74, d (6.9) |
| 1′ | 136.0 | C | |
| 2′, 6′ | 131.4 | CH | 7.16, d (8.8) |
| 3′, 5′ | 114.7 | CH | 6.92, d (8.8) |
| 4′ | 160.2 | C | |
| 4′-OCH3 | 55.8 | CH3 | 3.82, s |
| COCH3 | 21.0 | CH3 | 1.95, s |
| OCO | 173.1 | C | |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Ahn, J.; Pei, Y.; Chae, H.-S.; Kim, S.-H.; Kim, Y.-M.; Choi, Y.H.; Lee, J.; Chang, M.; Song, Y.S.; Rodriguez, R.; et al. Spiroketones and a Biphenyl Analog from Stems and Leaves of Larrea nitida and Their Inhibitory Activity against IL-6 Production. Molecules 2018, 23, 302. https://doi.org/10.3390/molecules23020302
Ahn J, Pei Y, Chae H-S, Kim S-H, Kim Y-M, Choi YH, Lee J, Chang M, Song YS, Rodriguez R, et al. Spiroketones and a Biphenyl Analog from Stems and Leaves of Larrea nitida and Their Inhibitory Activity against IL-6 Production. Molecules. 2018; 23(2):302. https://doi.org/10.3390/molecules23020302
Chicago/Turabian StyleAhn, Jongmin, Yihua Pei, Hee-Sung Chae, Seong-Hwan Kim, Young-Mi Kim, Young Hee Choi, Joongku Lee, Minsun Chang, Yun Seon Song, Roberto Rodriguez, and et al. 2018. "Spiroketones and a Biphenyl Analog from Stems and Leaves of Larrea nitida and Their Inhibitory Activity against IL-6 Production" Molecules 23, no. 2: 302. https://doi.org/10.3390/molecules23020302
APA StyleAhn, J., Pei, Y., Chae, H.-S., Kim, S.-H., Kim, Y.-M., Choi, Y. H., Lee, J., Chang, M., Song, Y. S., Rodriguez, R., Oh, D.-C., Kim, J., Choi, S., Joo, S. H., & Chin, Y.-W. (2018). Spiroketones and a Biphenyl Analog from Stems and Leaves of Larrea nitida and Their Inhibitory Activity against IL-6 Production. Molecules, 23(2), 302. https://doi.org/10.3390/molecules23020302








