Insight into Rapid DNA-Specific Identification of Animal Origin Based on FTIR Analysis: A Case Study
Abstract
1. Introduction
2. Results and Discussion
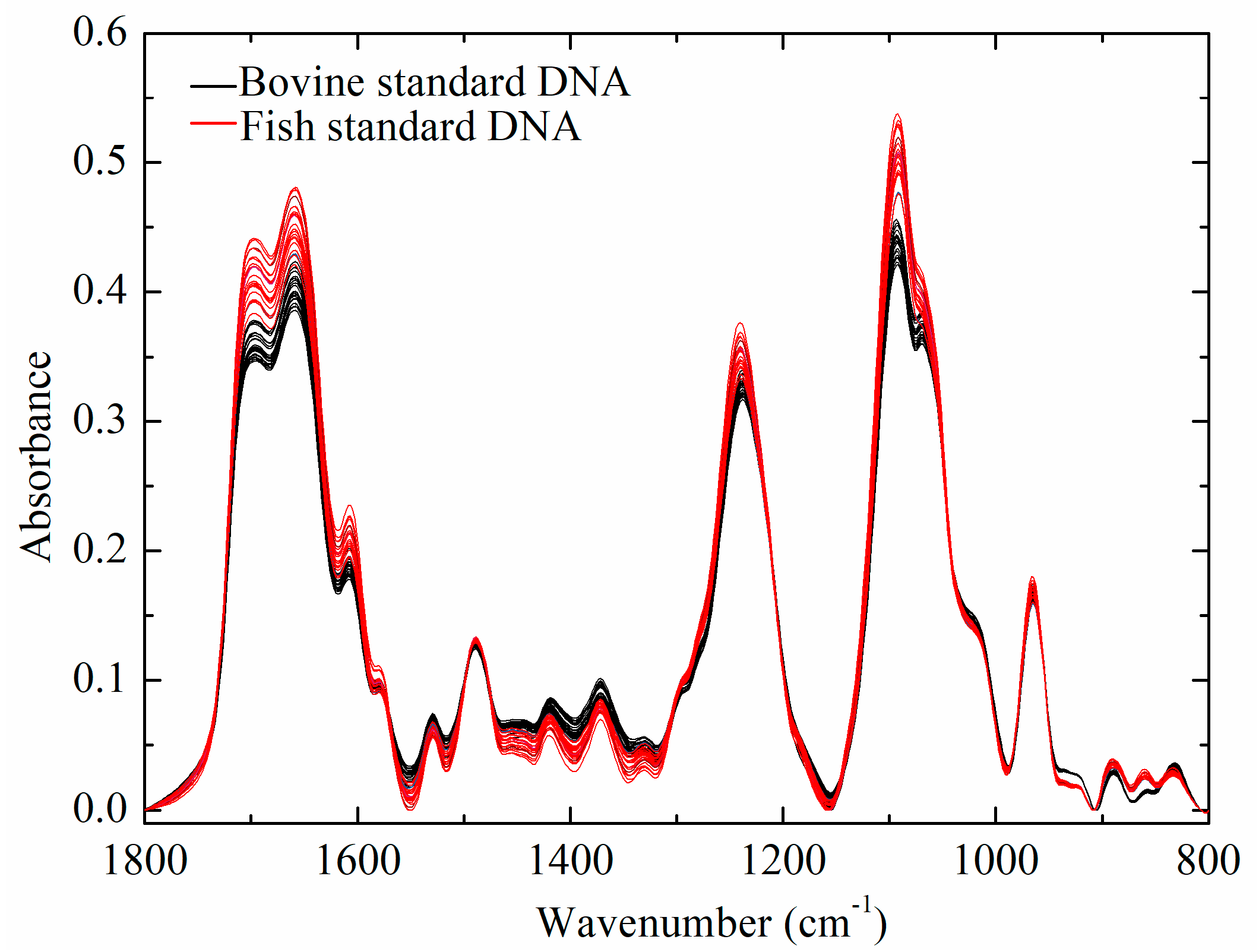
2.1. Infrared Spectral Characteristics of Bovine and Fish Standard DNA Samples
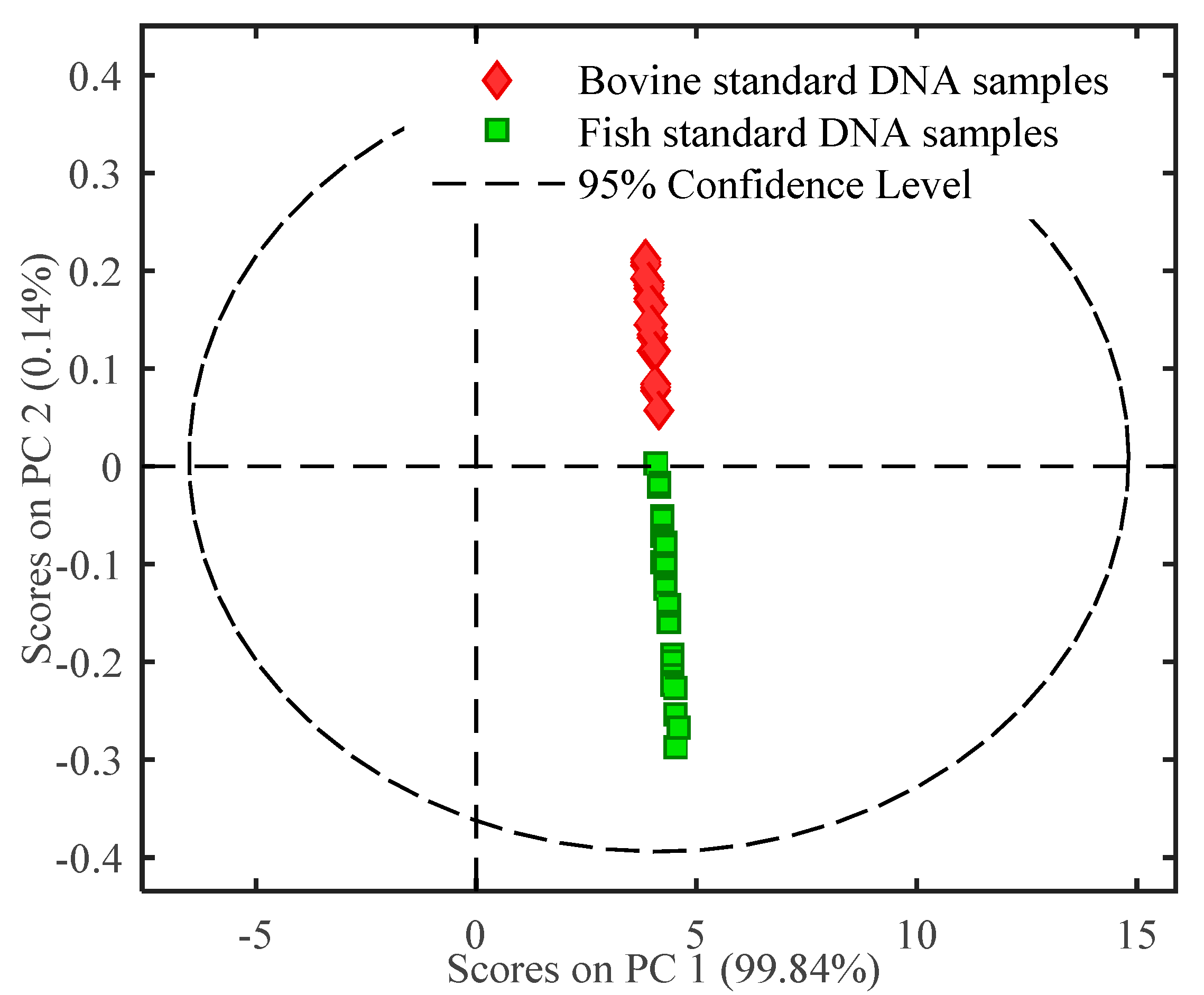
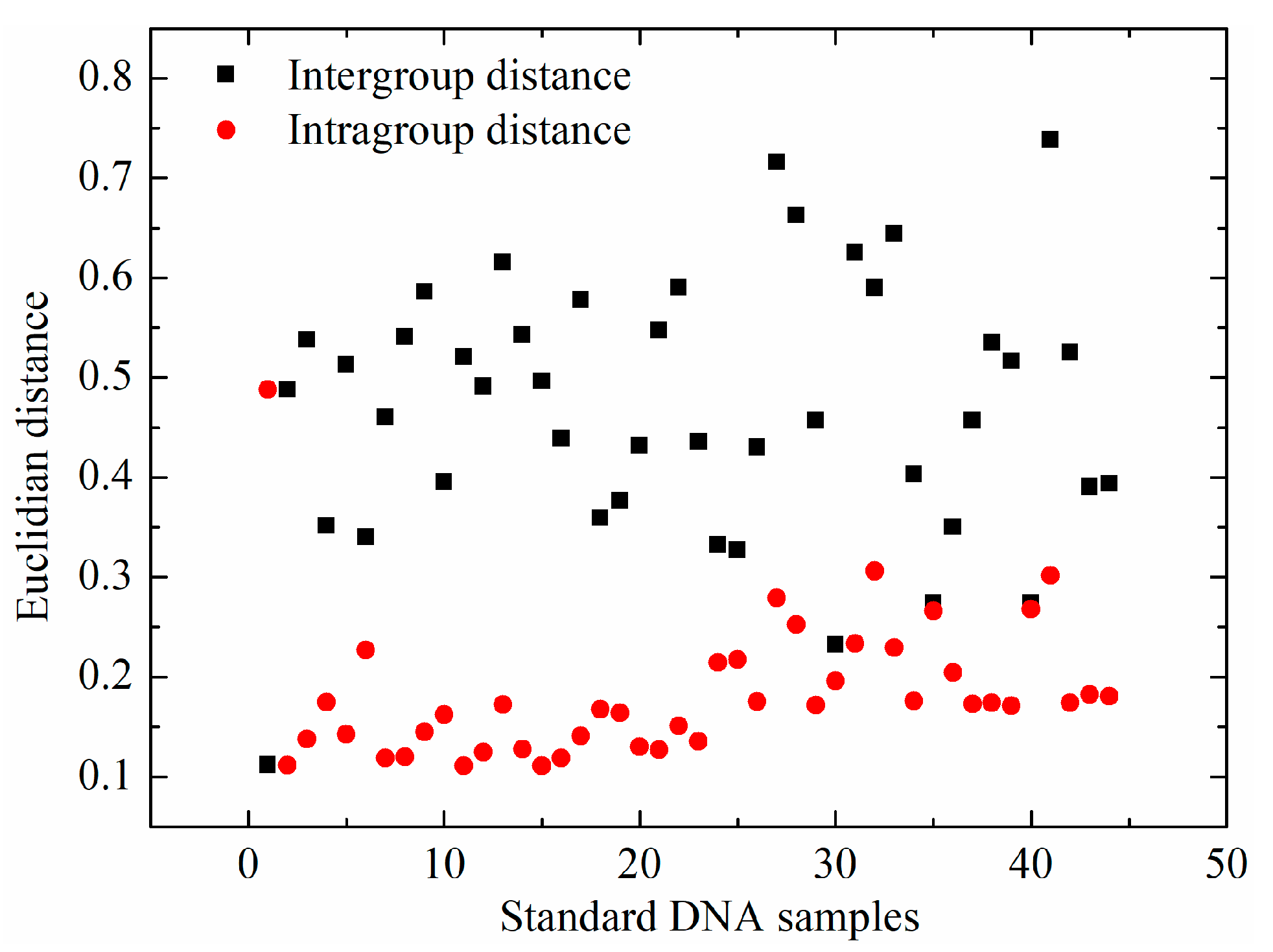
2.2. Multivariate Analysis of FTIR Spectral Data
2.3. Discussion on the Mechanism of the Discrimination Model
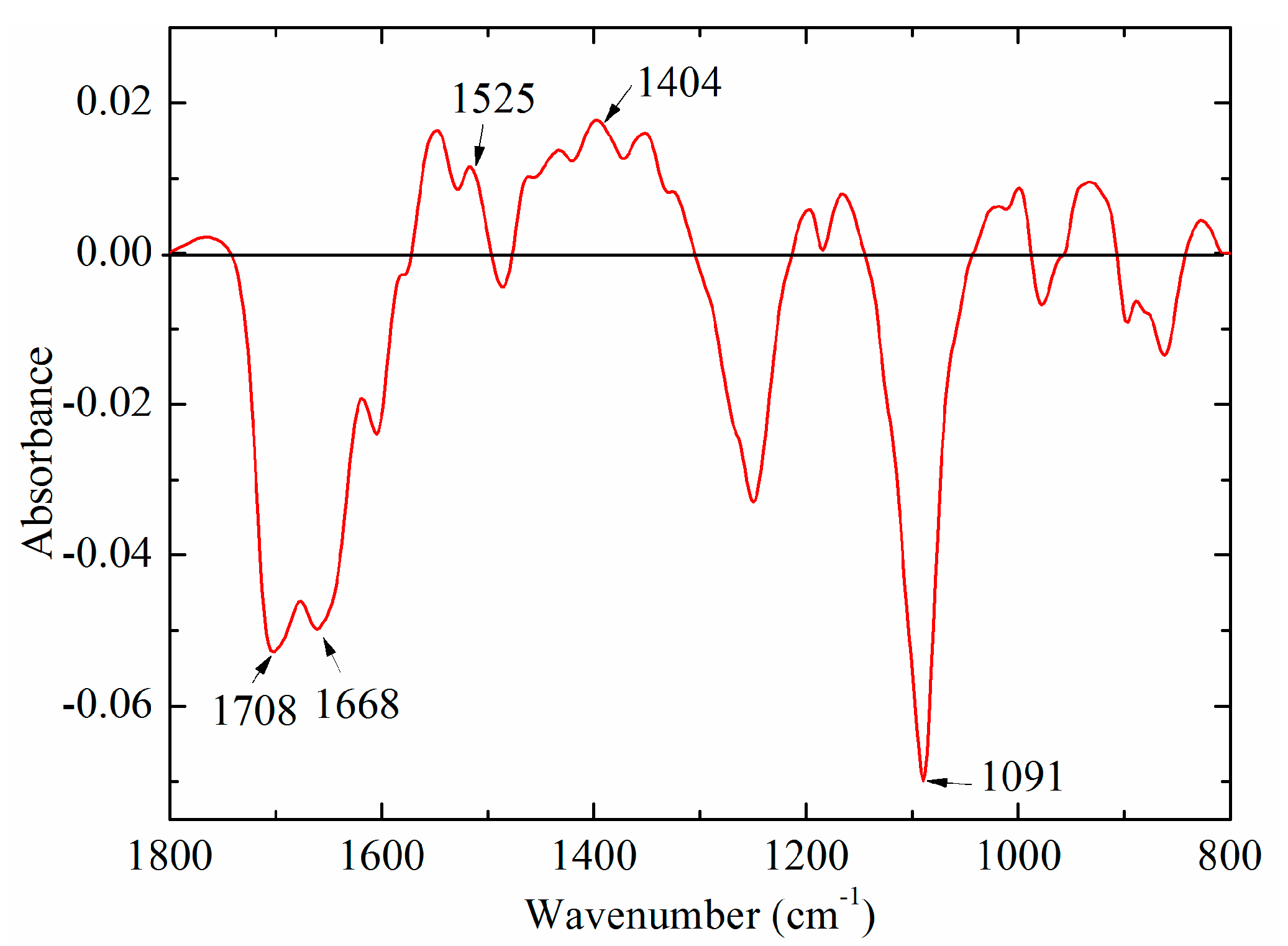
2.3.1. The Average-Difference Profile Analysis
2.3.2. Comparison of Spectral Characteristics of DNA of Two Groups
2.3.3. The Contribution of IR Spectral Characteristic Peaks of DNA
2.4. Methodology Validation of Market Meat Samples
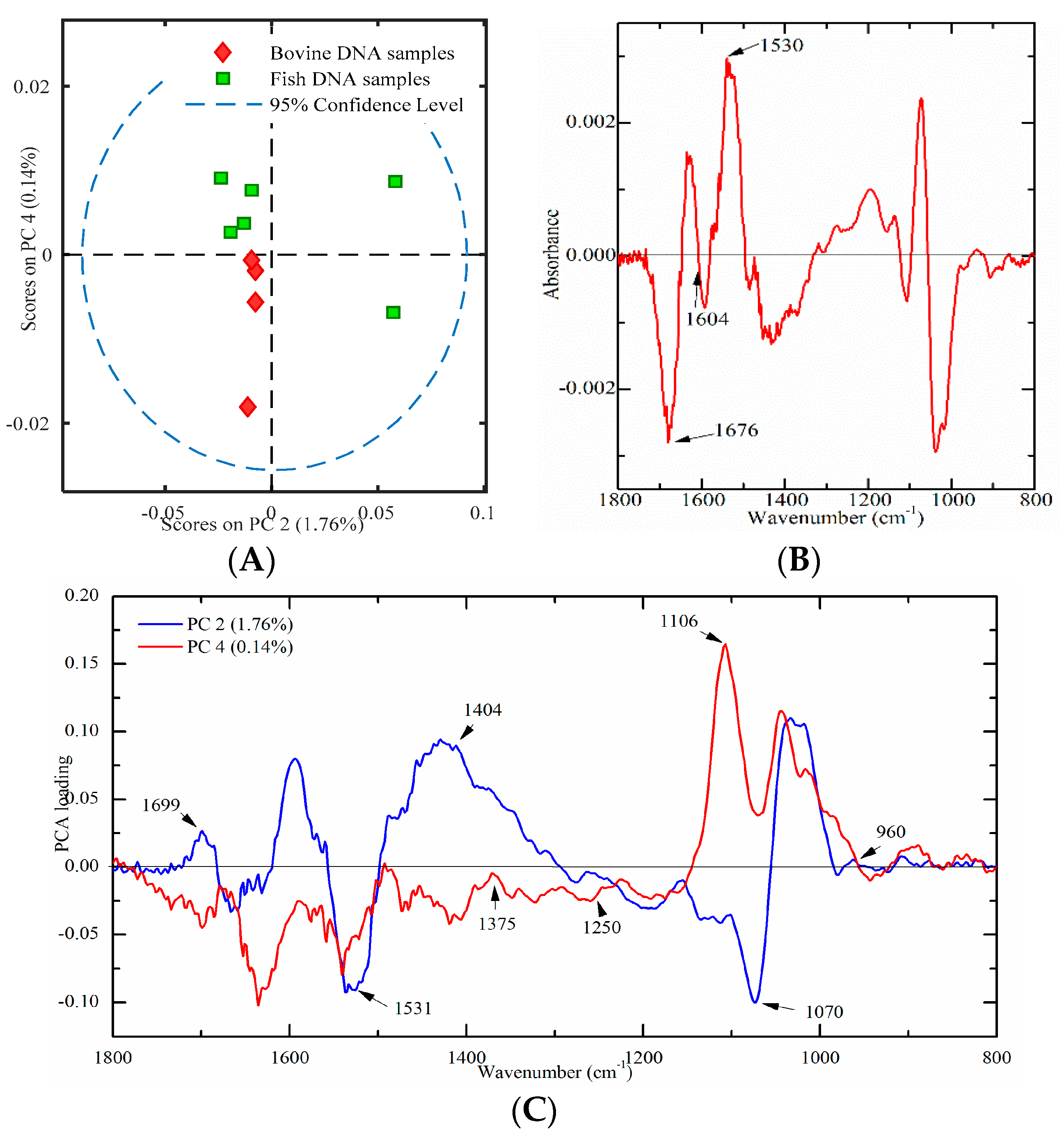
2.4.1. Bovine and Fish Samples
2.4.2. Bovine and Porcine Samples
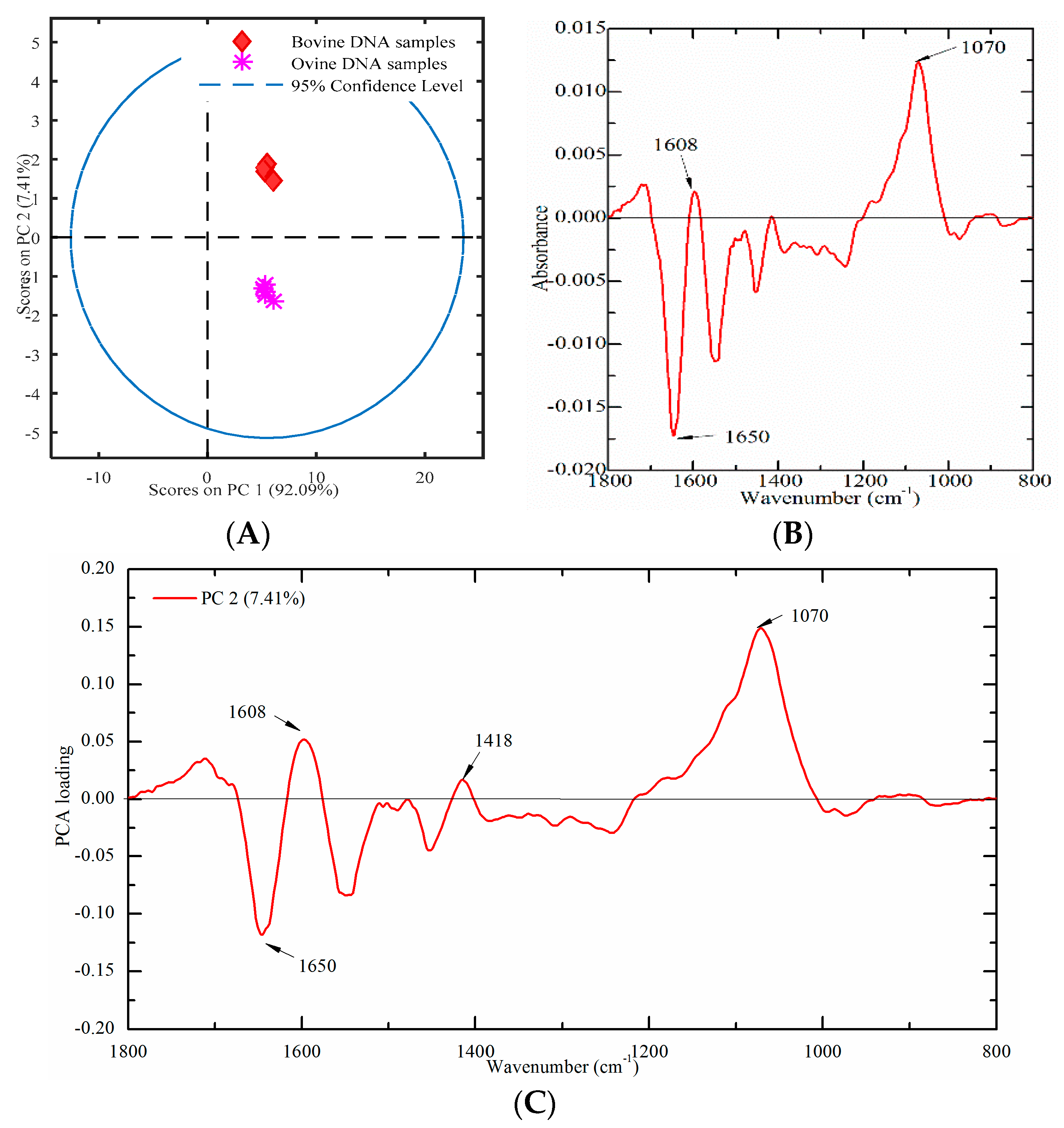
2.4.3. Bovine and Ovine Samples
3. Material and Methods
3.1. Materials
3.2. Preparation of DNA Samples
3.3. HT-FTIR Measurements
3.4. Data Processing and Multivariate Analysis
4. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Fang, X.; Zhang, C. Detection of adulterated murine components in meat products by TaqMan© real-time PCR. Food Chem. 2016, 192, 485–490. [Google Scholar] [CrossRef] [PubMed]
- Jansson, D.; Wolterink, A.; Bergwerff, L.; Hough, P.; Geukens, K.; Åstot, C. Source attribution profiling of five species of Amanita mushrooms from four European countries by high resolution liquid chromatography-mass spectrometry combined with multivariate statistical analysis and DNA-barcoding. Talanta 2018, 186, 636–644. [Google Scholar] [CrossRef] [PubMed]
- Pollack, S.J.; Kawalek, M.D.; Williams-Hill, D.M.; Hellberg, R.S. Evaluation of DNA barcoding methodologies for the identification of fish species in cooked products. Food Control 2018, 84, 297–304. [Google Scholar] [CrossRef]
- Hellberg, R.S.; Hernandez, B.C.; Hernandez, E.L. Identification of meat and poultry species in food products using DNA barcoding. Food Control 2017, 80, 23–28. [Google Scholar] [CrossRef]
- Dai, Z.; Qiao, J.; Yang, S.; Hu, S.; Zuo, J.; Zhu, W.; Huang, C. Species authentication of common meat based on PCR analysis of the mitochondrial COI gene. Appl. Biochem. Biotechnol. 2015, 176, 1770–1780. [Google Scholar] [CrossRef] [PubMed]
- Okuma, T.A.; Hellberg, R.S. Identification of meat species in pet foods using a real-time polymerase chain reaction (PCR) assay. Food Control 2015, 50, 9–17. [Google Scholar] [CrossRef]
- Lahiff, S.; Glennon, M.; Lyng, J.; Smith, T.; Maher, M.; Shilton, N. Species-specific PCR for the identification of ovine, porcine and chicken species in meat and bone meal (MBM). Mol. Cell. Probes 2001, 15, 27–35. [Google Scholar] [CrossRef] [PubMed]
- Luque-González, M.A.; Tabraue-Chávez, M.; López-Longarela, B.; Sánchez-Martín, R.M.; Ortiz-González, M.; Soriano-Rodríguez, M.; García-Salcedo, J.A.; Pernagallo, S.; Díaz-Mochón, J.J. Identification of Trypanosomatids by detecting Single Nucleotide Fingerprints using DNA analysis by dynamic chemistry with MALDI-ToF. Talanta 2018, 176, 299–307. [Google Scholar] [CrossRef] [PubMed]
- Lecrenier, M.; Ledoux, Q.; Berben, G.; Fumière, O.; Saegerman, C.; Baeten, V.; Veys, P. Determination of the ruminant origin of bone particles using fluorescence in situ hybridization (FISH). Sci. Rep. 2014, 4, 5730. [Google Scholar] [CrossRef] [PubMed]
- Emura, K.; Yamanaka, S.; Isoda, H.; Watanabe, K.N. Estimation for different genotypes of plants based on DNA analysis using near-infrared (NIR) and fourier-transform infrared (FT-IR) spectroscopy. Breed. Sci. 2006, 56, 399–403. [Google Scholar] [CrossRef]
- Song, S.Y.; Jie, E.Y.; Ahn, M.S.; Lee, I.H.; Nou, I.; Min, B.W.; Kim, S.W. Fourier transform infrared (FT-IR) spectroscopy of genomic DNA to discriminate F1 progenies from their paternal lineage of Chinese cabbage (Brassica rapa subsp. pekinensis). Mol. Breed. 2014, 33, 453–464. [Google Scholar]
- Qiu, L.; Wang, Z.; Liu, P.; Liu, R.; Cai, C.; Fan, S. Fourier Transform Infrared Spectroscopy of the DNA of the Chuxiong Population of Camellia reticulata Lindl. of China. Spectrosc. Lett. 2015, 48, 120–127. [Google Scholar] [CrossRef]
- Kirschner, C.; Maquelin, K.; Pina, P.; Ngo Thi, N.A.; Choo-Smith, L.P.; Sockalingum, G.D.; Sandt, C.; Ami, D.; Orsini, F.; Doglia, S.M.; et al. Classification and Identification of Enterococci: A Comparative Phenotypic, Genotypic, and Vibrational Spectroscopic Study. J. Clin. Microbiol. 2001, 39, 1763–1770. [Google Scholar] [CrossRef] [PubMed]
- Ami, D.; Natalello, A.; Zullini, A.; Doglia, S.M. Fourier transform infrared microspectroscopy as a new tool for nematode studies. FEBS Lett. 2004, 576, 297–300. [Google Scholar] [CrossRef] [PubMed]
- Regulation (EC). No 999/2001 of the European Parliament and of the Council laying down rules for the prevention, control and eradication of certain transmissible spongiform encephalopathies. Off. J. Eur. Union 2001, 31, L147. [Google Scholar]
- Staats, M.; Arulandhu, A.J.; Gravendeel, B.; Holst-Jensen, A.; Scholtens, I.; Peelen, T.; Prins, T.W.; Kok, E. Advances in DNA metabarcoding for food and wildlife forensic species identification. Anal. Bioanal. Chem. 2016, 408, 4615–4630. [Google Scholar] [CrossRef] [PubMed]
- Kitpipit, T.; Chotigeat, W.; Linacre, A.; Thanakiatkrai, P. Forensic animal DNA analysis using economical two-step direct PCR. Forensic Sci. Med. Pathol. 2014, 10, 29–38. [Google Scholar] [CrossRef] [PubMed]
- Han, Y.; Han, L.; Yao, Y.; Li, Y.; Liu, X. Key factors in FTIR spectroscopic analysis of DNA: The sampling technique, pretreatment temperature and sample concentration. Anal. Methods 2018, 10, 2436–2443. [Google Scholar] [CrossRef]
- Mello, M.L.S.; Vidal, B.C. Changes in the infrared microspectroscopic characteristics of DNA caused by cationic elements, different base richness and single-stranded form. PLoS ONE 2012, 7, e43169. [Google Scholar] [CrossRef] [PubMed]
- Jangir, D.K.; Tyagi, G.; Mehrotra, R.; Kundu, S. Carboplatin interaction with calf-thymus DNA: A FTIR spectroscopic approach. J. Mol. Struct. 2010, 969, 126–129. [Google Scholar] [CrossRef]
- Banyay, M.; Sarkar, M.; Gräslund, A. A library of IR bands of nucleic acids in solution. Biophys. Chem. 2003, 104, 477–488. [Google Scholar] [CrossRef]
- Ahmad, R.; Naoui, M.; Neault, J.F.; Diamantoglou, S.; Tajmir-Riahi, H.A. An FTIR spectroscopic study of calf-thymus DNA complexation with Al (III) and Ga (III) cations. J. Biomol. Struct. Dyn. 1996, 13, 795–802. [Google Scholar] [CrossRef] [PubMed]
- Taillandier, E.; Liquier, J. Infrared spectroscopy of DNA. Methods Enzymol. 1991, 211, 307–335. [Google Scholar]
- Alex, S.; Dupuis, P. FT-IR and Raman investigation of cadmium binding by DNA. Inorg. Chim. Acta 1989, 157, 271–281. [Google Scholar] [CrossRef]
- Brewer, S.H.; Anthireya, S.J.; Lappi, S.E.; Drapcho, D.L.; Franzen, S. Detection of DNA hybridization on gold surfaces by polarization modulation infrared reflection absorption spectroscopy. Langmuir 2002, 18, 4460–4464. [Google Scholar] [CrossRef]
- Blout, E.R.; Lenormant, H. Changes in the infrared spectra of solutions of deoxypentose nucleic acid in relation to its structure. Biochim. Biophys. Acta 1955, 17, 325–331. [Google Scholar] [CrossRef]
- Mello, M.L.S.; Vidal, B.C. Analysis of the DNA Fourier transform-infrared microspectroscopic signature using an all-reflecting objective. Micron 2014, 61, 49–52. [Google Scholar] [CrossRef] [PubMed]
- Adams, R.L. The Biochemistry of the Nucleic Acids; Springer Science & Business Media: New York, NY, USA, 2012. [Google Scholar]
- Hembram, K.P.S.S.; Rao, G.M. Studies on CNTs/DNA composite. Mater. Sci. Eng. C 2009, 29, 1093–1097. [Google Scholar] [CrossRef]
- Sivertsen, A.H.; Kimiya, T.; Heia, K. Automatic freshness assessment of cod (Gadus morhua) fillets by Vis/Nir spectroscopy. J. Food Eng. 2011, 103, 317–323. [Google Scholar] [CrossRef]
- Guckian, K.M.; Schweitzer, B.A.; Ren, R.X.; Sheils, C.J.; Tahmassebi, D.C.; Kool, E.T. Factors contributing to aromatic stacking in water: Evaluation in the context of DNA. J. Am. Chem. Soc. 2000, 122, 2213–2222. [Google Scholar] [CrossRef] [PubMed]
- Friedman, R.A.; Honig, B. A free energy analysis of nucleic acid base stacking in aqueous solution. Biophys. J. 1995, 69, 1528. [Google Scholar] [CrossRef]
- Egli, M.; Saenger, W. Principles of Nucleic Acid Structure; Springer Science & Business Media: New York, NY, USA, 2013. [Google Scholar]
- Nelson, H.C.; Finch, J.T.; Luisi, B.F.; Klug, A. The structure of an oligo (dA)·oligo (dT) tract and its biological implications. Nature 1987, 330, 221–226. [Google Scholar] [CrossRef] [PubMed]
- Cheng, X.; He, W.; Huang, F.; Huang, M.; Zhou, G. Multiplex real-time PCR for the identification and quantification of DNA from duck, pig and chicken in Chinese blood curds. Food Res. Int. 2014, 60, 30–37. [Google Scholar] [CrossRef]
- Yalçınkaya, B.; Yumbul, E.; Mozioğlu, E.; Akgoz, M. Comparison of DNA extraction methods for meat analysis. Food Chem. 2017, 221, 1253–1257. [Google Scholar] [CrossRef] [PubMed]
- Steward, K.F.; Robinson, C.; Waller, A.S. Transcriptional changes are involved in phenotype switching in Streptococcus equi subspecies equi. Mol. Biosyst. 2016, 12, 1194–1200. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.; Wei, J.; Xiang, L.; Cai, Z. Mass spectrometry investigation of DNA adduct formation from bisphenol A quinone metabolite and MCF-7 cell DNA. Talanta 2018, 182, 583–589. [Google Scholar] [CrossRef] [PubMed]
- Kondepati, V.R.; Heise, H.M.; Oszinda, T.; Mueller, R.; Keese, M.; Backhaus, J. Detection of structural disorders in colorectal cancer DNA with Fourier-transform infrared spectroscopy. Vib. Spectrosc. 2008, 46, 150–157. [Google Scholar] [CrossRef]
- Dina, N.E.; Muntean, C.M.; Leopold, N.; Fălămaș, A.; Halmagyi, A.; Coste, A. Structural changes induced in grapevine (Vitis vinifera L.) DNA by femtosecond IR laser pulses: A surface-enhanced Raman spectroscopic study. Nanomaterials 2016, 6, 96. [Google Scholar] [CrossRef] [PubMed]
- Abbas, O.; Fernández Pierna, J.A.; Codony, R.; von Holst, C.; Baeten, V. Assessment of the discrimination of animal fat by FT-Raman spectroscopy. J. Mol. Struct. 2009, 924–926, 294–300. [Google Scholar] [CrossRef]
- Correia, R.M.; Tosato, F.; Domingos, E.; Rodrigues, R.R.T.; Aquino, L.F.M.; Filgueiras, P.R.; Lacerda, V.; Romão, W. Portable near infrared spectroscopy applied to quality control of Brazilian coffee. Talanta 2018, 176, 59–68. [Google Scholar] [CrossRef] [PubMed]
- Gao, F.; Xu, L.; Zhang, Y.; Yang, Z.; Han, L.; Liu, X. Analytical Raman spectroscopic study for discriminant analysis of different animal-derived feedstuff: Understanding the high correlation between Raman spectroscopy and lipid characteristics. Food Chem. 2018, 240, 989–996. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Samples of the compounds are not available from the authors. |


| Assignment | Comment | Frequencies | References | |
|---|---|---|---|---|
| Bovine Standard DNA | Fish Standard DNA | |||
| Thymine | C=O stretching | 1712 | 1711 | [19,21,23,24] |
| Thymine | C=O stretching | 1659 | 1659 | [19,20,22] |
| Adenine | Base/in-plane vibration | 1608 | 1608 | [20,22] |
| Cytosine, Guanine | Base/in-plane vibration | 1529 | 1529 | [19] |
| Adenine, Guanine | Ring vibration, C=N | 1489 | 1489 | [21] |
| Adenine, Guanine | Base/in-plane vibration | 1420 | 1420 | [19] |
| Adenine, Guanine | dA, dG anti | 1371 | 1371 | [23] |
| Backbone-A form | Vas PO2− | 1238 | 1240 | [22] |
| Backbone | Vs PO2− | 1094 | 1092 | [20] |
| Backbone | O-P-O bending | 964 | 964 | [19] |
| Deoxyribose | Deoxyribose ring vibration | 889 | 889 | [21] |
| Deoxyribose | Deoxyribose-phosphate, B-marker | 831 | 833 | [19] |
| Bovine DNA Standard Samples | Fish DNA Standard Samples | |
|---|---|---|
| Sensitivity (Cal) | 1.00 | 1.00 |
| Specificity (Cal) | 1.00 | 1.00 |
| Classification error (Cal) | 0.00 | 0.00 |
| Sensitivity (CV) | 1.00 | 1.00 |
| Specificity (CV) | 1.00 | 1.00 |
| Classification error (CV) | 0.00 | 0.00 |
| Sensitivity (Val) | 1.00 | 1.00 |
| Specificity (Val) | 1.00 | 1.00 |
| Classification error (Val) | 0.00 | 0.00 |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Han, Y.; Jian, L.; Yao, Y.; Wang, X.; Han, L.; Liu, X. Insight into Rapid DNA-Specific Identification of Animal Origin Based on FTIR Analysis: A Case Study. Molecules 2018, 23, 2842. https://doi.org/10.3390/molecules23112842
Han Y, Jian L, Yao Y, Wang X, Han L, Liu X. Insight into Rapid DNA-Specific Identification of Animal Origin Based on FTIR Analysis: A Case Study. Molecules. 2018; 23(11):2842. https://doi.org/10.3390/molecules23112842
Chicago/Turabian StyleHan, Yahong, Lin Jian, Yumei Yao, Xinlei Wang, Lujia Han, and Xian Liu. 2018. "Insight into Rapid DNA-Specific Identification of Animal Origin Based on FTIR Analysis: A Case Study" Molecules 23, no. 11: 2842. https://doi.org/10.3390/molecules23112842
APA StyleHan, Y., Jian, L., Yao, Y., Wang, X., Han, L., & Liu, X. (2018). Insight into Rapid DNA-Specific Identification of Animal Origin Based on FTIR Analysis: A Case Study. Molecules, 23(11), 2842. https://doi.org/10.3390/molecules23112842





