Thermodynamic Model for B-Z Transition of DNA Induced by Z-DNA Binding Proteins
Abstract
1. Introduction
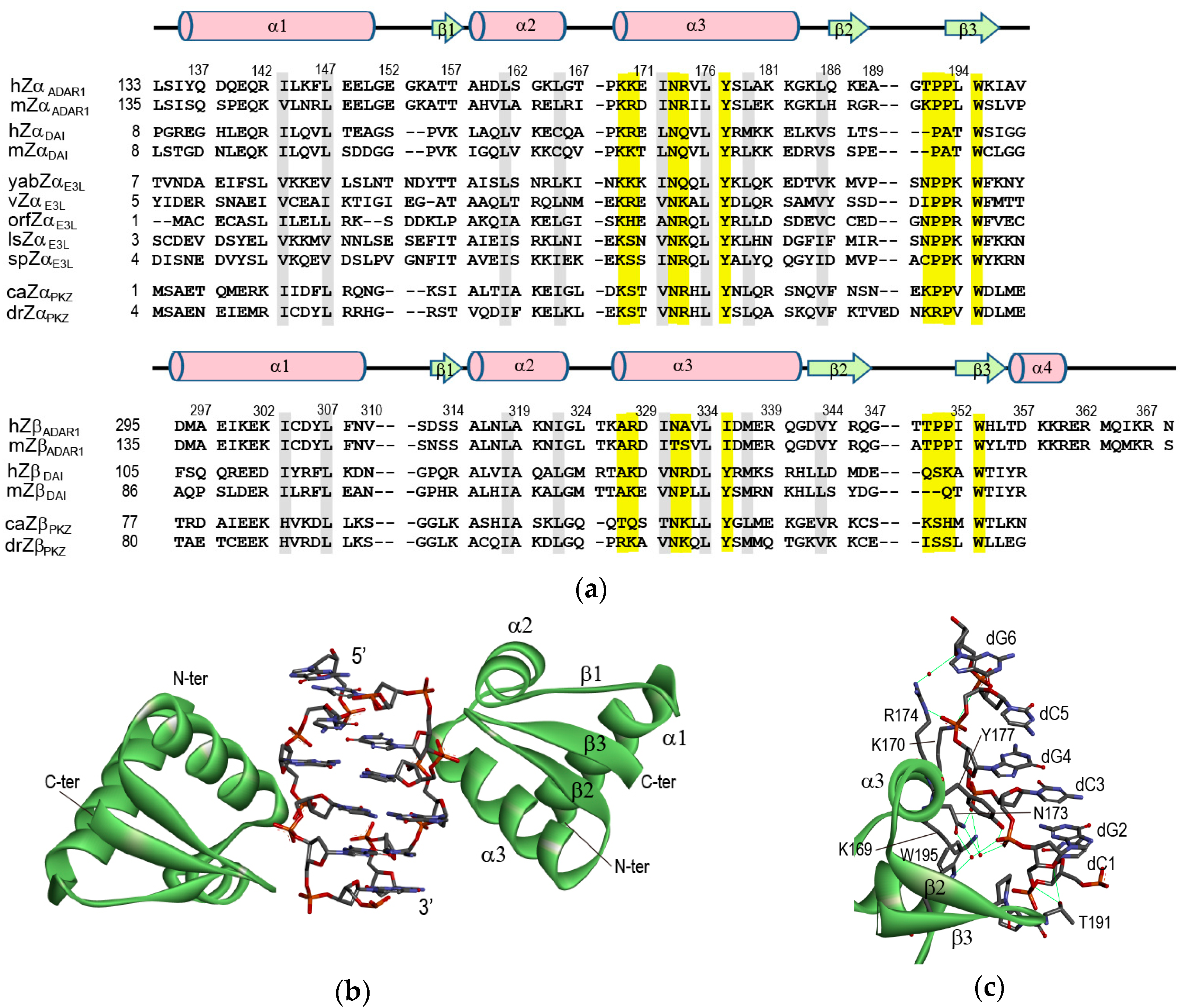
2. Crystal Structures of ZBPs Complexed with DNA Duplexes
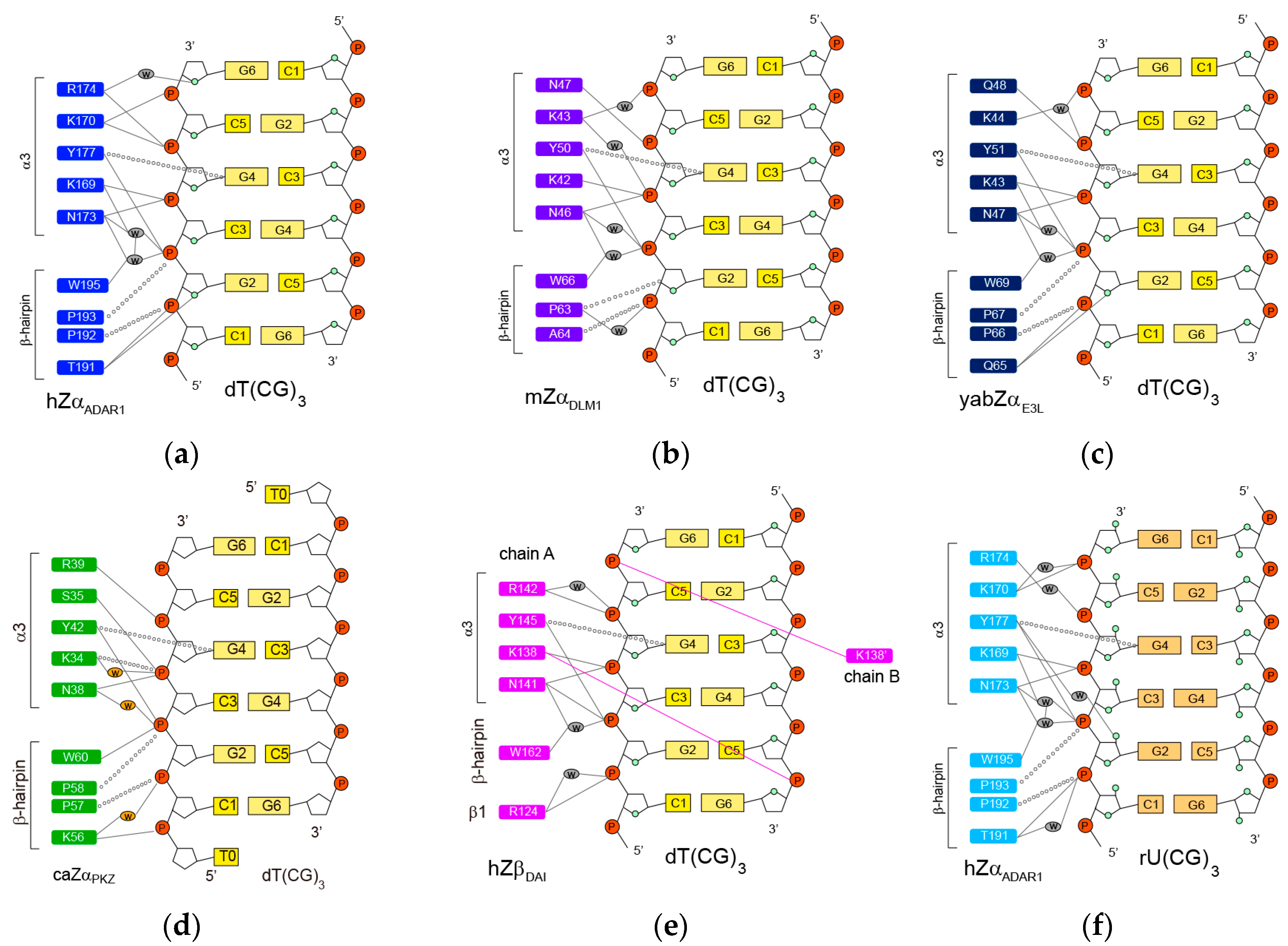
2.1. hZαADAR1-Z-DNA Complex
2.2. mZαDLM1-Z-DNA Complex
2.3. yabZαE3L-Z-DNA Complex
2.4. caZαPKZ-Z-DNA Complex
2.5. hZβDAI-Z-DNA Complex
2.6. hZαADAR1-Z-RNA Complex
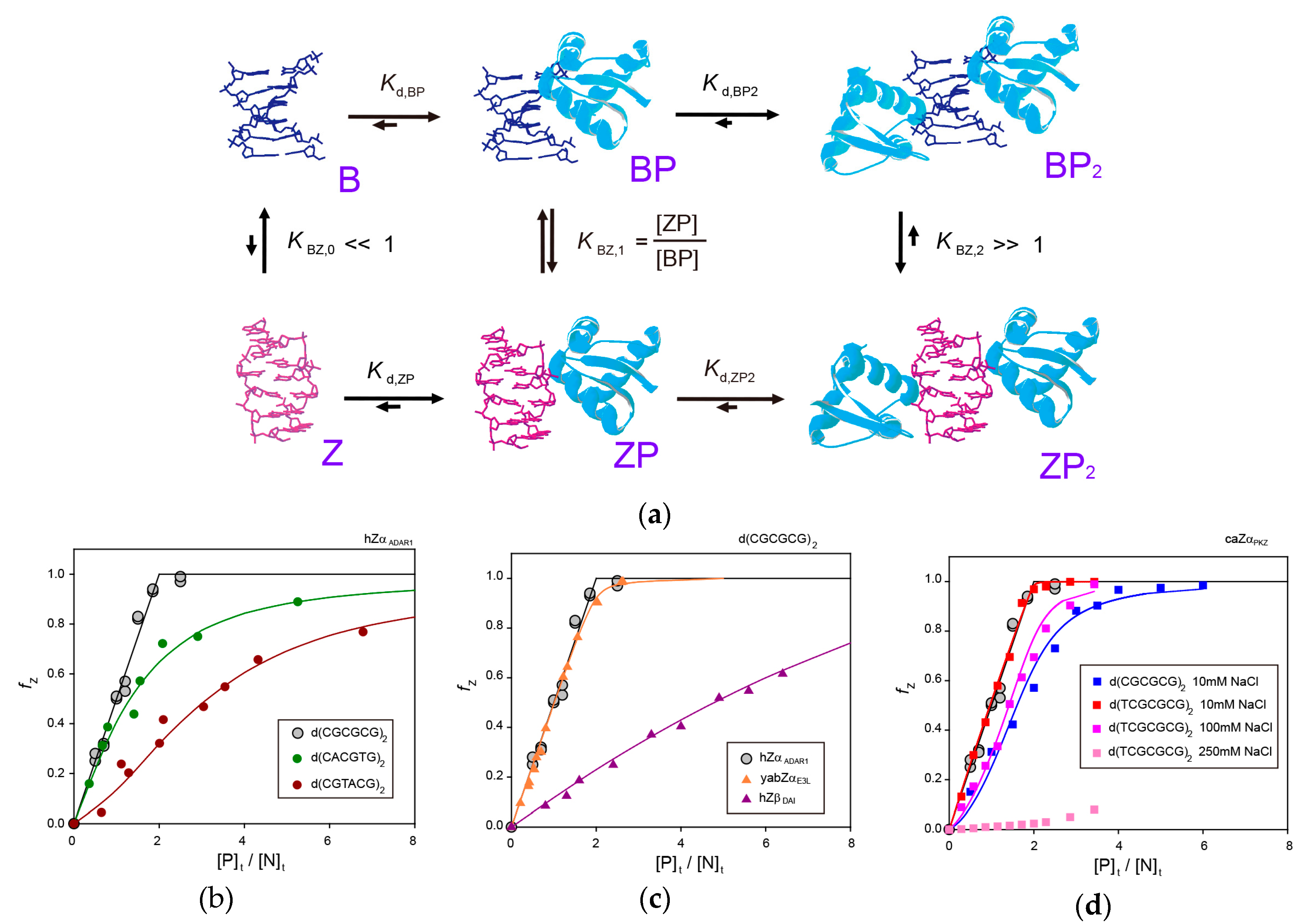
3. Molecular Mechanism of B-Z Transition of 6-bp DNA Induced by ZBPs
3.1. B-Z Transition of a 6-bp CG-Repeat DNA by hZαADAR1
3.2. B-Z Transition of a 6-bp Non-CG-Repeat DNA by hZαADAR1
3.3. B-Z Transition of a 6-bp DNA by yabZαE3L
3.4. B-Z Transition of a 6-bp DNA by hZβDAI
3.5. B-Z Transition of a 6-bp DNA by caZαPKZ
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Pohl, F.M.; Jovin, T.M. Salt-induced co-operative conformational change of a synthetic DNA: Equilibrium and kinetic studies with poly(dG-dC). J. Mol. Biol. 1972, 67, 375–396. [Google Scholar] [CrossRef]
- Wang, A.H.; Quigley, G.J.; Kolpak, F.J.; Crawford, J.L.; van Boom, J.H.; van der Marel, G.; Rich, A. Molecular structure of a left-handed double helical DNA fragment at atomic resolution. Nature 1979, 282, 680–686. [Google Scholar] [CrossRef] [PubMed]
- Herbert, A.; Rich, A. The biology of left-handed Z-DNA. J. Biol. Chem. 1996, 271, 11595–11598. [Google Scholar] [CrossRef] [PubMed]
- Herbert, A.; Rich, A. Left-handed Z-DNA: Structure and function. Genetica 1999, 106, 37–47. [Google Scholar] [CrossRef] [PubMed]
- Liu, L.F.; Wang, J.C. Supercoiling of the DNA template during transcription. Proc. Natl. Acad. Sci. USA 1987, 84, 7024–7027. [Google Scholar] [CrossRef] [PubMed]
- Herbert, A.; Lowenhaupt, K.; Spitzner, J.; Rich, A. Chicken double-stranded RNA adenosine deaminase has apparent specificity for Z-DNA. Proc. Natl. Acad. Sci. USA 1995, 92, 7550–7554. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, T.; Rould, M.A.; Lowenhaupt, K.; Herbert, A.; Rich, A. Crystal structure of the Zα domain of the human editing enzyme ADAR1 bound to left-handed Z-DNA. Science 1999, 284, 1841–5948. [Google Scholar] [CrossRef] [PubMed]
- Rich, A.; Zhang, S. Z-DNA: The long road to biological function. Nat. Rev. Genet 2003, 4, 566–572. [Google Scholar] [CrossRef] [PubMed]
- Schade, M.; Turner, C.J.; Kühne, R.; Schmieder, P.; Lowenhaupt, K.; Herbert, A.; Rich, A.; Oschkinat, H. The solution structure of the Zα domain of the human RNA editing enzyme ADAR1 reveals a prepositioned binding surface for Z-DNA. Proc. Natl. Acad. Sci. USA 1999, 96, 12465–12470. [Google Scholar] [CrossRef] [PubMed]
- Herbert, A.G.; Rich, A. A method to identify and characterize Z-DNA binding proteins using a linear oligodeoxynucleotide. Nucleic Acids Res. 1993, 21, 2669–2672. [Google Scholar] [CrossRef] [PubMed]
- Herbert, A.; Alfken, J.; Kim, Y.G.; Mian, I.S.; Nishikura, K.; Rich, A. A Z-DNA binding domain present in the human editing enzyme, double-stranded RNA adenosine deaminase. Proc. Natl. Acad. Sci. USA 1997, 94, 8421–8426. [Google Scholar] [CrossRef] [PubMed]
- Herbert, A.; Schade, M.; Lowenhaupt, K.; Alfken, J.; Schwartz, T.; Shlyakhtenko, L.S.; Lyubchenko, Y.L.; Rich, A. The Zα domain from human ADAR1 binds to the Z-DNA conformer of many different sequences. Nucleic Acids Res. 1998, 26, 3486–3493. [Google Scholar] [CrossRef] [PubMed]
- Yanai, H.; Savitsky, D.; Tamura, T.; Taniguchi, T. Regulation of the cytosolic DNA-sensing system in innate immunity: A current view. Curr. Opin. Immunol. 2009, 21, 17–22. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Choi, M.K.; Ban, T.; Yanai, H.; Negishi, H.; Lu, Y.; Tamura, T.; Takaoka, A.; Nishikura, K.; Taniguchi, T. Regulation of innate immune responses by DAI (DLM-1/ZBP1) and other DNA-sensing molecules. Proc. Natl. Acad. Sci. USA 2008, 105, 5477–5482. [Google Scholar] [CrossRef] [PubMed]
- Pham, H.T.; Park, M.Y.; Kim, K.K.; Kim, Y.-G.; Ahn, J.H. Intracellular localization of human ZBP1: Differential regulation by the Z-DNA binding domain, Zα, in splice variants. Biochem. Biophys. Res. Commun. 2006, 348, 145–152. [Google Scholar] [CrossRef] [PubMed]
- Deigendesch, N.; Koch-Nolte, F.; Rothenburg, S. ZBP1 subcellular localization and association with stress granules is controlled by its Z-DNA binding domains. Nucleic Acids Res. 2006, 34, 5007–5020. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.-G.; Muralinath, M.; Brandt, T.; Pearcy, M.; Hauns, K.; Lowenhaupt, K.; Jacobs, B.L.; Rich, A. A role for Z-DNA binding in vaccinia virus pathogenesis. Proc. Natl. Acad. Sci. USA 2003, 100, 6974–6979. [Google Scholar] [CrossRef] [PubMed]
- Kim, Y.-G.; Lowenhaupt, K.; Oh, D.-B.; Kim, K.K.; Rich, A. Evidence that vaccinia virulence factor E3L binds to Z-DNA in vivo: Implications for development of a therapy for poxvirus infection. Proc. Natl. Acad. Sci. USA 2004, 101, 1514–1518. [Google Scholar] [CrossRef] [PubMed]
- Kwon, J.-A.; Rich, A. Biological function of the vaccinia virus Z-DNA binding protein E3L: Gene transactivation and antiapoptotic activity in HeLa cells. Proc. Natl. Acad. Sci. USA 2005, 102, 12759–12764. [Google Scholar] [CrossRef] [PubMed]
- Williams, B.R. PKR: A sentinel kinase for cellular stress. Oncogene 1999, 18, 6112–6120. [Google Scholar] [CrossRef] [PubMed]
- Wu, S.; Kaufman, R.J. A model for the double-stranded RNA (dsRNA)-dependent dimerization and activation of the dsRNA-activated protein kinase PKR. J. Biol. Chem. 1997, 272, 1291–1296. [Google Scholar] [CrossRef] [PubMed]
- Tan, S.L.; Gale, M.J.; Katze, M.G. Double-stranded RNA-independent dimerization of interferon-induced protein kinase PKR and inhibition of dimerization by the cellular P58IPK inhibitor. Mol. Cell. Biol. 1998, 18, 2431–2443. [Google Scholar] [CrossRef] [PubMed]
- Su, J.; Zhu, Z.; Wang, Y. Molecular cloning, characterization and expression analysis of the PKZ gene in rare minnow Gobiocypris rarus. Fish Shellfish Immunol. 2008, 25, 106–113. [Google Scholar] [CrossRef] [PubMed]
- Bergan, V.; Jagus, R.; Lauksund, S.; Kileng, O.; Robertsen, B. The Atlantic salmon Z-DNA binding protein kinase phosphorylates translation initiation factor 2α and constitutes a unique orthologue to the mammalian dsRNA-activated protein kinase R. FEBS J. 2008, 275, 184–197. [Google Scholar] [CrossRef] [PubMed]
- Rothenburg, S.; Deigendesch, N.; Dittmar, K.; Koch-Nolte, F.; Haag, F.; Lowenhaupt, K.; Rich, A. A PKR-like eukaryotic initiation factor 2α kinase from zebrafish contains Z-DNA binding domains instead of dsRNA binding domains. Proc. Natl. Acad. Sci. USA 2005, 102, 1602–1607. [Google Scholar] [CrossRef] [PubMed]
- Hu, C.Y.; Zhang, Y.B.; Huang, G.P.; Zhang, Q.Y.; Gui, J.F. Molecular cloning and characterization of a fish PKR-like gene from cultured CAB cells induced by UV-inactivated virus. Fish Shellfish Immunol. 2004, 17, 353–366. [Google Scholar] [CrossRef] [PubMed]
- Schade, M.; Turner, C.J.; Lowenhaupt, K.; Rich, A.; Herbert, A. Structure–function analysis of the Z-DNA-binding domain Zα of dsRNA adenosine deaminase type I reveals similarity to the (α + β) family of helix–turn–helix proteins. EMBO J. 1999, 18, 470–479. [Google Scholar] [CrossRef] [PubMed]
- Jeong, M.; Lee, A.-R.; Kim, H.-E.; Choi, Y.-G.; Choi, B.-S.; Lee, J.-H. NMR study of the Z-DNA binding mode and B-Z transition activity of the Zα domain of human ADAR1 when perturbed by mutation on the α3 helix and β-hairpin. Arch. Biochem. Biophys. 2014, 558, 95–103. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, T.; Behlke, J.; Lowenhaupt, K.; Heinemann, U.; Rich, A. Structure of the DLM-1–Z-DNA complex reveals a conserved family of Z-DNA-binding proteins. Nat. Struct. Biol. 2001, 8, 761–765. [Google Scholar] [CrossRef] [PubMed]
- Ha, S.C.; Lokanath, N.K.; Quyen, D.V.; Wu, C.A.; Lowenhaupt, K.; Rich, A.; Kim, Y.-G.; Kim, K.K. A poxvirus protein forms a complex with left-handed Z-DNA: Crystal structure of a Yatapoxvirus Zα bound to DNA. Proc. Natl. Acad. Sci. USA 2004, 101, 14367–14372. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Hur, J.; Park, K.; Bae, S.; Shin, D.; Ha, S.C.; Hwang, H.-Y.; Hohng, S.; Lee, J.-H.; Lee, S.; et al. Distinct Z-DNA binding mode of a PKR-like protein kinase containing a Z-DNA binding domain (PKZ). Nucleic Acids Res. 2014, 42, 5937–5948. [Google Scholar] [CrossRef] [PubMed]
- Ha, S.C.; Kim, D.; Hwang, H.-Y.; Rich, A.; Kim, Y.-G.; Kim, K.K. The crystal structure of the second Z-DNA binding domain of human DAI (ZBP1) in complex with Z-DNA reveals an unusual binding mode to Z-DNA. Proc. Natl. Acad. Sci. USA 2008, 105, 20671–20676. [Google Scholar] [CrossRef] [PubMed]
- Placido, D.; Brown II, B.A.; Lowenhaupt, K.; Rich, A.; Athanasiadis, A. A left-handed RNA double helix bound by the Zα domain of the RNA-editing enzyme ADAR1. Structure 2007, 15, 395–404. [Google Scholar] [CrossRef] [PubMed]
- Ha, S.C.; Choi, J.; Hwang, H.-Y.; Rich, A.; Kim, Y.-G.; Kim, K.K. The structures of non-CG-repeat Z-DNAs co-crystallized with the Z-DNA-binding domain, hZαADAR1. Nucleic Acids Res. 2009, 37, 629–637. [Google Scholar] [CrossRef] [PubMed]
- Athanasiadis, A.; Placido, D.; Maas, S.; Brown II, B.A.; Lowenhaupt, K.; Rich, A. The crystal structure of the Zβ domain of the RNA-editing enzyme ADAR1 reveals distinct conserved surfaces among Z-domains. J. Mol. Biol. 2005, 351, 496–507. [Google Scholar] [CrossRef] [PubMed]
- Wu, C.X.; Wang, S.J.; Lin, G.; Hu, C.Y. The Zα domain of PKZ from Carassius auratus can bind to d(GC)(n) in negative supercoils. Fish Shellfish Immunol. 2010, 28, 783–788. [Google Scholar] [CrossRef] [PubMed]
- Kim, D.; Hwang, H.Y.; Kim, Y.G.; Kim, K.K. Crystallization and preliminary X-ray crystallographic studies of the Z-DNA-binding domain of a PKR-like kinase (PKZ) in complex with Z-DNA. Acta Crystallogr. F Struct. Biol. Cryst. Commun. 2009, 65, 267–270. [Google Scholar] [CrossRef] [PubMed]
- Kim, K.; Khayrutdinov, B.I.; Lee, C.-K.; Cheong, H.-K.; Kang, S.W.; Park, H.; Lee, S.; Kim, Y.-G.; Jee, J.G.; Rich, A.; et al. Solution structure of the Zβ domain of human DNA-dependent activator of IFN-regulatory factors and its binding modes to B- and Z-DNAs. Proc. Natl. Acad. Sci. USA 2011, 108, 6921–6926. [Google Scholar] [CrossRef] [PubMed]
- Koeris, M.; Funke, L.; Shrestha, J.; Rich, A.; Maas, S. Modulation of ADAR1 editing activity by Z-RNA in vitro. Nucleic Acids Res. 2005, 33, 5362–5370. [Google Scholar] [CrossRef] [PubMed]
- Brown, B.A., II; Lowenhaupt, K.; Wilbert, C.M.; Hanlon, E.B.; Rich, A. The Zα domain of the editing enzyme dsRNA adenosine deaminase binds left-handed Z-RNA as well as Z-DNA. Proc. Natl. Acad. Sci. USA 2000, 97, 13532–13536. [Google Scholar] [CrossRef] [PubMed]
- Kang, Y.-M.; Bang, J.; Lee, E.-H.; Ahn, H.-C.; Seo, Y.-J.; Kim, K.K.; Kim, Y.-G.; Choi, B.-S.; Lee, J.-H. NMR spectroscopic elucidation of the B-Z transition of a DNA double helix induced by the Zα domain of human ADAR1. J. Am. Chem. Soc. 2009, 131, 11485–11491. [Google Scholar] [CrossRef] [PubMed]
- Seo, Y.-J.; Ahn, H.-C.; Lee, E.-H.; Bang, J.; Kang, Y.-M.; Kim, H.-E.; Lee, Y.-M.; Kim, K.; Choi, B.-S.; Lee, J.-H. Sequence discrimination of the Zα domain of human ADAR1 during B-Z transition of DNA duplexes. FEBS Lett. 2010, 584, 4344–4350. [Google Scholar] [CrossRef] [PubMed]
- Lee, E.-H.; Seo, Y.-J.; Ahn, H.-C.; Kang, Y.-M.; Kim, H.-E.; Lee, Y.-M.; Choi, B.-S.; Lee, J.-H. NMR study of hydrogen exchange during the B-Z transition of a DNA duplex induced by the Zα domain of yatapoxvirus E3L. FEBS Lett. 2010, 584, 4453–4457. [Google Scholar] [CrossRef] [PubMed]
- Kim, H.-E.; Ahn, H.-C.; Lee, Y.-M.; Lee, E.-H.; Seo, Y.-J.; Kim, Y.-G.; Kim, K.K.; Choi, B.-S.; Lee, J.-H. The Zβ domain of human DAI binds to Z-DNA via a novel B-Z transition pathway. FEBS Lett. 2011, 585, 772–778. [Google Scholar] [CrossRef] [PubMed]
- Lee, A.-R.; Park, C.-J.; Cheong, H.-K.; Ryu, K.-S.; Park, J.-W.; Kwon, M.-Y.; Lee, J.; Kim, K.K.; Choi, B.-S.; Lee, J.-H. Solution structure of Z-DNA binding domain of PKR-like protein kinase from Carassius auratus and quantitative analyses of intermediate complex during B-Z transition. Nucleic Acids Res. 2016, 44, 2936–2948. [Google Scholar] [CrossRef] [PubMed]
- Lee, A.-R.; Kim, H.-E.; Lee, Y.-M.; Jeong, M.; Choi, K.-H.; Park, J.-W.; Choi, Y.-G.; Ahn, H.-C.; Choi, B.-S.; Lee, J.-H. NMR dynamics study of the Z-DNA binding domain of human ADAR1 bound to various DNA duplexes. Biochem. Biophys. Res. Commun. 2012, 428, 137–141. [Google Scholar] [CrossRef] [PubMed]
- Lee, A.-R.; Seo, Y.-J.; Choi, S.-R.; Ryu, K.-S.; Cheong, H.-K.; Lee, S.S.; Katahira, M.; Park, C.-J.; Lee, J.-H. NMR elucidation of reduced B-Z transition activity of PKZ protein kinase at high NaCl concentration. Biochem. Biophys. Res. Commun. 2017, 482, 335–340. [Google Scholar] [CrossRef] [PubMed]
| ZBP | DNA | α 1 | KBZ,1 | Kd,BP (μM) | Kd,ZP2 (μM) | Rerefences |
|---|---|---|---|---|---|---|
| hZαADAR1 | d(CGCGCG)2 | 1.15 × 10−2 | ~1 | <0.1 | <0.1 | [41] |
| hZαADAR1 | d(CACGTG)2 | 1.42 | 0.4 | 260 ± 87 | 180 ± 62 | [42] |
| hZαADAR1 | d(CGTACG)2 | 13.9 | 6.3 | 400 ± 144 | 29 ± 11 | [42] |
| yabZαE3L | d(CGCGCG)2 | 0.154 | 1.02 | n.d. 2 | n.d. 2 | [43] |
| ZBP | DNA | pH | [NaCl] | KBZ,1 | Kd,BP (μM) | Kd,ZP2 (μM) | Rerefences |
|---|---|---|---|---|---|---|---|
| caZαPKZ | d(TCGCGCG)2 | 6.0 | 10 mM | 0.87 ± 0.03 | 0.028 ± 0.017 | 0.345 ± 0.079 | [45] |
| caZαPKZ | d(TCGCGCG)2 | 6.0 | 100 mM | 0.19 ± 0.02 | 16.4 ± 0.8 | 8.76 ± 0.67 | [45] |
| caZαPKZ | d(TCGCGCG)2 | 6.0 | 250 mM | ~0.01 | 64.1 ± 8.3 | 9.57 ± 0.85 | [47] |
| caZαPKZ | d(TCGCGCG)2 | 8.0 | 10 mM | 1.18 ± 0.03 | 0.157 ± 0.021 | 0.129 ± 0.074 | [45] |
| caZαPKZ | d(TCGCGCG)2 | 8.0 | 100 mM | 0.18 ± 0.02 | 5.41 ± 0.66 | 2.41 ± 0.37 | [45] |
| caZαPKZ | d(CGCGCG)2 | 8.0 | 10 mM | 0.11 ± 0.05 | 5.18 ± 2.43 | 1.79 ± 0.95 | [45] |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Lee, A.-R.; Kim, N.-H.; Seo, Y.-J.; Choi, S.-R.; Lee, J.-H. Thermodynamic Model for B-Z Transition of DNA Induced by Z-DNA Binding Proteins. Molecules 2018, 23, 2748. https://doi.org/10.3390/molecules23112748
Lee A-R, Kim N-H, Seo Y-J, Choi S-R, Lee J-H. Thermodynamic Model for B-Z Transition of DNA Induced by Z-DNA Binding Proteins. Molecules. 2018; 23(11):2748. https://doi.org/10.3390/molecules23112748
Chicago/Turabian StyleLee, Ae-Ree, Na-Hyun Kim, Yeo-Jin Seo, Seo-Ree Choi, and Joon-Hwa Lee. 2018. "Thermodynamic Model for B-Z Transition of DNA Induced by Z-DNA Binding Proteins" Molecules 23, no. 11: 2748. https://doi.org/10.3390/molecules23112748
APA StyleLee, A.-R., Kim, N.-H., Seo, Y.-J., Choi, S.-R., & Lee, J.-H. (2018). Thermodynamic Model for B-Z Transition of DNA Induced by Z-DNA Binding Proteins. Molecules, 23(11), 2748. https://doi.org/10.3390/molecules23112748





