A Chemical Genetics Strategy That Identifies Small Molecules Which Induce the Triple Response in Arabidopsis
Abstract
1. Introduction
2. Results
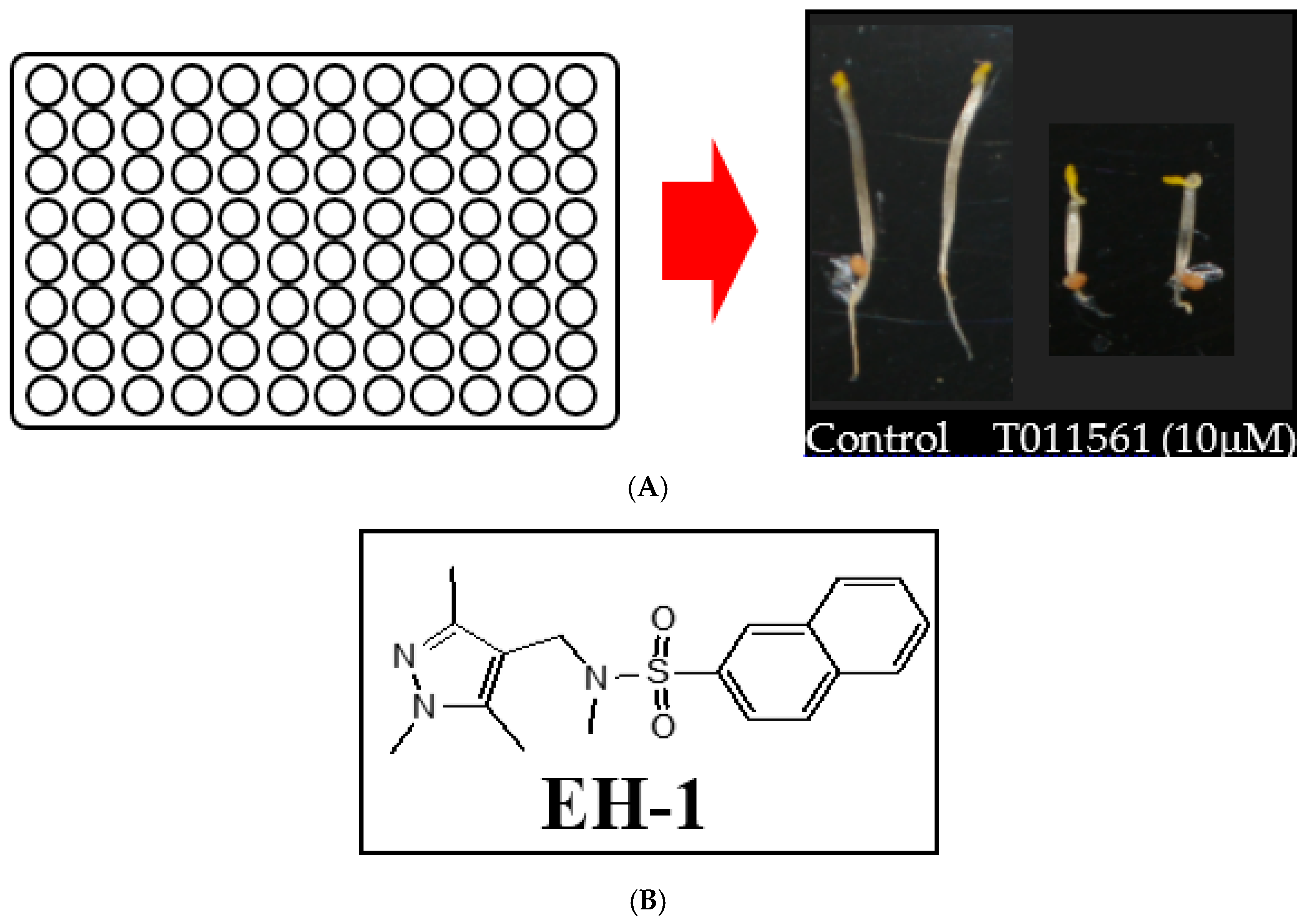
2.1. Screening of a Chemical Library for Small Molecules with Triple Response Induction Activity
2.2. Identification of EH-1
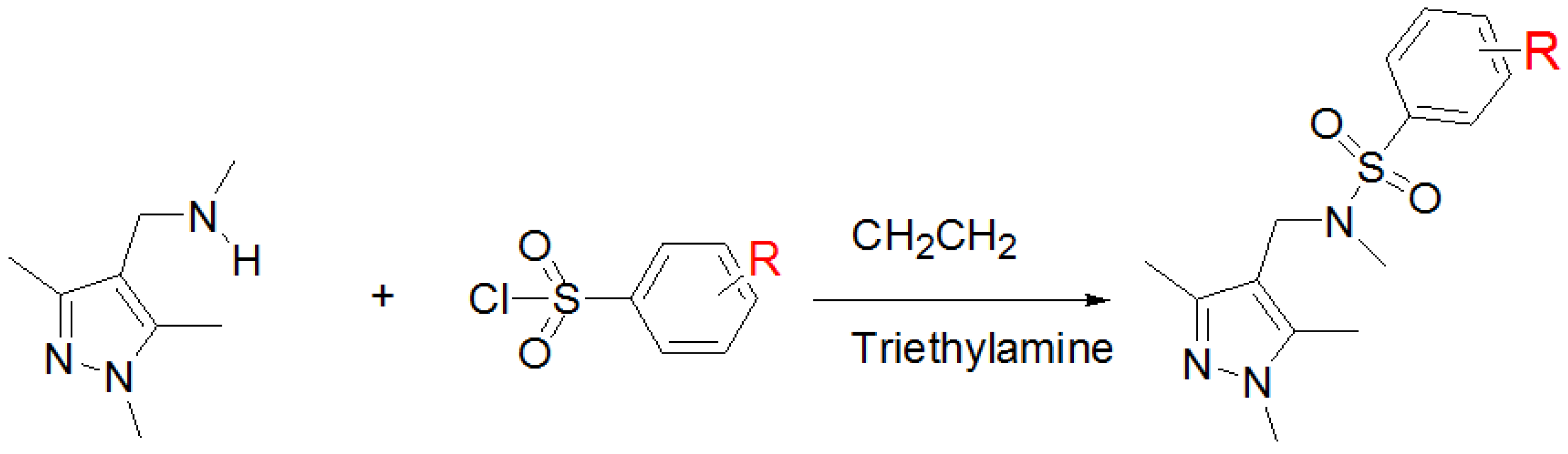
2.3. Chemical Synthesis of EH-1 Analogues
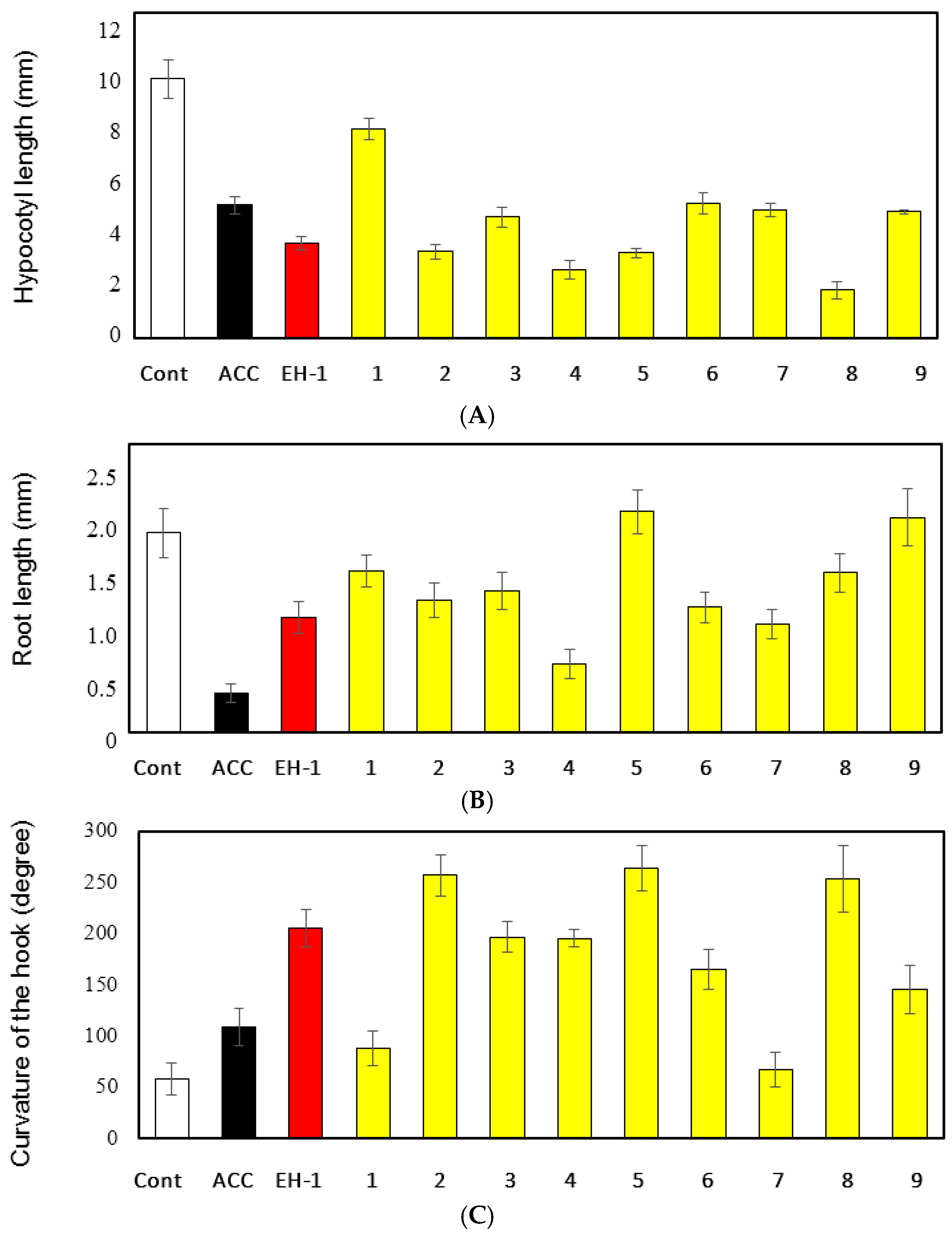
2.4. Triple Response Effect of EH-1 Analogues in Arabidopsis Seedlings
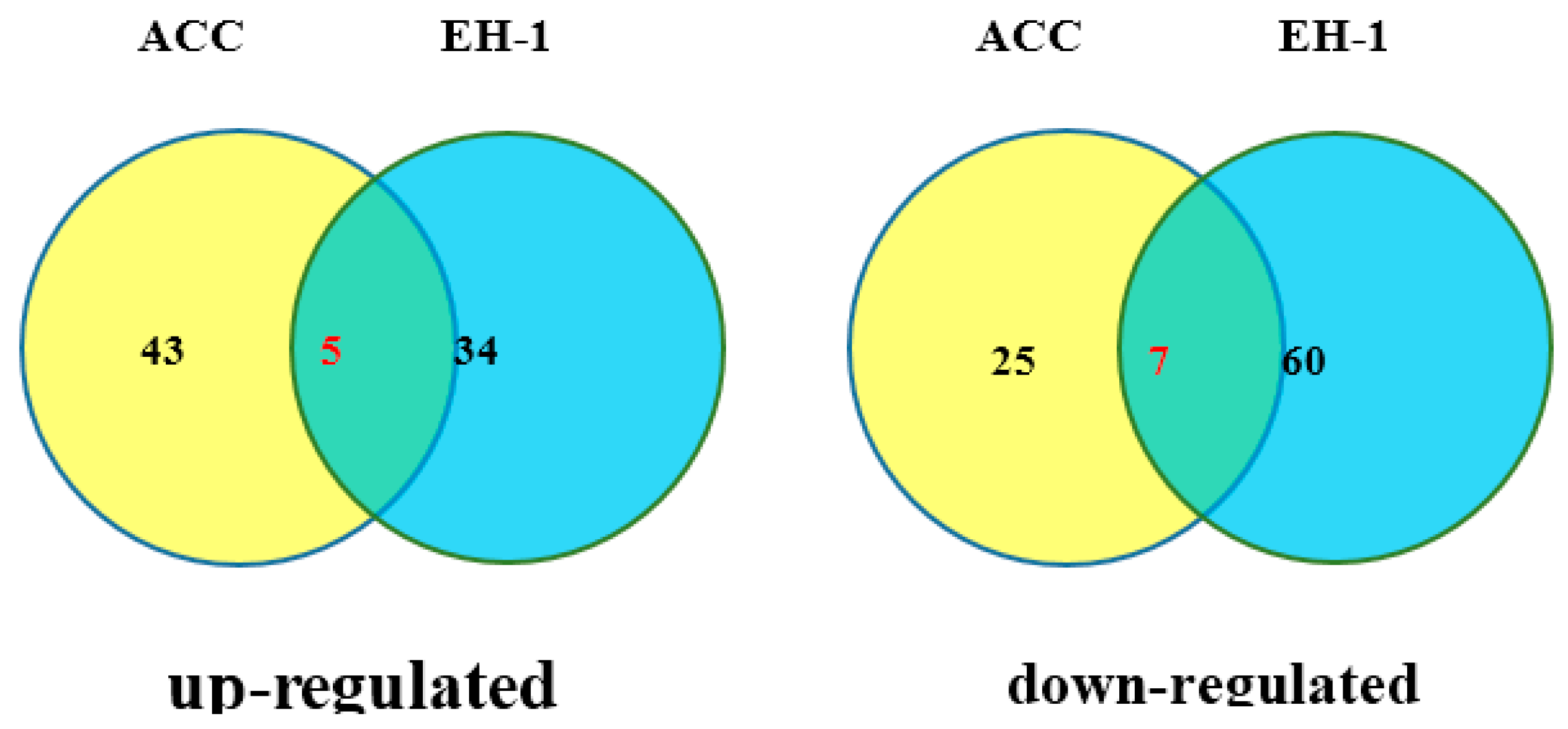
2.5. RNA-Seq Analysis of the Effect of EH-1 on Arabidopsis Seedlings
2.6. Quantitative RT-PCR Analysis of the Genes Induced by EH-1 and ACC
3. Discussion
4. Materials and Methods
4.1. General
4.2. Chemical Compound Library and Chemicals for Biological Studies
4.3. Chemical Synthesis: General Procedure of Preparation of N-Methyl-N-[(1,3,5-trimethyl-1H-pyrazol-4-yl)methyl]-2-naphthalenesulfonamide (EH-1)
4.4. Plant Materials and Growth Conditions
4.5. Isolation of Total-RNA
4.6. RNA-Seq Analysis
4.7. qRT-PCR Analysis
4.8. Statistical Analysis
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Waggoner, P.E.; Dimond, A.E. Non-specificity of the triple response for ethylene. Science 1954, 119, 123–124. [Google Scholar] [CrossRef] [PubMed]
- Bleecker, A.B.; Kende, H. Ethylene: A gaseous signal molecule in plants. Annu. Rev. Cell Dev. Biol. 2000, 16, 1–18. [Google Scholar] [CrossRef] [PubMed]
- Abeles, F.B.; Morgan, P.W.; Saltveit, M.E. Ethylene in Plant Biology; Academic Press: Cambridge, MA, USA, 1992; ISBN 9780120414512. [Google Scholar]
- Vriezen, W.H.; Achard, P.; Harberd, N.P.; Van Der Straeten, D. Ethylene-mediated enhancement of apical hook formation in etiolated Arabidopsis thaliana seedlings is gibberellin dependent. Plant J. 2004, 37, 505–516. [Google Scholar] [CrossRef] [PubMed]
- Van Doorn, W.G.; Kamdee, C. Flower opening and closure: An update. J. Exp. Bot. 2014, 65, 5749–5757. [Google Scholar] [CrossRef] [PubMed]
- Van de Poel, B.; Smet, D.; Van Der Straeten, D. Ethylene and hormonal cross talk in vegetative growth and development. Plant Physiol. 2015, 169, 61–72. [Google Scholar] [CrossRef] [PubMed]
- Liu, M.; Pirrello, J.; Chervin, C.; Roustan, J.P.; Bouzayen, M. Ethylene control of fruit ripening: Revisiting the complex network of transcriptional regulation. Plant Physiol. 2015, 169, 2380–2390. [Google Scholar] [CrossRef] [PubMed]
- Broekaert, W.F.; Delauré, S.L.; De Bolle, M.F.; Cammue, B.P. The role of ethylene in host-pathogen interactions. Annu. Rev. Phytopathol. 2006, 44, 393–416. [Google Scholar] [CrossRef] [PubMed]
- Sasidharan, R.; Voesenek, L.A. Ethylene-Mediated Acclimations to Flooding Stress. Plant Physiol. 2015, 169, 3–12. [Google Scholar] [CrossRef] [PubMed]
- Hopper, D.W.; Ghan, R.; Schlauch, K.A.; Cramer, G.R. Transcriptomic network analyses of leaf dehydration responses identify highly connected ABA and ethylene signaling hubs in three grapevine species differing in drought tolerance. BMC Plant Biol. 2016, 16, 118. [Google Scholar] [CrossRef] [PubMed]
- Yang, S.F. Ethylene evolution from 2-chloroethylphosphonic acid. Plant Physiol. 1969, 44, 1203–1204. [Google Scholar] [CrossRef] [PubMed]
- Ferrara, G.; Mazzeo, A.; Matarrese, A.M.; Pacucci, C.; Trani, A.; Fidelibus, M.W.; Gambacorta, G. Ethephon as a potential abscission agent for table grapes: Effects on pre-harvest abscission, fruit quality, and residue. Front. Plant Sci. 2016, 7, 620. [Google Scholar] [CrossRef] [PubMed]
- Webb, T.R. Current directions in the evolution of compound libraries. Curr. Opin. Drug Discov. Dev. 2005, 8, 303–308. [Google Scholar]
- Chen, I.J.; Lo, W.S.; Chuang, J.Y.; Cheuh, C.M.; Fan, Y.S.; Lin, L.C.; Wu, S.J.; Wang, L.C. A chemical genetics approach reveals a role of brassinolide and cellulose synthase in hypocotyl elongation of etiolated Arabidopsis seedlings. Plant Sci. 2013, 209, 46–57. [Google Scholar] [CrossRef] [PubMed]
- Hu, Y.; Callebert, P.; Vandemoortel, I.; Nguyen, L.; Audenaert, D.; Verschraegen, L.; Vandenbussche, F.; Van Der Straeten, D. TR-DB: An open-access database of compounds affecting the ethylene-induced triple response in Arabidopsis. Plant Physiol. Biochem. 2014, 75, 128–137. [Google Scholar] [CrossRef] [PubMed]
- Guzmán, P.; Ecker, J.R. Exploiting the triple response of Arabidopsis to identify ethylene-related mutants. Plant Cell 1990, 6, 513–523. [Google Scholar] [CrossRef] [PubMed]
- Cancel, J.D.; Larsen, P.B. Loss-of-function mutations in the ethylene receptor ETR1 cause enhanced sensitivity and exaggerated response to ethylene in Arabidopsis. Plant Physiol. 2002, 129, 1557–1567. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Gao, Z.; Wen, C.K.; Binder, B.M.; Chen, Y.F.; Chang, J.; Chiang, Y.H.; Kerris, R.J.; Chang, C.; Schaller, G.E. Heteromeric interactions among ethylene receptors mediate signaling in Arabidopsis. J. Biol. Chem. 2008, 283, 23801–23810. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Chang, C.; Kwok, S.F.; Bleecker, A.B.; Meyerowitz, E.M. Arabidopsis ethylene-response gene ETR1: Similarity of product to two-component regulators. Science 1993, 262, 539–544. [Google Scholar] [CrossRef] [PubMed]
- Hua, J.; Sakai, H.; Nourizadeh, S.; Chen, Q.G.; Bleecker, A.B.; Ecker, J.R.; Meyerowitz, E.M. EIN4 and ERS2 are members of the putative ethylene receptor gene family in Arabidopsis. Plant Cell 1998, 10, 1321–1332. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Sakai, H.; Hua, J.; Chen, Q.G.; Chang, C.; Medrano, L.J.; Bleecker, A.B.; Meyerowitz, E.M. ETR2 is an ETR1-like gene involved in ethylene signaling in Arabidopsis. Proc. Natl. Acad. Sci. USA 1998, 95, 5812–5817. [Google Scholar] [CrossRef] [PubMed] [PubMed Central]
- Kieber, J.J.; Rothenber, M.; Roman, G.; Feldmann, K.A.; Ecker, J.R. CTR1, a negative regulator of the ethylene response pathway in Arabidopsis, encodes a member of the Raf family of protein kinases. Cell 1993, 72, 427–441. [Google Scholar] [CrossRef]
- Chemical Library. Available online: http://www.ddi.u-tokyo.ac.jp/en/#5 (accessed on 15 December 2017).
- Oh, K.; Matsumoto, T.; Hoshi, T.; Yoshizawa, Y. Discovery of a new lead compound for plant growth retardants through compound library screening. J. Pestic. Sci. 2014, 39, 159–161. [Google Scholar] [CrossRef]
- Wang, Z.; Gerstein, M.; Snyder, M. RNA-Seq: A revolutionary tool for transcriptomics. Nat. Rev. Genet. 2009, 10, 57–63. [Google Scholar] [CrossRef] [PubMed]
- Ridge, I.; Osborne, D.J. Hydroxyproline and peroxidases in cell walls of Pisum sativum: Regulation by ethylene. J. Exp. Bot. 1970, 21, 843–856. [Google Scholar] [CrossRef]
- Zhong, G.Y.; Burns, J.K. Profiling ethylene-regulated gene expression in Arabidopsis thaliana by microarray analysis. Plant Mol. Biol. 2003, 53, 117–131. [Google Scholar] [CrossRef] [PubMed]
- Trapnell, C.; Roberts, A.; Goff, L.; Pertea, G.; Kim, D.; Kelley, D.R.; Pimentel, H.; Salzberg, S.L.; Rinn, J.L.; Pachter, L. Differential gene and transcript expression analysis of RNA-seq experiments with TopHat and Cufflinks. Nat. Protoc. 2012, 7, 562–578. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Samples of the compounds are available from the authors upon request. |
| Compound No. | Chemical Structure | Compound No. | Chemical Structure |
|---|---|---|---|
| EH-1 | 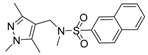 | 5 |  |
| 1 |  | 6 | 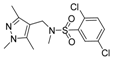 |
| 2 | 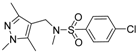 | 7 | 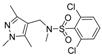 |
| 3 | 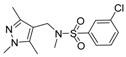 | 8 | 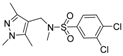 |
| 4 | 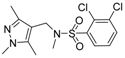 | 9 | 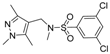 |
| A. Genes Up-Regulated by EH-1 | ||||
| Gene ID | Gene | Fold Change RNA-Seq (EH-1/Cont) | Fold Change qRT-PCR (EH-1/Cont) | Name of the Gene |
| AT1G72290 | AT1G72290 | 134.32 | 97.13 ± 49.08 | At1g72290; T9N14.19 |
| AT1G15520 | ABCG40 | 69.63 | 33.51 ± 15.26 | ABCG40 |
| AT4G12870 | AT4G12870 | 48.45 | 23.36 ± 15.19 | GILT reductase family protein |
| AT1G17180 | GSTU25 | 46.71 | 14.65 ± 7.47 | Glutathione S-transferase U25 |
| AT3G22640 | PAP85 | 41.20 | 39.69 ± 17.62 | AT3g22640/MWI23_1 |
| AT5G44120 | CRA1 | 36.33 | 47.93 ± 18.96 | 12S seed storage protein CRA1 |
| B. Genes Up-Regulated by ACC | ||||
| Gene ID | Gene | Fold Change RNA-Seq (ACC/Cont) | Fold Change qRT-PCR (ACC/Cont) | Name of the Gene |
| AT5G42530 | AT5G42530 | 103.72 | 78.96 ± 12.28 | At5g42530 |
| AT1G72290 | AT1G72290 | 85.83 | 73.50 ± 24.30 | At1g72290 |
| AT4G12480 | EARLI1 | 48.43 | 49.84 ± 13.56 | At1g72290 |
| AT2G41230 | ORS1 | 47.02 | 35.98 ± 2.72 | Lipid transfer protein EARLI 1 |
| AT2G05510 | AT2G05510 | 44.77 | 43.67 ± 11.78 | Protein ORGAN SIZE RELATED 1 |
| AT5G21120 | EIL2 | 32.40 | 3.99 ± 2.64 | At2g05510 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Oh, K.; Hoshi, T.; Tomio, S.; Ueda, K.; Hara, K. A Chemical Genetics Strategy That Identifies Small Molecules Which Induce the Triple Response in Arabidopsis. Molecules 2017, 22, 2270. https://doi.org/10.3390/molecules22122270
Oh K, Hoshi T, Tomio S, Ueda K, Hara K. A Chemical Genetics Strategy That Identifies Small Molecules Which Induce the Triple Response in Arabidopsis. Molecules. 2017; 22(12):2270. https://doi.org/10.3390/molecules22122270
Chicago/Turabian StyleOh, Keimei, Tomoki Hoshi, Sumiya Tomio, Kenji Ueda, and Kojiro Hara. 2017. "A Chemical Genetics Strategy That Identifies Small Molecules Which Induce the Triple Response in Arabidopsis" Molecules 22, no. 12: 2270. https://doi.org/10.3390/molecules22122270
APA StyleOh, K., Hoshi, T., Tomio, S., Ueda, K., & Hara, K. (2017). A Chemical Genetics Strategy That Identifies Small Molecules Which Induce the Triple Response in Arabidopsis. Molecules, 22(12), 2270. https://doi.org/10.3390/molecules22122270




