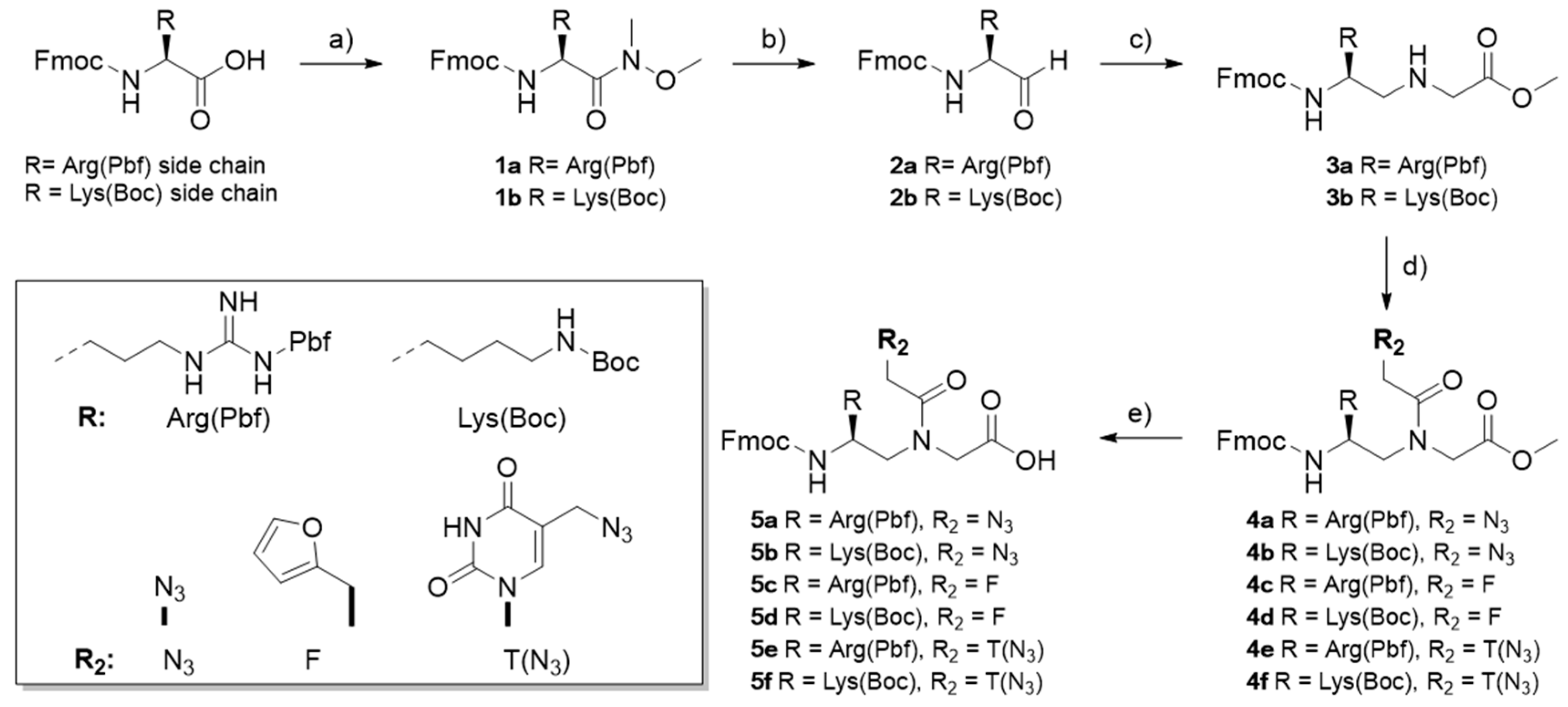
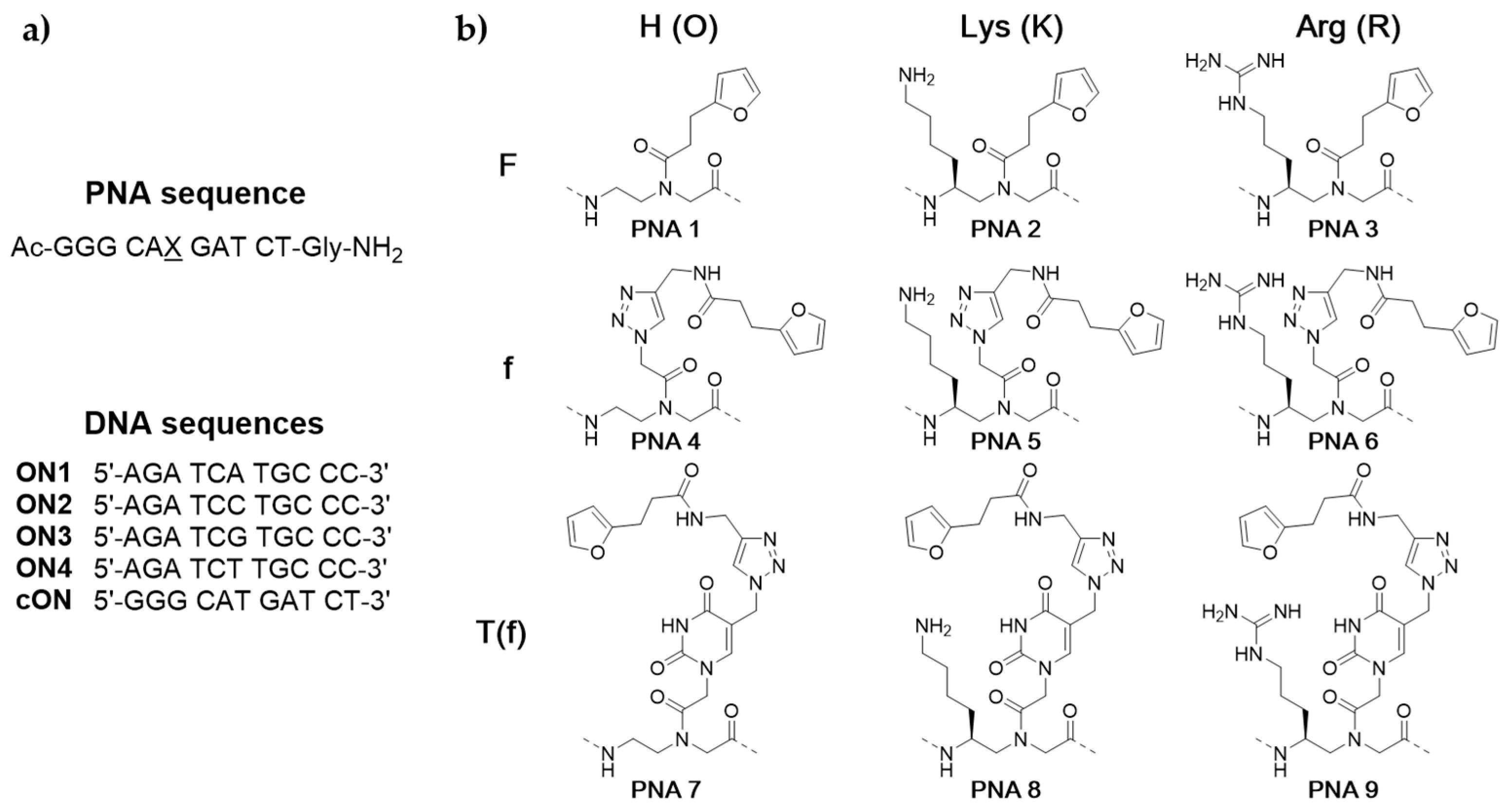
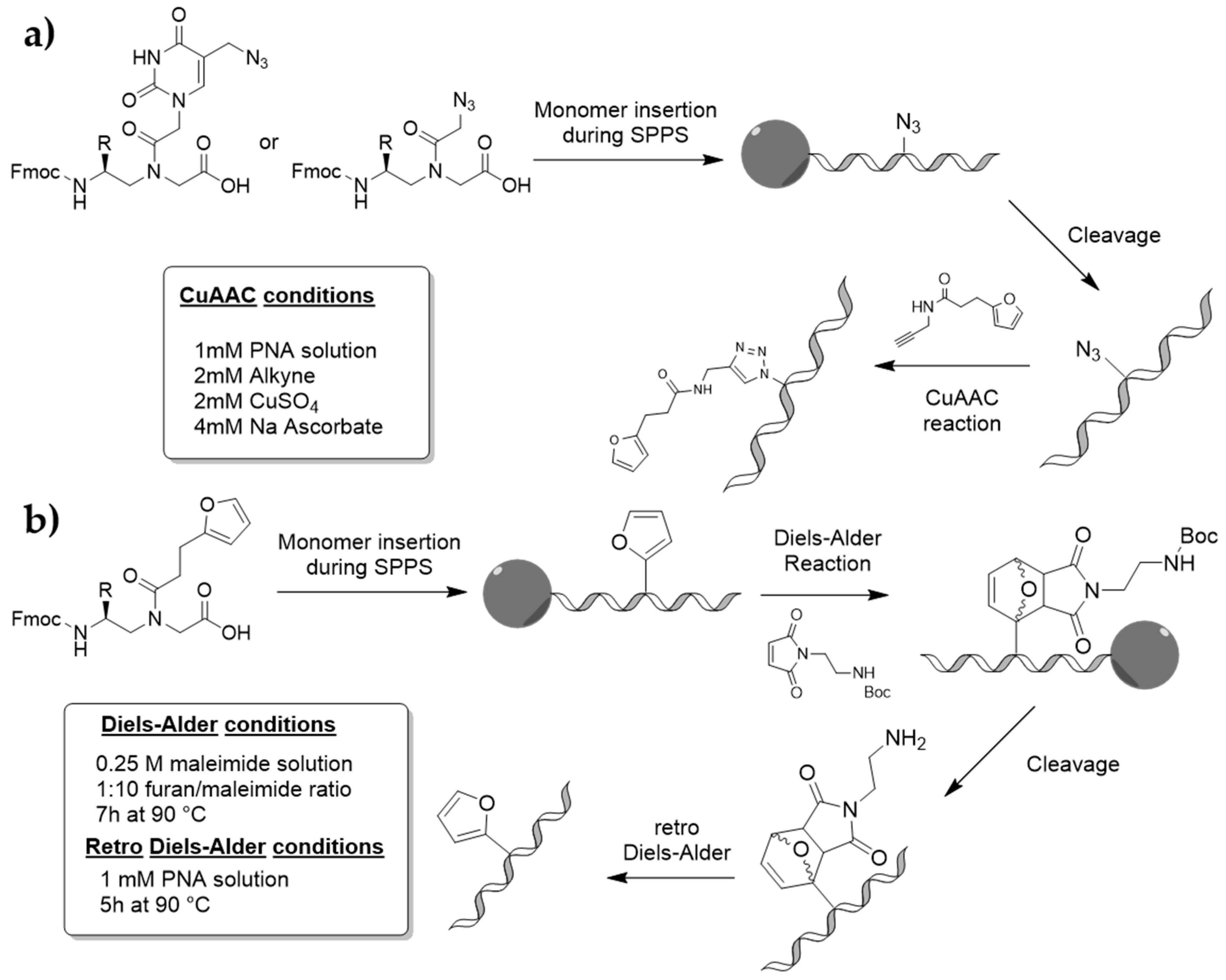
3.2. Monomer Synthesis
Synthesis of N-methyl-N-methoxy-Nα-Fmoc-Nω-Pbf-l-arginine-amide (1a). In a round-bottom flask equipped with a CaCl2 valve, Fmoc-l-Arg(Pbf)-OH (4.00 g, 6.2 mmol, 1 eq) was dissolved in 20 mL of dry DMF together with N,N,N′,N′-Tetramethyl-O-(1H-benzotriazol-1-yl)uronium hexafluorophosphate (HBTU, 3.52 g, 9.3 mmol, 1.5 eq). Subsequently, the solution was cooled to 0 °C with an ice bath, N,N-Diisopropylethylamine (DIPEA, 3.8 mL, 21.7 mmol, 3.5 eq) was added, and the mixture was allowed to react for 30 min. Next, N,O-dimethylhydroxylamine hydrochloride (1.21 g, 12.4 mmol, 2 eq) was added to the solution. The reagents were allowed to react for 10 min at 0 °C and 3 h 20 min at room temperature. Next, the reaction mixture was diluted with 200 mL EtOAc and washed with saturated KHSO4 (2 × 200 mL), saturated NaHCO3 (2 × 200 mL), and brine (200 mL). The organic phase was dried over Na2SO4 and the solvent was evaporated under reduced pressure. 1a was obtained as a pale yellow foam (6.189 mmol, 99.8% yield). TLC (AcOEt) Rf: 0.29, MP (°C): 106.2–108.3 °C; 1H-NMR (CDCl3, 400 MHz) δ (ppm): 7.77 (d, J = 7.6 Hz, 2H), 7.59 (d, J = 6.7 Hz, 2H), 7.41 (t, J = 7.6 Hz, 2H), 7.31 (t, J = 6.6 Hz, 2H), 6.09 (3H, br s), 5.84 (d, J = 8.4 Hz, 1H), 4.80–4.65 (1H, m), 4.52–4.31 (1H, m), 4.20 (1H, t, J = 6.9 Hz), 3.74 (3H, s), 3.21 (3H, s), 2.94 (2H, s), 2.60 (3H, s), 2.53 (3H, s), 2.10 (3H, s), 1.84–1.54 (6H, m), 1.46 (6H, s); 13C-NMR (CDCl3, 100 MHz) δ (ppm): 172.1 (31), 158.8, 156.8, 156.0, 143.7, 141.3, 138.4, 132.9, 132.4, 127.8, 127.1, 125.1, 124.6, 120.0, 117.5, 86.4, 67.2, 61.7, 50.2, 47.1, 43.2, 41.0, 32.1, 30.8, 28.6, 24.7, 19.3, 17.9, 12.5; MS (ESI, MeOH): m/z calcd. for C36H45N5O7S [M]: 691.30397, found: 692.3 [M + H]+, 1383.5 [2M + H]+; HRMS (LTQ Orbitrap, MeOH) m/z found: 692.31076 [C36H46N5O7S]+; FT-IR (ATR) ν (cm−1): 3322.4 (w), 2952.0 (m), 2936.1 (m), 2356.7 (s), 2334.6 (s), 1700.0 (s), 1652.4 (s), 1615.5 (s), 1543.3 (s), 1448.5 (s), 1247.7 (m), 1104.6 (s), 1088.6 (s), 993.7 (s), 783.4 (s), 757.3 (s).
Synthesis of N-methyl-N-methoxy-Nα-Fmoc-Nω-Boc-l-lysine-amide (1b). In a round-bottom flask equipped with a CaCl2 valve, Fmoc-l-Lys(Boc)-OH (3.0 g, 6.4 mmol, 1 eq) was dissolved in 20 mL of dry DMF together with HBTU (3.64 g, 9.6 mmol, 1.5 eq). Subsequently, the solution was cooled to 0 °C with an ice bath, DIPEA (3.9 mL, 22.4 mmol, 3.5 eq) was added, and the mixture was allowed to react for 30 min. Next, N,O-dimethylhydroxylamine hydrochloride (1.25 g, 12.8 mmol, 2 eq) was added to the solution. The reagents were allowed to react for 10 min at 0 °C and 3 h 20 min at room temperature. Next, the reaction mixture was diluted with 200 mL EtOAc and washed with saturated KHSO4 (2 × 200 mL), saturated NaHCO3 (2 × 200 mL), and brine (200 mL). The organic phase was dried over Na2SO4 and the solvent was evaporated under reduced pressure. 1b was obtained as a white foam in quantitative yield. TLC (AcOEt) Rf: 0.29, MP (°C): 50.2–50.5 °C; 1H-NMR (CDCl3, 400 MHz) δ (ppm): 7.78 (2H, d, J = 7.6 Hz), 7.63 (2H, t, J = 7.5 Hz), 7.42 (2H, t, J = 7.5 Hz), 7.34 (2H, tdd, J = 7.4 Hz, 2.9 Hz, 1.1 Hz), 5.55 (1H, d, J = 9.1 Hz), 4.81–4.72 (1H, br s), 4.65–4.53 (1H, s br), 4.39 (2H, d, J = 7.2 Hz), 4.24 (1H, t, J = 7.0 Hz), 3.80 (3H, s), 3.25 (3H, s), 3.14 (2H, t, J = 6.7), 1.86–1.70 (2H, m), 1.69–1.58 (1H, m), 1.56–1.49 (2H, m), 1.49–1.36 (10H, m); 13C-NMR (CDCl3, 100 MHz) δ (ppm): 172.6 (22), 156.2, 156.0, 144.0, 141.3, 127.7, 127.0, 125.1, 119.9, 79.1, 67.0, 61.6, 50.7, 47.2, 40.3, 32.5, 32.1, 29.6, 28.4, 22.5; MS (ESI, MeOH): m/z calcd. for C28H37N3O6 [M]: 511.26824, found: 512.2 [M + H]+, 1023.4 [2M + H]+ 534.2 [M + Na]+, 550.2 [M + K]+; HRMS (LTQ-Orbitrap, MeOH) m/z found: 534.25746 [C28H37N3O6Na]+; FT-IR (ATR) ν (cm−1): 3316.1 (w), 2972.7 (m), 2938.8 (m), 2863.8 (m), 2359.3 (s), 1700.0 (m), 1655.0 (s), 1513.1 (s), 1450.0 (s), 1390.9 (s), 1365.0 (s), 1246.2 (s), 1165.1 (s), 1040.4 (s), 990.9 (s), 860.9 (m), 758.7 (s), 738.9 (s).
Synthesis of Nα-Fmoc-Nω-Pbf-l-argininal (2a). In a three-neck flask connected with a nitrogen line through a bubble counter, 1a (1.0 g, 1.44 mmol, 1 eq) was dissolved in 20 mL dry THF and cooled down to 0 °C with an ice bath. LiAlH4 (60.33 mg, 1.58 mmol, 1.1 eq) was dissolved in 1.5 mL dry THF and slowly added to the solution. The reaction was allowed to react for 30 min under stirring. Next, the reaction was quenched by addition of 20 mL saturated KHSO4. The organic fraction was evaporated under reduced pressure, after which the resulting water phase was diluted with 200 mL EtOAc and 200 mL saturated KHSO4. The organic phase was washed with saturated KHSO4 (2 × 100 mL) and brine (2 × 100 mL). The organic phase was dried with Na2SO4 and the solvent removed under reduced pressure. 2a was obtained as a white foam (1.28 mmol, 88.6% yield). TLC (AcOEt) Rf: 0.56, MP (°C): 95.5–97.7 °C; 1H-NMR (CDCl3, 400 MHz) δ (ppm): 9.53 (1H, s), 7.77 (2H, d, J = 7.6 Hz), 7.60 (2H, d, J = 8.4 Hz), 7.41 (2H, t, J = 6.5 Hz), 7.32 (2H, m), 6.56 (3H, br s), 5.67 (1H, br s), 5.26 (1H, d, J = 9.9 Hz), 4.39 (2H, d, J = 7.2 Hz), 4.21 (1H, t, J = 7.2 Hz), 3.77–3.63 (1H, m), 3.19 (1H, m), 2.96 (2H, s), 2.57 (3H, s), 2.51 (3H, s), 2.11 (3H, s), 1.89–1.50 (6H, m), 1.47 (6H, s); 13C-NMR (CDCl3, 100 MHz) δ (ppm): 158.0, 155.9, 155.3, 144.4, 144.3, 141.2, 138.0, 134.3, 132.0, 128.1, 127.5, 125.8, 124.9, 120.6, 116.9, 86.8, 66.1, 55.4, 51.7, 47.1, 42.9, 28.8, 24.3, 19.4, 18.1, 12.7; MS (ESI, MeOH): m/z calcd. for C34H40N4O6S [M]: 632.26686, found: 633.3[M + H]+, 1265.6 [2M + H]+; HRMS (LTQ-Orbitrap, MeOH) m/z found: 633. [C34H41N4O6S]+; FT-IR (ATR) ν (cm−1): 3414.2 (m), 3341.4 (m), 2967.8 (m), 2936.1 (m), 1704.6 (s), 1684.8 (s), 1628.8 (m), 1576.9 (s), 1510.0 (s), 1448.5 (s), 1241.0 (m), 1086.9 (m), 990.2 (s), 869.7 (s), 852.4 (s), 818.9 (s), 758.8 (s), 738.1 (s).
Synthesis of Nα-Fmoc-Nω-Boc-l-lysinal (2b). In a three-neck flask connected with a nitrogen line through a bubble counter, 1b (1.44 g, 2.8 mmol, 1 eq) was dissolved in 20 mL dry THF and cooled down to 0 °C with an ice bath. LiAlH4 (117.7 mg, 3.1 mmol, 1.1 eq) was dissolved in 1.5 mL dry THF and slowly added to the solution. The reaction was allowed to react for 32 min under stirring. Next, the reaction was quenched by addition of 20 mL saturated KHSO4. The organic fraction was evaporated under reduced pressure, after which the resulting water phase was diluted with 200 mL EtOAc and 200 mL saturated KHSO4. The organic phase was washed with saturated KHSO4 (2 × 100 mL), and brine (2 × 100 mL). The organic phase was dried with Na2SO4 and the solvent removed under reduced pressure. 2b was obtained as a white foam (1.92 mmol, 68.7% yield). TLC (AcOEt) Rf: 0.56; 1H-NMR (CDCl3, 400 MHz) δ (ppm): 9.61 (1H, s), 7.80 (2H, d, J = 7.5Hz), 7.63 (2H, d, J = 7.4 Hz), 7.43 (2H, t, J = 7.3 Hz), 7.38–7.31 (2H, m), 4.80–4.55 (2H, m), 4.44 (2H, d, J = 7.0 Hz), 4.26 (1H, t, J = 6.9 Hz), 3.43–3.36 (2H, m), 3.15 (1H, br s), 1.91–1.37 (15H, m); 13C-NMR (CDCl3, 100 MHz) δ (ppm): 197.0, 156.5, 143.8, 141.3, 127.7, 127.1, 125.0, 120.0, 80.4, 66.9, 57.7, 49.7, 47.1, 42.8, 28.7, 28.4, 24.00; MS (ESI, MeOH): m/z calcd. for C26H32N2O5 [M]: 452.23112, found: 453.2 [M + H]+; 475.2 [M + Na]+, 491.1 [M + K]+; HRMS (LTQ-Orbitrap, MeOH) m/z found: 507.24656 [C27H36N2O6Na]+.
Synthesis of Nα-Fmoc-Ψ(l-arginine-(Nω-Pbf))-glycine methyl ester (3a). In a 50 mL round-bottom flask, 2a (200 mg, 0.316 mmol, 1 eq) was dissolved in 10 mL dry MeOH, together with glycine methyl ester hydrochloride (79.4 mg, 0.632 mmol, 2 eq) and reacted for 15 min at 0 °C and 15 min at room temperature. Next, NaBH3CN (41.7 mg, 0.66 mmol, 2.1 eq) and acetic acid (36.9 μL, 0.66 mmol, 2.1 eq) were added. The pH of the solution was checked to be around 4.5. The reaction mixture was stirred overnight at room temperature. Next, the organic solvent was removed, and the crude was taken up with 20 mL EtOAc and washed with saturated NaHCO3 (2 × 20 mL) and brine (2 × 20 mL). The organic phase was dried over Na2SO4 and evaporated under reduced pressure. The product was purified using flash chromatography (DCM:MeOH 96:4 to DCM:MeOH 94:6), which yielded 3a as a white foam (0.184 mmol, 58.3% yield). TLC (AcOEt) Rf: 0.28, MP (°C): 69.5–71.4 °C; 1H-NMR (CDCl3, 400 MHz) δ (ppm): 7.76 (2H, d, J = 7.0 Hz), 7.60 (2H, d, J = 8.0 Hz), 7.40 (2H, t, J = 7.2 Hz), 7.31 (2H, m), 6.13 (3H, s), 5.51 (1H, d, J = 7.3 Hz), 4.41 (2H, d, J = 4.4 Hz), 4.20 (1H, t, J = 6.7 Hz), 3.73 (3H, s), 3.70–3.62 (1H, m), 3.46 (1H, d, J = 17.6 Hz), 3.38 (1H, d, J = 16.7 Hz), 3.33–3.11 (2H, m), 2.94 (2H, s), 2.73–2.62 (2H, m), 2.60 (3H, s), 2.53 (3H, s), 2.09 (3H, s), 1.86 (1H, s), 1.70–1.47 (4H, m), 1.45 (6H, s); MS (ESI, MeOH): m/z calcd. for C37H47N5O7S [M]: 705.31962, found: 706.3 [M + H]+, 1411.7 [2M + H]+; HRMS (LTQ-Orbitrap, MeOH) m/z found: 706.3258 [C37H48N5O7S]+; FT-IR (ATR) ν (cm−1): 3325.6 (m), 2936.1 (m), 2917.2 (m), 1718.4 (m), 1700.8 (m), 1622.6 (m), 1560.7 (s), 1546.8 (s), 1448.8 (s), 1242.3 (m), 1153.9 (s), 1089.3 (s), 1033.6 (s), 993.1 (s), 851.7 (s), 807.7 (s), 781.3 (s), 760.1 (s), 739.4 (s).
Synthesis of Nα-Fmoc-Ψ(l-lysine-(Nω-Boc))-glycine methyl ester (3b). In a 50 mL round-bottom flask, 2b (861 mg, 1.90 mmol, 1 eq) was dissolved in 10 mL dry MeOH, together with glycine methyl ester hydrochloride (477.8 mg, 3.81 mmol, 2 eq) and reacted for 15 min at 0 °C and 15 min at room temperature. Next, NaBH3CN (251.1 mg, 4.00 mmol, 2.1 eq) and acetic acid (228 μL, 4.00 mmol, 2.1 eq) were added. The pH of the solution was checked to be around 4.5. The reaction mixture was stirred overnight at room temperature. Next, the organic solvent was removed, and the crude was taken up with 20 mL EtOAc and washed with saturated NaHCO3 (2 × 20 mL) and brine (2 × 20 mL). The organic phase was dried over Na2SO4 and evaporated under reduced pressure. The crude was purified using flash chromatography (DCM to DCM:MeOH 96:4), which yielded 3b as a white foam (1.66 mmol, 87.2% yield). TLC (AcOEt) Rf: 0.26; 1H-NMR (CDCl3, 400 MHz) δ (ppm): 7.79 (2H, d, J = 7.5 Hz), 7.63 (2H, d, J = 7.1 Hz), 7.42 (2H, t, J = 7.4 Hz), 7.34 (2H, t, J = 7.4 Hz), 5.13 (1H, d, J = 7.6 Hz), 4.61 (1H, br s), 4.43 (2H, d, J = 8.8 Hz), 4.24 (1H, t, J = 6.7 Hz), 3.78–3.63 (4H, m),3.49 (1H, d, J = 15.3 Hz), 3.40 (1H, d, J = 16.8 Hz), 3.19–3.03 (2H, m), 2.80–2.64 (2H, m), 1.86 (1H, br s), 1.59–1.30 (15H, m); 13C-NMR (CDCl3, 100 MHz) δ (ppm): 172.8, 156.4, 156.1, 144.0, 141.3, 127.7, 127.0, 125.1, 119.9, 79.1, 66.5, 53.4, 52.8, 51.8, 50.7, 47.4, 40.2, 32.6, 29.8, 28.4, 23.0; MS (ESI, MeOH): m/z calcd. for C29H39N3O6 [M]: 525.28389, found: 526.2 [M + H]+, 548.3 [M + Na]+, 1051.5 [2M + H]+; HRMS (LTQ-Orbitrap, MeOH) m/z found: 526.29116 [C29H40N3O6]+.
Synthesis of Fmoc-PNA(5L-Arg(Pbf))-N3-OMe (4a). In a round-bottom flask, 2-azidoacetic acid (63.6 μL, 0.85 mmol, 2 eq) and 3-hydroxy-1,2,3-benzotriazin-4 (3 h)-one (DhBtOH) (138.7 mg, 0.85 mmol, 2 eq) were dissolved in 5 mL of dry DMF and cooled to 0 °C. Then, 1-(3-dimethylaminopropyl)-3-ethylcarbodiimide hydrochloride (EDC·HCl) (162.9 mg, 0.85 mmol, 2 eq) and DIPEA (148.2 μL, 0.85 mmol, 2 eq) were added and stirred for 10 min at 0 °C, followed by 20 min at room temperature. Next, the reaction was cooled again to 0 °C and 3a was added (300.5 mg, 0.425 mmol, 1 eq). The reaction was stirred for 5 min at 0 °C and for 5 h at room temperature. The reaction mixture was then diluted with EtOAc (100 mL) and washed with saturated KHSO4 (2 × 100 mL), saturated NaHCO3 (2 × 100 mL), and brine (100 mL). The organic phase was dried over Na2SO4 and the solvent was evaporated under reduced pressure. The crude was purified using flash chromatography (AcOEt:Hexane 8:2 to AcOEt) and yielded 4a as a white foam (0.272 mmol, 63.9% yield). TLC (AcOEt) Rf: 0.28, MP (°C): 85.1–85.3; 1H-NMR (CDCl3, 400 MHz, main rotamer) δ (ppm): 7.77 (2H, d, J = 7.6 Hz), 7.59 (2H, d, J = 7.3 Hz), 7.40 (2H, t, J = 7.5 Hz), 7.31 (2H, m), 6.28 (3H, s), 5.62 (1H, d, J = 7.6 Hz), 4.43–4.30 (2H, m), 4.22–4.13 (1H, m), 4.06 (2H, s), 3.86 (2H, s), 3.82–3.77 (1H, m), 3.74 (3H, s), 3.44–3.12 (4H, m), 2.94 (2H, s), 2.60 (3H, s), 2.53 (3H, s), 2.10 (3H, s), 1.75–1.50 (4H, m), 1.45 (6H, s); 13C-NMR (CDCl3, 100 MHz, main rotamer) δ (ppm): 169.2, 168.6, 159.2, 157.0, 155.9, 143.8, 141.3, 138.7, 132.7, 127.8, 127.1, 125.1, 124.9, 120.0, 117.8, 114.7, 86.6, 66.8, 52.9, 52.4, 51.5, 50.4, 49.8, 47.2, 43.2, 41.0, 28.6, 25.2, 25.0, 19.3, 17.5, 12.5 (21); MS (ESI, MeOH): m/z calcd. for C39H48N8O8S [M]: 788.33158, found: 789.2 [M + H]+; HRMS (LTQ-Orbitrap, MeOH) m/z found: 789.33886 [C39H49N8O8S]+; FT-IR (ATR) ν (cm−1): 3449.1 (m), 3426.9 (m), 3340.3 (w), 2939.3 (w), 2104.1 (s), 1745.9 (s), 1714.5 (s), 1653.8 (s), 1616.2 (s), 1547.4 (s), 1450.1 (s), 1369.3 (s), 1241.1 (m), 1217.8 (m), 1154.4 (s), 1103.5 (s), 1088.7 (s), 992.2 (s), 808.4 (m), 758.5 (s), 740.0 (s).
Synthesis of Fmoc-PNA(5L-Lys(Boc))-N3-OMe (4b). In a round-bottomed flask, 2-azidoacetic acid (72.6 μL, 0.97 mmol, 2 eq) and DhBtOH (158.2 mg, 0.97 mmol, 2 eq) were dissolved in 5 mL of dry DMF and cooled to 0 °C. Then, EDC·HCl (185.9 mg, 0.97 mmol, 2 eq) and DIPEA (169.2 μL, 0.97 mmol, 2 eq) were added and stirred for 10 min at 0 °C, followed by 20 min at room temperature. Next, the reaction was cooled again to 0 °C and 3b was added (255.0 mg, 0.48 mmol, 1 eq). The reaction was stirred for 5 min at 0 °C and for 3 h at room temperature. The reaction mixture was then diluted with EtOAc (100 mL) and washed with saturated KHSO4 (2 × 100 mL), saturated NaHCO3 (2 × 100 mL), and brine (100 mL). The organic phase was dried over Na2SO4 and the solvent was evaporated under reduced pressure. The crude was purified using flash chromatography (AcOEt:Hexane 6:4) and yielded 4b as a pale yellow foam (0.360 mmol, 74.2% yield). TLC (AcOEt) Rf: 0.28, MP (°C): 51.3–51.6; 1H-NMR (CDCl3, 400 MHz, main rotamer) δ (ppm): 7.79 (2H, d, J = 7.4 Hz), 7.61 (2H, d, J = 7.2 Hz), 7.43 (2H, t, J = 7.3 Hz), 7.35 (2H, t, J = 7.3 Hz), 5.05 (1H, d, J = 5.7 Hz), 4.60 (1H, br s), 4.52–4.39 (2H, m), 4.22 (1H, d, J = 6.8 Hz), 4.01 (2H, s), 3.82 (2H, s), 3.78 (3H, s), 3.72–3.62 (1H, m), 3.29–3.18 (2H, m), 3.17–3.07 (2H, m), 1.58–1.37 (15H, m); 13C-NMR (CDCl3, 100 MHz, main rotamer) δ (ppm): 169.1, 168.4, 156.5, 156.1, 143.8, 141.3, 127.7, 127.1, 125.0, 120.0, 79.1, 66.6, 52.8, 52.4, 50.5, 49.9, 49.2, 47.3, 40.1, 32.4, 29.8, 28.4, 22.8; MS (ESI, MeOH): m/z calcd. for C31H40N6O7 [M]: 608.29585, found: 648.39 [M + K]+; HRMS (LTQ-Orbitrap, MeOH) m/z found: 631.28507 [C31H40N6O7Na]+; FT-IR (ATR) ν (cm−1): 3350.5 (w), 2941.5 (w), 2103.9 (s), 1747.8 (s), 1699.8 (m), 1662.1 (m), 1521.4 (s), 1450.4 (s), 1365.3 (s), 1247.5 (s), 1210.8 (s), 1168.4 (s), 1079.8 (s), 1022.9 (m), 759.2 (s), 739.4 (s).
Synthesis of Fmoc-PNA(5L-Arg(Pbf))-F-OMe (4c). In a round-bottomed flask, 3-(2-furyl)propionic acid (119.1 mg, 0.85 mmol, 2 eq) and DhBtOH (138.6 mg, 0.85 mmol, 2 eq) were dissolved in 5 mL of dry DMF and cooled to 0 °C. Then, EDC·HCl (162.9 mg, 0.85 mmol, 2 eq) and DIPEA (148.2 μL, 0.85 mmol, 2 eq) were added and stirred for 10 min at 0 °C, followed by 20 min at room temperature. Next, the reaction was cooled again to 0 °C and 3a was added (300.0 mg, 0.43 mmol, 1 eq). The reaction was stirred for 5 min at 0 °C and for 3 h at room temperature. The reaction mixture was then diluted with EtOAc (100 mL) and washed with saturated KHSO4 (2 × 100 mL), saturated NaHCO3 (2 × 100 mL), and brine (100 mL). The organic phase was dried over Na2SO4 and the solvent was evaporated under reduced pressure. The crude was purified using flash chromatography (AcOEt:Hexane 8:2 to AcOEt) and yielded 4c as a white solid (0.347 mmol, 81.6% yield). TLC (AcOEt) Rf: 0.28, MP (°C): 75.1–75.3; 1H-NMR (CDCl3, 400 MHz, major rotamer) δ (ppm): 7.76 (2H, d, J = 7.6 Hz), 7.59 (2H, d, J = 7.5 Hz), 7.39 (2H, t, J = 7.5 Hz), 7.32–7.21 (3H, m), 6.33–6.23 (4H, m), 6.02–5.96 (1H, m), 5.66 (1H, d, J = 7.4 Hz), 4.36 (2H, d, J = 7.3 Hz), 4.20–4.10 (1H, m), 4.06 (2H, s), 3.73 (3H, s), 3.68–3.58 (1H, m), 3.35–3.24 (2H, m), 3.21–3.09 (2H, m), 3.00–2.89 (4H, m), 2.60 (3H, s), 2.57–2.50 (5H, m, 36), 2.10 (3H, s), 1.70–1.48 (4H, m), 1.45 (6H, s); 13C-NMR (CDCl3, 100 MHz, major rotamer) δ (ppm): 173.9, 169.6, 159.0, 157.0, 155.9, 154.3, 143.8, 141.3, 141.1, 138.6, 132.6, 127.7, 127.1, 125.1, 124.7, 120.0, 117.6, 110.3, 105.4, 86.5, 66.7, 52.7, 52.2, 51.1, 50.4, 47.2, 43.2, 41.0, 31.4, 28.6, 25.6, 25.1, 23.5, 19.3, 18.0, 12.5; MS (ESI, MeOH): m/z calcd. for C44H53N5O9S [M]: 827.35640, found: 828.2 [M + H]+, 850.21 [M + Na]+; HRMS (LTQ-Orbitrap, MeOH) m/z found: 828.36368 [C44H54N5O9S]+; FT-IR (ATR) ν (cm−1): 3341.4 (m), 2933.0 (m), 1718.2 (m), 1623.7 (m), 1543.8 (m), 1449.6 (m), 1241.3 (m), 1212.4 (m), 1151.5 (m), 1087.6 (s), 1006.0 (s), 810.7 (s), 736.6 (s).
Synthesis of Fmoc-PNA(5L-Lys(Boc))-F-OMe (4d). In a round-bottom flask, 3-(2-furyl)propionic acid (135.9 mg, 0.97 mmol, 2 eq) and DhBtOH (158.2 mg, 0.97 mmol, 2 eq) were dissolved in 5 mL of dry DMF and cooled to 0 °C. Then, EDC·HCl (185.9 mg, 0.97 mmol, 2 eq) and DIPEA (169.2 μL, 0.97 mmol, 2 eq) were added and stirred for 10 min at 0 °C, followed by 20 min at room temperature. Next, the reaction was cooled again to 0 °C and 3b was added (255.0 mg, 0.49 mmol, 1 eq). The reaction was stirred for 5 min at 0 °C and for 3 h at room temperature. The reaction mixture was then diluted with EtOAc (100 mL) and washed with saturated KHSO4 (2 × 100 mL), saturated NaHCO3 (2 × 100 mL), and brine (100 mL). The organic phase was dried over Na2SO4 and the solvent was evaporated under reduced pressure. The crude was purified using flash chromatography (AcOEt:Hexane 8:2) and yielded 4d as a white solid (0.364 mmol, 75.0% yield). TLC (AcOEt) Rf: 0.39, MP (°C): 47.7–48.0; 1H-NMR (CDCl3, 400 MHz, major rotamer) δ (ppm): 7.78 (2H, d, J = 7.6 Hz), 7.61 (2H, d, J = 8.6 Hz), 7.42 (2H, td, J = 7.4 Hz, 2.8 Hz), 7.33 (2H, t, J = 7.4 Hz), 7.28 (1H), 6.26 (1H, dd, J = 4.4 Hz, 2.5 Hz), 6.00 (1H, dd, J = 12.1 Hz, 3.0 Hz), 5.22 (1H, d, J = 7.8 Hz), 4.60 (1H, br s), 4.41–4.34 (2H, m), 4.23–4.17 (1H, m), 4.07–3.95 (2H, m), 3.81–3.60 (4H, m), 3.53–3.30 (2H, m), 3.18–3.06 (2H, m), 3.02–2.92 (2H, m), 2.54 (2H, t, J = 7.6 Hz), 1.62–1.40 (15H, m); 13C-NMR (CDCl3, 100 MHz, major rotamer) δ (ppm): 173.4, 169.6, 156.6, 156.1, 154.6, 143.9, 141.3, 141.1, 127.7, 127.1, 125.3, 120.0, 110.4, 105.4, 79.1, 66.6, 53.0, 52.6, 52.2, 49.7, 47.3, 39.8, 32.5, 31.3, 29.8, 28.4, 23.6, 22.8; MS (ESI, MeOH): m/z calcd. for C36H45N3O8 [M]: 647.32067, found: 648.3 [M + H]+; HRMS (LTQ-Orbitrap, MeOH) m/z found: 670.30989 [C36H45N3O8Na]+; FT-IR (ATR) ν (cm−1): 3312.4 (w), 2977.3 (m), 2934.5 (m), 2857.0 (s), 1748.5 (s), 1701.2 (m), 1641.5 (m), 1705.5 (m), 1450.0 (m), 1365.1 (s), 1246.4 (s), 1210.6 (s), 1169.8 (s), 1076.9 (s), 1012.8 (s), 884.4 (s), 862.9 (m), 738.5 (s).
Synthesis of Fmoc-PNA(5L-Arg(Pbf))-T(N3)-OMe (4e). In a round-bottom flask, 2-((5-azidomethyl)uracil-1-yl)acetic acid (134.0 mg, 0.59 mmol, 1.1 eq) and DhBtOH (380 mg, 0.59 mmol, 1.1 eq) were dissolved in 2 mL of dry DMF and cooled to 0 °C. Then, EDC·HCl (72.1 mg, 0.59 mmol, 1.1 eq) and DIPEA (103.1 μL, 0.96 mmol, 1.1 eq) were added and stirred for 10 min at 0 °C, followed by 20 min at room temperature. Next, the reaction was cooled again to 0 °C and 3a was added (380 mg, 0.48 mmol, 1 eq). The reaction was stirred for 5 min at 0 °C and overnight at room temperature. The reaction mixture was then diluted with EtOAc (100 mL) and washed with saturated KHSO4 (2 × 100 mL), saturated NaHCO3 (2 × 100 mL), and brine (100 mL). The organic phase was dried over Na2SO4 and the solvent was evaporated under reduced pressure. The crude was purified using flash chromatography (AcOEt) and 4e was obtained as a yellowish solid (0.28 mmol, 74.7% yield). TLC (AcOEt) Rf: 0.18, 1H-NMR (400 MHz, DMSO-d6, major rotamer) δ 11.58 (1H, s), 7.89 (2H, d, J = 7.5 Hz), 7.67 (2H, t, J = 7.5 Hz), 7.60 (1H, d, J = 10.2 Hz), 7.41 (2H, t, J = 7.3 Hz), 7.37–7.24 (3H, m), 6.66 (1H, br s), 6.40 (1H, br s), 4.83 (1H, d, J = 16.6 Hz), 4.68 (1H, d, J = 16.7 Hz), 4.40–4.20 (3H, m), 4.03 (2H, s), 3.70 (2H, s), 3.61 (2H, s), 3.39–3.35 (1H, m), 3.04 (2H, s), 2.94 (2H, s), 2.48 (3H, s), 2.43 (3H, s), 2.00 (3H, s), 1.55–1.28 (10H, m); 13C-NMR (100 MHz, DMSO-d6) δ 169.7, 167.9, 163.9, 157.9, 156.5, 156.4, 151.1, 146.0, 144.5, 144.3, 144.2, 141.2, 138.8, 137.7, 134.7, 132.4, 131.9, 128.1, 127.5, 125.6, 124.8, 120.6, 117.7, 116.7, 107.8, 86.8, 65.7, 52.7, 52.2, 51.8, 50.0, 48.6, 48.6, 48.4, 47.3, 42.9, 28.7, 26.2, 19.4, 18.1, 12.7; MS (ESI, MeOH): m/z calcd. for C44H52N10O10S [M]: 912.35886, found: 913.7 [M + H]+, 935.7 [M + Na]+, 951.7 [M + K]+.
Synthesis of Fmoc-PNA(5L-Lys(Boc))-T(N3)-OMe (4f). In a round-bottom flask, 2-((5-azidomethyl)uracil-1-yl)acetic acid (215.0 mg, 0.96 mmol, 2 eq) and DhBtOH (155.8 mg, 0.96 mmol, 2 eq) were dissolved in 5 mL of dry DMF and cooled to 0 °C. Then, EDC·HCl (183.1 mg, 0.96 mmol, 2 eq) and DIPEA (166.6 μL, 0.96 mmol, 2 eq) were added and stirred for 10 min at 0 °C, followed by 20 min at room temperature. Next, the reaction was cooled again to 0 °C and 3b was added (251.0 mg, 0.48 mmol, 1 eq). The reaction was stirred for 5 min at 0 °C and for 3 h at room temperature. The reaction mixture was then diluted with EtOAc (100 mL) and washed with saturated KHSO4 (2 × 100 mL), saturated NaHCO3 (2 × 100 mL), and brine (100 mL). The organic phase was dried over Na2SO4 and the solvent was evaporated under reduced pressure. The crude was purified using flash chromatography (AcOEt:Hexane 8:2 to AcOEt) and yielded 4f as a pale yellow foam (0.339 mmol, 71.0% yield). TLC (AcOEt) Rf: 0.41, MP (°C): 74.1–74.4; 1H-NMR (CDCl3, 400 MHz, major rotamer) δ (ppm): 9.04 (1H, s), 7.78 (2H, d, J = 7.4 Hz), 7.62 (2H, d, J = 7.5 Hz), 7.41 (2H, t, J = 7.4 Hz), 7.33 (2H, m), 7.19 (1H, s), 5.11 (1H, d, 8.2 Hz), 4.81–4.58 (2H, m), 4.58–4.40 (3H, m), 4.36–4.29 (1H, m), 4.13–3.94 (4H, m), 3.81 (3H, s), 3.82–3.74 (1H, m), 3.64–3.27 (2H, m), 3.21–3.06 (2H, m), 1.59–1.32 (15 H, m); 13C-NMR (CDCl3, 100 MHz, major rotamer) δ (ppm): 169.5, 167.9, 162.7, 156.4, 150.5, 144.1, 143.8, 141.3, 127.8, 127.1, 125.1, 120.0, 109.4, 79.3, 66.5, 52.9, 52.4, 51.8, 50.4, 49.5, 47.9, 47.1, 40.0, 31.8, 29.7, 28.4, 22.9; MS (ESI, MeOH): m/z calcd. for C36H44N8O9 [M]: 732.32313, found: 733.4 [M + H]+; 755.4 [M + Na]+; 771.4 [M + K]+; HRMS (LTQ-Orbitrap, MeOH) m/z found: 755.31235 [C36H44N8O9Na]+; FT-IR (ATR) ν (cm−1): 3331.9 (m), 2974.1 (s), 2945.6 (s), 2929.8 (s), 2102.1 (s), 1672.4 (m), 1517.8 (m), 1449.5 (m), 1364.9 (s), 1245.0 (s), 1212.6 (s), 1167.3 (s), 1103.5 (s), 1078.4 (s), 891.7 (m), 863.9 (m), 833.4 (s), 758.6 (s), 740.4 (s).
Synthesis of Fmoc-PNA(5L-Arg(Pbf))-N3-OH (5a). Here, 4a (214.3 mg, 0.27 mmol, 1 eq) was solubilized in 11 mL THF and cooled to 0 °C with an ice bath. Next, a solution of Ba(OH)2·8H2O (128.5 mg, 0.41 mmol, 1.5 eq) in 11 mL H2O was added and left to react for 30 min at 0 °C and a further 35 min at room temperature. The reaction was quenched with 1M HCl (0.95 mL, 0.95 mmol, 2.5 eq), the organic fraction was evaporated under reduced pressure, and the pH of the solution was adjusted to 3.5. The product was allowed to precipitate for 2 h at 4 °C. Then, 5a was collected as white solid through Buchner filtration and washed with H2O (0.186 mmol, 68.6% yield). TLC (AcOEt) Rf: 0.17, MP (°C): 101.4–101.7; 1H-NMR (DMSO-d6, 400 MHz, major rotamer) δ (ppm): 8.69–8.50 (1H, br s), 7.89 (2H, d, J = 7.0 Hz), 7.66 (2H, d, J = 7.9 Hz), 7.41 (2H, t, J = 6.4 Hz), 7.33 (2H, m), 7.21 (1H, d, J = 8.3 Hz), 6.58–6.33 (3H, br s), 4.29 (2H, m), 4.21 (1H, m), 3.97 (1H, d, J = 17.4 Hz), 3.87 (1H, d, J = 17.6 Hz), 3.67 (1H, m), 3.61 (2H, s), 3.21 (2H, m), 3.02 (2H, s), 2.95 (2H, s), 2.49 (3H, s), 2.44 (3H, s), 2.00 (3H, s), 1.77 (2H, s), 1.40 (8H, s); 13C-NMR (DMSO-d6, 100 MHz, major rotamer) δ (ppm): 170.7, 168.5, 158.1, 156.6, 156.4,144.3, 141.2, 137.7, 134.7, 131.9, 128.1, 127.5, 125.7, 124.8, 120.6, 116.7, 86.8, 67.5, 51.8, 50.0, 49.6, 48.1, 47.2, 42.9, 40.4, 28.7, 25.6, 19.4, 18.1, 12.7; MS (ESI, MeOH): m/z calcd. for C38H46N8O8S [M]: 774.31593, found: 775.2 [M + H]+; HRMS (LTQ-Orbitrap, MeOH) m/z found: 773.30756 [C38H45N8O8S]−; FT-IR (ATR) ν (cm−1): 3339.2 (w), 2971.0 (m), 2929.8 (m), 2106.0 (s), 1704.0 (m), 1617.3 (m), 1547.0 (s), 1449.8 (s), 1419.3 (m), 1241.8 (m), 1153.5 (s), 1089.7 (s), 1033.7 (s), 993.3 (s), 969.0 (s), 814.1 (m), 782.7 (s), 760.0 (s), 739.6 (s).
Synthesis of Fmoc-PNA(5L-Lys(Boc))-N3-OH (5b). Here, 4b (203.2 mg, 0.33 mmol, 1 eq) was solubilized in 15 mL THF and cooled to 0 °C with an ice bath. Next, a solution of Ba(OH)2·8H2O (157.8 mg, 0.50 mmol, 1.5 eq) in 15 mL H2O was added and left to react for 30 min at 0 °C and further 11 min at room temperature. The reaction was quenched with 1M HCl (1.2 mL, 1.2 mmol, 3.5 eq) and the organic fraction was evaporated under reduced pressure and the pH of the solution was adjusted to 3.5. The product was allowed to precipitate overnight at 4 °C. Then, 5b was collected as beige solid through Buchner filtration and washed with H2O (0.209 mmol, 63.3% yield). TLC (AcOEt) Rf: 0.19, MP (°C): 62.7–63.0; 1H-NMR (DMSO-d6, 400 MHz, major rotamer) δ (ppm): 13.01–12.45 (1H, br s), 7.89 (2H, d, J = 7.5 Hz), 7.65 (2H, d, J = 7.7 Hz), 7.42 (2H, t, J = 7.4 Hz), 7.34 (2H, td, J = 7.4 Hz, 1.9 Hz), 7.19 (1H, d, J = 9.1 Hz), 6.76 (1H, s), 4.37–4.25 (2H, m), 4.23–4.15 (2H, m), 4.04 (1H, d, J = 17.2 Hz), 3.99 (1H, d, J = 38.3 Hz), 3.87 (1H, d, J = 17.3 Hz), 3.70–3.56 (1H, m), 3.28–2.97 (2H, m), 2.94–2.82 (2H, m), 1.46–1.12 (15H, m) 13C-NMR (DMSO-d6, 100 MHz, major rotamer) δ (ppm): 170.7, 168.5, 156.6, 156.0, 144.3, 141.2, 128.1, 127.5, 125.6, 120.6, 77.8, 65.8, 52.0, 49.9, 49.8, 48.1, 47.3, 39.8, 31.7, 28.8, 25.6, 23.3; MS (ESI, MeOH): m/z calcd. for C30H38N6O7 [M]: 594.28020, found: 595.4 [M + H]+; HRMS (LTQ-Orbitrap, MeOH) m/z found: 593.27182 [C30H37N6O7]−; FT-IR (ATR) ν (cm−1): 2926.7 (m), 2111.0 (s), 1714.3 (s), 1699.9 (s), 1683.7 (s), 1655.1 (s), 1648.3 (s), 1527.5 (s), 1506.7 (s), 1449.1 (s), 1419.7 (s), 1366.0 (s), 1244.9 (m), 1163.7 (m), 1102.7 (s), 1079.8 (s), 1020.7 (m), 949.7 (s), 799.2 (s), 759.0 (s), 737.4 (s).
Synthesis of Fmoc-PNA(5L-Arg(Pbf))-F-OH (5c). 4c (286.8 mg, 0.35 mmol, 1 eq) was solubilized in 15 mL THF and cooled to 0 °C with an ice bath. Next, a solution of Ba(OH)2·8H2O (164.0 mg, 0.52 mmol, 1.5 eq) in 15 mL H2O was added and left to react for 30 min at 0 °C and further 46 min at room temperature. The reaction was quenched with 1M HCl (1.2 mL, 1.2 mmol, 3.5 eq), the organic fraction was evaporated under reduced pressure, and the pH of the solution was adjusted to 3.5. The product was allowed to precipitate for 2 h at 4 °C. Then, 5c was collected as a white solid through Buchner filtration and washed with H2O (0.223 mmol, 64.4% yield). TLC (AcOEt) Rf: 0.20, MP (°C): 108.2–108.4; 1H-NMR (DMSO-d6, 400 MHz, major rotamer) δ (ppm): 8.81–8.59 (1H, br s), 7.88 (2H, d, J = 7.5 Hz), 7.68 (2H, d, J = 6.8 Hz), 7.47 (1H, d, J = 10.6 Hz), 7.43–7.36 (2H, m), 7.33–7.24 (2H, m), 7.14 (1H, d, J = 9.5 Hz), 6.60–6.40 (3H, br s), 6.34–6.27 (1H, m), 6.13–6.03 (1H, m), 4.35–4.24 (2H, m), 4.23–4.13 (1H, m), 4.01 (1H, d, J = 16.8 Hz), 3.83 (1H, d, J = 17.8 Hz), 3.70–3.54 (3H, m), 3.33–3.21 (2H, m), 3.08–2.96 (2H, m), 2.94 (2H, s), 2.83–2.71 (2H, m), 2.47–2.51 (3H, s) 2.43 (3H, s), 2.00 (3H, s), 1.80–1.73 (2H, m), 1.39 (6H, s), 1.37–1.20 (2H, m); 13C-NMR (DMSO-d6, 100 MHz, major rotamer) δ (ppm): 172.3, 171.8, 157.9, 156.6, 156.4, 155.3, 144.3, 141.6, 141.2, 137.7, 134.8, 131.9, 128.1, 127.5, 125.6, 124.8, 120.6, 116.7, 110.8, 105.5, 86.7, 67.5, 52.4, 49.9, 48.2, 47.3, 42.9, 40.4, 31.1, 28.7, 25.6, 23.5, 19.4, 18.1, 12.7; MS (ESI, MeOH): m/z calcd. for C43H51N5O9S [M]: 813.34075, found: 814.2 [M + H]+, 836.1 [M + Na]+; HRMS (LTQ-Orbitrap, MeOH) m/z found: 814.34803 [C43H52N5O9S]+; FT-IR (ATR) ν (cm−1): 3329.0 (w), 2967.8 (m), 2936.0 (m), 1715.1 (s), 1617.1 (m), 1545.7 (m), 1449.0 (m), 1406.0 (s), 1242.9 (m), 1151.9 (m), 1088.5 (m), 1012.4 (s), 904.6 (s), 851.4 (s), 810.7 (s), 782.6 (s), 760.6 (s), 733.9 (s).
Synthesis of Fmoc-PNA(5L-Lys(Boc)-F-OH (5d). Here, 4d (219.6 mg, 0.34 mmol, 1 eq) was solubilized in 15 mL THF and cooled to 0 °C with an ice bath. Next, a solution of Ba(OH)2·8H2O (160.4 mg, 0.51 mmol, 1.5 eq) in 15 mL H2O was added and left to react for 30 min at 0 °C and further 46 min at room temperature. The reaction was quenched with 1M HCl (1.2 mL, 1.2 mmol, 3.5 eq) and the organic fraction was evaporated under reduced pressure and the pH of the solution was adjusted to 3.5. The product was allowed to precipitate overnight at 4 °C. Then, 5d was collected as beige solid through Buchner filtration and washed with H2O (0.142 mmol, 42% yield). TLC (AcOEt) Rf: 0.19; MP (°C): 51.4–51.8; 1H-NMR (DMSO-d6, 400 MHz, major rotamer) δ (ppm): 8.17–8.12 (1H, br s), 7.88 (2H, d, J = 7.4 Hz), 7.65 (2H, d, J = 7.1 Hz), 7.52–7.36 (3H, m), 7.36–7.25 (2H, m), 7.21 (1H, d, J = 9.2 Hz), 6.75 (1H, s), 6.36–6.25 (1H, m), 6.10 (1H, d, J = 2.6 Hz), 4.28 (2H, m), 4.18 (1H, m), 4.12–3.78 (2H, m), 3.74–3.62 (1H, s), 3.42–3.21 (2H, m), 2.87 (2H, s), 2.75 (2H, m), 2.58–2.46 (2H, m), 1.36 (15H, m); 13C-NMR (DMSO-d6, 100 MHz, major rotamer) δ (ppm): 171.9, 171.2, 156.5, 156.0, 155.2, 144.3, 141.6, 141.2, 128.0, 127.5, 125.5, 120.5, 110.8, 105.6, 77.8, 66.7, 52.6, 50.2, 48.1, 47.3, 40.4, 31.6, 30.7, 29.7, 28.7, 23.6, 23.3; MS (ESI, MeOH): m/z calcd. for C35H43N3O8 [M]: 633.30502, found: 634.4 [M + H]+; 672.3 [M + K]+; HRMS (LTQ-Orbitrap, MeOH) m/z found: 656.29424 [C35H43N3O8Na]+; FT-IR (ATR) ν (cm−1): 3314.8 (w), 2937.2 (m), 2357.5 (s), 2344.2 (s), 1701.0 (m), 1636.7 (m), 1507.6 (m), 1449.8 (s), 1365.9 (s), 1246.6 (m), 1166.2 (m), 1077.1 (s), 1014.1 (s), 859.5 (m), 759.6 (s), 736.3 (s).
Synthesis of Fmoc-PNA(5L-Arg(Pbf))-T(N3)-OH (5e). Here, 4c (256.6 mg, 0.28 mmol, 1 eq) was solubilized in 10 mL THF and cooled to 0 °C with an ice bath. Next, a solution of Ba(OH)2·8H2O (133.0 mg, 0.42 mmol, 1.5 eq) in 15 mL H2O was added and left to react for 30 min at 0 °C and then a further 46 min at room temperature. The reaction was quenched with 1M HCl (0.84 mL, 0.84 mmol, 3.5 eq) and the organic fraction was evaporated under reduced pressure and the pH of the solution was adjusted to 3.5. The product was allowed to precipitate for 2 h at 4 °C. Then, 5c was collected as white solid through Buchner filtration and washed with H2O (0.271 mmol, 96.3% yield). TLC (EtOAc) Rf: 0.1; 1H-NMR (400 MHz, DMSO-d6, major rotamer) δ 11.53 (1H, s), 7.89 (2H, d, J = 7.4 Hz), 7.67 (2H, t, J = 8.3 Hz), 7.60 (1H, d, J = 6.9 Hz), 7.41 (2H, t, J = 7.4 Hz), 7.40–7.25 (3H, m), 7.06 (1H, 1 br s), 6.82 (1H, br s), 6.54 (1H, br s), 4.52 (1H, s), 4.44–4.28 (2H, m), 4.22 (1H, d, J = 5.9 Hz), 4.03 (2H, s), 3.70–3.53 (3H, m), 3.04 (2H, s), 2.94 (2H, s), 2.49 (3H, s), 2.42 (3H, s), 2.00 (3H, s), 1.55–1.20 (10H, m); 13C-NMR (100 MHz, DMSO-d6) δ 171.4, 168.0, 163.9, 157.9, 156.5, 156.3, 151.1, 146.4, 144.3, 141.2, 137.7, 134.7, 131.9, 130.1, 129.4, 129.0, 128.0, 127.5, 125.7, 124.8, 120.5, 116.7, 107.6, 86.7, 65.6, 51.7, 49.9, 49.4, 48.6, 47.3, 42.9, 28.7, 25.6, 19.4, 18.1, 12.7; MS (ESI, MeOH): m/z calcd. for C43H50N10O10S [M]: 898.34321, found: 897.7 [M − H]−, 935.7 [M + Na]+, 951.7 [M + K]+; HRMS (LTQ-Orbitrap, MeOH) m/z found: 897.3360 [C43H49N10O10S]−.
Synthesis of Fmoc-PNA(5L-Lys(Boc))-T(N3)-OH (5f). Here, 4f (232.8 mg, 0.32 mmol, 1 eq) was solubilized in 14 mL THF and cooled to 0 °C with an ice bath. Next, a solution of Ba(OH)2·8H2O (151.1 mg, 0.46 mmol, 1.5 eq) in 14 mL H2O was added and left to react for 30 min at 0 °C and a further 10 min at room temperature. The reaction was quenched with 1M HCl (1.1 mL, 1.1 mmol, 3.5 eq) and the organic fraction was evaporated under reduced pressure and the pH of the solution was adjusted to 3.5. The product was allowed to precipitate overnight at 4 °C. 5f was then collected as white solid through Buchner filtration and washed with H2O (0.244 mmol, 77% yield). TLC (AcOEt) Rf: 0.21; MP (°C): 97.4–97.7; 1H-NMR (DMSO-d6, 400M Hz, major rotamer) δ (ppm): 11.57 (1H, s), 8.10 (1H, m), 7.90 (2H, d, J = 7.5 Hz), 7.69 (2H, d, J = 7.0 Hz), 7.60 (1H, s), 7.42 (2H, t, J = 7.4 Hz), 7.33 (2H, t, J = 7.4 Hz), 7.27 (1H, d, J = 8.7 Hz), 6.81–6.72 (1H, m), 4.74 (2H, dd, J = 57.4 Hz, 16.7 Hz), 4.40–4.28 (2H, m), 4.26–4.19 (1H, m), 4.13 (1H, d, J = 7.7 Hz), 4.17–3.87 (4H, m), 3.76–3.64 (1H, m), 3.63–3.50 (1H, m), 3.01–2.82 (2H, m), 1.56–1.14 (15H, m); 13C-NMR (DMSO-d6, 100 MHz, major rotamer) δ (ppm): 170.6, 167.7, 163.9, 156.6, 156.0, 151.1, 146.1, 144.4, 141.2, 128.1, 127.5, 125.7, 120.6, 107.7, 77.8, 65.8, 51.9, 51.5, 50.3, 48.6, 47.3, 47.2, 40.4, 32.0, 29.8, 28.7, 23.3; MS (ESI, MeOH): m/z calcd. for C35H42N8O9 [M]: 718.30747, found: 719.4 [M + H]+; 741.4 [M + Na]+; 757.4 [M + K]+; HRMS (LTQ-Orbitrap, MeOH) m/z found: 741.29822 [C35H42N8O9Na]+; FT-IR (ATR) ν (cm−1): 3317.9 (m), 2932.8 (m), 2100.8 (m), 1669.5 (m), 1517.9 (m), 1450.4 (m), 1389.0 (s), 1365.3 (s), 1338.8 (s), 1246.0 (m), 1165.9 (m), 1104.4 (s), 1080.1 (s), 948.7 (m), 864.4 (m), 760.2 (s), 740.4 (s).