Abstract
Acetylserotonin methyltransferase (ASMT) is the last enzyme of melatonin biosynthesis and may play a rate-limiting role in the melatonin production of plants. In this study, systematic analysis of the ASMT gene family in tomato (Solanum lycopersicum Mill) has been presented by the integration of the structural features, phylogenetic relationships, exon/intron configuration, and expression profile during growth and development, as well as biotic stresses. The results revealed that the tomato genome encoded a minimum of 14 members, containing three probable encoded pseudogenes. Chromosome mapping indicated that the family had probably expanded via tandem duplication events. Genome-wide RNA-seq and qRT-PCR based gene expression analysis revealed that almost half of the SlASMT genes were expressed in at least one of the experimental stages studied and also showed differential accumulation. Furthermore, the tandem duplicated SlASMT genes showed differential expression levels, which indicated probable functional divergence during the course of the evolution. Finally, this study also determined that some SlASMT genes were induced by multiple pathogens. The results suggested that these genes could be involved in tomato plant response to biotic stresses.
1. Introduction
Melatonin (N-acetyl-5-methoxytryptamine) has been identified as an indoleamine, with an unstable form. It widely exists in various tissues of higher plants, such as the roots, stems, leaves, flowers, fruit, and seeds [1,2,3,4,5,6]. Following the first report in 1995, it was determined that melatonin exists in almost every plant species [7,8]. Moreover, these studies have shown that melatonin plays an important role in regards to broad-spectrum antioxidants [9,10], biotic and abiotic stresses [11,12,13,14,15,16,17], seed dormancy, and the growth and development of plants [18,19,20]. Meanwhile, plants treated with exogenous melatonin in specific concentrations also have the ability to appropriately regulate resistance mechanisms and metabolic processes during growth and development stages [9,21,22,23].
In recent years, various studies have confirmed that melatonin synthesis in plants is involved with four different enzymes, including tryptophan decarboxylase (TrpDC), tryptamine 5-hydroxylase (T5H), serotonin N-acetyltransferase (SNAT), and acetylserotonin-O-methyltransferase (ASMT) [24,25]. However, when compared with that in animals, plant serotonin is purportedly synthesized first by the catalysis of tryptophan decarboxylase (TrpDC), followed by tryptamine 5-hydroxylase (T5H), rather than tryptophan 5-hydroxylase (Trp5H), and aromatic l-amino acid decarboxylase (AADC) as found in animals [24,25]. In plant species, previous results have shown that the over-expression of apple MzASMT improves melatonin production (2 to 4 times higher than that in the wild type)and enhances the drought tolerance of transgenic Arabidopsis thaliana plants [26]. In 2011, the first rice ASMT was cloned through the in vivo analysis of melatonin in recombinant Escherichia coli, and it was found that rice ASMT mRNA was induced upon senescence. Also, its induction closely paralleled the melatonin production in rice leaves. Also, rice ASMT mRNA was shown to be induced in abscisic acid (ABA), salt, and copper treatments, which suggested the possible involvement of melatonin in oxidative stress [27]. In 2013, Park et al. further determined that the over-expression of rice ASMT1, ASMT2, and ASMT3 independently enhanced enzyme activity [28]. Moreover, ASMT1 and ASMT2 transcripts were highly expressed in the stems and flowers. However, ASMT3 was barely detectable in any of the plant organs. All three ASMT mRNAs were simultaneously induced in the treatments with abscisic and methyl jasmonic acids [28]. In tomato plants, the transgenic plants’ over-expression of homologous sheep oHIOMT genes had higher melatonin levels, as well as enhanced drought tolerance, which provided new proof for the important role played by ASMT in plant melatonin synthesis [29]. Therefore, the accumulated evidence has demonstrated that ASMT is a very important enzyme for the biosynthesis of melatonin.
The tomato, as a model plant, has become an excellent material for the research interpretation of the various life activities of plants. Previously, according to an enzyme-linked immunosorbent assay, Okazaki et al. detected melatonin in the roots, stems, leaves, flowers, fruits, seedlings, and seeds, which ranged from 1.5 to 66.6 ng/g of the tomato plants’ fresh weight [2]. Sun et al. reported that an exogenous melatonin treatment significantly promoted ripening and improved tomato fruit quality post-harvest [30]. Arnao and Hernández-Ruiz reported that tomato plants under variable conditions exhibited a higher melatonin content [31]. Under abiotic stress, melatonin can enhance thermotolerance and cadmium phytotoxicity [32,33]. In addition, Li et al. reported that melatonin mediated selenium-induced tolerance to cadmium stress in tomato plants [34]. Subsequently, researchers found that HsfA1 not only induces drought tolerance but only upregulates melatonin biosynthesis to confer cadmium tolerance to tomato plants [35,36]. Overall, melatonin plays a very important role in regulating the growth and development, as well as controlling the environmental adaptation of tomato plants.
In this study, the ASMT gene family in tomato plants was characterized by the integration of structural features, phylogenetic relationships, chromosome localization, and the expression patterns in various tissues, as well as the responses to biotic stresses. This study’s in silico search identified a total of 14 putative ASMT genes in the S. lycopersicum genome. Evolutionary analysis revealed that tandem duplication events largely contributed to the expansion of the SlASMT gene family. A detailed expression analysis of the SlASMT genes, based on RNA-seq and qRT-PCR, showed distinct patterns of expression in the different organs and identified a number of SlASMT genes that were regulated by multiple pathogen infections. This study not only contributes to the understanding of the evolutionary patterns of the SlASMT genes in plants but also lays a foundation for deciphering the important functions of SlASMT in regulating melatonin biosynthesis in tomato plants.
2. Results
2.1. Identification of SlASMT Genes in the Tomato Genome
In previous research, three rice ASMT genes (ASMT1, ASMT2 and ASMT3) were identified with different functions [27,28], leading to the hypothesis that plant species may contain a family of ASMT genes. In this current study, the BlastP tool was used to search the tomato genome database of the Sol Genomics Network (SGN, http://solgenomics.net/) using an amino acid sequence of the O-methyltransferase conserved domain as the query. Based on previous reports [26,37], we selected tomato ASMT homologs with more than 31% amino acid identities since the rice ASMT gene (a query) has about 31% aa identity to tomato caffeic acid O-methyltransferase (COMT). A total of fourteen non-redundant SlASMT genes candidates were identified (SlASMT01 to SlASMT14). Each of the three SlASMT genes (SlASMT06, SlASMT09, and SlASMT13) was missing significant portions of their coding regions. Therefore, it was most likely encoded truncated protein, or possibly pseudogenes. These three members were subsequently excluded from the phylogenetic and expression analysis. The characteristics of these SlASMT genes from the tomato plants were illustrated (Table 1), including the gene name, ID, and location, along with the number of exons, protein size, molecular weight (MW), and isoelectric point (pI).

Table 1.
Tomato ASMT gene family members.
2.2. Analysis of the SlASMT Gene Phylogeny and Exon-Intron Structure
A Neighbor-Joining phylogenetic tree with 18SlASMT genes was constructed for the purpose of exploring the phylogenetic relationship of the SlASMTs in tomato plants using the MEGA5.0 program (Figure 1A). According to the phylogenetic tree topology, these genes could be divided into three groups (including Groups I to III). In order to examine the structural features of the SlASMT genes, the exon/intron configurations of SlASMT genes in the tomato plants were compared. The view of exon/intron structures was obtained using an online Gene Structure Display Server (GSDS: http://gsds.cbi.pku.edu.cn), with both coding sequences (CDS) and genomic sequences. Figure 1B provides a detailed illustration of the relative length of the introns and the conservation of the corresponding exon sequences within each of the SlASMTs. The results confirmed that various numbers of introns were found, and each of the SlASMTs contained fewer introns (0–3). In Group I, only one intron was observed in each member. The members of Group II included two introns, with the exception of the SlASMT12, which had no introns. In Group III, three introns were found in SlASMT02, and the remaining SlASMT03 and SlASMT04 had two introns. The results revealed relatively few introns in the process of the evolution of the tomato plants.
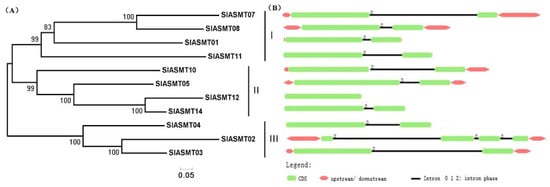
Figure 1.
Phylogenetic analysis and intron/exon configurations of SlASMT genes in tomato plants. (A) Phylogenetic tree of the SlASMT genes was constructed using MEGA 5.0; (B) The introns and exons were drawn to scale with the full encoding regions of their respective genes. The boxes indicate the exons, and lines indicate the introns: 0 = intron phrase 0; 1 = intron phrase 1; 2 = intron phrase 2.
2.3. Genomic Distribution and Gene Duplication
In this study, chromosomal mapping was performed to examine the genome distribution of the SlASMT genes in the tomato plants. As the results indicated, multiple genes were located on seven of the twelve chromosomes and exhibited uneven distributions (Figure 2). A maximum number of three genes was mapped on chromosomes 02, 06 and 12, while only one gene was located on chromosomes 01, 03 and 09. In contrast, chromosomes 10 contained two members (SlASMT10 and SlASMT11).
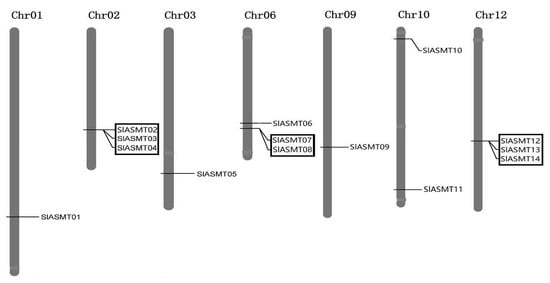
Figure 2.
Position of SlASMT genes on the tomato chromosomes. The chromosome numbers are indicated at the top end of the chromosome. The tandem duplicated gene clusters are indicated by the black boxes.
It is well known that tandem and segmental duplications play roles in gene family expansion during plant evolution [38,39]. In this study, 10 (41.7%) of the SlASMT genes were clustered into three groups (A, B and C), and seemed to be produced from tandem duplications (Figure 3; Table 2). These three groups were juxtaposed, with no intervening gene (Table 2). The distance between these genes ranged from 0.128 kb to 139.18 kb (Table 2), and the overall similarity of the encoding sequences of these genes ranged from 38.2% to 60.2%. There were two genes in tandem in group B while groups A and C contained three genes each. However, no evidence of segmental genome duplications was identified for these SlASMT genes in the tomato plants. Therefore, these results suggested that tandem duplication events largely contributed to the expansion of the SlASMT gene family in the tomato plants.
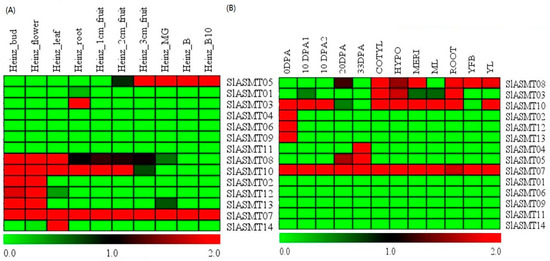
Figure 3.
Expression profiles of SlASMT genes based on RNA-Seq in tomato plants. (A) Cultivated tomato (S. lycopersicum); (B) Wild tomato (S. pinpinellifolium); all RNA-Seq datasets were obtained from the Tomato Functional Genomics Database (http://ted.bti.cornell.edu/), and a detailed description of the samples is available from the Tomato Functional Genomics Database. Then, the log2-transformed RPKM values were used to obtain a heatmap using Multi-Experiment Viewer software. The blocks with colors indicate low (green) or high (red) transcript accumulation, relative to the respective control. MG is the mature green fruit; B represents the breaker stages (early ripening); and B10 represents the 10-day post breaker stages (red ripe).

Table 2.
Tandem duplicated SlASMT genes.
2.4. SlASMTs Genes Have Specific Expression Patterns in Different Tomato Genotypes
All of the available RNA-Seq data in the Tomato Functional Genomics Database (http://ted.bti.cornell.edu/) were downloaded in order to decipher the expression pattern of the SlASMT genes among the various tissues of the tomato plants. The normalized gene expression values were estimated by the reads per kilo bases per million reads mapped (RPKM). Subsequently, the log2-transformed RPKM values were used to draw heatmaps using Mev4.9 software, and the results are shown in Figure 3. In this study, an in silico expression analysis was performed on various tissues in S. lycopersicum. As shown in Figure 3A, the results revealed that 9 of the 14 genes were expressed in all of the tested tissues. The transcripts of SlASMT07 consistently appeared in all of the various tissues, while two of the genes (SlASMT03, and SlASMT14) showed tissue-specific expression profiles. Three of the genes (SlASMT02, SlASMT12 and SlASMT13) showed similar expression profiles and were expressed in the buds and flowers. The gene SlASMT05 was expressed exclusively in fruit. Expression profiles of the SlASMT08 and SlASMT10 genes were similar and were found under multiple tested conditions, including the bud, flower, leaf, root, 1 cm-fruit, 2 cm-fruit and 3 cm-fruit.
To further expand on the present knowledge of the expression profiles of the SlASMT genes in the different tissues, the expression patterns of the SlASMT genes in the different tissues of S. Lycopersicum and S. pinpinellifolium were compared (Figure 3B). The results showed that 9 of 14 genes were expressed in the tested tissues of S. pinpinellifolium. Seven of the genes (SlASMT02, SlASMT07, SlASMT08, SlASMT05, SlASMT10, SlASMT12, and SlASMT13) showed similar expression profiles in the different tissues of S. lycopersicum and S. pinpinellifolium. Two of the genes (SlASMT03 and SlASMT04) showed different expression profiles in the different tissues of the two species.
2.5. SlASMT Genes are Regulated by Multiple Pathogens
Subsequently, the expression patterns of the SlASMTs were further analyzed in response to multiple pathogens, including Pseudomonas syringae tomato DC3000, Pseudomonas fluorescens, Pseudomonas putida, and Agrobacterium tumefaciens. The results showed that the expression abundances of 10 of the 14 genes were not detected under these biotic stresses. However, the remaining 4 genes were induced by these biotic stress factors (Figure 4). The SlASMT03 and SlASMT07 genes showed similar expression profiles and were up-regulated by the three pathogens, DC3000, Pseudomonas fluorescens, and Pseudomonas putida. The SlASMT08 gene was weakly non-regulated under all of the tested biotic stress conditions, while SlASMT10 genes were only up-regulated by Pseudomonas fluorescens, not Pseudomonas syringae tomato DC3000, Pseudomonas putida, or Agrobacterium tumefaciens.
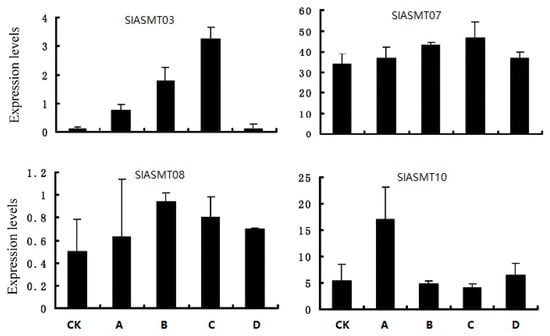
Figure 4.
Expression patterns of SlASMTs in response to different pathogens. The axis of abscissas A, B, C and D represent Pseudomonas syringae tomato DC3000, Pseudomonas fluorescens, Pseudomonas putida and Agrobacterium tumefaciens.
2.6. Expression Divergence of the Tandem Duplicated SlASMT Genes in Tomato Plants
In this study, the expression patterns of the tandem duplicated SlASMT genes were examined. The RPKM values were available for three groups in the RNA-seq data under the different conditions in this study. Genes in the first group (SlASMT02, SlASMT03 and SlASMT04) showed divergent expression profiles. For example, the SlASMT04 gene was not expressed at significant levels in all tested tissues (Figure 5A), which indicated that pseudo-functionalization after duplication could occur. Of these three genes, only the SlASMT03 gene was induced by multiple pathogens (Figure 5B). The SlASMT07 and SlASMT08 genes of the second group showed similar expression profiles in different tissues and development stages (Figure 5C), while only the former was found to be induced by multiple pathogens (Figure 5D). In the third group (SlASMT12, SlASMT13 and SlASMT14), the SlASMT12 and SlASMT13 genes revealed similar expression profiles, while the SlASMY14 gene was expressed in the leaves (Figure 5E). Moreover, these three genes were not induced by the four pathogens (Figure 5F).
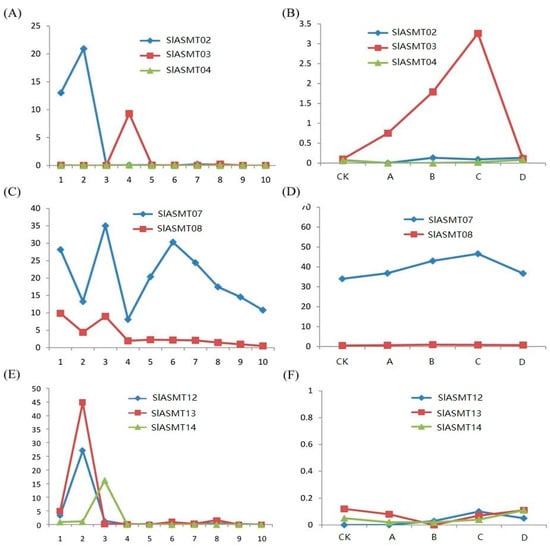
Figure 5.
Expression patterns of SlASMT genes presented as tandem duplicates. The X-axis represents the developmental stages as shown in supplemental Table S2. The Y-axis represents the raw expression values obtained from RNA-seq.
2.7. RT-qPCR Analysis
In order to confirm the results obtained by the RNA-Seq and to attempt to quantify the expression levels, qRT-PCR was performed, and the results were compared. In this study, the expressions of eleven SlASMT genes were analyzed for eight different tissues, including the roots, stems, leaves, flowers, immature green fruit, mature green fruit, breaker fruit, and 10-day fruit post breaker stages (red ripe). Among these eleven SlASMT genes, four (SlASMT01, SlASMT04, SlASMT08 and SlASMT11) were not detected in any tissues. The remaining seven genes were expressed in at least one of the tested tissues (Figure 6). However, the expression levels of these genes displayed obvious differences. Three of the genes (SlASMT02, SlASMT03 and SlASMT12) showed tissue-specific expression patterns. Of these seven SlASMTs, five were expressed in the roots, with the exception of the SlASMT02 gene, which was expressed in the flowers. SlASMT07 was ubiquitously expressed at similar levels in most of the tested organs. Additionally, the SlASMT14 gene was expressed in the leaves based on the RNA-seq method. However, in this study, it was only detected in the roots and flowers when using qRT-PCR. Overall, these results were consistent with the expression of the SlASMT genes obtained by the RNA-Seq data.
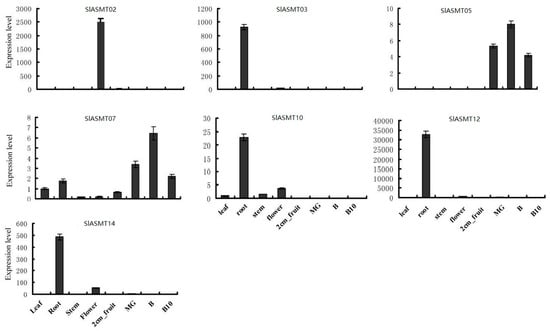
Figure 6.
Expression patterns of SlASMT genes in different organs and developmental stages.
3. Discussion
In previous studies, acetylserotonin-O-methyltransferase from rice, the last enzyme involved in melatonin biosynthesis, was cloned. It possesses partial homologies to other plant O-methyltransferases [28] and is localized in the cytosol [40]. Subsequently, two different genes (ASMT2 and ASMT3), which were expressed at different levels, have been also reported in rice [41]. Given the multiple number of putative ASMT gene members identified in rice, it is possible that the ASMT gene family may exist in different plant genomes. Therefore, in this present study, in silico identification of the ASMT genes was performed by searching the S. lycopersicum genomic database (http://solgenomics.net/). The results showed a total of fourteen candidate non-redundant SlASMT genes, which indicated that the tomato genome encoded a family of SlASMT genes. Surprisingly, a large number of the pseudogenes (3/14 = 20%) was found in the tomato genome. The presence of such a high proportion of pseudogenes indicated that these genes could potentially play a role in the course of the plants’ evolution.
It is well known that gene duplications (tandem and segmental duplications) are one of the primary driving forces in the evolution of gene families with variations in size and distribution [38,39]. In tomato plants, the SlASMT gene family comprises a diverse family of proteins (Table 1). A total of 8 duplicated SlASMT genes were found to be present in the tomato chromosomes (Figure 2), and these were only involved in tandem duplications. Therefore, the gene duplication and subsequent expansion of the SlASMT genes seemed to have occurred throughout evolution. However, segmental duplication events were not observed in these genes. Therefore, in the present study, tandem duplication events have been determined to contribute significantly to the evolution of the SlASMT genes in tomato plants.
In addition, it was also found that the expression pattern of the tandem duplicated genes was highly variable within the three groups (Figure 5). For example, the sequence comparison between the SlASMT02 and SlASMT03 in the first group revealed a reduced level of homology at the amino acid level (56.27%), indicating that these genes might have undergone significant diversification after duplication, resulting in neo-functionalization. Also, in regards to the SlASMT04 gene in the first group of the tandem duplicated genes, it did not show expression at significant levels in any of the tested tissues, which indicated that a pseudo-functionalization could have occurred after the duplication. This result may have been due to the fact that the genes with low levels of expression slowly lose their functions during the course of the evolutionary process. Therefore, the functional divergence of the SlASMTs occurred during the course of the evolution.
In order to investigate the possible functional differences of the SlASMT genes, their expression patterns were analyzed based on RNA-Seq and qRT-PCR technology. The results demonstrated different types of expression patterns among these SlASMTs (Figure 3 and Figure 6). Three of these genes (SlASMT08 and SlASMT10) were expressed in the buds, flowers, leaves, roots, and non-mature fruit, which suggested an important role in the vegetative growth, as well as the early stages of reproductive growth. The expression of seven of the SlASMT genes showed tissue-specific patterns, including flowers and buds (SlASMT02, SlASMT12 and SlASMT13), leaves (SlASMT14), and roots (SlASMT03). These results indicated a vital role in the growth and development of the flowers, leaves, and roots, respectively. In addition, it was also found that the SlASMT05 gene was expressed in the mature fruit, which indicated that it could potentially play vital roles in the vegetative growth, as well as the latter stages of the fruit development. Finally, the expression of the SlASMT07 gene was detected in all of the tested tissues and was involved in all growth and development stages of the tomato plants. In regards to the remaining five genes, their expressions were not detected in any of the tissues. These results suggested that the expression levels of these genes were too low to be detected in the tested tissues, or that they were not expressed to any significant degree.
The expression patterns of the SlASMTs in response to multiple pathogens were further analyzed, including Pseudomonas syringae tomato DC3000, Pseudomonas fluorescens, Pseudomonas putida, and Agrobacterium tumefaciens. The results showed that the expression of 10 of 14 genes was not detected under these biotic stresses. However, the remaining 4 genes were induced by these biotic stress factors (Figure 4). The SlASMT03 and SlASMT07 genes showed similar expression profiles and were up-regulated by three pathogens (Pseudomonas syringae tomato DC3000, Pseudomonas fluorescens, and Pseudomonas putida). SlASMT08 was weakly non-regulated under all of the tested biotic stress conditions. In addition, the SlASMT16 genes were only up-regulated by Pseudomonas fluorescens. All the results suggested that these four genes are involved in tomato plant responses to biotic stresses.
4. Materials and Methods
4.1. Retrieval and Identification of the SlASMT Gene Family
In the present study, two methods were used to identify the potential AMST homologs in the tomato genome. First, a TBLASTN search was performed using the protein sequences of the ASMT genes from rice [27,28] as the query against the SGN tomato genome database (http://solgenomics.net/). Then, a HMM profile of the ASMT conserved domain (Pfam: PF00282) was downloaded from the Pfam protein family database (http://pfam.sanger.ac.uk/). For the purpose of identifying the SlAMST genes, BlastP was used in the S. lycopersicum genome. The default parameters were employed. All of the non-redundant gene sequences were searched against the tomato genome data of the SGN (http://solgenomics.net/). The e-value used was 1 × 10−5. The conserved NBS domain of these predicted NBS-encoding proteins was determined by Pfam version 22.0 (http://pfam.janelia.org). Subsequently, the molecular weights and the isoelectric point of the SlAMSTs deduced proteins were predicted using the online tool ExPASy (http://web.expasy.org/protparam/).
4.2. Mapping of the SlASMT Genes on the S. lycopersicum Genome and the Determination of the Exon-Intron Structure
In order to determine the location of the SlASMT genes on the 12 tomato chromosomes, the data concerning the gene positions were extracted from the tomato Genome Sequence ITAG Release 2.4 in the SGN (http://solgenomics.net). The software MapDraw 2.2 was used to visualize the locations of the SlASMT genes on the tomato chromosomes. The tandem duplicated genes were defined as being closely related genes on a single chromosome, with a maximum of ten intervening genes [42]. The segmentally duplicated genes were detected by a CoGeSynMap program (http://genomevolution.org/CoGe/SynMap.pl). Additionally, in order to further investigate the structural characteristics of the SlAMSTs genes family, the genome sequence, coding sequence (CDS), and protein sequence of the homologous genes of the SlAMSTs in the tomato plants were downloaded from the SGN (http://solgenomics.net). A schematic diagram of the intron-exon structure of the SlAMSTs genes was depicted by the online tool Gene Structure Display Sever (version 2.0) (http://gsds.cbi.pku.edu.cn/).
4.3. Phylogenetic Tree Constructions
In order to determine the phylogenetic relationships of the SlAMST genes in tomato plants, multiple sequence alignments of the SlAMST protein sequences were performed using Clustal software (version 2.0) [43] and were then encoded by a BioEdit Sequence Alignment Editor [44]. Then, a phylogenetic tree was constructed using MEGA 5.0 software and a Neighbor-Joining method [45,46]. Bootstrap analysis was performed using 1000 resampling replications, and the branch lengths were assigned through the pair-wise calculations of the genetic distances. The missing data were treated as pair-wise deletions of the gaps.
4.4. Expression Analysis of the SlASMT Genes Based on RNA-seq
The widespread application of RNA-seq data was convenient for detecting the differential expression of the genes [47]. In this study, for the purpose of deciphering the expression pattern of the SlAMST gene family in the tomato plants’ various tissues, as well as the responses to biotic stresses, all available transcriptome data of the SlAMSTs genes were obtained from the Tomato Functional Genomics Database (http://ted.bti.cornell.edu/). This was followed by the obtained expression data being submitted to a Multiple Experiment Viewer (Verision Mev 4.9) software program with a log2 transformation in order to generate a heat map [48]. The obtained data were hierarchically clustered, based on Pearson’s correlation distance with average linkage, and a cluster analysis was performed on the rows of the expression values.
4.5. RT-qPCR Analysis
At this point, to further verify the expression pattern of the SlASMTs, eight tissues samples were obtained. These tissue samples included roots, stems, leaves, flowers, immature green fruit, mature green fruit, breaker fruit, and red fruit, obtained from S. lycopersicum L. var zhefen702, which had been grown in a controlled environment chamber at the Zhejiang Academy of Agricultural Sciences. Then, the total RNA was extracted, and the first-strand cDNA was synthesized using a RNA simple Total RNA Kit (Tiangen, Beijing, China), and a TIANScript cDNA Synthesize Kit (Tiangen Biotech, Beijing, China), respectively, both in accordance with the manufacturer’s instructions. Supplemental Table S1 lists the gene-specific primers of the SlASMTs for the qRT-PCR. The real-time PCR reactions were carried out in a total volume of 20 μL, containing 10 μL of SuperMix, 0.4 μL of each primer, 1 μL of template (10× the diluted cDNA from the samples), and 7.8 μL of sterile distilled water. The thermal conditions were as follows: 95 °C for 30 s; followed by 40 cycles at 95 °C for 5 s; 55 °C for 15 s; and 72 °C for 10 s. The relative gene expression values were calculated using the 2−ΔΔCt method. GAPDH was used as a reference gene for the expression analysis of the SlASMT genes in the tomato plants [49], and three independent replicates were then performed.
5. Conclusions
Here, we tried to characterize the ASMT gene family in tomato plants by the integration of structural features, phylogenetic relationships, chromosome localization, and the expression patterns in various tissues, as well as the responses to biotic stresses. In summary, this study identified fourteen members of the SlASMT gene family in tomato plants, and elaborated their phylogenetic relationships and evolutionary modes. Further analysis determined that several SlASMT genes displayed tissue-specific expression profiles. The results of this study lay the foundation for the deciphering of the function of the SlASMT family members in tomato plants.
Supplementary Materials
The following are available online, Table S1: Primers used to detect the expression of tomato SlASMT genes in this study. Table S2: The developmental stages and leaves infected by different pathogens are represented on the X-axis.
Acknowledgments
This research was partially supported by the National Natural Science Foundation of China (31301774), Zhejiang Provincial Natural Science Foundation of China (LY18C150008, LQ15C150002, LY18C030001, LQ13C030002), State Key Laboratory Breeding Base for Zhejiang Sustainable Pest and Disease Control (No. 2010DS700124-KF15168), the Earmarked Fund for China Agriculture Research System (CARS-23-G44), National Science and Technology basic Resources survey Social (2017FY100700), Science and Technology Plan Zhejiang Province (2018C35077), Wenzhou Municipal Science and Technology Plan Project (N20170001, S20160004, N20140046), Zhejiang Provincial major Agricultural Science and Technology Projects of New Varieties Breeding (2016C02051), and National key research and development program (2017YFD0101902).
Author Contributions
W.H., Z.C., L.W. and D.Z. participated in the design of the study, helped to interpret the results and draft the manuscript. L.W. and D.Z. performed part of the experimental work; Y.C. carried out the retrieval and identification of the SlASMT gene family. X.P. directed the expression analysis of the SlASMT genes based on RNA-seq; C.C. performed the mapping of the SlASMT genes in the S. lycopersicum genome and the determined the exon-intron structure; L.W constructed the phylogenetic tree and performed the RT-qPCR analysis. D.Z. wrote the paper.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Manchester, L.C.; Tan, D.X.; Reiter, R.J.; Park, W.; Monis, K.; Qi, W. High levels of melatonin in the seeds of edible plants: Possible function in germ tissue protection. Life Sci. 2000, 67, 3023–3029. [Google Scholar] [CrossRef]
- Okazaki, M.; Ezura, H. Profiling of melatonin in the model tomato (Solanum lycopersicum L.) cultivar Micro-Tom. J. Pineal Res. 2009, 46, 338–343. [Google Scholar] [CrossRef] [PubMed]
- Murch, S.J.; Alan, A.R.; Cao, J.; Saxena, P.K. Melatonin and serotonin in flowers and fruits of Daturametel L. J. Pineal Res. 2009, 47, 277–283. [Google Scholar] [CrossRef] [PubMed]
- Zhao, Y.; Tan, D.X.; Lei, Q.; Chen, H.; Wang, L.; Li, Q.T.; Gao, Y.; Kong, J. Melatonin and its potential biological functions in the fruits of sweet cherry. J. Pineal Res. 2013, 55, 79–88. [Google Scholar] [CrossRef] [PubMed]
- Burkhardt, S.; Tan, D.X.; Manchester, L.C.; Hardeland, R.; Reiter, R.J. Detection and quantification of the antioxidant melatonin in Montmorency and balaton tart cherries (Prunus cerasus). J. Agric. Food Chem. 2001, 49, 4898–4902. [Google Scholar] [CrossRef] [PubMed]
- González-Gómez, D.; Lozano, M.; Fernández-León, M.F.; Ayuso, M.C.; Bernalte, M.J.; Rodríguez, A.B. Detection and quantification of melatonin and serotonin in eight Sweet Cherry cultivars (Prunus avium L.). Eur. Food Res. Technol. 2009, 229, 223–229. [Google Scholar] [CrossRef]
- Hattori, A.; Migitaka, H.; Iigo, M.; Itoh, M.; Yamamoto, K.; Ohtani-Kaneko, R.; Hara, M.; Suzuki, T.; Reiter, R.J. Identification of melatonin in plants and its effects on plasma melatonin levels and binding to melatonin receptors in vertebrates. Biochem. Mol. Biol. Int. 1995, 35, 627–634. [Google Scholar] [PubMed]
- Dubbels, R.; Reiter, R.J.; Klenke, E.; Goebel, A.; Schnakenberg, E.; Ehlers, C.; Schiwara, H.W.; School, W. Melatonin in edible plants identified by radioimmunoassay and by high performance liquid chromatography-mass spectrometry. J. Pineal Res. 1995, 18, 28–31. [Google Scholar] [CrossRef] [PubMed]
- Kolár, J.; Machácková, I. Melatonin in higher plants: Occurrence and possible functions. J. Pineal Res. 2005, 39, 333–341. [Google Scholar] [CrossRef] [PubMed]
- Tan, D.X.; Hardeland, R.; Manchester, L.C.; Korkmaz, A.; Ma, S.; Rosales-Corral, S.; Reiter, R.J. Functional roles of melatonin in plants, and perspectives in nutritional and agricultural science. J. Exp. Bot. 2012, 63, 577–597. [Google Scholar] [CrossRef] [PubMed]
- Zhang, N.; Zhao, B.; Zhang, H.J.; Weeda, S.; Yang, C.; Yang, Z.C.; Ren, S.; Gut, Y.D. Melatonin promotes water-stress tolerance, lateral root formation, and seed germination in cucumber (Cucumis sativus L.). J. Pineal Res. 2013, 54, 15–23. [Google Scholar] [CrossRef] [PubMed]
- Bajwa, V.S.; Shukla, M.R.; Sherif, S.M.; Murch, S.J.; Saxena, P.K. Role of melatonin in alleviating cold stress in Arabidopsis thaliana. J. Pineal Res. 2014, 56, 238–245. [Google Scholar] [CrossRef] [PubMed]
- Yin, L.; Wang, P.; Li, M.; Ke, X.; Li, C.; Liang, D.; Wu, S.; Ma, X.; Li, C.; Zou, Y.; Ma, F. Exogenous melatonin improves Malus resistance to Marssonina apple blotch. J. Pineal Res. 2013, 54, 426–434. [Google Scholar] [CrossRef] [PubMed]
- Weeda, S.; Zhang, N.; Zhao, X.; Ndip, G.; Guo, Y.; Buck, G.A.; Fu, C.; Ren, S. Arabidopsis transcriptome analysis reveals key roles of melatonin in plant defense systems. PLoS ONE 2014, 9, e93462. [Google Scholar] [CrossRef] [PubMed]
- Zhao, H.B.; Xu, L.F.; Su, T.; Jiang, Y.; Hu, L.Y.; Ma, F.W. Melatonin regulates carbohydrate metabolism and defenses against Pseudomonas syringae pv. tomato DC3000 infection in Arabidopsis thaliana. J. Pineal Res. 2015, 59, 109–119. [Google Scholar] [CrossRef] [PubMed]
- Shi, H.T.; Qian, Y.Q.; Tan, D.X.; Reiter, R.J.; He, C.Z. Melatonin induces the transcripts of CBF/DREB1s and their involvement in both abiotic and biotic stresses in Arabidopsis. J. Pineal Res. 2015, 59, 334–342. [Google Scholar] [CrossRef] [PubMed]
- Shi, H.T.; Qian, Y.Q.; Tan, D.X.; Reiter, R.J.; Chan, Z.L.; He, C.Z. Melatonin induces nitric oxide and the potential mechanisms relate to innate immunity against bacterial pathogen infection in Arabidopsis. J. Pineal Res. 2015, 59, 102–108. [Google Scholar] [CrossRef] [PubMed]
- Arnao, M.B.; Hernández-Ruiz, J. Melatonin promotes adventitious-and lateral root regeneration in etiolated hypocotyls of Lupinusalbus L. J. Pineal Res. 2007, 42, 147–152. [Google Scholar] [CrossRef] [PubMed]
- Hernández-Ruiz, J.; Cano, A.; Arnao, M.B. Melatonin: A growth-stimulating compound present in lupin tissues. Planta 2004, 220, 140–144. [Google Scholar] [CrossRef] [PubMed]
- Hernández-Ruiz, J.; Cano, A.; Arnao, M.B. Melatonin acts as a growth-stimulating compound in some monocot species. J. Pineal Res. 2005, 39, 137–142. [Google Scholar] [CrossRef] [PubMed]
- Kolář, J.; Johnson, C.H.; Macháčková, I. Exogenously applied melatonin (N-acetyl-5-methoxytryptamine) affects flowering of the short-day plant Chenopodium rubrum. Physiol. Plant. 2003, 118, 605–612. [Google Scholar] [CrossRef]
- Chen, Q.; Qi, W.B.; Reiter, R.J.; Wei, W.; Wang, B.M. Exogenously applied melatonin stimulates root growth and raises endogenous indoleacetic acid in roots of etiolated seedlings of etiolated seedlings of Brassica juncea. J. Plant Physiol. 2009, 166, 324–328. [Google Scholar] [CrossRef] [PubMed]
- Wang, P.; Sun, X.; Li, C.; Wei, Z.; Liang, D.; Ma, F. Long-term exogenous application of melatonin delays drought-induced leaf senescence in apple. J. Pineal Res. 2013, 54, 292–302. [Google Scholar] [CrossRef] [PubMed]
- Zhang, N.; Sun, Q.; Zhang, H.; Cao, Y.; Weeda, S.; Ren, S.; Guo, Y.D. Roles of melatonin in abiotic stress resistance in plants. J. Exp. Bot. 2015, 66, 647–656. [Google Scholar] [CrossRef] [PubMed]
- Hardeland, R. Melatonin in plants and other phototrophs: Advances and gaps concerning the diversity of functions. J. Exp. Bot. 2015, 66, 627–646. [Google Scholar] [CrossRef] [PubMed]
- Zuo, B.X.; Zheng, X.D.; He, P.L.; Wang, L.; Lei, Q.; Zhou, J.; Li, Q.; Han, Z.; Kong, J. Overexpression of MzASMT improves melatonin production and enhances drought tolerance in transgenic Arabidopsis thaliana plants. J. Pineal Res. 2014, 57, 408–417. [Google Scholar] [CrossRef] [PubMed]
- Kang, K.; Kong, K.; Park, S.; Natsagdorj, U.; Kim, Y.S.; Back, K. Molecular cloning of a plant N-acetylserotonin methyltransferase and its expression characteristics in rice. J. Pineal Res. 2011, 50, 304–309. [Google Scholar] [CrossRef] [PubMed]
- Park, S.; Byeon, Y.; Back, K. Functional analyses of three ASMT gene family members in rice plants. J. Pineal Res. 2013, 55, 409–415. [Google Scholar] [CrossRef] [PubMed]
- Wang, L.; Zhao, Y.; Reiter, R.J.; He, C.; Liu, G.; Lei, Q.; Zhou, J.; Li, Q.; Han, Z.; Kong, J. Changes in melatonin levels in transgenic ‘Micro-Tom’ tomato overexpressing ovine AANAT and ovine HIOMT genes. J. Pineal Res. 2014, 56, 134–142. [Google Scholar] [CrossRef] [PubMed]
- Sun, Q.Q.; Zhang, N.; Wang, J.F.; Zhang, H.J.; Li, D.B.; Shi, J.; Li, R.; Weeda, S.; Zhao, B.; Ren, S.X.; et al. Melatonin promotes ripening and improves quality of tomato fruit during postharvest life. J. Exp. Bot. 2015, 66, 657–668. [Google Scholar] [CrossRef] [PubMed]
- Arnao, M.B.; Hernández-Ruiz, J. Growth conditions influence the melatonin content of tomato plants. Food Chem. 2013, 138, 1212–1214. [Google Scholar] [CrossRef] [PubMed]
- Xu, W.; Cai, S.Y.; Zhang, Y.; Wang, Y.; Ahammed, G.J.; Xia, X.J.; Shi, K.; Zhou, Y.H.; Yu, J.Q.; Reiter, R.J.; et al. Melatonin enhances thermotolerance by promoting cellular protein protection in tomato plants. J. Pineal Res. 2016, 61, 457–469. [Google Scholar] [CrossRef] [PubMed]
- Hasan, M.K.; Ahammed, G.J.; Yin, L.; Shi, K.; Xia, X.; Zhou, Y.; Yu, J.; Zhou, J. Melatonin mitigates cadmium phytotoxicity through modulation of phytochelatins biosynthesis, vacuolar sequestration, and antioxidant potential in Solanum lycopersicum L. Front. Plant Sci. 2015, 6, 601. [Google Scholar] [CrossRef] [PubMed]
- Li, M.; Hasan, M.K.; Li, C.; Ahammed, G.J.; Xia, X.; Shi, K.; Zhou, Y.; Reiter, R.J.; Yu, J.; Xu, M.; et al. Melatonin mediates selenium-induced tolerance to cadmium stress in tomato plants. J. Pineal Res. 2016, 61, 291–302. [Google Scholar] [CrossRef] [PubMed]
- Wang, Y.; Cai, S.; Yin, L.; Shi, K.; Xia, X.; Zhou, Y.; Yu, J.; Zhou, J. Tomato HsfA1a plays a critical role in plant drought tolerance by activating ATG genes and inducing autophagy. Autophagy 2015, 11, 2033–2047. [Google Scholar] [CrossRef] [PubMed]
- Cai, S.; Zhang, Y.; Xu, Y.; Qi, Z.; Li, M.; Ahammed, G.J.; Xia, X.; Shi, K.; Zhou, Y.; Reiter, R.J.; et al. HsfA1a upregulates melatonin biosynthesis to confer cadmium tolerance in tomato plants. J. Pineal Res. 2017, 62, e12387. [Google Scholar] [CrossRef] [PubMed]
- Byeon, Y.; Lee, H.J.; Lee, H.Y.; Back, K. Cloning and functional characterization of the Arabidopsis N-acetylserotonin O-methyltransferase responsible for melatonin synthesis. J. Pineal Res. 2016, 60, 65–73. [Google Scholar] [CrossRef] [PubMed]
- Moore, R.C.; Purugganan, M.D. The early stages of duplicate gene evolution. Proc. Natl. Acad. Sci. USA 2003, 100, 15682–15687. [Google Scholar] [CrossRef] [PubMed]
- Cannon, S.B.; Mitra, A.; Baumgarten, A.; Young, N.D.; May, G. The roles of segmental and tandem gene duplication in the evolution of large gene families in Arabidopsis thaliana. BMC Plant Biol. 2004, 4, 10. [Google Scholar] [CrossRef] [PubMed]
- Byeon, Y.; Back, K. An increase in melatonin in transgenic rice causes pleiotropic phenotypes, including enhanced seedling growth, delayed flowering, and low grain yield. J. Pineal Res. 2014, 56, 408–414. [Google Scholar] [CrossRef] [PubMed]
- Park, S.; Byeon, Y.; Kim, Y.S.; Back, K. Kinetic analysis if purified recombinant rice N-acetylserotonin methyltransferase and peak melatonin production in etiolated rice shoots. J. Pineal Res. 2013, 54, 139–144. [Google Scholar] [CrossRef] [PubMed]
- Wei, K.F.; Wang, Y.M.; Xie, D.X. Identification and expression profile analysis of the protein kinase gene superfamily in maize development. Mol. Breed. 2014, 33, 155–172. [Google Scholar] [CrossRef]
- Larkin, M.A.; Blackshields, G.; Brown, N.P.; Chenna, R.; McGettigan, P.A.; McWilliam, H.; Valentin, F.; Wallace, I.M.; Wilm, A.; Lopez, R.; et al. Clustal W and Clustal X version 2.0. Bioinformatics 2007, 23, 2947–2948. [Google Scholar] [CrossRef] [PubMed]
- Lange, B.M.; Ghassemian, M. Genome organization in Arabidopsis thaliana: A survey for genes involved in isoprenoid and chlorophyll metabolism. Plant Mol. Biol. 2003, 51, 925–948. [Google Scholar] [CrossRef] [PubMed]
- Saitou, N.; Nei, M. The neighbor-joining method: A new method for reconstructing phylogenetic trees. Mol. Biol. Evol. 1987, 4, 406–425. [Google Scholar] [PubMed]
- Tamura, K.; Peterson, D.; Peterson, N.; Stecher, G.; Nei, M.; Kumar, S. MEGA5: Molecular evolutionary genetics analysis using maximum likelihood, evolutionary distance, and maximum parsimony methods. Mol. Biol. Evol. 2011, 28, 2731–2739. [Google Scholar] [CrossRef] [PubMed]
- Wang, Z.; Gerstein, M.; Snyder, M. RNA-Seq: A revolutionary tool for transcriptomics. Nat. Rev. Genet. 2009, 10, 57–63. [Google Scholar] [CrossRef] [PubMed]
- Saeed, A.I.; Sharov, V.; White, J.; Li, J.; Liang, W.; Bhagabati, N.; Braisted, J.; Klapa, M.; Currier, T.; Thiagarajan, M.; et al. TM4: A free, open-source system for microarray data management and analysis. BioTechniques 2003, 34, 374–378. [Google Scholar] [PubMed]
- Expósito-Rodríguez, M.; Borges, A.A.; Borges-Pérez, A.; Pérez, J.A. Selection of internal control genes for quantitative real-time RT-PCR studies during tomato development process. BMC Plant Biol. 2008, 8, 131. [Google Scholar] [CrossRef] [PubMed]
Sample Availability: Not available. |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).