Characterization, Function, and Transcriptional Profiling Analysis of 3-Hydroxy-3-methylglutaryl-CoA Synthase Gene (GbHMGS1) towards Stresses and Exogenous Hormone Treatments in Ginkgo biloba
Abstract
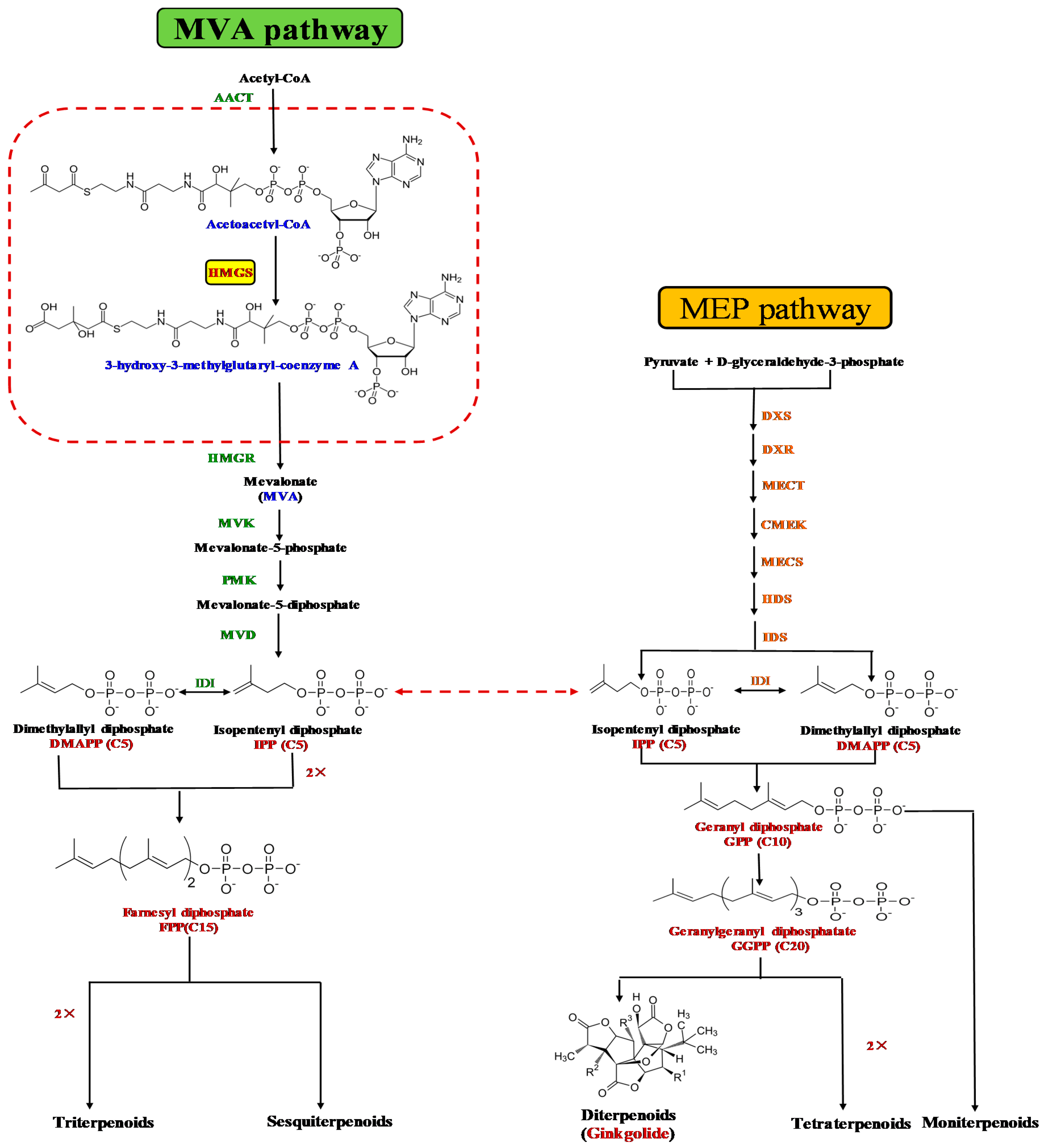
:1. Introduction
2. Results
2.1. Identification and Characterization of GbHMGS1
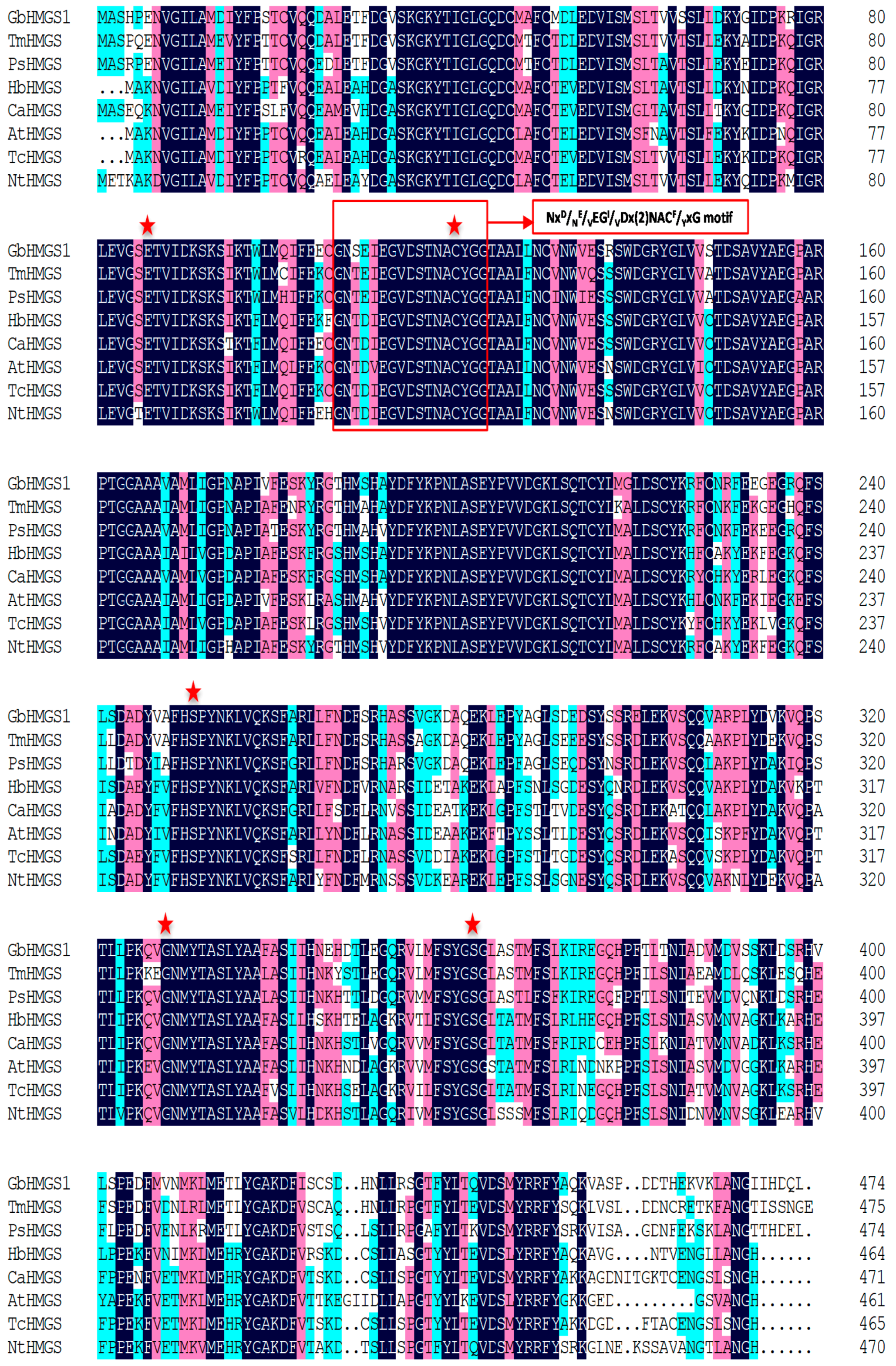
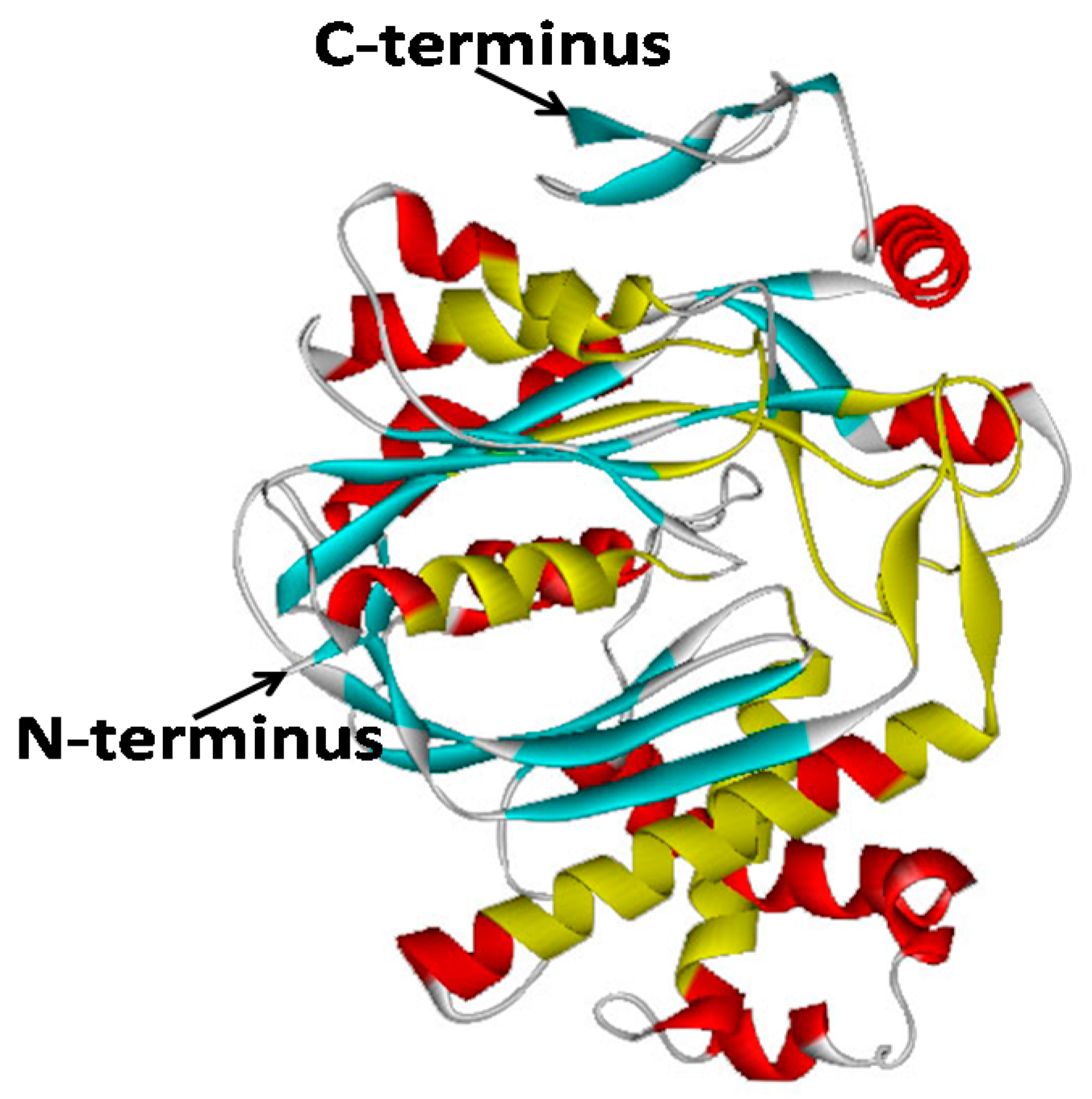
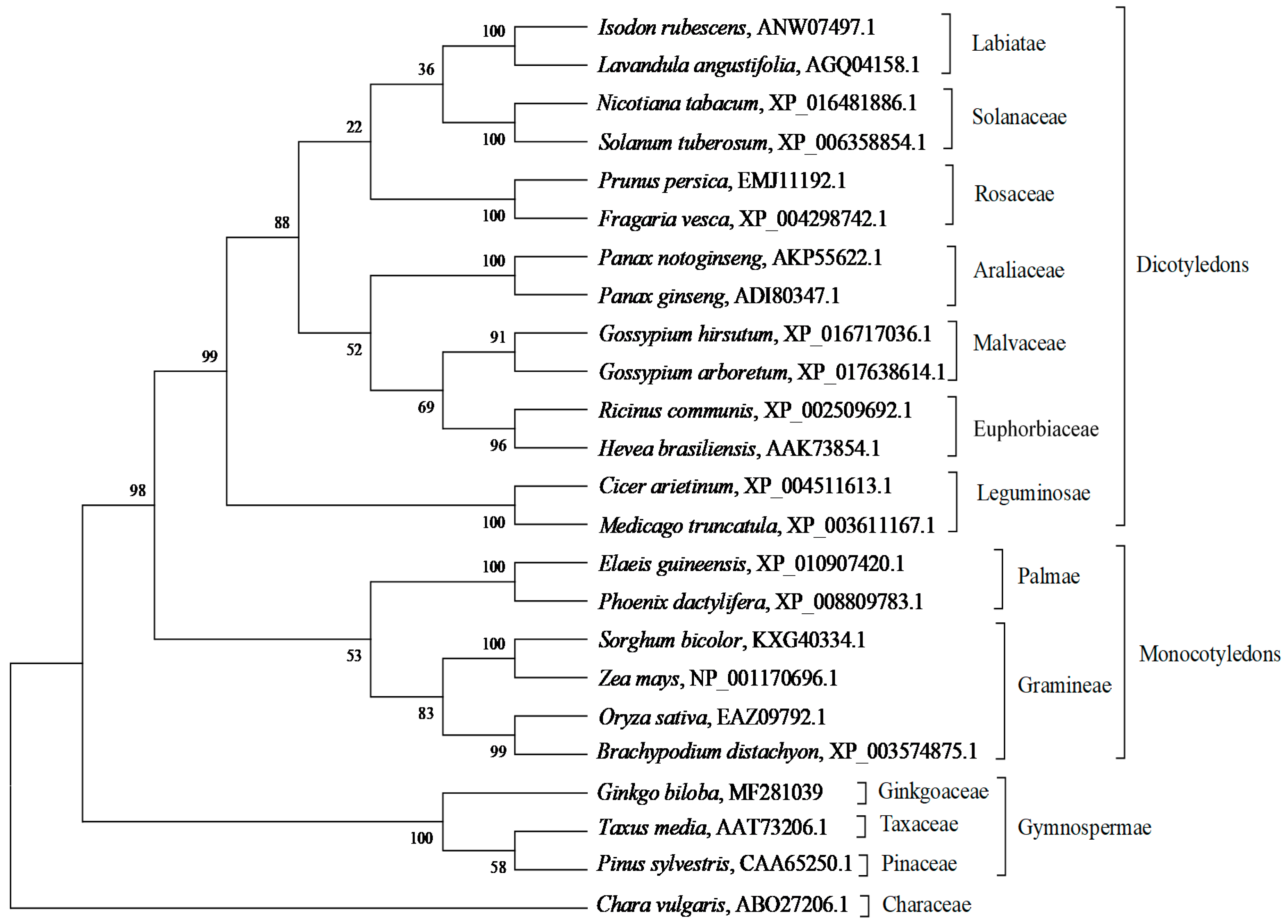
2.2. Bioinformatic Analysis of GbHMGS1 Protein
2.3. Prokaryotic Expression and In Vitro Enzyme Activity Analysis of GbHMGS1
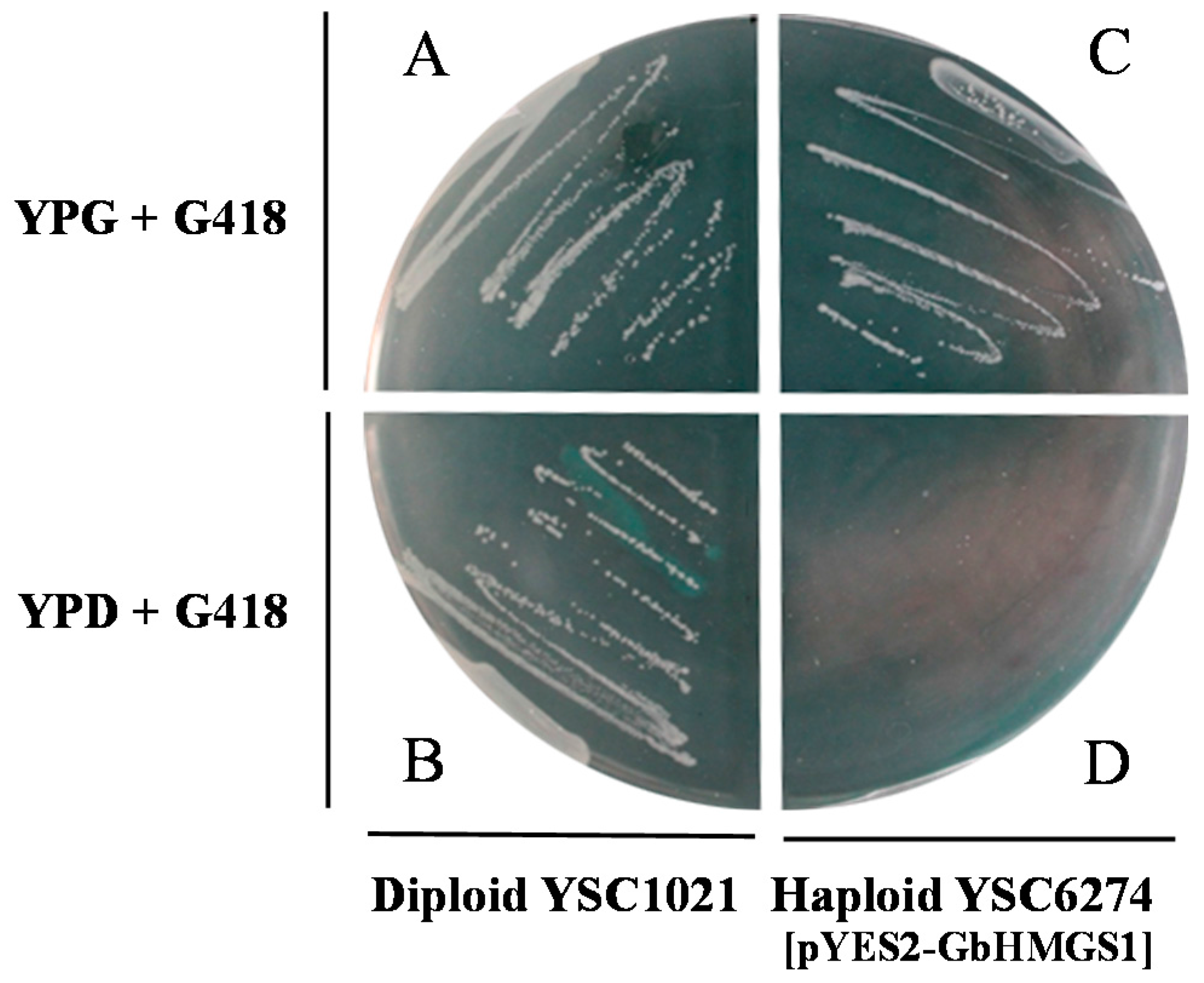
2.4. Functional Complementation of GbHMGS1 in Saccharomyces cerevisiae
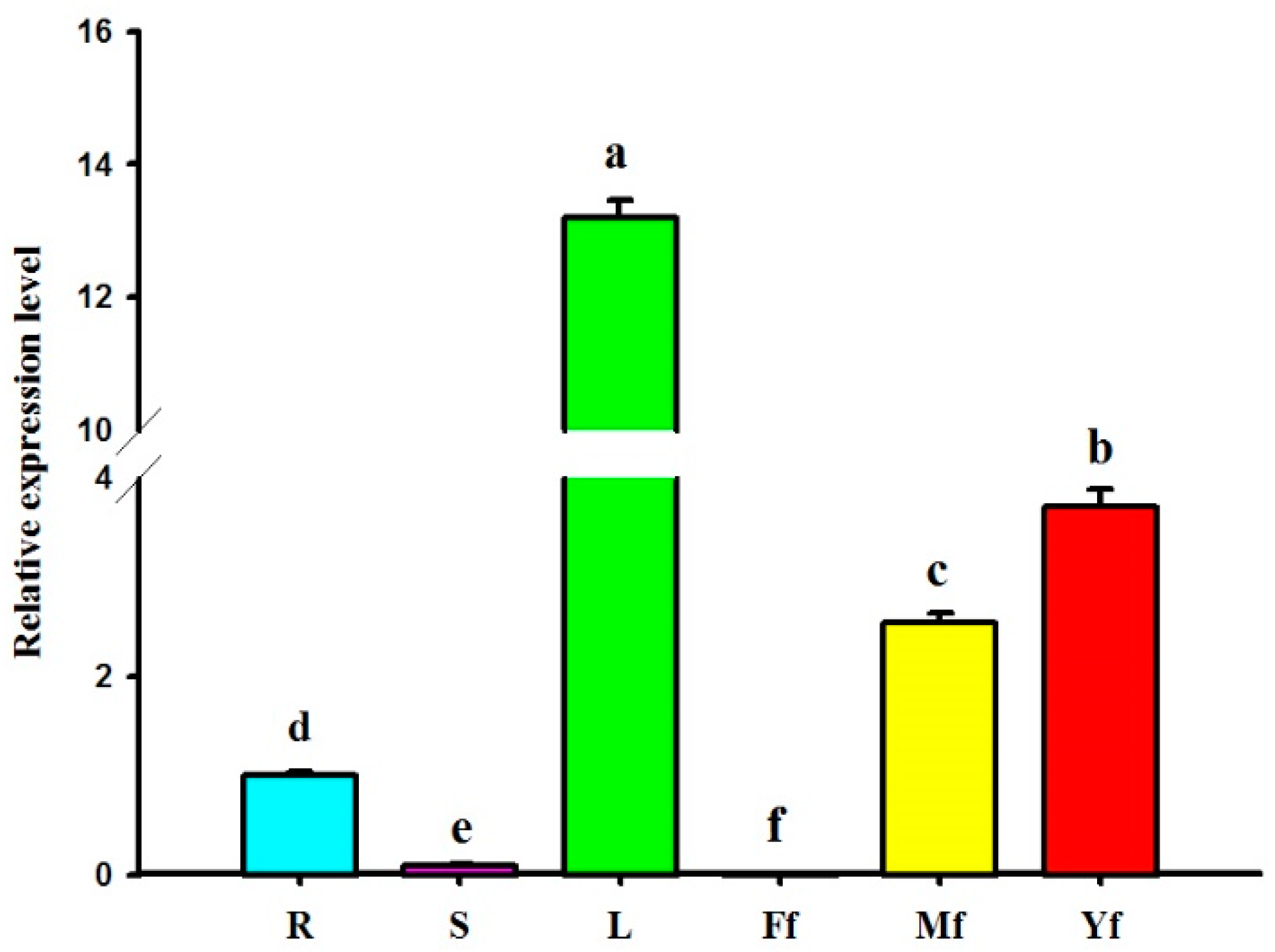
2.5. Differential Expression Profiles of GbHMGS1 in Various Tissues of G. biloba
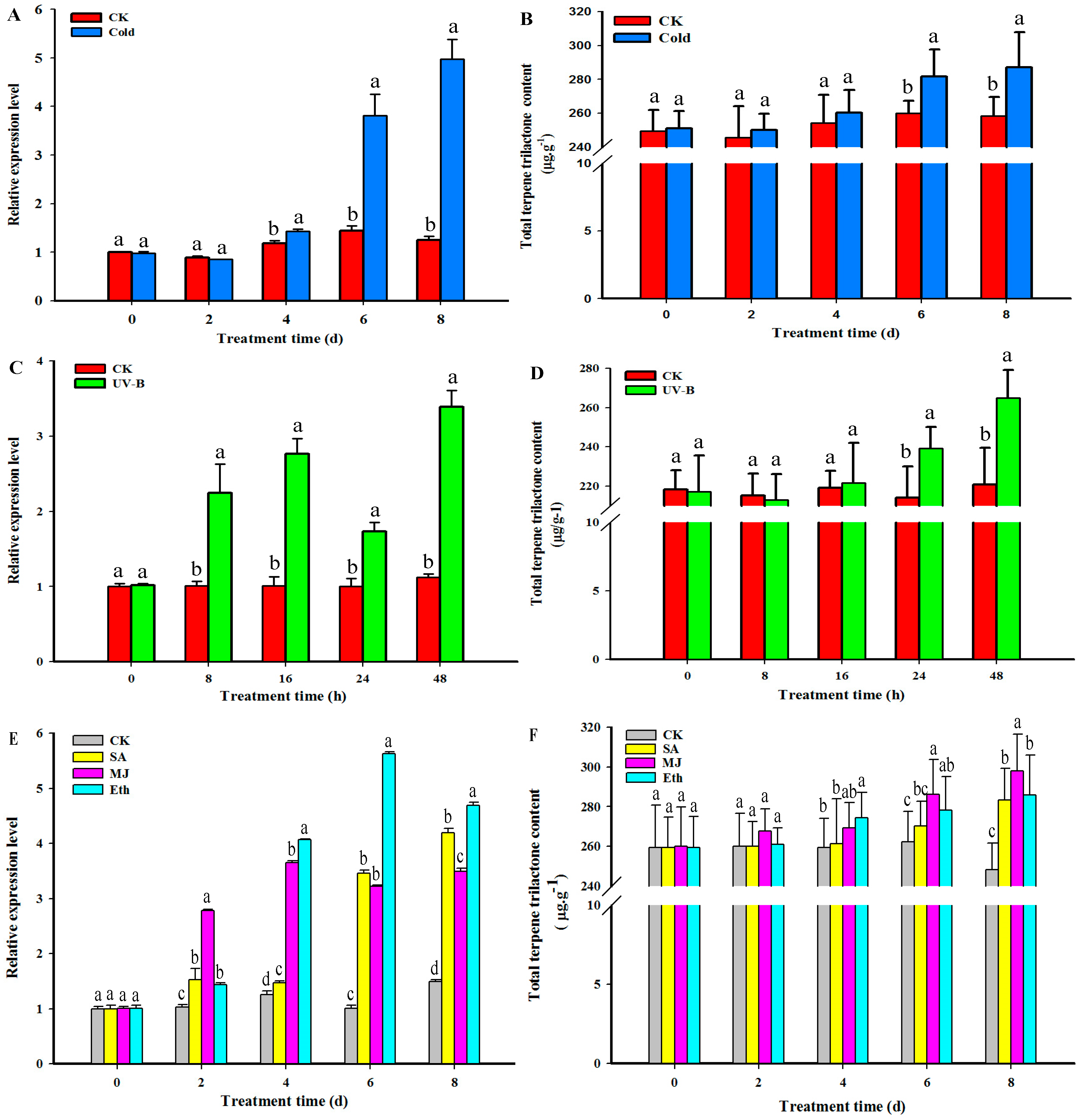
2.6. GbHMGS1 Transcript Level and TTL Content Changes in G. biloba under the Induction of UV-B, Cold, MJ, Eth, SA Elicitor
3. Discussion
3.1. GbHMGS1 is a Member of HMGS Gene Family
3.2. Expression Pattern of GbHMGS1 in Different Tissues of G. biloba
3.3. Transcript Level of GbHMGS1 and TTL Content Changes under Different Elicitor Treatments in Ginkgo Seedlings
4. Materials and Methods
4.1. Plant Materials and Multiple Stress Treatments
4.2. Cloning of the Full-length cDNA of GbHMGS1
4.3. Bioinformatics and Molecular Evolution Analysis
4.4. Prokaryotic Expression and In Vitro Enzyme Activity Analysis of GbHMGS1
4.5. Functional Complementation of GbHMGS1 in Yeast
4.6. Quantitative RT-PCR Analysis of GbHMGS1 Expression Levels
4.7. Determination of TTL Content in Ginkgo under Induction of Abiotic Stresses
4.8. Statistical Analysis
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Carrier, D.J.; van Beek, T.A.; van der Heijden, R.; Verpoorte, R. Distribution of ginkgolides and terpenoid biosynthetic activity in Ginkgo biloba. Phytochemistry 1998, 48, 89–92. [Google Scholar] [CrossRef]
- Mohanta, T.K.; Tamboli, Y.T.P.K. Phytochemical and medicinal importance of Ginkgo biloba L. Nat. Prod. Res. 2004, 28, 746–752. [Google Scholar] [CrossRef] [PubMed]
- Liao, H.J.; Zheng, Y.F.; Li, H.Y.; Peng, G.P. Two new ginkgolides from the leaves of Ginkgo biloba. Planta Med. 2011, 77, 1818–1821. [Google Scholar] [CrossRef] [PubMed]
- Nakanishi, K. Terpene trilactones from Gingko biloba: From ancient times to the 21st century. Bioorgan. Med. Chem. 2005, 13, 4987–5000. [Google Scholar] [CrossRef] [PubMed]
- Diamond, B.J.; Bailey, M.R. Ginkgo biloba: Indications, mechanisms, and safety. Psychiatr. Clin. North. Am. 2013, 36, 73–83. [Google Scholar] [CrossRef] [PubMed]
- Serranogarcia, N.; Pedrazachaverri, J.; Maressamano, J.J.; Orozcoibarra, M.; Cruzsalgado, A.; Jiménezanguiano, A.; Sotelo, J.; Trejosolís, C. Antiapoptotic effects of EGb 761. Evid-based Compl. Alt. 2013, 2013, 495703. [Google Scholar]
- Beek, T.A.V.; Montoro, P.; Xie, P.S.; Van Beek, T.A. Chemical analysis and quality control of Ginkgo biloba leaves, extracts, and phytopharmaceuticals. J. Chromatogr. A 2009, 1216, 2002–2032. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.M.; Kim, Y.B.; Kuzuyama, T.; Kim, S. Two copies of 4-(cytidine 50-diphospho)-2-C-methyl-d-erythritol kinase (CMK) gene in Ginkgo biloba: Molecular cloning and functional characterization. Planta 2008, 228, 941–950. [Google Scholar] [CrossRef] [PubMed]
- Kim, J.H.; Lee, K.I.; Chang, Y.J.; Kim, S.U. Developmental pattern of Ginkgo biloba levopimaradiene synthase (GbLPS) as probed by promoter analysis in Arabidopsis thaliana. Plant Cell Rep. 2012, 31, 1119–1127. [Google Scholar] [CrossRef] [PubMed]
- Liao, Y.L.; Xu, F.; Huang, X.H.; Zhang, W.W.; Cheng, H.; Wang, X.H.; Cheng, S.Y.; Shen, Y.B. Characterization and transcriptional profiling of Ginkgo biloba mevalonate diphosphate decarboxylase gene (GbMVD) promoter towards light and exogenous hormone treatments. Plant Mol. Biol. Rep. 2016, 34, 566–581. [Google Scholar] [CrossRef]
- Tholl, D. Terpene synthases and the regulation, diversity and biological roles of terpene metabolism. Curr. Opin. Plant Biol. 2006, 9, 297–304. [Google Scholar] [CrossRef] [PubMed]
- Chadwick, M.; Trewin, H.; Gawthrop, F.; Wagstaff, C. Sesquiterpenoids lactones: Benefits to plants and people. Int. J. Mol. Sci. 2013, 14, 12780–12805. [Google Scholar] [CrossRef] [PubMed]
- Shen, G.A.; Pang, Y.Z.; Wu, W.S.; Liao, Z.H.; Zhao, L.X.; Sun, X.F.; Tang, K.X. Cloning and characterization of a root-specific expressing gene encoding 3-hydroxy-3-methylglutaryl coenzyme a reductase from Ginkgo biloba. Mol. Biol. Rep. 2006, 33, 117–127. [Google Scholar] [CrossRef] [PubMed]
- Schwarz, M.; Arigoni, D. Ginkgolide biosynthesis. Compr. Nat. Prod. Chem. 1999, 2, 367–399. [Google Scholar]
- Hemmerlin, A.; Hoeffler, J.F.; Meyer, O.; Tritsch, D.; Kagan, I.A.; Grosdemange-Billiard, C.; Rohmer, M.; Bach, T.J. Cross-talk between the cytosolic mevalonate and the plastidial alonate and the plastidial methylerythritol phosphate pathways in tobacco bright yellow-2 cells. J. Biol. Chem. 2003, 278, 26666–26676. [Google Scholar] [CrossRef] [PubMed]
- Patra, B.; Schluttenhofer, C.; Wu, Y.; Pattanaik, S.; Yuan, L. Transcriptional regulation of secondary metabolite biosynthesis in plants. BBA-Gene Regul. Mech. 2013, 1829, 1236–1247. [Google Scholar] [CrossRef] [PubMed]
- Vranová, E.; Coman, D.; Gruissem, W. Network analysis of the MVA and MEP pathways for isoprenoid synthesis. Annu. Rev. Plant Biol. 2013, 64, 665–700. [Google Scholar] [CrossRef] [PubMed]
- Suvachittanont, W.; Wititsuwannakul, R. 3-Hydroxy 3-methylglutaryl-CoA synthase in Hevea brasiliensis. Phytochemistry 1995, 40, 757–761. [Google Scholar] [CrossRef]
- Ren, A.; Ouyang, X.; Shi, L.; Jiang, A.L.; Mu, D.S.; Li, M.J.; Han, Q.; Zhao, M.W. Molecular characterization and expression analysis of GlHMGS, a gene encoding hydroxymethylglutaryl-CoA synthase from Ganoderma lucidum (Ling-zhi) in ganoderic acid biosynthesis pathway. World J. Microb. Biot. 2013, 29, 523–531. [Google Scholar] [CrossRef] [PubMed]
- Wang, H.; Nagegowda, D.A.; Rawat, R.; Bouvier-Nave, P.; Guo, D.; Bach, T.J.; Chye, M.L. Overexpression of Brassica juncea wildtype and mutant HMG-CoA synthase 1 in Arabidopsis up-regulates genes in sterol biosynthesis and enhances sterol production and stress tolerance. Plant Biotechnol. J. 2012, 10, 31–42. [Google Scholar] [CrossRef] [PubMed]
- Parveen, I.; Wang, M.; Zhao, J.P.; Chittiboyina, A.G.; Tabanca, N.; Ali, A.; Baerson, S.R.; Techen, N.; Chappell, J.; Khan, I.A.; et al. Investigating sesquiterpene biosynthesis in Ginkgo biloba: Molecular cloning and functional characterization of (E,E)-farnesol and α-bisabolene synthases. Plant Mol. Biol. 2015, 89, 451–462. [Google Scholar] [CrossRef] [PubMed]
- Gong, Y.F.; Liao, Z.H.; Guo, B.H.; Sun, X.F.; Tang, K.X. Molecular cloning and expression profile analysis of Ginkgo biloba DXS gene encoding 1-deoxy-d-xylulose 5-phosphate synthase, the first committed enzyme of the 2-C-methyl-d-erythritol 4-phosphate pathway. Planta Med. 2006, 72, 329–335. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.M.; Kuzuyama, T.; Chang, Y.J.; Song, K.S.; Kim, S.U. Identification of class 2 1-deoxy-d-xylulose 5-phosphate synthase and 1-deoxy-d-xylulose 5-phosphate reductoisomerase genes from Ginkgo biloba and their transcription in embryo culture with respect to ginkgolide biosynthesis. Planta Med. 2006, 72, 234–240. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.M.; Kuzuyama, T.; Chang, Y.J.; Kwon, H.J.; Kim, S.U. Cloning and functional characterization of 2-C-methyl-d-erythritol 4-phosphate cytidyltransferase (GbMECT) gene from Ginkgo biloba. Phytochemistry 2006, 67, 1435–1441. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.M.; Zuyama, T.; Chang, Y.J.; Kim, S.U. Cloning and characterization of 2-C-methyl-d-erythritol 2,4-cyclodiphosphate synthase (MECS) gene from Ginkgo biloba. Plant Cell Rep. 2006, 25, 829–835. [Google Scholar] [CrossRef] [PubMed]
- Kim, S.M.; Kim, S.U. Characterization of 1-hydroxy-2-methyl-2-(E)-butenyl-4-diphosphate synthase (HDS) gene from Ginkgo biloba. Mol. Biol. Rep. 2010, 37, 973–979. [Google Scholar] [CrossRef]
- Kim, S.M.; Kuzuyama, T.; Kobayashi, A.; Sando, T.; Chang, Y.J.; Kim, S.U. 1-Hydroxy-2-methyl-2-(E)-butenyl 4-diphosphate reductase (IDS) is encoded by multicopy genes in gymnosperms Ginkgo biloba and Pinus taeda. Planta 2008, 227, 287–298. [Google Scholar] [CrossRef] [PubMed]
- Lichtenthaler, H.K.; Rohmer, M.; Schwender, J. Two independent biochemical pathways for isopentenyl diphosphate and isoprenoid biosynthesis in higher plants. Physiol. Plantarum 1997, 101, 643–652. [Google Scholar] [CrossRef]
- Liao, Y.L.; Xu, F.; Huang, X.H.; Zhang, W.W.; Cheng, H.; Li, L.L.; Cheng, S.Y.; Shen, Y.B. Promoter analysis and transcriptional profiling of Ginkgo biloba 3-hydroxy-3-methylglutaryl coenzyme a reductase (GbHMGR) gene in abiotic stress responses. Not. Bot. Horti Agrobot. 2015, 43, 25–34. [Google Scholar] [CrossRef]
- Chen, Q.W.; Yan, J.P.; Meng, X.X.; Xu, F.; Zhang, W.W.; Liao, Y.L.; Q, J.W. Molecular cloning, characterization, and functional analysis of acetyl-CoA C-acetyltransferase and mevalonate kinase genes involved in terpene trilactone biosynthesis from Ginkgo biloba. Molecules 2017, 22, 74. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; Yan, X.; Wang, J.; Li, S.; Liao, P.; Kai, G. Molecular cloning and expression analysis of a new putative gene encoding 3-hydroxy-3-methylglutaryl-CoA synthase from Salvia miltiorrhiza. Acta Physiol. Plant 2011, 33, 953–961. [Google Scholar] [CrossRef]
- Kai, G.; Li, S.; Wang, W.; Lu, Y.; Wang, J.; Liao, P.; Cui, L. Molecular cloning and expression analysis of a gene encoding 3-hydroxy-3-methylglutaryl-CoA synthase from Camptotheca acuminata. Russ. J. Plant Physiol. 2013, 60, 131–138. [Google Scholar] [CrossRef]
- Liu, Y.J.; Zhao, Y.J.; Zhang, M.; Su, P.; Wang, X.J.; Zhang, X.N.; Gao, W.; Huang, L.Q. Cloning and characterisation of the gene encoding 3-hydroxy-3-methylglutaryl-CoA synthase in Tripterygium wilfordii. Molecules 2014, 19, 19696–19707. [Google Scholar] [CrossRef] [PubMed]
- Cheng, S.; Wang, X.; Xu, F.; Chen, Q.; Tao, T.; Lei, J.; Zhang, W.; Liao, Y.; Chang, J.; Li, X. Cloning, expression profiling and functional analysis of CnHMGS, a gene encoding 3-hydroxy-3-methylglutaryl coenzyme A synthase from Chamaemelum nobile. Molecules 2016, 21, 316. [Google Scholar] [CrossRef] [PubMed]
- Campobasso, N.; Patel, M.; Wilding, I.E.; Kallender, H.; Rosenberg, M.; Gwynn, M.N. Staphylococcus aureus 3-hydroxy-3-methylglutaryl-CoA synthase: Crystal structure and mechanism. J. Biol. Chem. 2004, 279, 44883–44888. [Google Scholar] [CrossRef] [PubMed]
- Schwede, T.; Kopp, J.; Guex, N.; Peitsch, M.C. SWISS-MODEL: An automated protein homology-modeling server. Nucleic Acids Res. 2003, 31, 3381–3385. [Google Scholar] [CrossRef] [PubMed]
- Theisen, M.J.; Misra, I.; Saadat, D.; Campobasso, N.; Miziorko, H.M.; Harrison, D.H. 3-hydroxy-3-methylglutaryl-CoA synthase intermediate complex observed in “real-time”. Proc. Natl. Acad. Sci. USA 2004, 101, 16442–16447. [Google Scholar] [CrossRef] [PubMed]
- Sirinupong, N.; Suwanmanee, P.; Doolittle, R.F.; Suvachitanont, W. Molecular cloning of a new cDNA and expression of 3-hydroxy-3-methylglutaryl-CoA synthase gene from Hevea brasiliensis. Planta 2005, 221, 502–512. [Google Scholar] [CrossRef] [PubMed]
- Chang, J.; Ning, Y.; Xu, F.; Cheng, S.; Li, X. Research advance of 3-hydroxy-3-methylglutaryl-coenzyme a synthase in plant isoprenoid biosynthesis. J. Anim. Plant Sci. 2015, 25, 1441–1450. [Google Scholar]
- Holton, T.A.; Brugliera, F.; Tanaka, Y. Cloning and expression of flavonol synthase from Petunia hybrida. Plant J. 1993, 4, 1003–1010. [Google Scholar] [CrossRef] [PubMed]
- Lin, G.; Lian, Y.; Ryu, J.; Sung, M.; Park, J.; Park, H.; Park, B.; Shin, J.; Lee, M.; Cheon, C. Expression and purification of His-tagged flavonol synthase of Camellia sinensis from Escherichia coli. Protein Expres. Purif. 2007, 55, 287–292. [Google Scholar] [CrossRef] [PubMed]
- Owens, D.K.; Alerding, A.B.; Crosby, K.C.; Bandara, A.B.; Westwood, J.H.; Winkel, B.S.J. Functional analysis of a predicted flavonol synthase gene family in Arabidopsis. Plant Physiol. 2008, 147, 1046–1061. [Google Scholar] [CrossRef] [PubMed]
- Misra, I.; Wang, C.Z.; Miziorko, H.M. The influence of conserved aromatic residues in 3-hydroxy-3-methylglutaryl-CoA synthase. J. Biol. Chem. 2003, 278, 26443–26449. [Google Scholar] [CrossRef] [PubMed]
- Cartayrade, A.; Neau, E.; Sohier, C.; Balz, J.P.; Carde, J.P.; Walter, J. Ginkgolide and bilobalide biosynthesis in Ginkgo biloba. 1. Sites of synthesis, translocation and accumulation of ginkgolides and bilobalide. Plant Physiol. Biochem. 1997, 35, 859–868. [Google Scholar]
- Lu, X.; Yang, H.; Liu, X.; Shen, O.; Wang, N.; Qi, L.W.; Li, P. Combining metabolic profiling and gene expression analysis to reveal the biosynthesis site and transport of ginkgolides in Ginkgo biloba L. Front. Plant Sci. 2017, 8, 872. [Google Scholar] [CrossRef] [PubMed]
- Alex, D.; Bach, T.J.; Chye, M.L. Expression of Brassica juncea 3-hydroxy-3-methylglutaryl CoA synthase is developmentally regulated and stress-responsive. Plant J. 2000, 22, 415–426. [Google Scholar] [CrossRef] [PubMed]
- Xu, F.; Cai, R.; Cheng, S.; Du, H.; Wang, Y.; Cheng, S. Molecular cloning, characterization and expression of phenylalanine ammonialyase gene from Ginkgo biloba. Afr. J. Biotechnol. 2008, 7, 721–729. [Google Scholar]
- Xu, F.; Cheng, H.; Cai, R.; Li, L.L.; Chang, J.; Zhu, J.; Zhang, F.X.; Chen, L.J.; Wang, Y.; Cheng, S.H.; et al. Molecular cloning and function analysis of an anthocyanidin synthase gene from Ginkgo biloba, and its expression in abiotic stress responses. Mol. Cells 2008, 26, 536–547. [Google Scholar] [PubMed]
- Xu, F.; Li, L.; Zhang, W.; Cheng, H.; Sun, N.; Cheng, S.; Wang, Y. Isolation, characterization, and function analysis of a flavonol synthase gene from Ginkgo biloba. Mol. Biol. Rep. 2012, 39, 2285–2296. [Google Scholar] [CrossRef] [PubMed]
- Cheng, H.; Li, L.; Xu, F.; Cheng, S.; Cao, F.; Wang, Y.; Yuan, H.; Jiang, D.; Wu, C. Expression patterns of a cinnamyl alcohol dehydrogenase gene involved in lignin biosynthesis and environmental stress in Ginkgo biloba. Mol. Biol. Rep. 2013, 40, 707–721. [Google Scholar] [CrossRef] [PubMed]
- Cheng, H.; Li, L.; Cheng, S.; Cao, F.; Wang, Y.; Yuan, H. Molecular cloning and function assay of a chalcone isomerase gene (GbCHI) from Ginkgo biloba. Plant Cell Rep. 2011, 30, 49–62. [Google Scholar] [CrossRef] [PubMed]
- Robert-Seilaniantz, A.; Grant, M.; Jones, J.D.G. Hormone crosstalk in plant disease and defense: More than just jasmonate-salicylate antagonism. Annu. Rev. Phytopathol. 2011, 49, 317–343. [Google Scholar] [CrossRef] [PubMed]
- Pauwels, L.; Morreel, K.; Witte, E.D.; Lammertyn, F.; Montagu, M.V.; Boerjan, W.; Inzée, D.; Goossens, A. Mapping methyl jasmonate-mediated transcriptional reprogramming of metabolism and cell cycle progression in cultured Arabidopsis cells. Proc. Natl. Acad. Sci. USA 2008, 105, 1380–1385. [Google Scholar] [CrossRef] [PubMed]
- Dong, J.; Wan, G.; Liang, Z. Accumulation of salicylic acid-induced phenolic compounds and raised activities of secondary metabolic and antioxidative enzymes in Salvia miltiorrhiza cell culture. J. Biotechnol. 2010, 148, 99–104. [Google Scholar] [CrossRef] [PubMed]
- Gaffney, T.; Friedrich, L.; Vernooij, B.; Negrotto, D.; Nye, D.; Uknes, S.; Ward, E.; Kessmann, H.; Ryals, J. Requirement of salicylic acid for the induction of systemic acquired resistance. Science 1993, 261, 754–756. [Google Scholar] [CrossRef] [PubMed]
- Kang, S.M.; Min, J.Y.; Kim, Y.D. Effects of methyl jasmonate and salicylic acid on the production of bilobalide and ginkgolides in cell cultures of Ginkgo biloba. In Vitro Cell. Dev. Biol. Plant 2006, 42, 44–49. [Google Scholar] [CrossRef]
- Kang, M.K.; Nargis, S.; Kim, S.M.; Kim, S.U. Distinct expression patterns of two Ginkgo biloba, 1-hydroxy-2-methyl-2-(E)-butenyl-4-diphosphate reductase/isopentenyl diphospahte synthase (HDR/IDS) promoters in Arabidopsis model. Plant Physiol. Biochem. 2013, 62, 47–53. [Google Scholar] [CrossRef] [PubMed]
- Wang, Q.J.; Zheng, L.P.; Zhao, P.F.; Zhao, Y.L.; Wang, J.W. Cloning and characterization of an elicitor-responsive gene encoding 3-hydroxy-3-methylglutaryl coenzyme A reductase involved in 20-hydroxyecdysone production in cell cultures of Cyanotis arachnoidea. Plant Physiol. Biochem. 2014, 84, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Martin, D.; Tholl, D.; Gershenzon, J.; Bohlmann, J. Methyl jasmonate induces traumatic resin ducts, terpenoid resin biosynthesis, and terpenoid accumulation in developing xylem of Norway spruce stems. Plant Physiol. 2002, 129, 1003–1018. [Google Scholar] [CrossRef] [PubMed]
- Keeling, C.I.; Bohlmann, J. Diterpene resin acids in conifers. Phytochemistry 2006, 67, 2415–2423. [Google Scholar] [CrossRef] [PubMed]
- Dai, Z.; Cui, G.; Zhou, S.F.; Zhang, X.; Huang, L. Cloning and characterization of a novel 3-hydroxy-3-methylglutaryl coenzyme A reductase gene from Salvia miltiorrhiza involved in diterpenoid tanshinone accumulation. J. Plant Physiol. 2011, 168, 148–157. [Google Scholar] [CrossRef] [PubMed]
- Bae, K.H.; Choi, Y.; Shin, C.G.; Kim, Y.Y.; Kim, Y.S. Enhanced ginsenoside productivity by combination of ethephon and methyl jasmoante in ginseng (Panax ginseng C.A. Meyer) adventitious root cultures. Biotechnol. Lett. 2006, 28, 1163. [Google Scholar] [CrossRef] [PubMed]
- Huang, X.H. Molecular Cloning and Function Analysis of Promoters of the Terpene Lactones Synthase Key Gene from Ginkgo biloba, and Their Expressions in Abiotic Stress Responses. M.D. Thesis, Yangtze University, Jingzhou, China, 2013. [Google Scholar]
- Woolf, A.B.; Clemens, J.; Plummer, J.A. Leaf maturity and temperature affect the selective removal of floral buds from Camellia with ethephon. Laryngoscope 1995, 123, 2176–2179. [Google Scholar]
- Cai, R.; Xu, F.; Chen, L.; Cheng, S. Modification of total RNA isolation method from different Ginkgo biloba organs. Biotechnology 2007, 17, 38–41. [Google Scholar]
- Lowry, O.H.; Rosebrough, N.J.; Farr, A.L.; Randall, R.J. Protein measurement with the Folin phenol reagent. J. Biol. Chem. 1951, 193, 265. [Google Scholar] [PubMed]
- Xu, F.; Ning, Y.J.; Zhang, W.W.; Liao, Y.L.; Li, L.L.; Chengm, H.; Cheng, S.Y. An R2R3-MYB transcription factor as a negative regulator of the flavonoid biosynthesis pathway in Ginkgo biloba. Funct. Integr. Genom. 2014, 14, 177–189. [Google Scholar] [CrossRef] [PubMed]
- Schmittgen, T.D.; Livak, K.J. Analyzing real-time PCR data by the comparative CT method. Nat. Protoc. 2008, 3, 1101–1108. [Google Scholar] [CrossRef] [PubMed]
- Liao, Y.; Xu, F.; Zhu, J.; Wang, Y.; Cheng, S. Separation and determination of terpene trilactiones by gas chromatography with wide bore capillary column. Acta Agric. Boreali-occidentalis Sin. 2008, 17, 146–149. [Google Scholar]
Sample Availability: Samples of Ginkgo biloba and cDNAs of GbHMGS1 are available from the authors. |

| Enzyme | kcat (min−1) | Km (μM) | kcat/Km (μM−1·min−1) | Specific Activity (μmol·min−1·mg−1) | Vmax (nM·min−1) |
|---|---|---|---|---|---|
| No HMGS in pET32 | ND | ND | ND | 3.4 ± 0.5 | ND |
| GbHMGS1 | 195.4 ± 13.1 | 689 ± 32.6 | 0.28 ± 0.01 | 15.5 ± 4.8 | 37.1 ± 5.4 |
| HvHMGS | ND | 530 ± 50 | ND | 41.3 ± 2.9 | 130 ± 30 |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Meng, X.; Song, Q.; Ye, J.; Wang, L.; Xu, F. Characterization, Function, and Transcriptional Profiling Analysis of 3-Hydroxy-3-methylglutaryl-CoA Synthase Gene (GbHMGS1) towards Stresses and Exogenous Hormone Treatments in Ginkgo biloba. Molecules 2017, 22, 1706. https://doi.org/10.3390/molecules22101706
Meng X, Song Q, Ye J, Wang L, Xu F. Characterization, Function, and Transcriptional Profiling Analysis of 3-Hydroxy-3-methylglutaryl-CoA Synthase Gene (GbHMGS1) towards Stresses and Exogenous Hormone Treatments in Ginkgo biloba. Molecules. 2017; 22(10):1706. https://doi.org/10.3390/molecules22101706
Chicago/Turabian StyleMeng, Xiangxiang, Qiling Song, Jiabao Ye, Lanlan Wang, and Feng Xu. 2017. "Characterization, Function, and Transcriptional Profiling Analysis of 3-Hydroxy-3-methylglutaryl-CoA Synthase Gene (GbHMGS1) towards Stresses and Exogenous Hormone Treatments in Ginkgo biloba" Molecules 22, no. 10: 1706. https://doi.org/10.3390/molecules22101706
APA StyleMeng, X., Song, Q., Ye, J., Wang, L., & Xu, F. (2017). Characterization, Function, and Transcriptional Profiling Analysis of 3-Hydroxy-3-methylglutaryl-CoA Synthase Gene (GbHMGS1) towards Stresses and Exogenous Hormone Treatments in Ginkgo biloba. Molecules, 22(10), 1706. https://doi.org/10.3390/molecules22101706




