A Novel Insecticidal Peptide SLP1 Produced by Streptomyces laindensis H008 against Lipaphis erysimi
Abstract
:1. Introduction
2. Results
2.1. Microorganisms
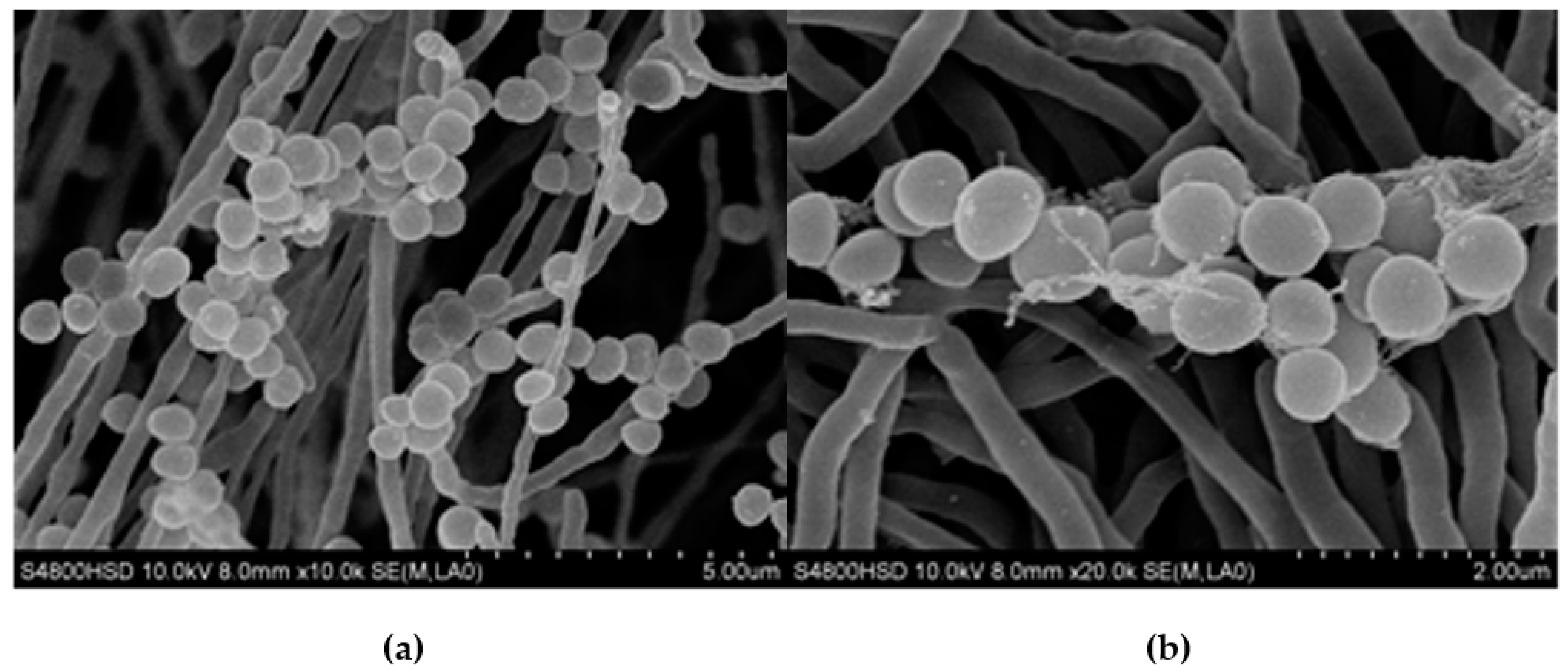
2.1.1. Morphological and Physiological Traits of Strain H008
2.1.2. Molecular Biological Identification of Strain H008
2.2. Insecticidal Activity
2.3. Purification and Identification of Insecticidal Peptide
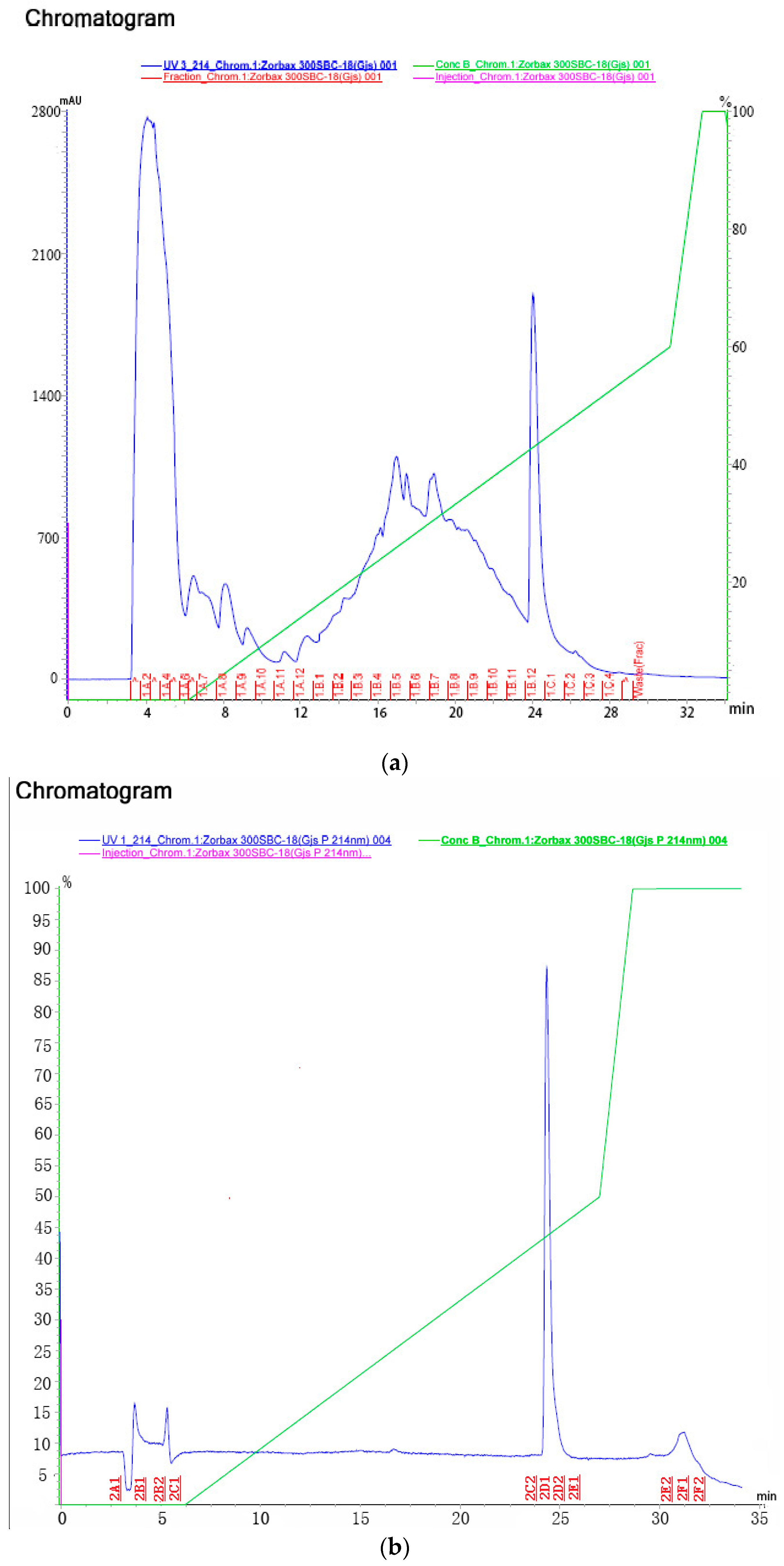
2.3.1. Purification
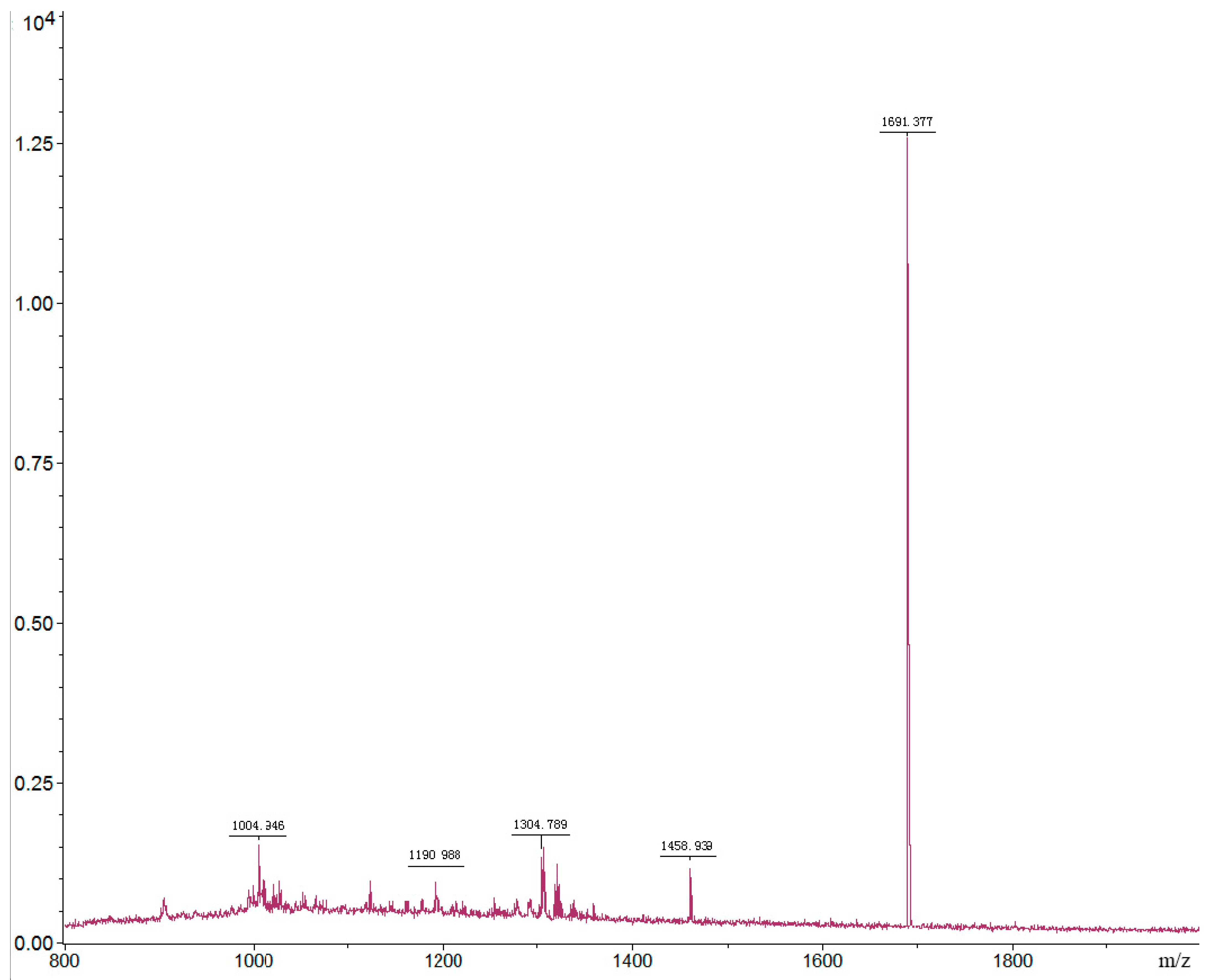
2.3.2. Identification
3. Discussion
4. Materials and Methods
4.1. Microorganism
4.1.1. Isolation
4.1.2. Morphological and Physiological Traits
4.1.3. Molecular Biological Identification
4.1.4. Fermentation
4.2. Insecticidal Activity
4.3. Purification and Identification of Insecticidal Peptide
5. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Barka, E.A.; Vatsa, P.; Sanchez, L.; Gaveau-Vaillant, N.; Jacquard, C.; Klenk, H.P.; Clement, C.; Ouhdouch, Y.; van Wezel, G.P. Taxonomy, Physiology, and Natural Products of Actinobacteria. Microbiol. Mol. Biol. Rev. 2015, 80, 1–43. [Google Scholar] [CrossRef] [PubMed]
- Katz, L.; Baltz, R.H. Natural product discovery: Past, present, and future. J. Ind. Microbiol. Biot. 2016, 43, 155–176. [Google Scholar] [CrossRef] [PubMed]
- Clardy, J.; Fischbach, M.A.; Walsh, C.T. New antibiotics from bacterial natural products. Nat. Biotechnol. 2006, 24, 1541–1550. [Google Scholar] [CrossRef] [PubMed]
- Hwang, K.S.; Kim, H.U.; Charusanti, P.; Palsson, B.O.; Lee, S.Y. Systems biology and biotechnology of Streptomyces species for the production of secondary metabolites. Biotechnol. Adv. 2014, 32, 255–268. [Google Scholar] [CrossRef] [PubMed]
- Blackman, R.L.; Eastop, V.E. Aphids on the World’s Crops. An Identification and Information Guide, 2nd ed.; John Wiley and Sons Ltd.: Chichester, UK, 2000. [Google Scholar]
- Koramutla, M.K.; Aminedi, R.; Bhattacharya, R. Comprehensive evaluation of candidate reference genes for qRT-PCR studies of gene expression in mustard aphid, Lipaphis erysimi (Kalt). Sci. Rep. 2016, 6, 25883. [Google Scholar] [CrossRef] [PubMed]
- Whetstone, P.A.; Hammock, B.D. Delivery methods for peptide and protein toxins in insect control. Toxicon 2007, 49, 576–596. [Google Scholar] [CrossRef] [PubMed]
- Lamberth, C. Naturally occurring amino acid derivatives with herbicidal, fungicidal or insecticidal activity. Amino Acids 2016, 48, 929–940. [Google Scholar] [CrossRef] [PubMed]
- Urushibata, I.; Isogai, A.; Matsumoto, S.; Suzuki, A. Respirantin, a Novel Insecticidal Cyclodepsipeptide from Streptomyces. J. Antibiot. 1993, 46, 701–703. [Google Scholar] [CrossRef] [PubMed]
- Ortiz-Urquiza, A.; Luo, Z.B.; Keyhani, N.O. Improving mycoinsecticides for insect biological control. Appl. Microbiol. Biot. 2015, 99, 1057–1068. [Google Scholar] [CrossRef] [PubMed]
- Tresner, H.D.; Hayes, J.A.; Backus, E.J. Differential tolerance of streptomycetes to sodium chloride as a taxonomic aid. Appl. Microbiol. 1968, 16, 1134–1136. [Google Scholar] [PubMed]
- Williams, S.T.; Goodfellow, M.; Alderson, G.; Wellington, E.M.; Sneath, P.H.; Sackin, M.J. Numerical classification of Streptomyces and related genera. J. Gen. Microbiol. 1983, 129, 1743–1813. [Google Scholar] [CrossRef] [PubMed]
- Kämpfer, P.; Kroppenstedt, R.M.; Dott, W. A numerical classification of the genera Streptomyces and Streptoverticillium using miniaturized physiological tests. Microbiology 1991, 137, 1831–1891. [Google Scholar] [CrossRef]
- Kuster, E. Simple Working Key for the Classification and Identification of Named Taxa Included in the International Streptomyces Project 1, 2. Int. J. Syst. Evol. Microbiol. 1972, 22, 139–148. [Google Scholar] [CrossRef]
- Shirling, E.B.; Gottlieb, D. Cooperative description of type cultures of Streptomyces. IV. Species descriptions from the second, third and fourth studies. Int. J. Syst. Evol. Microbiol. 1969, 19, 391–512. [Google Scholar] [CrossRef]
- Lewer, P.; Chapin, E.L.; Graupner, P.R.; Gilbert, J.R.; Peacock, C. Tartrolone C: A novel insecticidal macrodiolide produced by Streptomyces sp. CP1130. J. Nat. Prod. 2003, 66, 143–145. [Google Scholar] [CrossRef] [PubMed]
- Kaur, T.; Manhas, R.K. Antifungal, insecticidal, and plant growth promoting potential of Streptomyces hydrogenans DH16. J. Basic Microb. 2014, 54, 1175–1185. [Google Scholar] [CrossRef] [PubMed]
- Kaur, T.; Vasudev, A.; Sohal, S.K.; Manhas, R.K. Insecticidal and growth inhibitory potential of Streptomyces hydrogenans DH16 on major pest of India, Spodoptera litura (Fab.) (Lepidoptera: Noctuidae). BMC Microbiol. 2014, 14, 227. [Google Scholar] [CrossRef] [PubMed]
- Koul, O.; Singh, G.; Singh, R.; Walia, S.; Kaul, V.K. Comparative bioefficacy of biorational ethylene glycol diesters and sucrose octanoate against Lipaphis erysimi (Homoptera: Aphididae). J. Appl. Entomol. 2009, 133, 682–688. [Google Scholar] [CrossRef]
- Zeng, Y.; Zhang, Y.M.; Weng, Q.F.; Hu, M.Y.; Zhong, G.H. Cytotoxic and Insecticidal Activities of Derivatives of Harmine, a Natural Insecticidal Component Isolated from Peganum harmala. Molecules 2010, 15, 7775–7791. [Google Scholar] [CrossRef] [PubMed]
- Ge, Y.; Liu, P.P.; Yang, R.; Zhang, L.; Chen, H.X.; Camara, I.; Liu, Y.Q.; Shi, W.P. Insecticidal Constituents and Activity of Alkaloids from Cynanchum mongolicum. Molecules 2015, 20, 17483–17492. [Google Scholar] [CrossRef] [PubMed]
- Sarkar, P.; Jana, J.; Chatterjee, S.; Sikdar, S.R. Functional characterization of Rorippa indica defensin and its efficacy against Lipaphis erysimi. Springerplus 2016, 5, 511. [Google Scholar] [CrossRef] [PubMed]
- Banerjee, N.; Sengupta, S.; Roy, A.; Ghosh, P.; Das, K.; Das, S. Functional Alteration of a Dimeric Insecticidal Lectin to a Monomeric Antifungal Protein Correlated to Its Oligomeric Status. PLoS ONE 2011, 6, e18593. [Google Scholar] [CrossRef] [PubMed]
- Williams, S.; Cross, T. Chapter XI Actinomycetes. Meth. Microbiol. 1971, 4, 295–334. [Google Scholar]
- Shirling, E.B.; Gottlieb, D. Methods for characterization of Streptomyces species. Int. J. Syst. Bacteriol. 1966, 16, 313–340. [Google Scholar] [CrossRef]
- Bozzola, J.J.; Russell, L.D. Electron Microscopy: Principles and Techniques for Biologists, 1st ed.; Jones & Bartlett Learning: Woods Hole, MA, USA, 1999. [Google Scholar]
- Gottlieb, D. An Evaluation of Criteria and Procedures Used in the Description and Characterization of the Streptomycetes A Cooperative Study. Appl. Microbiol. 1961, 9, 55–65. [Google Scholar] [PubMed]
- Rainey, F.A.; Ward-Rainey, N.; Kroppenstedt, R.M.; Stackebrandt, E. The genus Nocardiopsis represents a phylogenetically coherent taxon and a distinct actinomycete lineage: proposal of Nocardiopsaceae fam. nov. Int. J. Syst. Evol. Microbiol. 1996, 46, 1088–1092. [Google Scholar] [CrossRef] [PubMed]
- Edwards, U.; Rogall, T.; Blöcker, H.; Emde, M.; Böttger, E.C. Isolation and direct complete nucleotide determination of entire genes. Characterization of a gene coding for 16S ribosomal RNA. Nucleic Acids Res. 1989, 17, 7843–7853. [Google Scholar] [CrossRef] [PubMed]
- Mehling, A.; Wehmeier, U.F.; Piepersberg, W. Nucleotide sequences of streptomycete 16S ribosomal DNA: Towards a specific identification system for streptomycetes using PCR. Microbiology 1995, 141, 2139–2147. [Google Scholar] [CrossRef] [PubMed]
- Tuntiwachwuttikul, P.; Taechowisan, T.; Wanbanjob, A.; Thadaniti, S.; Taylor, W.C. Lansai A–D, secondary metabolites from Streptomyces sp. SUC1. Tetrahedron 2008, 64, 7583–7586. [Google Scholar] [CrossRef]
- Jørgensen, H.; Fjærvik, E.; Hakvåg, S.; Bruheim, P.; Bredholt, H.; Klinkenberg, G.; Ellingsen, T.E.; Zotchev, S.B. Candicidin biosynthesis gene cluster is widely distributed among Streptomyces spp. isolated from the sediments and the neuston layer of the Trondheim fjord, Norway. Appl. Environ. Microbiol. 2009, 75, 3296–3303. [Google Scholar] [CrossRef] [PubMed]
- Kumar, S.; Stecher, G.; Tamura, K. MEGA7: Molecular Evolutionary Genetics Analysis version 7.0 for bigger datasets. Mol. Biol. Evol. 2016, 33, 1870–1874. [Google Scholar] [CrossRef] [PubMed]
- Tulla-Puche, J.; El-Faham, A.; Galanis, A.S.; de Oliveira, E.; Zompra, A.A.; Albericio, F. Methods for the Peptide Synthesis and Analysis. In Peptide Chemistry and Drug Design; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2015; pp. 11–73. [Google Scholar]
- Aguilar, M.-I. HPLC of Peptides and Proteins. In HPLC of Peptides and Proteins: Methods and Protocols; Springer New York: Totowa, NJ, USA, 2004; Volume 251, pp. 3–8. [Google Scholar]
- Rudewicz, P.J. Mass Spectrometry for the Study of Peptide Drug Metabolism. In Protein and Peptide Mass Spectrometry in Drug Discovery; John Wiley & Sons, Inc.: Hoboken, NJ, USA, 2011; Volume 15, pp. 435–447. [Google Scholar]
- Sample Availability: Samples of S. laindensis H008 is available from the authors.


| Culture Media | Growth | Aerial Mycelium | Substrate Mycelium | Soluble Pigment |
|---|---|---|---|---|
| Tryptone | Abundant | Trace (Pale gray) | Brown | None |
| Yeast malt agar (ISP 2) | Abundant | Gray | Dark brown | None |
| Oatmeal agar (ISP 3) | Moderate | Gray | Colorless | None |
| Inorganic salt starch agar (ISP 4) | Moderate | Mouse gray | Colorless | None |
| Glycerol asparagine agar (ISP 5) | Poor | Trace (Pale gray) | Light brown | None |
| Peptone yeast extract iron agar (ISP 6) | Abundant | Gray | Brown | Yellow brown |
| Tyrosine agar (ISP 7) | Moderate | Gray | Colorless | None |
| Malt extract agar | Abundant | Gray | Dark brown | None |
| Maltose tryptone agar | Abundant | Gray | Brown | None |
| Nutrient agar | Moderate | Mouse gray | Colorless | None |
| Test Items | Results | Utilization of Carbon Source | Results |
|---|---|---|---|
| Melanoid pigments | + | d-fructose | + |
| Production of H2S | + | d-glucose | + |
| Liquefaction of gelatin | + | d-galactose | + |
| Starch hydrolysis | + | d-mannitol | + |
| Coagulation of milk | − | d-xylose | + |
| Peptonization of milk | − | Inositol | + |
| Nitrate reduction | + | l-arabinose | − |
| Degradation of cellulose | − | l-rhamnose | − |
| Citrate utilization | + | Sucrose | + |
| Tolerance to NaCl | Up to 7% | Raffinose | − |
| Step | Fraction No. | Mortality | Adjust Mortality |
|---|---|---|---|
| Step 1 | 1A2-1A6 | 10% | 5.3% |
| 1A7 | 15% | 10.5% | |
| 1A8-1A9 | 10% | 5.3% | |
| 1A10 | 10% | 5.3% | |
| 1A11-1A12 | 15% | 10.5% | |
| 1B1-1B3 | 10% | 5.3% | |
| 1B4-1B6 | 5% | 0 | |
| 1B7-1B8 | 15% | 10.5% | |
| 1B9-1B11 | 5% | 0 | |
| 1B12-1C1 | 95% | 94.7% | |
| 1C2-1C4 | 10% | 5.3% | |
| Control | 5% | 0 | |
| Step 2 | 2A1 | 10% | 5.3% |
| 2B1 | 5% | 0 | |
| 2B2 | 10% | 5.3% | |
| 2C1-2C2 | 5% | 0 | |
| 2D1-2D2 | 100% | 100% | |
| 2E1-2E2 | 10% | 5.3% | |
| 2F1-2F2 | 15% | 10.5% | |
| Control | 5% | 0 |
© 2016 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC-BY) license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xu, L.; Liang, K.; Duan, B.; Yu, M.; Meng, W.; Wang, Q.; Yu, Q. A Novel Insecticidal Peptide SLP1 Produced by Streptomyces laindensis H008 against Lipaphis erysimi. Molecules 2016, 21, 1101. https://doi.org/10.3390/molecules21081101
Xu L, Liang K, Duan B, Yu M, Meng W, Wang Q, Yu Q. A Novel Insecticidal Peptide SLP1 Produced by Streptomyces laindensis H008 against Lipaphis erysimi. Molecules. 2016; 21(8):1101. https://doi.org/10.3390/molecules21081101
Chicago/Turabian StyleXu, Lijian, Kangkang Liang, Bensha Duan, Mengdi Yu, Wei Meng, Qinggui Wang, and Qiong Yu. 2016. "A Novel Insecticidal Peptide SLP1 Produced by Streptomyces laindensis H008 against Lipaphis erysimi" Molecules 21, no. 8: 1101. https://doi.org/10.3390/molecules21081101
APA StyleXu, L., Liang, K., Duan, B., Yu, M., Meng, W., Wang, Q., & Yu, Q. (2016). A Novel Insecticidal Peptide SLP1 Produced by Streptomyces laindensis H008 against Lipaphis erysimi. Molecules, 21(8), 1101. https://doi.org/10.3390/molecules21081101







