Identification of Eupatilin from Artemisia argyi as a Selective PPARα Agonist Using Affinity Selection Ultrafiltration LC-MS
Abstract
:1. Introduction
2. Results and Discussion
2.1. Validation of the Ultrafiltration LC-MS Assay for PPAR Screening
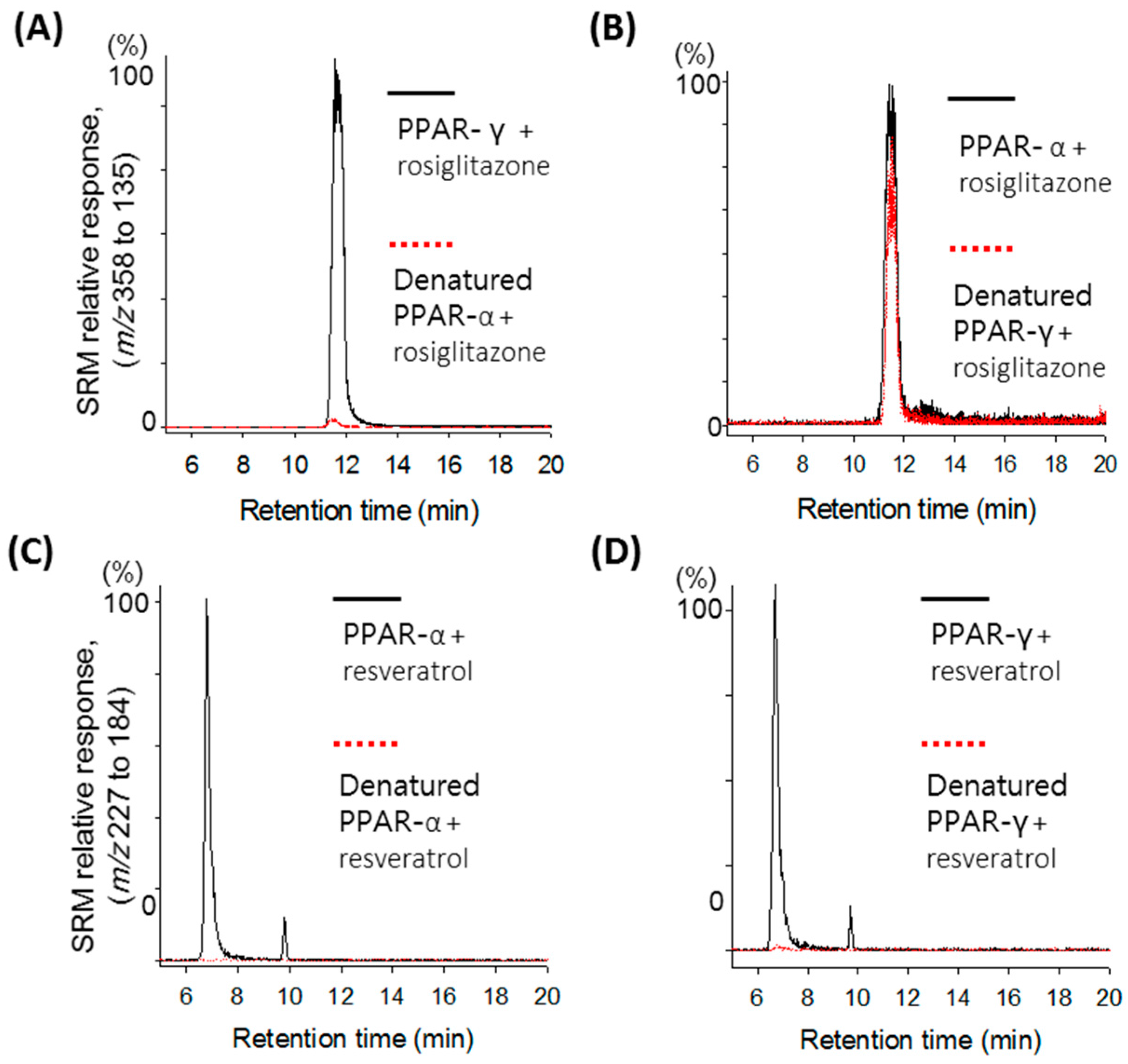
2.2. Screening of the A. argyi Extract Using the Ultrafiltration LC-MS Assay
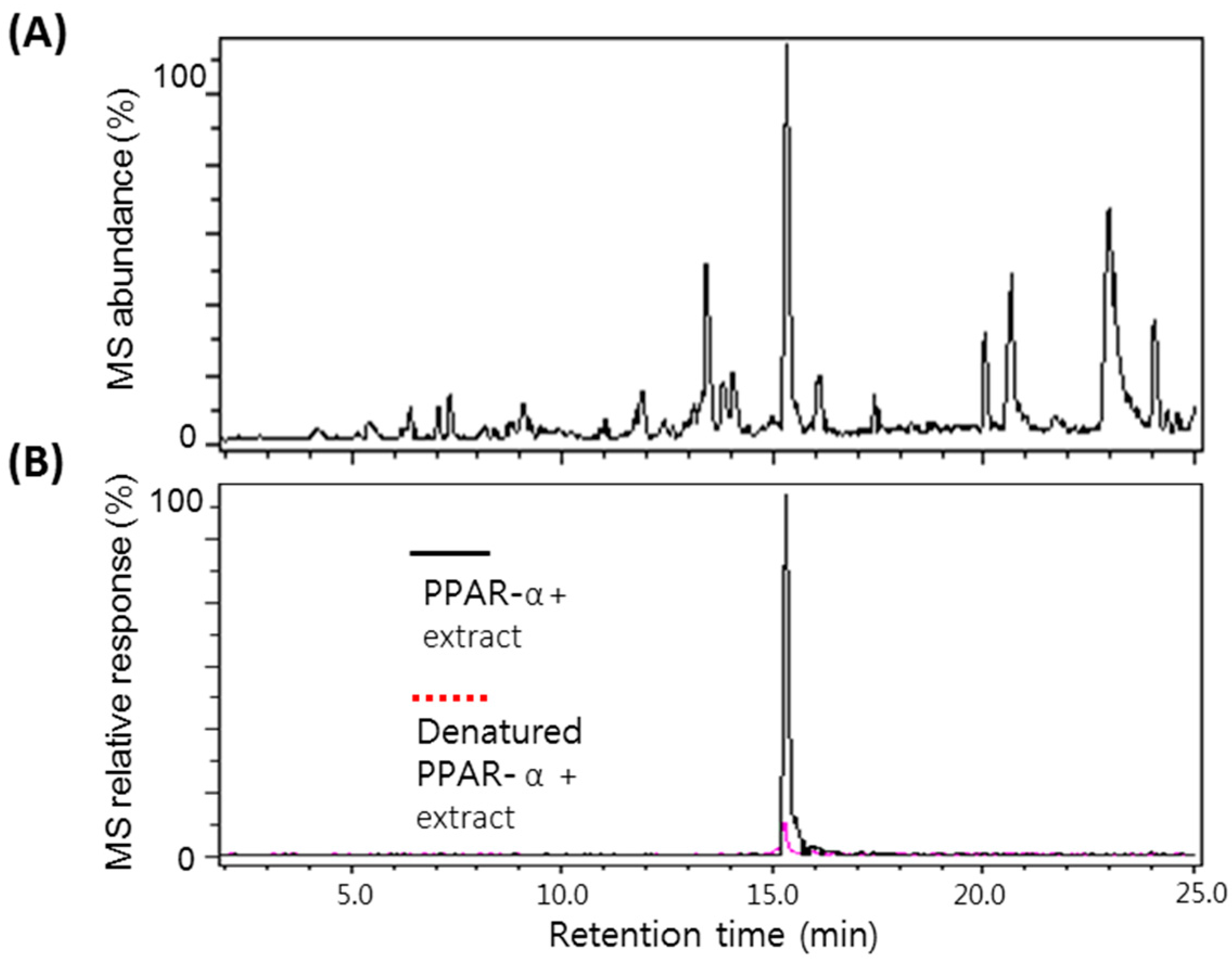
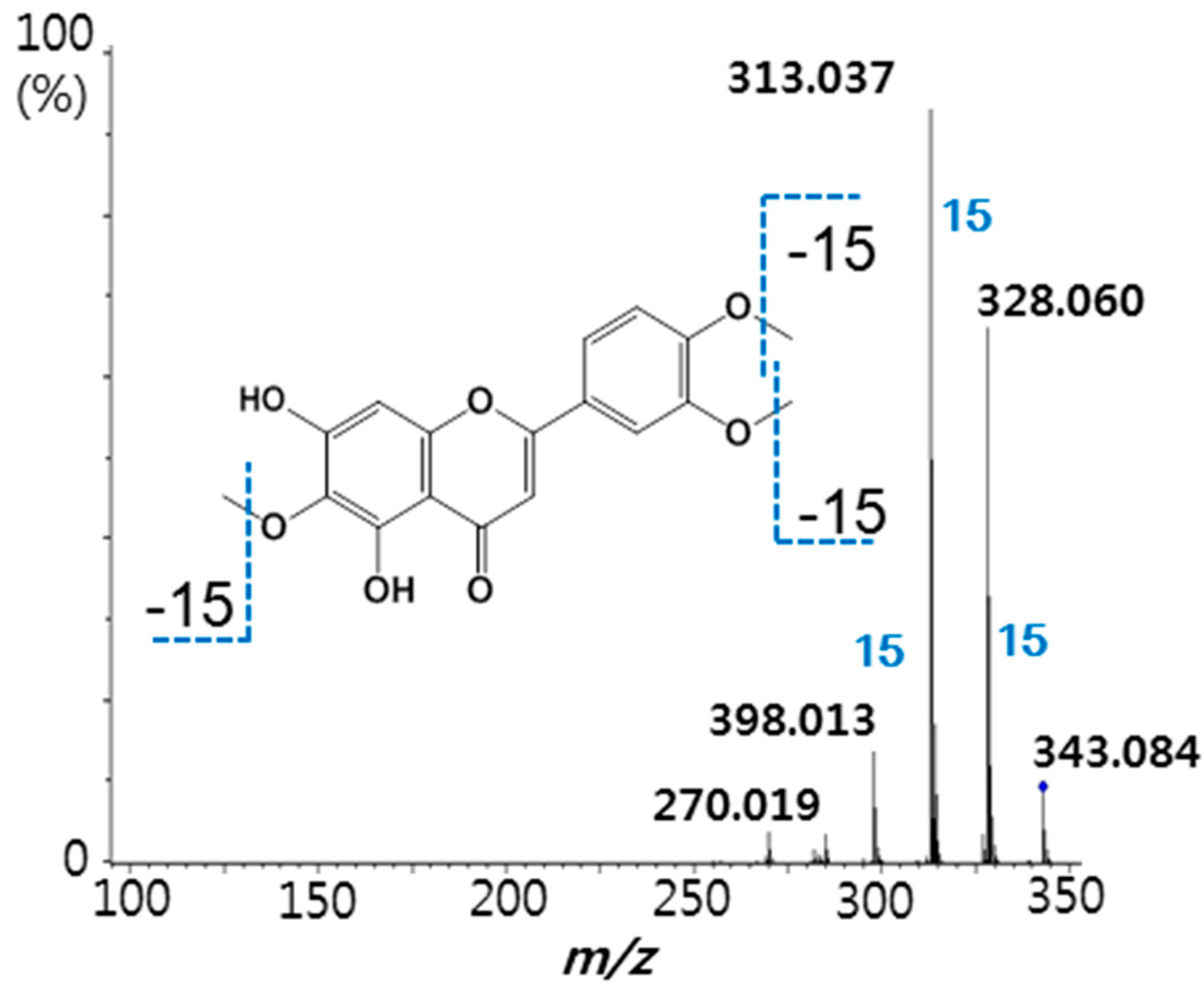
2.3. Binding Affinity of Eupatilin and Functional Cell-Based Assay
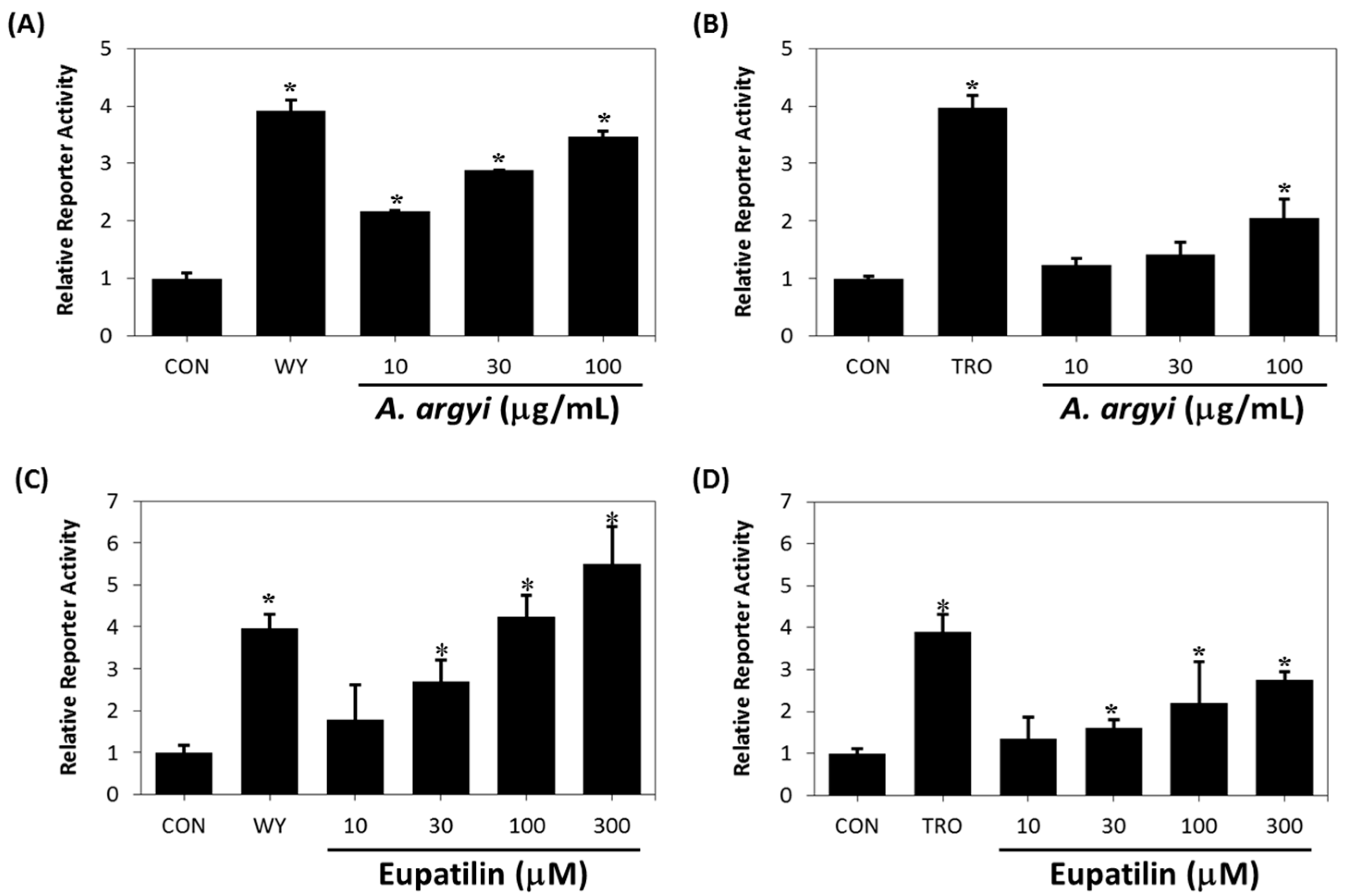
| Compound | IC50 | |
|---|---|---|
| PPARα | PPARγ | |
| GW6471 | 226.9 nM | - |
| GW9662 | - | 24.3 nM |
| Eupatilin | 1.18 μM | >100 μM |
3. Experimental Section
3.1. Chemicals and Reagents
3.2. Binding to PPARα or PPARγ and Ultrafiltration
3.3. LC-MS and LC-MS/MS
| Compound | SRM Transition (m/z) | DP | EP | CE | CXP |
|---|---|---|---|---|---|
| Resveratrol | 227 to 185 | −105 | −10 | −26 | −9 |
| Rosiglitazone | 358 to 135 | 116 | 10 | 39 | 8 |
3.4. Cell-Based Transactivation Assay
3.5. Ligand Binding Assay
3.6. Statistical Analyses
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Michalik, L.; Auwerx, J.; Berger, J.P.; Chatterjee, V.K.; Glass, C.K.; Gonzalez, F.J.; Grimaldi, P.A.; Kadowaki, T.; Lazar, M.A.; O’Rahilly, S.; et al. International union of pharmacology. LXI. peroxisome proliferator-activated receptors. Pharmacol. Rev. 2006, 58, 726–741. [Google Scholar] [CrossRef] [PubMed]
- Bajaj, M.; Suraamornkul, S.; Hardies, L.J.; Glass, L.; Musi, N.; DeFronzo, R.A. Effects of peroxisome proliferator-activated receptor (PPAR)-α and PPAR-γ agonists on glucose and lipid metabolism in patients with type 2 diabetes mellitus. Diabetologia 2007, 50, 1723–1731. [Google Scholar] [CrossRef] [PubMed]
- Jeong, D.; Yi, Y.S.; Sung, G.H.; Yang, W.S.; Park, J.G.; Yoon, K.; Yoon, D.H.; Song, C.; Lee, Y.; Rhee, M.H.; et al. Anti-inflammatory activities and mechanisms of Artemisia asiatica ethanol extract. J. Ethnopharmacol. 2014, 152, 487–496. [Google Scholar] [CrossRef] [PubMed]
- Gervois, P.; Fruchart, J.C.; Staels, B. Inflammation, dyslipidaemia, diabetes and PPARs: Pharmacological interest of dual PPARα and PPARγ agonists. Int. J. Clin. Pract. Suppl. 2004, 58, 22–29. [Google Scholar] [CrossRef]
- Bhattacharyya, N.; Hu, X.; Chen, C.Z.; Mathews Griner, L.A.; Zheng, W.; Inglese, J.; Austin, C.P.; Marugan, J.J.; Southall, N.; Neumann, S.; et al. A high throughput screening assay system for the identification of small molecule inhibitors of gsp. PLoS ONE 2014, 9, e90766. [Google Scholar] [CrossRef] [PubMed]
- Christopeit, T.; Overbo, K.; Danielson, U.H.; Nilsen, I.W. Efficient screening of marine extracts for protease inhibitors by combining FRET based activity assays and surface plasmon resonance spectroscopy based binding assays. Mar. Drugs 2013, 11, 4279–4293. [Google Scholar] [CrossRef] [PubMed]
- Miyagi, T.; Shiotani, B.; Miyoshi, R.; Yamamoto, T.; Oka, T.; Umezawa, K.; Ochiya, T.; Takano, M.; Tahara, H. DSE-FRET: A new anticancer drug screening assay for DNA binding proteins. Cancer Sci. 2014, 105, 870–874. [Google Scholar] [CrossRef] [PubMed]
- Lee, H.; Zhu, T.; Patel, K.; Zhang, Y.Y.; Truong, L.; Hevener, K.E.; Gatuz, J.L.; Subramanya, G.; Jeong, H.Y.; Uprichard, S.L.; et al. High-throughput screening (HTS) and hit validation to identify small molecule inhibitors with activity against NS3/4A proteases from multiple hepatitis C virus genotypes. PLoS ONE 2013, 8, e75144. [Google Scholar] [CrossRef] [PubMed]
- Liu, L. Efficient hit and lead compound evaluation strategy based on off-rate screening by surface plasmon resonance. J. Med. Chem. 2014, 57, 2843–2844. [Google Scholar] [CrossRef] [PubMed]
- Olaru, A.; Bala, C.; Jaffrezic-Renault, N.; Aboul-Enein, H.Y. Surface plasmon resonance (SPR) biosensors in pharmaceutical analysis. Crit. Rev. Anal. Chem. 2015, 45, 97–105. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.; Jermihov, K.; Nam, S.J.; Sturdy, M.; Maloney, K.; Qiu, X.; Chadwick, L.R.; Main, M.; Chen, S.N.; Mesecar, A.D.; et al. Screening natural products for inhibitors of quinone reductase-2 using ultrafiltration LC-MS. Anal. Chem. 2011, 83, 1048–1052. [Google Scholar] [CrossRef] [PubMed]
- Choi, Y.; van Breemen, R.B. Development of a screening assay for ligands to the estrogen receptor based on magnetic microparticles and LC-MS. Comb. Chem. High Throughput Screen. 2008, 11, 1–6. [Google Scholar] [PubMed]
- Sun, Y.; Gu, C.; Liu, X.; Liang, W.; Yao, P.; Bolton, J.L.; van Breemen, R.B. Ultrafiltration tandem mass spectrometry of estrogens for characterization of structure and affinity for human estrogen receptors. J. Am. Soc. Mass Spectrom. 2005, 16, 271–279. [Google Scholar] [CrossRef] [PubMed]
- Liu, D.; Guo, J.; Luo, Y.; Broderick, D.J.; Schimerlik, M.I.; Pezzuto, J.M.; van Breemen, R.B. Screening for ligands of human retinoid X receptor-α using ultrafiltration mass spectrometry. Anal. Chem. 2007, 79, 9398–9402. [Google Scholar] [CrossRef] [PubMed]
- Chen, Z.; Vigueira, P.A.; Chambers, K.T.; Hall, A.M.; Mitra, M.S.; Qi, N.; McDonald, W.G.; Colca, J.R.; Kletzien, R.F.; Finck, B.N. Insulin resistance and metabolic derangements in obese mice are ameliorated by a novel peroxisome proliferator-activated receptor γ-sparing thiazolidinedione. J. Biol. Chem. 2012, 287, 23537–23548. [Google Scholar] [CrossRef] [PubMed]
- Calleri, E.; Pochetti, G.; Dossou, K.S.; Laghezza, A.; Montanari, R.; Capelli, D.; Prada, E.; Loiodice, F.; Massolini, G.; Bernier, M.; et al. Resveratrol and its metabolites bind to PPARs. ChemBioChem 2014, 15, 1154–1160. [Google Scholar] [CrossRef] [PubMed]
- Atanasov, A.G.; Wang, J.N.; Gu, S.P.; Bu, J.; Kramer, M.P.; Baumgartner, L.; Fakhrudin, N.; Ladurner, A.; Malainer, C.; Vuorinen, A.; et al. Honokiol: A non-adipogenic PPARγ agonist from nature. Biochim. Biophys. Acta 2013, 1830, 4813–4819. [Google Scholar] [CrossRef] [PubMed]
- Dat, N.T.; Lee, K.; Hong, Y.S.; Kim, Y.H.; Minh, C.V.; Lee, J.J. A peroxisome proliferator-activated receptor-γ agonist and other constituents from Chromolaena odorata. Planta Med. 2009, 75, 803–807. [Google Scholar] [CrossRef] [PubMed]
- Hsu, W.H.; Lee, B.H.; Chang, Y.Y.; Hsu, Y.W.; Pan, T.M. A novel natural Nrf2 activator with PPARγ-agonist (monascin) attenuates the toxicity of methylglyoxal and hyperglycemia. Toxicol. Appl. Pharmacol. 2013, 272, 842–851. [Google Scholar] [CrossRef] [PubMed]
- Salam, N.K.; Huang, T.H.; Kota, B.P.; Kim, M.S.; Li, Y.; Hibbs, D.E. Novel PPAR-γ agonists identified from a natural product library: A virtual screening, induced-fit docking and biological assay study. Chem. Biol. Drug Des. 2008, 71, 57–70. [Google Scholar] [CrossRef] [PubMed]
- Zhang, M.L.; Irwin, D.; Li, X.N.; Sauriol, F.; Shi, X.W.; Wang, Y.F.; Huo, C.H.; Li, L.G.; Gu, Y.C.; Shi, Q.W. PPARγ agonist from Chromolaena odorata. J. Nat. Prod. 2012, 75, 2076–2081. [Google Scholar] [CrossRef] [PubMed]
- Kang, J.; Hick, L.A.; Price, W.E. A fragmentation study of isoflavones in negative electrospray ionization by MSn ion trap mass spectrometry and triple quadrupole mass spectrometry. Rapid Commun. Mass Spectrom. 2007, 21, 857–868. [Google Scholar] [CrossRef] [PubMed]
- Hajdu, Z.; Hohmann, J.; Forgo, P.; Mathe, I.; Molnar, J.; Zupko, I. Antiproliferative activity of Artemisia asiatica extract and its constituents on human tumor cell lines. Planta Med. 2014, 80, 1692–1697. [Google Scholar] [CrossRef] [PubMed]
- Khlifi, D.; Sghaier, R.M.; Amouri, S.; Laouini, D.; Hamdi, M.; Bouajila, J. Composition and anti-oxidant, anti-cancer and anti-inflammatory activities of Artemisia herba-alba, Ruta chalpensis L. and Peganum harmala L. Food Chem. Toxicol. 2013, 55, 202–208. [Google Scholar] [CrossRef] [PubMed]
- Willoughby, J.A., Sr.; Sundar, S.N.; Cheung, M.; Tin, A.S.; Modiano, J.; Firestone, G.L. Artemisinin blocks prostate cancer growth and cell cycle progression by disrupting Sp1 interactions with the cyclin-dependent kinase-4 (CDK4) promoter and inhibiting CDK4 gene expression. J. Biol. Chem. 2009, 284, 2203–2213. [Google Scholar] [CrossRef] [PubMed]
- Park, J.M.; Hahm, K.B.; Kwon, S.O.; Kim, E.H. The anti-inflammatory effects of acidic polysaccharide from Artemisia capillaris on Helicobacter pylori infection. J. Cancer Prev. 2013, 18, 161–168. [Google Scholar] [CrossRef] [PubMed]
- Vandanmagsar, B.; Haynie, K.R.; Wicks, S.E.; Bermudez, E.M.; Mendoza, T.M.; Ribnicky, D.; Cefalu, W.T.; Mynatt, R.L. Artemisia dracunculus L. extract ameliorates insulin sensitivity by attenuating inflammatory signalling in human skeletal muscle culture. Diabetes Obes. Metab. 2014, 16, 728–738. [Google Scholar] [CrossRef] [PubMed]
- Zeng, K.W.; Wang, S.; Dong, X.; Jiang, Y.; Tu, P.F. Sesquiterpene dimer (DSF-52) from Artemisia argyi inhibits microglia-mediated neuroinflammation via suppression of NF-κB, JNK/p38 MAPKs and Jak2/Stat3 signaling pathways. Phytomedicine 2014, 21, 298–306. [Google Scholar] [CrossRef] [PubMed]
- Kirk-Ballard, H.; Wang, Z.Q.; Acharya, P.; Zhang, X.H.; Yu, Y.; Kilroy, G.; Ribnicky, D.; Cefalu, W.T.; Floyd, Z.E. An extract of Artemisia dracunculus L. inhibits ubiquitin-proteasome activity and preserves skeletal muscle mass in a murine model of diabetes. PLoS ONE 2013, 8, e57112. [Google Scholar] [CrossRef] [PubMed]
- Obanda, D.N.; Hernandez, A.; Ribnicky, D.; Yu, Y.M.; Zhang, X.H.; Wang, Z.Q.; Cefalu, W.T. Bioactives of Artemisia dracunculus L. mitigate the role of ceramides in attenuating insulin signaling in rat skeletal muscle cells. Diabetes 2012, 61, 597–605. [Google Scholar] [CrossRef] [PubMed]
- Lee, W.; Ham, J.; Kwon, H.C.; Kim, Y.K.; Kim, S.N. Anti-diabetic effect of amorphastilbol through PPARαα/γ dual activation in db/db mice. Biochem. Biophys. Res. Commun. 2013, 432, 73–79. [Google Scholar] [CrossRef] [PubMed]
- Sample Availability: Samples of the compounds are not available from the authors.
© 2015 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Choi, Y.; Jung, Y.; Kim, S.-N. Identification of Eupatilin from Artemisia argyi as a Selective PPARα Agonist Using Affinity Selection Ultrafiltration LC-MS. Molecules 2015, 20, 13753-13763. https://doi.org/10.3390/molecules200813753
Choi Y, Jung Y, Kim S-N. Identification of Eupatilin from Artemisia argyi as a Selective PPARα Agonist Using Affinity Selection Ultrafiltration LC-MS. Molecules. 2015; 20(8):13753-13763. https://doi.org/10.3390/molecules200813753
Chicago/Turabian StyleChoi, Yongsoo, Yujung Jung, and Su-Nam Kim. 2015. "Identification of Eupatilin from Artemisia argyi as a Selective PPARα Agonist Using Affinity Selection Ultrafiltration LC-MS" Molecules 20, no. 8: 13753-13763. https://doi.org/10.3390/molecules200813753
APA StyleChoi, Y., Jung, Y., & Kim, S.-N. (2015). Identification of Eupatilin from Artemisia argyi as a Selective PPARα Agonist Using Affinity Selection Ultrafiltration LC-MS. Molecules, 20(8), 13753-13763. https://doi.org/10.3390/molecules200813753







