Abstract
The C-type lectin-like receptors include the Nkrp1 protein family that regulates the activity of natural killer (NK) cells. Rat Nkrp1a was reported to bind monosaccharide moieties in a Ca2+-dependent manner in preference order of GalNac > GlcNAc >> Fuc >> Gal > Man. These findings established for rat Nkrp1a have been extrapolated to all additional Nkrp1 receptors and have been supported by numerous studies over the past two decades. However, since 1996 there has been controversy and another article showed lack of interactions with saccharides in 1999. Nevertheless, several high affinity saccharide ligands were synthesized in order to utilize their potential in antitumor therapy. Subsequently, protein ligands were introduced as specific binders for Nkrp1 proteins and three dimensional models of receptor/protein ligand interaction were derived from crystallographic data. Finally, for at least some members of the NK cell C-type lectin-like proteins, the “sweet story” was impaired by two reports in recent years. It has been shown that the rat Nkrp1a and CD69 do not bind saccharide ligands such as GlcNAc, GalNAc, chitotetraose and saccharide derivatives (GlcNAc-PAMAM) do not directly and specifically influence cytotoxic activity of NK cells as it was previously described.
1. Introduction
Natural killer (NK) cells are an essential component of the innate immunity system that play a key role against virally infected or tumor cells. The specificity of NK cells for target cells is determined by a sophisticated repertoire of activating and inhibitory receptors expressed on the surface of NK cells [1]. Besides the eradication of a variety of different pathological cells, NK cells produce several classes of immunoregulatory cytokines, which help to shape the immune response. Interactions between these molecules are involved in maintaining homeostasis within the immune system [2]. Furthermore, NK cells are also involved in transplantation immunity [3], autoimmunity [4], allergic diseases [5] or reproduction [6].
The cytolytic activity of NK cells is determined by a dynamic interplay between activating and inhibitory signals [7,8,9]. According to the missing self-recognition, the inhibitory receptors recognize ligands, such as MHC I molecules, which are broadly expressed by healthy cells but their expression is significantly downregulated during pathological circumstances. On the other hand, according to the induced self-recognition, activating receptors recognize ligands, such as MICA [10], which are minimally expressed by healthy cells but their expression is strongly upregulated during cellular stress, and infectious or tumorigenic processes.
NK receptors belong to two distinct structural classes: the immunoglobulin-like receptors and the C-type lectin-like receptors. The first group includes Killer Cell Immunoglobulin-Like Receptors (KIR), Leukocyte Immunoglobulin-Like Receptors (LILR) and Natural Cytotoxicity Receptors (NCR) protein families, the second one Ly49, NKG2D, CD94/NKG2 and Nkrp1 protein families.
The Nkrp1 protein family contains an important set of both activating and inhibitory receptors. The first identified Nkrp1 receptors were originally classified as members of the C-type animal lectin family [11,12]. Moreover, the ability to bind saccharides was extended to other Nkrp1 receptors and several high affinity saccharide ligands with antitumor potential were synthesized. However, due to irreproducibility of binding experiments the saccharide binding activity of Nkrp1 receptors became a matter of controversy and most of the NK receptor research field dismissed the saccharide binding properties of Nkrp1. This review is focused on Nkrp1 protein family and their biological functions. Particularly, we discuss discrepancies concerning Nkrp1 ligand binding specificity accumulated during 20 years of the research in the field.
2. Nkrp1 Family
The first cell-surface antigen specific for mouse NK cells was described in 1977 [13]. Later, it was named NK1.1 and classified as a member of the Nkrp1 family [14], Nkrp1c [12,15]. Since then several other Nkrp1 (Klrb1) genes have been identified.
In the mouse genome, members of the Klrb1 family (Klrb1a, b/d, c, e, f and g [12,16,17,18] encoding Nkrp1a, b/d, c, f and g proteins) are located in the Natural Killer Cell (NKC) gene complex and map to the chromosome 6 [19]. Klrb1b and d genes represent two divergent alleles of the same locus and Klrb1e is a pseudogene [16]. Four Klrb1 genes located on chromosome 4 [20] were defined in rat: Klrb1a [21,22], b/c [20,23], f [24] and g [17] (Klrb1b and c are also divergent alleles [17]) encoding Nkrp1a, b/c, f and g proteins. There is only one KLRB1 gene encoding NKRP1A in the human genome, which is located on chromosome 12 [15]. Besides NK cells, Nkrp1 receptors are also expressed on the surface of NKT cells, T lymphocytes and dendritic cells [24,25].
3. Structure of Nkrp1 Receptors
From a structural point of view, Nkrp1 receptors are type II transmembrane proteins containing an N-terminal cytoplasmic domain, a transmembrane region, and an ectodomain with the C-type lectin-like domain (CTLD), which is connected by a stalk region to the transmembrane segment. Several members of the Nkrp1 family have been demonstrated to exist as disulfide linked homodimers [11,12,26], in which cysteines in the stalk region mediate intermolecular disulfide bonds. Nevertheless, the existence of Nkrp1 proteins as disulfide linked homodimers requires further investigation as some members of the Nkrp1 family show variability in the number of cysteines in the stalk region [27].
In addition, the CTLD shares structural homology with the C-type lectin domain, but contrary to the C-type lectin domain, CTLD lacks the residues involved in calcium binding and recognizes other ligands than saccharides. The CTLD consists of two α-helices (α1 and α2) and two antiparallel β-sheets (Figure 1A). Numbering of secondary structural elements is usually based on rat MBP-A [28]. In most of the known three-dimensional structures of CTLD, the two β-sheets are formed by β0, β1 and β5 strands, and β2, β2’, β3 and β4 strands, respectively. Four cysteine residues, which are the most conserved residues in CTLD, adopt a disulfide pattern of C1-C4 and C2-C3. On the other hand, arrangement of the loops connecting the secondary structural elements differs to a large extent between the particular CTLDs [29].
In terms of Nkrp1 receptors, initial work on their structure demonstrated that Nkrp1 receptors adopt the fold typical for the CTLD with increased flexibility of a long loop region, which corresponds to the region between strands β2 and β3. Moreover, the sequence differences in the loop suggested that the long loop region might encode the ligand specificity [30]. Indeed, the crystal structure of mouse Nkrp1a, which is the only known structure among Nkrp1 receptors so far, revealed that Nkrp1a adopts a fold similar to the CTLD with the exception of the extended long loop region pointing away from the CTLD (Figure 1B). In the crystal, the extended loop tightly interacted with a neighboring loop of another molecule using a domain swapping effect [31]. The position of the extended loop pointing away from the CTLD has not been described in any NK receptors. However, this conformation has been observed in other C-type lectin-like proteins important in the immune system such as the macrophage mannose receptor [32] and in snake venoms [33]. On the other hand, structural analysis of mouse Nkrp1a and Nkrp1c in solution using mass spectrometric techniques unveiled that the loop region was not extended in solution, but instead collapsed and closely interacted with the core of the molecule [34,35]. Moreover, the recently determined structure of NKp65 in complex with its ligand KACL, which is the first report of CTLD recognition by another CTLD, suggests that the extended loop of Nkrp1a is sterically incompatible with the NKp65 mode of ligand binding and probably represents a crystallization artifact [36].
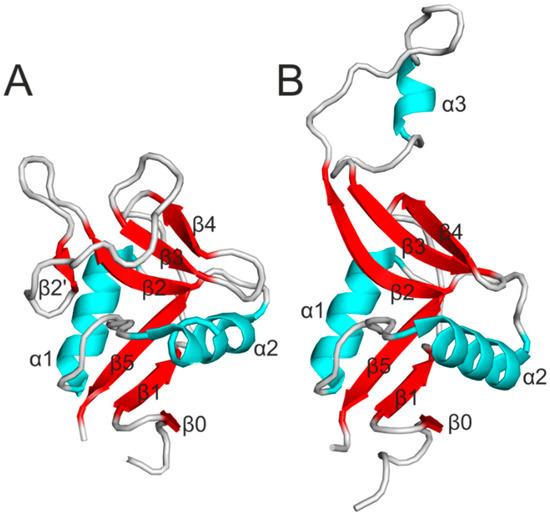
Figure 1.
(A) Ribbon diagram of a typical CTLD. Secondary structure elements are labeled. α-helices are cyan, β-strands red and loops are gray; (B) Structure of the mouse Nkrp1a with a loop region pointing away from the central core.
4. Signaling
Signals from many different NK receptors enter the cell simultaneously and need to be integrated. Through the inhibitory receptors, signaling is commonly mediated via tyrosine phosphorylation of an Immunoreceptor Tyrosine-based Inhibitory Motif (ITIM), which is present in the cytoplasmic part of the NK receptor and is a component of mouse and rat Nkrp1b/d [37,38] or Nkrp1g receptors [18].
Phosphorylation of Immunoreceptor Tyrosine-based Activation Motifs (ITAM), typically in adaptor proteins (e.g., FcRγ or CD3ζ) is crucial for signal transduction of activating receptors. Receptors usually contain positively charged amino acids in their transmembrane region interacting with negatively charged amino acids in associated adaptors [39,40]. The most important motif for a subsequent signaling cascade in the cytoplasmic domain of all murine NK receptors, except the Nkrp1g isoform, is CxCP (Cys-X-Cys-Pro), which represents a binding site for a kinase of the Src family, Lck [41]. Lck phosphorylates tyrosines in ITAM/ITIM sequences as was described for mouse Nkrp1c and Nkrp1b [40]. In case of an activating receptor, phosphorylation leads to recruitment of the Syk kinase [42], that triggers a downstream cascade. Conversely, phosphorylation of ITIM creates a docking template for the phosphatase SHP-1 [43,44], which dephosphorylates substrates of kinases functioning in activating pathways [45].
Human NKRP1A represents an exception in such signaling. Although the receptor has an inhibitory function, it does not possess an ITIM sequence or Lck binding motif in its cytoplasmic domain. Signaling via human NKRP1A involves association with acid sphingomyelinase, production of ceramide, and PI3K-dependent activation of the Akt and ERK pathways [46].
5. Saccharide Ligands of Nkrp1—20 Years of Controversy
The ligands for rat Nkrp1 have been a matter of controversy for 20 years. The first report showing Nkrp1 as a lectin binding monosaccharides was published by Bezouska et al., in 1994. The binding of monosaccharides was reported to be strongly Ca2+-dependent in preference order of GalNac > GlcNAc >> Fuc >> Gal > Man [47]. These results were extended by the identification of high affinity oligosaccharides of the blood group family, the ganglio family and glycosaminoglycans with IC50 in the range 10−9–10−12 M. These high affinity oligosaccharide ligands localized at the cell membrane were reported to lead to cytolysis of tumor cells suggesting a new and particularly promising antitumor therapy [48].
However, a correction was published in 1996 by six out of the ten original authors, who wished to retract that article owing to their inability to reproduce the results [49]. Furthermore, Nkrp1 monosaccharide binding activity was re-evaluated in 1999 as well, showing no data indicating specific binding to any of the monosaccharides as described in the previous report [50].
Despite all the controversies and contradictions, many studies have been accumulated using the binding paradigm from 1994 in data interpretations. Simultaneously, other ligands for Nkrp1 were described. New glycomimetics (saccharide molecules with some variations in the structure, modifying biological properties) on GlcNAc basis were synthesized since GlcNAc had been shown in previous articles to be the best ligand among the monosaccharides tested [47]. Very high affinity was observed for a number of chitooligomers, particularly chitotetraose, and thus several chitotetraose derivatives were synthesized and reported as high affinity ligands of Nkrp1 [51,52,53,54,55]. A next generation of glycomimetics with strong antitumor response was based on highly branched GlcNAc-terminated glycoclusters such as GlcNAc coated octavalent glycodendrimers (PAMAM-GlcNAc8). In addition, the considerable immune response was explained by a multivalency effect of these derivatives [56,57,58,59].
Further studies revealed disaccharides such as ManNAc(1→4)Glc [60] or monovalent and bivalent LacdiNAc as ligands with exceptional affinity to Nkrp1 [61,62]. Moreover, several negatively charged oligosaccharides and glycosides [63,64,65], or deoxynorijimycin with its hexosaminyl derivatives [66], as well as glycosyl-1H-1,2,3-triazoles [67] were demonstrated as effective Nkrp1 ligands too. Furthermore, series of calixarenes substituted with GlcNAc were reported as high affinity ligands of Nkrp1 and described as a next generation of saccharide derivatives with a strong antitumor effect [68].
Direct binding between Nkrp1a and saccharide ligands was extended by biological assays in mouse models. Indeed, treatment with PAMAM-GlcNAc8 or GlcNAc-coated calixarene dendrimer was reported to reduce tumor growth in melanoma-bearing C57BL/6 mice and prolonged survival time of experimental animals in several studies. Additionally, high levels of IgG as a result of increased antibody-dependent cell-mediated cytotoxicity (ADCC) and upregulated synthesis of IFN-γ by NK cells was observed [69,70,71].
Besides the rat and mouse Nkrp1 receptors, interactions of the synthesized saccharide ligands were tested also with human CD69, a C-type lectin-like receptor structurally similar to Nkrp1, and most of them showed binding patterns similar to rat Nkrp1 [72,73,74,75]. The majority of the above mentioned descriptions of the Nkrp1 ligand specificity were provided by a limited number of laboratories, with some methodologies performed and interpreted only by a single scientist.
In 2012 Karel Bezouska was investigated by the Ethical committee of the Institute of Microbiology and the Charles University in Prague. After investigation the committee announced that Bezouska had repeatedly and for several years committed scientific misconduct, and asked the coauthors of all respective papers to re-evaluate their experiments and interpretations [76].
In the context of later findings, the binding activity of Nkrp1 described before was re-examined over the past two years. The interaction between rat Nkrp1a and multiple ligands was re-evaluated using two independent biophysical techniques, isothermal titration calorimetry and NMR titration. Though the same experimental conditions (buffer, temperature, calcium concentration, a positive and a negative control sample etc.) were used as described before, no binding was detected [77]. Moreover, the interaction analysis using a neoglycoconjugate binding assay revealed no affinity between rat Nkrp1a or mouse Nkrp1c and the various neoglycoproteins such as N-acetylglucosamine36BSA. Furthermore, no significant changes in serum cytokine patterns relevant to anti-cancer treatment, or changes in Klrb1 gene expression were observed in animal models after injection with PAMAM-GlcNAc8 [78]. These findings raised serious doubts about the credibility of several previous papers. Indeed, most of them have been already retracted while the others are still in the retraction process.
However, the lack of interactions with the tested saccharide samples does not imply, that Nkrp1 might not bind other saccharide moieties. Interestingly, several C-type lectin-like receptors, including the inhibitory receptors Ly49A and C, have been reported to recognize sulfated polysaccharides [79]. In addition, glycosylation plays an important role in NK cell activation and regulation of their effector functions. Hartmann et al. published that glycosylation of the stalk domain of the NK receptor NKp30 influences receptor-ligand interactions and intracellular signaling via an adaptor protein [80]. Similarly, binding of inhibitory receptors Ly49 to MHC class I molecules is affected by glycosylation, but contrary to NKp30, glycosylation of Ly49 receptors lowers the affinity to their ligands [81]. Further, altered glycosylation on the cell surface of cancer cells may result in modulation of surface expression of NKD2D ligands indicating that the surface expression of MICA/B is dependent on N-linked glycosylation [82].
6. The Clr Ligands
Meantime, protein ligands for several members of the Nkrp1 receptor family have been identified (Figure 2). They represent the C-type lectin-related (Clr) protein family, alternatively known as the osteoclast inhibitory lectin (Ocil). Intriguingly, Clr ligands share 40% amino acid identity with the CTLD of CD69 [83,84] and are encoded by genes (Clec2) intermingled within NKC (Natural Killer Cell gene complex) with Klrb1 genes encoding their receptors [26].
In mouse, binding pairs include Nkrp1b/d:Clr-b, Nkrp1f:Clr-c,-d,-g, and Nkrp1g:Clr-d,-f,-g; ligands for Nkrp1a and Nkrp1c remain still elusive [18,26,31,34,35,41,83,85,86,87]. In rat, Nkrp1a and Nkrp1b recognize Clr11, whereas Nkrp1f and Nkrp1g recognize Clr-2,-3,-4,-6,-7 and Clr-2,-6,-7, respectively. Interestingly, both mouse and rat Nkrp1f and Nkrp1g show high promiscuity with an overlapping set of ligands [17].
In human, the only receptor in the Nkrp1 family, NKRP1A, binds LLT1, which is structurally similar to mouse Clr proteins [88,89]. Surface plasmon analysis of the interaction revealed that the extracellular domain of NKRP1A binds LLT1 with fast kinetics and low affinity with Kd = 48 µM. Binding is entropically and enthalpically driven with a small heat capacity. In addition, these binding properties are typical features for cell-cell recognition molecules. Experiments employing mutagenesis of the proteins indicated a new structural model suggesting a dimer-dimer interaction [86]. Nevertheless, the crystal structure of any Nkrp1:Clr complex remains to be determined.
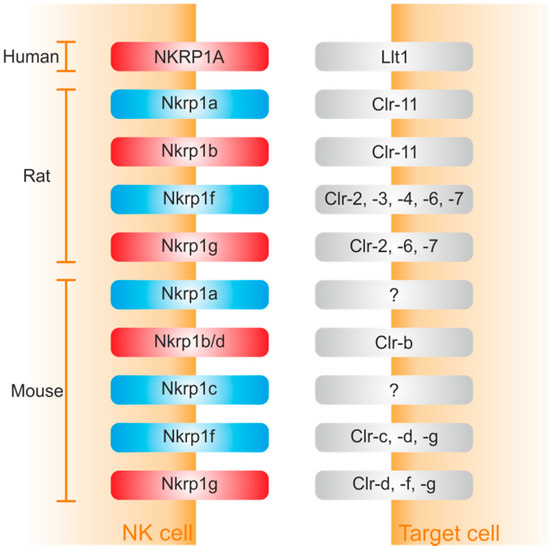
Figure 2.
Human, rat and mouse Nkrp1-Clr interactions showing the combinatorial nature of the Nkrp1 repertoire. Inhibitory receptors are in red; activating in blue.
7. Biological Relevance of Nkrp1:Clr Interactions
In general, inhibitory C-type lectin-like receptors interact with MHC class I or MHC class I-like molecules. Classical MHC I molecules are recognized by members of the mouse Ly49 receptor family, whereas non-classical MHC I molecules associate with the heterodimeric CD94/NKG2 receptor [90,91,92] and with the activating NKG2D receptor, respectively [93].
Interestingly, inhibitory receptors Nkrp1b/d recognize MHC-independent ligands. Clr-b, which is expressed on most hematopoietic and several non-hematopoietic cell types, displays an expression pattern comparable to classical MHC I antigens [94]. Moreover, it is downregulated in poxvirus-infected [95] and tumor cells, which contributes to higher susceptibility of target cells to NK cell-mediated cytotoxicity, supporting the “missing self” hypothesis [85].
Additionally, a rapid downregulation of Clr-b at both RNA and protein levels under chemotherapy-induced genotoxic stress has been shown in some leukemia cells resulting in decreased NK-cell inhibition mediated by the Nkrp1b:Clr-b receptor-ligand pair. Further on, the levels of MHC class I molecules remained normal [96], suggesting that the functional effect of the Nkrp1b:Clr-b interaction is strictly regulated by Clr-b expression levels [25,85].
In the mouse interaction between Nkrp1f and Clr-g resulted in stimulation of TCR/CD28-mediated T cell proliferation and IL-2 production in in vitro and in vivo experiments, respectively [97]. However, since only basic studies about the surface expression of both interaction partners and their specific binding are available, further detailed functional analysis is required.
Moreover, the role of Nkrp1b:Clr-b interactions in innate immunity and viral infections has been investigated. It was described that rat cytomegalovirus (RCMV) has developed a mechanism to avoid NK cell killing through the expression of a viral protein RCTL on the surface of an infected cell; RCTL is a homolog of rat Clr-b interacting with Nkrp1b on NK cells [98]. In mice with a knocked-out Ly49H gene, encoding another NK cell activating receptor, mouse cytomegalovirus (MCMV) infection causes extensive viral expansion and initiates effector cytokine secretion [99]. Moreover, in some rat strains, Nkrp1b has numerous allelic variations which abort the viral protein-Nkrp1b interaction and support host Nkrp1b:Clr-b recognition [23,38].
The NKRP1A:LLT1 interaction in human causes inhibition of cytotoxicity and IFN-γ secretion in NKRP1+ NK cells [89]. In contrast, LLT1 engaging NKRP1A on T cells provides T cell costimulation and increases cytokine production [100]. Furthermore, expression of LLT1 on TLR-stimulated plasmacytoid dendritic cells and monocyte-derived dendritic cells might explain the signal transduction pathways between NK cells and dendritic cells during an immune response to pathogens [101]. While the NKRP1A:LLT1 interactions on NK and T cells have different effects in vitro, the immunological relevance of this engagement in vivo is not well understood. A recent study has shown the connection between respiratory-virus infection and proinflammatory cytokine production, where LLT1 expressed on epithelial cells may represent a part of a regulatory feedback mechanism [102].
Generally, structural analysis is obligatory to understand the functions of proteins or protein-protein interactions. Nonetheless, structural characterization of the molecular details of the Nkrp1:Clr interaction is at the beginning. Until now there is only a limited number of relevant receptor or ligand structures solved, namely for mouse Nkrp1a [31,103] and Clr-g proteins [87]. The first crystal structure of the CTLD in complex with another CTLD, the NKp65:KACL complex, has been suggested to be a model structure for the rest of the known rodent Nkrp1:Clr and human NKRP1A:LLT1 receptor-ligand pairs, respectively [36].
8. Conclusions
Nkrp1 receptors were originally classified as members of the C-type animal lectin family with well documented ligand binding specificity. There were several reasons for this notion, e.g., sequence homology to well described lectin molecules or striking and well documented data published in high ranking journals. It was temping for the NK-cell receptor community to extrapolate data from one model protein (rat Nkrp1a) to a broader set of molecules encoded in different species (mouse, human), establishing a paradigm in the field. It is not surprising that lectin-based interpretation of the primary data led to misleading results, which accumulated during the years, even in the situation when early after publishing the seminal article in Nature a correction and publications questioning the relevance and the reproducibility of the results, appeared. Over the years more and more data challenging the lectin-based concept—mutations in the calcium binding site, originally saccharide binding site, turned out to better fit a protein-recognition module and finally, specific protein ligands for members of the Nkrp1 family were found and protein-protein interactions were characterized in molecular details. Whatever kind of a mistake there was at the beginning, it sent at least part of the researchers in a wrong direction for decades. It is satisfactory that the evidence coming from different sources finally results in the change of the paradigm and clarifies the field of NK-cell receptor biology.
Acknowledgments
This work has been supported by Charles University (projects UNCE 356 204013/2012 and 204025/2012), Institutional Research Concept of the Institute of Microbiology (RVO61388971); grants from the Ministry of Education Youth and Sports of the Czech Republic and European Regional Development Funds (CZ.1.07/2.3.00/20.0055, CZ.1.07/2.3.00/30.0003 and BIOCEV CZ.1.05/1.1.00/02.0109); European Social Fund and the state budget of the Czech Republic. Project No. CZ.1.07/2.3.00/30.0022 and the Grant Agency of Charles University (800413).
Author Contributions
Daniel Rozbeský prepared article layout, covered structural issues and controversies in ligand binding focusing on original seminal publications, discussed all parts of the manuscript; Ljubina Ivanova was responsible for description of Nkrp1:Clr interactions; Lucie Hernychová wrote introduction to NK cells, Nkrp1 proteins and their signaling properties; Valéria Grobárová focused on controversial issues in ligand binding and in vivo experiments together with the description of the Clr ligands; Petr Novák and Jan Černý coordinated manuscript preparation, were responsible for critical reading and overall philosophy of the manuscript.
Abbreviation
| CTLD | C-type lectin-like domain |
| GalNAc | 2-acetamido-2-deoxy-d-galactopyranose |
| GlcNAc | 2-acetamido-2-deoxy-d-glucopyranose |
| ITAM | immunoreceptor tyrosine-based activation motif |
| ITIM | immunoreceptor tyrosine-based inhibitory motif |
| LacdiNAc | 2-acetamido-2-deoxy-β-d-galactopyranosyl-(1→4)-2-acetamido-2-deoxy-d-glucopyranose |
| ManNAc | 2-acetamido-d-mannopyranose |
| NKC | NK cell gene complex |
| PAMAM dendrimer | polyamidoamine dendrimer |
Conflicts of Interest
The authors declare no conflict of interest.
References
- Vivier, E.; Tomasello, E.; Baratin, M.; Walzer, T.; Ugolini, S. Functions of natural killer cells. Nat. Immunol. 2008, 9, 503–510. [Google Scholar] [CrossRef] [PubMed]
- Strowig, T.; Brilot, F.; Münz, C. Noncytotoxic functions of NK cells: Direct pathogen restriction and assistance to adaptive immunity. J. Immunol. 2008, 180, 7785–7791. [Google Scholar] [CrossRef] [PubMed]
- Villard, J. The role of natural killer cells in human solid organ and tissue transplantation. J. Innate Immun. 2011, 3, 395–402. [Google Scholar] [CrossRef] [PubMed]
- Tian, Z.; Gershwin, M.E.; Zhang, C. Regulatory NK cells in autoimmune disease. J. Autoimmun. 2012, 39, 206–215. [Google Scholar] [CrossRef] [PubMed]
- Mathias, C.B.; Guernsey, L.A.; Zammit, D.; Brammer, C.; Wu, C.A.; Thrall, R.S.; Aguila, H.L. Pro-inflammatory role of natural killer cells in the development of allergic airway disease. Clin. Exp. Allergy 2014, 44, 589–601. [Google Scholar] [CrossRef] [PubMed]
- Winger, E.E.; Reed, J.L. The multiple faces of the decidual natural killer cell. Am. J. Reprod. Immunol. 2013, 70, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Lanier, L.L. NK cell recognition. Annu. Rev. Immunol. 2005, 23, 225–274. [Google Scholar] [CrossRef] [PubMed]
- Kärre, K.; Ljunggren, H.G.; Piontek, G.; Kiessling, R. Selective rejection of H-2-deficient lymphoma variants suggests alternative immune defence strategy. Nature 1986, 319, 675–678. [Google Scholar] [CrossRef] [PubMed]
- Raulet, D.H.; Vance, R.E. Self-tolerance of natural killer cells. Nat. Rev. Immunol. 2006, 6, 520–531. [Google Scholar] [CrossRef] [PubMed]
- Bauer, S.; Groh, V.; Wu, J.; Steinle, A.; Phillips, J.H.; Lanier, L.L.; Spies, T. Activation of NK cells and T cells by NKG2D, a receptor for stress-inducible MICA. Science 1999, 285, 727–729. [Google Scholar] [CrossRef] [PubMed]
- Giorda, R.; Rudert, W.; Vavassori, C.; Chambers, W.H.; Hiserodt, J.C.; Trucco, M. NKR-P1, a signal transduction molecule on natural killer cells. Science 1990, 249, 1298–300. [Google Scholar] [CrossRef] [PubMed]
- Giorda, R.; Trucco, M. Mouse NKR-P1. A family of genes selectively coexpressed in adherent lymphokine-activated killer cells. J. Immunol. 1991, 147, 1701–1708. [Google Scholar] [PubMed]
- Glimcher, L.; Shen, F.W.; Cantors, H. Identification of a cell-surface expressed antigen selectively on the natural killer cell. J. Exp. Med. 1977, 145, 1–9. [Google Scholar] [CrossRef] [PubMed]
- Ryan, J.C.; Turck, J.; Niemi, E.C.; Yokoyama, W.M.; Seaman, W.E. Molecular cloning of the NK1.1 antigen, a member of the NKR-P1 family of natural killer cell activation molecules. J. Immunol. 1992, 149, 1631–1635. [Google Scholar] [PubMed]
- Lanier, L.L.; Chang, C.; Phillips, J.H. Human NKR-P1A. A disulfide-linked homodimer of the C-type lectin superfamily expressed by a subset of NK and T lymphocytes. J. Immunol. 1994, 153, 2417–2428. [Google Scholar] [PubMed]
- Plougastel, B.; Matsumoto, K.; Dubbelde, C.; Yokoyama, W.M. Analysis of a 1-Mb BAC contig overlapping the mouse Nkrp1 cluster of genes: Cloning of three new Nkrp1 members, Nkrp1d, Nkrp1e, and Nkrp1f. Immunogenetics 2001, 53, 592–598. [Google Scholar] [CrossRef] [PubMed]
- Kveberg, L.; Dai, K.-Z.; Westgaard, I.H.; Daws, M.R.; Fossum, S.; Naper, C.; Vaage, J.T. Two major groups of rat NKR-P1 receptors can be distinguished based on chromosomal localization, phylogenetic analysis and Clr ligand binding. Eur. J. Immunol. 2009, 39, 541–551. [Google Scholar] [CrossRef] [PubMed]
- Kveberg, L.; Dai, K.-Z.; Inngjerdingen, M.; Brooks, C.G.; Fossum, S.; Vaage, J.T. Phylogenetic and functional conservation of the NKR-P1F and NKR-P1G receptors in rat and mouse. Immunogenetics 2011, 63, 429–436. [Google Scholar] [CrossRef] [PubMed]
- Yokoyama, W.M.; Ryan, J.C.; Hunter, J.J.; Smith, H.R.C.; Stark, M.; Seamant, W.E. cDNA cloning of mouse NKR-P1 and genetic linkage with LY-49. Identification of a natural killer cell gene complex on chromosome 6. J. Immunol. 1991, 147, 3229–3236. [Google Scholar] [PubMed]
- Dissen, E.; Ryan, J.C.; Seaman, W.E.; Fossum, S. An Autosomal Dominant Locus, Nka, Mapping to the Ly-49 Region of a Rat Natural Killer (NK) Gene Complex, Controls NK Cell Lysis of Allogeneic Lymphocytes. J. Exp. Med. 1996, 183, 2197–2207. [Google Scholar] [CrossRef] [PubMed]
- Chambers, W.H.; Vujanovic, N.L.; DeLeo, A.B.; Olszowy, M.W.; Herberman, R.B.; Hiserodt, J.C. Monoclonal antibody to a triggering structure expressed on rat natural killer cells and adherent lymphokine-activated killer cells. J. Exp. Med. 1989, 169, 1373–1389. [Google Scholar] [CrossRef] [PubMed]
- Ryan, J.C.; Niemi, E.C.; Goldfien, R.D.; Hiserodt, J.C.; Seaman, W.E. NKR-P1, an activating molecule on rat natural killer cells, stimulates phosphoinositide turnover and a rise in intracellular calcium. J. Immunol. 1991, 3244–3250. [Google Scholar]
- Kveberg, L.; Bäck, C.J.; Dai, K.-Z.; Inngjerdingen, M.; Rolstad, B.; Ryan, J.C.; Vaage, J.T.; Naper, C. The novel inhibitory NKR-P1C receptor and Ly49s3 identify two complementary, functionally distinct NK cell subsets in rats. J. Immunol. 2006, 176, 4133–4140. [Google Scholar] [CrossRef] [PubMed]
- Appasamy, P.M.; Kenniston, T.W.; Brissette-Storkus, C.S.; Chambers, W.H. NKR-P1dim/TCR alpha beta + T cells and natural killer cells share expression of NKR-P1A and NKR-P1D. Nat. Immun. 1996, 15, 259–268. [Google Scholar] [PubMed]
- Aust, J.G.; Gays, F.; Mickiewicz, K.M.; Buchanan, E.; Brooks, C.G. The expression and function of the NKRP1 receptor family in C57BL/6 mice. J. Immunol. 2009, 183, 106–116. [Google Scholar] [CrossRef] [PubMed]
- Iizuka, K.; Naidenko, O.V.; Plougastel, B.F.M.; Fremont, D.H.; Yokoyama, W.M. Genetically linked C-type lectin-related ligands for the NKRP1 family of natural killer cell receptors. Nat. Immunol. 2003, 4, 801–807. [Google Scholar] [CrossRef] [PubMed]
- Rozbeský, D.; Kavan, D.; Chmelík, J.; Novák, P.; Vaněk, O.; Bezouška, K. High-level expression of soluble form of mouse natural killer cell receptor NKR-P1C(B6) in Escherichia coli. Protein Expr. Purif. 2011, 77, 178–184. [Google Scholar] [CrossRef] [PubMed]
- Weis, W.I.; Kahn, R.; Fourme, R.; Drickamer, K.; Hendrickson, W.A. Structure of the calcium-dependent lectin domain from a rat mannose-binding protein determined by MAD phasing. Science 1991, 254, 1608–1615. [Google Scholar] [CrossRef] [PubMed]
- Zelensky, A.N.; Gready, J.E. The C-type lectin-like domain superfamily. FEBS J. 2005, 272, 6179–6217. [Google Scholar] [CrossRef] [PubMed]
- Sovová, Z.; Kopecký, V.; Pazderka, T.; Hofbauerová, K.; Rozbeský, D.; Vaněk, O.; Bezouška, K.; Ettrich, R. Structural analysis of natural killer cell receptor protein 1 (NKR-P1) extracellular domains suggests a conserved long loop region involved in ligand specificity. J. Mol. Model. 2011, 17, 1353–1370. [Google Scholar] [CrossRef] [PubMed]
- Kolenko, P.; Rozbeský, D.; Vaněk, O.; Kopecký, V.; Hofbauerová, K.; Novák, P.; Pompach, P.; Hašek, J.; Skálová, T.; Bezouška, K.; et al. Molecular architecture of mouse activating NKR-P1 receptors. J. Struct. Biol. 2011, 175, 434–441. [Google Scholar] [CrossRef] [PubMed]
- Feinberg, H.; Park-Snyder, S.; Kolatkar, R.; Heise, C.T.; Taylor, M.E.; Weis, W.I. Structure of a C-type carbohydrate recognition domain from the macrophage mannose receptor. J. Biol. Chem. 2000, 275, 21539–21548. [Google Scholar] [CrossRef] [PubMed]
- Maita, N.; Nishio, K.; Nishimoto, E.; Matsui, T.; Shikamoto, Y.; Morita, T.; Sadler, J.E.; Mizuno, H. Crystal structure of von Willebrand factor A1 domain complexed with snake venom, bitiscetin: Insight into glycoprotein Ib alpha binding mechanism induced by snake venom proteins. J. Biol. Chem. 2003, 278, 37777–37781. [Google Scholar] [CrossRef] [PubMed]
- Rozbesky, D.; Man, P.; Kavan, D.; Chmelik, J.; Cerny, J.; Bezouska, K.; Novak, P. Chemical cross-linking and H/D exchange for fast refinement of protein crystal structure. Anal. Chem. 2012, 84, 867–870. [Google Scholar] [CrossRef] [PubMed]
- Rozbesky, D.; Sovova, Z.; Marcoux, J.; Man, P.; Ettrich, R.; Robinson, C.V.; Novak, P. Structural model of lymphocyte receptor NKR-P1C revealed by mass spectrometry and molecular modeling. Anal. Chem. 2013, 85, 1597–1604. [Google Scholar] [CrossRef] [PubMed]
- Li, Y.; Wang, Q.; Chen, S.; Brown, P.H.; Mariuzza, R.A. Structure of NKp65 bound to its keratinocyte ligand reveals basis for genetically linked recognition in natural killer gene complex. Proc. Natl. Acad. Sci. USA 2013, 110, 11505–11510. [Google Scholar] [CrossRef] [PubMed]
- Carlyle, J.R.; Martin, A.; Mehra, A.; Attisano, L.; Tsui, F.W.; Zúñiga-Pflücker, J.C. Mouse NKR-P1B, a novel NK1.1 antigen with inhibitory function. J. Immunol. 1999, 162, 5917–5923. [Google Scholar] [PubMed]
- Li, J.; Rabinovich, B.A.; Hurren, R.; Shannon, J.; Miller, R.G. Expression cloning and function of the rat NK activating and inhibitory receptors NKR-P1A and -P1B. Int. Immunol. 2003, 15, 411–416. [Google Scholar] [CrossRef] [PubMed]
- Lanier, L.L. Up on the tightrope: Natural killer cell activation and inhibition. Nat. Immunol. 2008, 9, 495–502. [Google Scholar] [CrossRef] [PubMed]
- Ljutic, B.; Carlyle, J.R.; Filipp, D.; Nakagawa, R.; Julius, M.; Zúñiga-Pflücker, J.C. Functional requirements for signaling through the stimulatory and inhibitory mouse NKR-P1 (CD161) NK cell receptors. J. Immunol. 2005, 174, 4789–4796. [Google Scholar] [CrossRef] [PubMed]
- Chen, P.; Bélanger, S.; Aguilar, O.; Zhang, Q.; St-Laurent, A.; Rahim, M.M.A.; Makrigiannis, A.P.; Carlyle, J.R. Analysis of the mouse 129-strain Nkrp1-Clr gene cluster reveals conservation of genomic organization and functional receptor-ligand interactions despite significant allelic polymorphism. Immunogenetics 2011, 63, 627–640. [Google Scholar] [CrossRef] [PubMed]
- Vivier, E.; Nunès, J.; Vély, F. Natural killer cell signaling pathways. Science 2004, 306, 1517–1519. [Google Scholar] [CrossRef] [PubMed]
- Burshtyn, D.N.; Yang, W.; Yi, T.; Long, E.O. A novel phosphotyrosine motif with a critical amino acid at position -2 for the SH2 domain-mediated activation of the tyrosine phosphatase SHP-1. J. Biol. Chem. 1997, 272, 13066–13072. [Google Scholar] [CrossRef] [PubMed]
- Tamir, I.; dal Porto, J.M.; Cambier, J.C. Cytoplasmic protein tyrosine phosphatases SHP-1 and SHP-2: Regulators of B cell signal transduction. Curr. Opin. Immunol. 2000, 12, 307–315. [Google Scholar] [CrossRef] [PubMed]
- Ravetch, J.V.; Lanier, L.L. Immune Inhibitory Receptors. Science 2000, 290, 84–89. [Google Scholar] [CrossRef] [PubMed]
- Pozo, D.; Valés-Gómez, M.; Mavaddat, N.; Williamson, S.C.; Chisholm, S.E.; Reyburn, H. CD161 (human NKR-P1A) signaling in NK cells involves the activation of acid sphingomyelinase. J. Immunol. 2006, 176, 2397–2406. [Google Scholar] [CrossRef] [PubMed]
- Bezouska, K.; Vlahas, G.; Horváth, O.; Jinochová, G.; Fiserová, A.; Giorda, R.; Chambers, W.H.; Feizi, T.; Pospísil, M. Rat natural killer cell antigen, NKR-P1, related to C-type animal lectins is a carbohydrate-binding protein. J. Biol. Chem. 1994, 269, 16945–16952. [Google Scholar] [PubMed]
- Bezouska, K.; Yuen, C.T.; O’Brien, J.; Childs, R.A.; Chai, W.; Lawson, A.M.; Drbal, K.; Fiserová, A.; Pospísil, M.; Feizi, T. Oligosaccharide ligands for NKR-P1 protein activate NK cells and cytotoxicity. Nature 1994, 372, 150–157. [Google Scholar] [CrossRef] [PubMed]
- Bezouska, K.; Yuen, C.T.; O’Brien, J.; Childs, R.A.; Chai, W.; Lawson, A.M.; Drbal, K.; Fiserová, A.; Pospisil, M.; Feizi, T. Correspondence: Oligosaccharide ligands for NKR-P1 protein activate NK cells and cytotoxicity. Nature 1996, 380, 559–559. [Google Scholar]
- Kogelberg, H.; Montero, E.; Bay, S.; Lawson, A.M.; Feizi, T. Re-evaluation of monosaccharide binding property of recombinant soluble carbohydrate recognition domain of the natural killer cell receptor NKR-P1A. J. Biol. Chem. 1999, 274, 30335–30336. [Google Scholar] [PubMed]
- Bezouska, K.; Sklenar, J.; Dvorakova, J.; Havlicek, V.; Pospisil, M.; Thiem, J.; Kren, V. NKR-P1A protein, an activating receptor of rat natural killer cells, binds to the chitobiose core ofuncompletely glycosylated N-linked glycans, and to linear chitooligomers. Biochem. Biophys. Res. Commun. 1997, 238, 149–153. [Google Scholar] [CrossRef] [PubMed]
- Krist, P.; Herkommerová-Rajnochová, E.; Rauvolfová, J.; Semenuk, T.; Vavrusková, P.; Pavlícek, J.; Bezouska, K.; Petrus, L.; Kren, V. Toward an optimal oligosaccharide ligand for rat natural killer cell activation receptor NKR-P1. Biochem. Biophys. Res. Commun. 2001, 287, 11–20. [Google Scholar] [CrossRef] [PubMed]
- Sedmera, P.; Přikrylová, V.; Bezouška, K.; Rajnochová, E.; Thiem, J.; Křen, V. Preparation of mannac containing chitooligomers by isomerisation and their binding to Nkr-P1 Protein. J. Carbohydr. Chem. 1998, 17, 1351–1357. [Google Scholar] [CrossRef]
- Kren, V.; Dvoráková, J.; Gambert, U.; Sedmera, P.; Havlícek, V.; Thiem, J.; Bezouska, K. beta-Glucosylation of chitooligomers by galactosyltransferase. Carbohydr. Res. 1997, 305, 517–523. [Google Scholar] [CrossRef] [PubMed]
- Semenuk, T.; Krist, P.; Pavlícek, J.; Bezouska, K.; Kuzma, M.; Novák, P.; Kren, V. Synthesis of chitooligomer-based glycoconjugates and their binding to the rat natural killer cell activation receptor NKR-P1. Glycoconj. J. 2001, 18, 817–826. [Google Scholar] [CrossRef] [PubMed]
- Bezouska, K.; Kren, V.; Kieburg, C.; Lindhorst, T.K. GlcNAc-terminated glycodendrimers form defined precipitates with the soluble dimeric receptor of rat natural killer cells, sNKR-P1A. FEBS Lett. 1998, 426, 243–247. [Google Scholar] [CrossRef] [PubMed]
- Pospisil, M.; Vannucci, L.; Fiserova, A.; Krausova, K.; Horvath, O.; Kren, V.; Mosca, F.; Lindhorst, T.K.; Sadalapure, K.; Bezouska, K. Glycodendrimeric ligands of c-type lectin receptors as therapeutic agents in experimental cancer. Adv. Exp. Med. Biol. 2001, 495, 343–347. [Google Scholar] [PubMed]
- Krist, P.; Vannucci, L.; Kuzma, M.; Man, P.; Sadalapure, K.; Patel, A.; Bezouska, K.; Pospísil, M.; Petrus, L.; Lindhorst, T.K.; Kren, V. Fluorescent labelled thiourea-bridged glycodendrons. Chembiochem 2004, 5, 445–452. [Google Scholar] [CrossRef] [PubMed]
- Vannucci, L.; Fiserová, A.; Sadalapure, K.; Lindhorst, T.K.; Kuldová, M.; Rossmann, P.; Horváth, O.; Kren, V.; Krist, P.; Bezouska, K.; et al. Effects of N-acetyl-glucosamine-coated glycodendrimers as biological modulators in the B16F10 melanoma model in vivo. Int. J. Oncol. 2003, 23, 285–296. [Google Scholar] [PubMed]
- Attolino, E.; Bonaccorsi, F.; Catelani, G.; D’Andrea, F.; Křenek, K.; Bezouška, K.; Křen, V. Improved preparation of β-d-ManNAc-(1→4)-D-Glc and β-d-TalNAc-(1→4)-d-Glc disaccharides and evaluation of their activating properties on the Natural Killer cells NKR-P1 and CD69 Receptors. J. Carbohydr. Chem. 2008, 27, 156–171. [Google Scholar] [CrossRef]
- Bojarová, P.; Krenek, K.; Wetjen, K.; Adamiak, K.; Pelantová, H.; Bezouska, K.; Elling, L.; Kren, V. Synthesis of LacdiNAc-terminated glycoconjugates by mutant galactosyltransferase-a way to new glycodrugs and materials. Glycobiology 2009, 19, 509–517. [Google Scholar] [CrossRef] [PubMed]
- Drozdová, A.; Bojarová, P.; Křenek, K.; Weignerová, L.; Henssen, B.; Elling, L.; Christensen, H.; Jensen, H.H.; Pelantová, H.; Kuzma, M.; et al. Enzymatic synthesis of dimeric glycomimetic ligands of NK cell activation receptors. Carbohydr. Res. 2011, 346, 1599–1609. [Google Scholar] [CrossRef] [PubMed]
- Bojarová, P.; Křenek, K.; Kuzma, M.; Petrásková, L.; Bezouška, K.; Namdjou, D.-J.; Elling, L.; Křen, V. N-Acetylhexosamine triad in one molecule: Chemoenzymatic introduction of 2-acetamido-2-deoxy-β-d-galactopyranosyluronic acid residue into a complex oligosaccharide. J. Mol. Catal. B Enzym. 2008, 50, 69–73. [Google Scholar] [CrossRef]
- Fialová, P.; Namdjou, D.-J.; Ettrich, R.; Přikrylová, V.; Rauvolfová, J.; Křenek, K.; Kuzma, M.; Elling, L.; Bezouška, K.; Křen, V. Combined application of galactose oxidase and β-N-acetylhexosaminidase in the synthesis of complex immunoactive N-acetyl-d-galactosaminides. Adv. Synth. Catal. 2005, 347, 997–1006. [Google Scholar] [CrossRef]
- Bojarová, P.; Slámová, K.; Křenek, K.; Gažák, R.; Kulik, N.; Ettrich, R.; Pelantová, H.; Kuzma, M.; Riva, S.; Adámek, D.; et al. Charged hexosaminides as new substrates for β-N-acetylhexosaminidase-catalyzed synthesis of immunomodulatory disaccharides. Adv. Synth. Catal. 2011, 353, 2409–2420. [Google Scholar] [CrossRef]
- Catelani, G.; D’Andrea, F.; Griselli, A.; Guazzelli, L.; Nemcová, P.; Bezouska, K.; Krenek, K.; Kren, V. Deoxynojirimycin and its hexosaminyl derivatives bind to natural killer cell receptors rNKR-P1A and hCD69. Bioorg. Med. Chem. Lett. 2010, 20, 4645–4648. [Google Scholar] [CrossRef] [PubMed]
- Slámová, K.; Marhol, P.; Bezouska, K.; Lindkvist, L.; Hansen, S.G.; Kren, V.; Jensen, H.H. Synthesis and biological activity of glycosyl-1H-1,2,3-triazoles. Bioorg. Med. Chem. Lett. 2010, 20, 4263–4265. [Google Scholar] [CrossRef] [PubMed]
- Krenek, K.; Kuldová, M.; Hulíková, K.; Stibor, I.; Lhoták, P.; Dudic, M.; Budka, J.; Pelantová, H.; Bezouska, K.; Fiserová, A.; et al. N-acetyl-d-glucosamine substituted calix[4]arenes as stimulators of NK cell-mediated antitumor immune response. Carbohydr. Res. 2007, 342, 1781–1792. [Google Scholar] [CrossRef] [PubMed]
- Hulikova, K.; Benson, V.; Svoboda, J.; Sima, P.; Fiserova, A. N-Acetyl-d-glucosamine-coated polyamidoamine dendrimer modulates antibody formation via natural killer cell activation. Int. Immunopharmacol. 2009, 9, 792–799. [Google Scholar] [CrossRef] [PubMed]
- Hulíková, K.; Grobárová, V.; Křivohlavá, R.; Fišerová, A. Antitumor activity of N-acetyl-d-glucosamine-substituted glycoconjugates and combined therapy with keyhole limpet hemocyanin in B16F10 mouse melanoma model. Folia Microbiol. (Praha). 2010, 55, 528–532. [Google Scholar] [CrossRef] [PubMed]
- Hulikova, K.; Svoboda, J.; Benson, V.; Grobarova, V.; Fiserova, A. N-acetyl-d-glucosamine-coated polyamidoamine dendrimer promotes tumor-specific B cell responses via natural killer cell activation. Int. Immunopharmacol. 2011, 11, 955–961. [Google Scholar] [CrossRef] [PubMed]
- Bezouska, K.; Nepovim, A.; Horvath, O.; Pospisil, M.; Hamann, J.; Feizi, T. CD69 antigen of human lymphocytes is a calcium-dependent carbohydrate-binding protein. Biochem. Biophys. Res. Commun. 1995, 208, 68–74. [Google Scholar] [CrossRef] [PubMed]
- Pavlicek, J.; Sopko, B.; Ettrich, R.; Kopecky, V.; Baumruk, V.; Man, P.; Havlicek, V.; Vrbacky, M.; Martinkova, L.; Kren, V.; et al. Molecular characterization of binding of calcium and carbohydrates by an early activation antigen of lymphocytes CD69. Biochemistry 2003, 42, 9295–9306. [Google Scholar] [CrossRef] [PubMed]
- Kavan, D.; Kubickova, M.; Bily, J.; Vanek, O.; Hofbauerova, K.; Mrazek, H.; Rozbesky, D.; Bojarova, P.; Kren, V.; Zidek, L.; et al. Cooperation between subunits is essential for high-affinity binding of N-acetyl-d-hexosamines to dimeric soluble and dimeric cellular forms of human CD69. Biochemistry 2010, 49, 4060–4067. [Google Scholar] [CrossRef] [PubMed]
- Kovalova, A.; Ledvina, M.; Saman, D.; Zyka, D.; Kubickova, M.; Zidek, L.; Sklenar, V.; Pompach, P.; Kavan, D.; Bily, J.; et al. Synthetic N-acetyl-d-glucosamine based fully branched tetrasaccharide, a mimetic of the endogenous ligand for CD69, activates CD69+ killer lymphocytes upon dimerization via a hydrophilic flexible linker. J. Med. Chem. 2010, 53, 4050–4065. [Google Scholar] [CrossRef] [PubMed]
- Report of the Joint Ethical Committee of the Institute of Microbiology, Prague and Charles University in Prague. English translation can be found in the Supplementary Material. Available online: http://www.biomed.cas.cz/mbu/doc/VyjadreniEK.PDF (accessed on 11 November 2014).
- Rozbeský, D.; Krejzová, J.; Křenek, K.; Prchal, J.; Hrabal, R.; Kožíšek, M.; Weignerová, L.; Fiore, M.; Dumy, P.; Křen, V.; et al. Re-evaluation of binding properties of recombinant lymphocyte receptors NKR-P1A and CD69 to chemically synthesized glycans and peptides. Int. J. Mol. Sci. 2014, 15, 1271–1283. [Google Scholar] [CrossRef] [PubMed]
- Grobárová, V.; Benson, V.; Rozbeský, D.; Novák, P.; Cerný, J. Re-evaluation of the involvement of NK cells and C-type lectin-like NK receptors in modulation of immune responses by multivalent GlcNAc-terminated oligosaccharides. Immunol. Lett. 2013, 156, 110–117. [Google Scholar] [CrossRef] [PubMed]
- Daniels, B.F.; Nakamura, M.C.; Rosen, S.D.; Yokoyama, W.M.; Seaman, W.E. Ly-49A, a receptor for H-2Dd, has a functional carbohydrate recognition domain. Immunity 1994, 1, 785–792. [Google Scholar] [CrossRef] [PubMed]
- Hartmann, J.; Tran, T.-V.; Kaudeer, J.; Oberle, K.; Herrmann, J.; Quagliano, I.; Abel, T.; Cohnen, A.; Gatterdam, V.; Jacobs, A.; et al. The stalk domain and the glycosylation status of the activating natural killer cell receptor NKp30 are important for ligand binding. J. Biol. Chem. 2012, 287, 31527–31539. [Google Scholar] [CrossRef] [PubMed]
- Mason, L.H.; Willette-Brown, J.; Anderson, S.K.; Alvord, W.G.; Klabansky, R.L.; Young, H.A.; Ortaldo, J.R. Receptor Glycosylation Regulates Ly-49 Binding to MHC Class I. J. Immunol. 2003, 171, 4235–4242. [Google Scholar] [CrossRef] [PubMed]
- Andresen, L.; Skovbakke, S.L.; Persson, G.; Hagemann-Jensen, M.; Hansen, K.A.; Jensen, H.; Skov, S. 2-deoxy d-glucose prevents cell surface expression of NKG2D ligands through inhibition of N-linked glycosylation. J. Immunol. 2012, 188, 1847–1855. [Google Scholar] [CrossRef] [PubMed]
- Plougastel, B.; Dubbelde, C.; Yokoyama, W.M. Cloning of Clr, a new family of lectin-like genes localized between mouse Nkrp1a and Cd69. Immunogenetics 2001, 53, 209–214. [Google Scholar] [CrossRef] [PubMed]
- Zhou, H.; Kartsogiannis, V.; Hu, Y.S.; Elliott, J.; Quinn, J.M.; McKinstry, W.J.; Gillespie, M.T.; Ng, K.W. A novel osteoblast-derived C-type lectin that inhibits osteoclast formation. J. Biol. Chem. 2001, 276, 14916–14923. [Google Scholar] [CrossRef] [PubMed]
- Carlyle, J.R.; Jamieson, A.M.; Gasser, S.; Clingan, C.S.; Arase, H.; Raulet, D.H. Missing self-recognition of Ocil/Clr-b by inhibitory NKR-P1 natural killer cell receptors. Proc. Natl. Acad. Sci. USA 2004, 101, 3527–3532. [Google Scholar] [CrossRef] [PubMed]
- Kamishikiryo, J.; Fukuhara, H.; Okabe, Y.; Kuroki, K.; Maenaka, K. Molecular basis for LLT1 protein recognition by human CD161 protein (NKRP1A/KLRB1). J. Biol. Chem. 2011, 286, 23823–23830. [Google Scholar] [CrossRef] [PubMed]
- Skálová, T.; Kotýnková, K.; Dušková, J.; Hašek, J.; Koval, T.; Kolenko, P.; Novák, P.; Man, P.; Hanč, P.; Vaněk, O.; et al. Mouse Clr-g, a ligand for NK cell activation receptor NKR-P1F: Crystal structure and biophysical properties. J. Immunol. 2012, 189, 4881–4889. [Google Scholar] [CrossRef] [PubMed]
- Rosen, D.B.; Bettadapura, J.; Alsharifi, M.; Mathew, P.; Warren, H.S.; Lanier, L.L. Cutting edge: Lectin-like transcript-1 is a ligand for the inhibitory human NKR-P1A receptor. J. Immunol. 2005, 175, 7796–7799. [Google Scholar] [CrossRef] [PubMed]
- Aldemir, H.; Prod’homme, V.; Dumaurier, M.-J.; Retiere, C.; Poupon, G.; Cazareth, J.; Bihl, F.; Braud, V.M. Cutting Edge: Lectin-Like Transcript 1 Is a Ligand for the CD161 Receptor. J. Immunol. 2005, 175, 7791–7795. [Google Scholar] [CrossRef] [PubMed]
- Borrego, F.; Ulbrecht, M.; Weiss, E.H.; Coligan, J.E.; Brooks, A.G. Recognition of human histocompatibility leukocyte antigen (HLA)-E complexed with HLA class I signal sequence-derived peptides by CD94/NKG2 confers protection from natural killer cell-mediated lysis. J. Exp. Med. 1998, 187, 813–818. [Google Scholar] [CrossRef] [PubMed]
- Braud, V.M.; Allan, D.S.; O’Callaghan, C.A.; Söderström, K.; D’Andrea, A.; Ogg, G.S.; Lazetic, S.; Young, N.T.; Bell, J.I.; Phillips, J.H.; et al. HLA-E binds to natural killer cell receptors CD94/NKG2A, B and C. Nature 1998, 391, 795–799. [Google Scholar] [CrossRef] [PubMed]
- Lee, N.; Llano, M.; Carretero, M.; Ishitani, A.; Navarro, F.; López-Botet, M.; Geraghty, D.E. HLA-E is a major ligand for the natural killer inhibitory receptor CD94/NKG2A. Proc. Natl. Acad. Sci. USA 1998, 95, 5199–5204. [Google Scholar] [CrossRef] [PubMed]
- Mistry, A.R.; O’Callaghan, C.A. Regulation of ligands for the activating receptor NKG2D. Immunology 2007, 121, 439–447. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Q.; Rahim, M.M.; Allan, D.S.J.; Tu, M.M.; Belanger, S.; Abou-Samra, E.; Ma, J.; Sekhon, H.S.; Fairhead, T.; Zein, H.S.; et al. Mouse Nkrp1-Clr gene cluster sequence and expression analyses reveal conservation of tissue-specific MHC-independent immunosurveillance. PLoS One 2012, 7, e50561. [Google Scholar] [CrossRef] [PubMed]
- Williams, K.J.N.; Wilson, E.; Davidson, C.L.; Aguilar, O.; Fu, L.; Carlyle, J.R.; Burshtyn, D.N. Poxvirus infection-associated downregulation of C-type lectin-related-b prevents NK cell inhibition by NK receptor protein-1B. J. Immunol. 2012, 188, 4980–4991. [Google Scholar] [CrossRef] [PubMed]
- Fine, J.H.; Chen, P.; Mesci, A.; Allan, D.S.J.; Gasser, S.; Raulet, D.H.; Carlyle, J.R. Chemotherapy-induced genotoxic stress promotes sensitivity to natural killer cell cytotoxicity by enabling missing-self recognition. Cancer Res. 2010, 70, 7102–7113. [Google Scholar] [CrossRef] [PubMed]
- Tian, W.; Nunez, R.; Cheng, S.; Ding, Y.; Tumang, J.; Lyddane, C.; Roman, C.; Liou, H.-C. C-type lectin OCILRP2/Clr-g and its ligand NKRP1f costimulate T cell proliferation and IL-2 production. Cell Immunol. 2005, 234, 39–53. [Google Scholar] [CrossRef] [PubMed]
- Voigt, S.; Mesci, A.; Ettinger, J.; Fine, J.H.; Chen, P.; Chou, W.; Carlyle, J.R. Cytomegalovirus evasion of innate immunity by subversion of the NKR-P1B:Clr-b missing-self axis. Immunity 2007, 26, 617–627. [Google Scholar] [CrossRef] [PubMed]
- Fodil-Cornu, N.; Lee, S.-H.; Belanger, S.; Makrigiannis, A.P.; Biron, C.A.; Buller, R.M.; Vidal, S.M. Ly49h-deficient C57BL/6 mice: A new mouse cytomegalovirus-susceptible model remains resistant to unrelated pathogens controlled by the NK gene complex. J. Immunol. 2008, 181, 6394–6405. [Google Scholar] [CrossRef] [PubMed]
- Germain, C.; Meier, A.; Jensen, T.; Knapnougel, P.; Poupon, G.; Lazzari, A.; Neisig, A.; Håkansson, K.; Dong, T.; Wagtmann, N.; et al. Induction of lectin-like transcript 1 (LLT1) protein cell surface expression by pathogens and interferon-γ contributes to modulate immune responses. J. Biol. Chem. 2011, 286, 37964–37975. [Google Scholar] [CrossRef] [PubMed]
- Rosen, D.B.; Cao, W.; Avery, D.T.; Tangye, S.G.; Liu, Y.-J.; Houchins, J.P.; Lanier, L.L. Functional consequences of interactions between human NKR-P1A and its ligand LLT1 expressed on activated dendritic cells and B cells. J. Immunol. 2008, 180, 6508–6517. [Google Scholar] [CrossRef] [PubMed]
- Satkunanathan, S.; Kumar, N.; Bajorek, M.; Purbhoo, M.; Culley, F.J. Respiratory syncytial virus infection, TLR3 ligands, and proinflammatory cytokines induce CD161 ligand LLT1 expression on the respiratory epithelium. J. Virol. 2014, 88, 2366–2373. [Google Scholar] [CrossRef] [PubMed]
- Kolenko, P.; Rozbeský, D.; Vaněk, O.; Bezouška, K.; Hašek, J.; Dohnálek, J. Structure of the H107R variant of the extracellular domain of mouse NKR-P1A at 2.3 Å resolution. Acta Crystallogr. Sect. F Struct. Biol. Cryst. Commun. 2011, 67, 1519–1523. [Google Scholar] [CrossRef] [PubMed]
© 2015 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/4.0/).