A Multi-Scale–Multi-Stable Model for the Rhodopsin Photocycle
Abstract
:1. Introduction
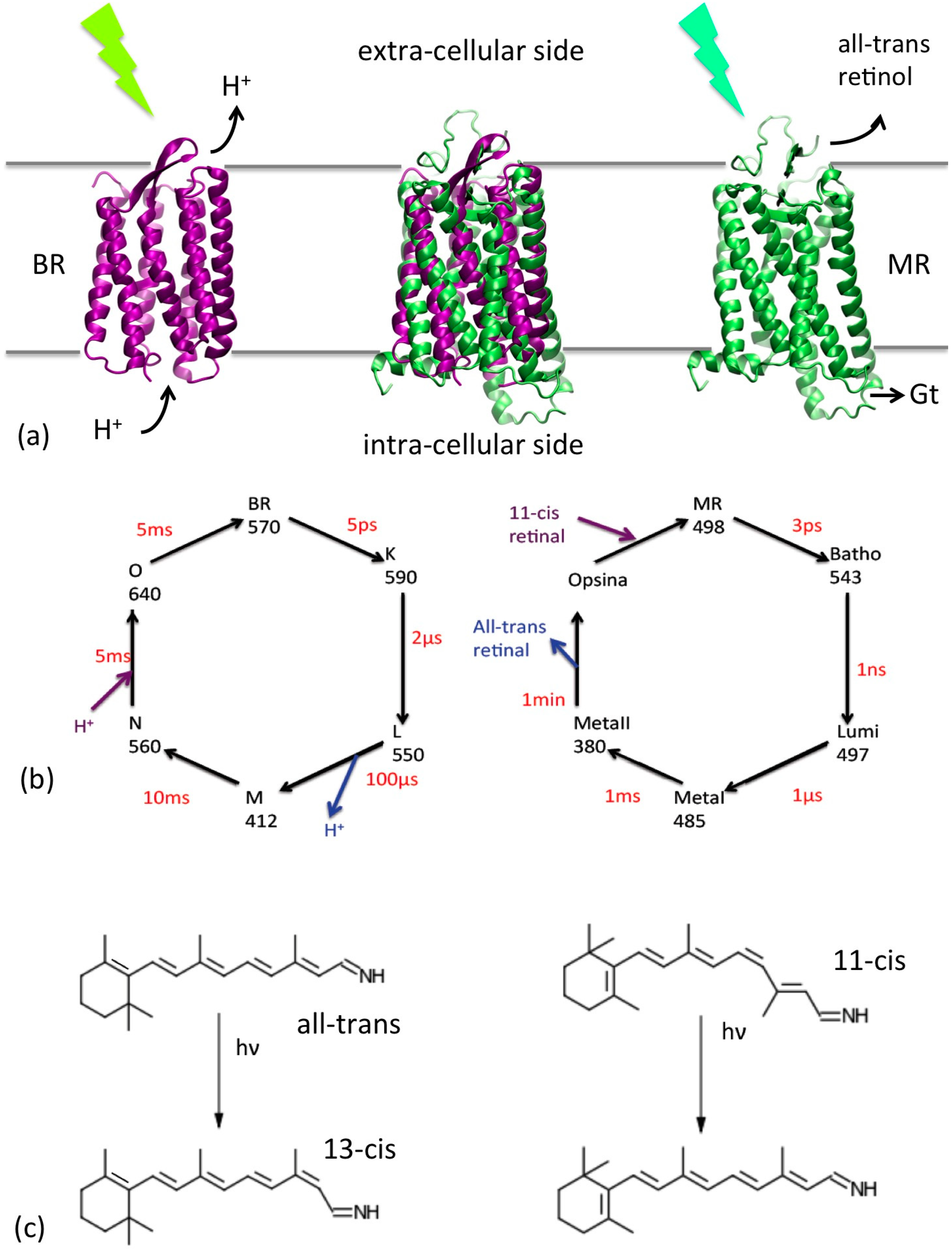
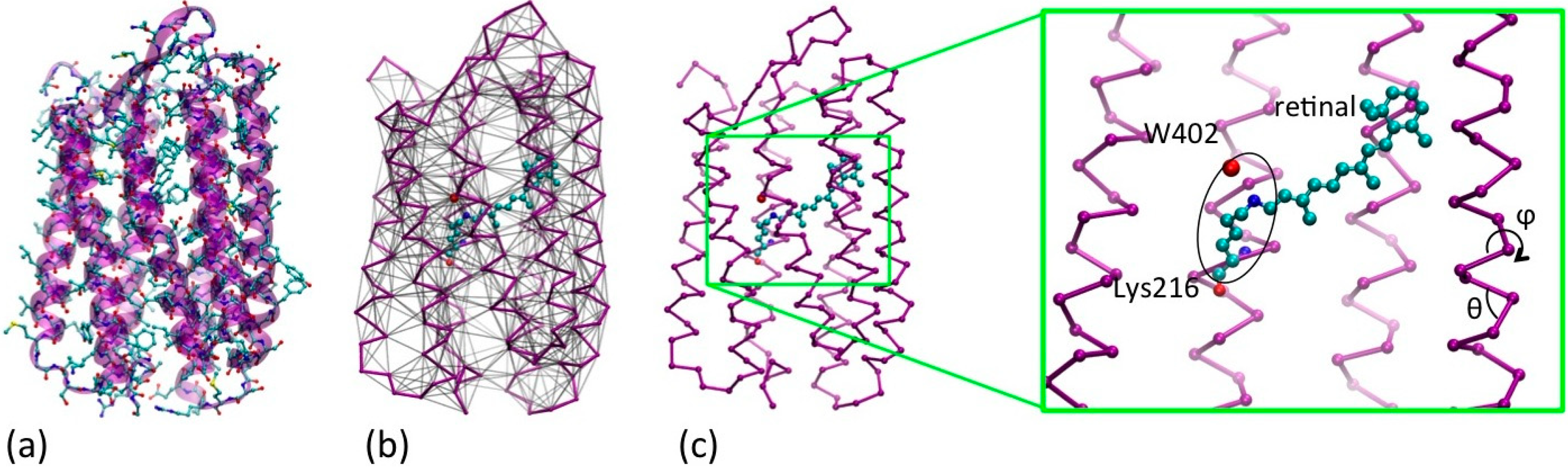
2. UA/CG Models of Rhodopsins
2.1. Model and Force Field Definition
2.2. Single States Multi-Scale Model Building and Parameterization
2.3. Multi-Stable Model Building
3. Results and Discussion
3.1. Single States Dynamics
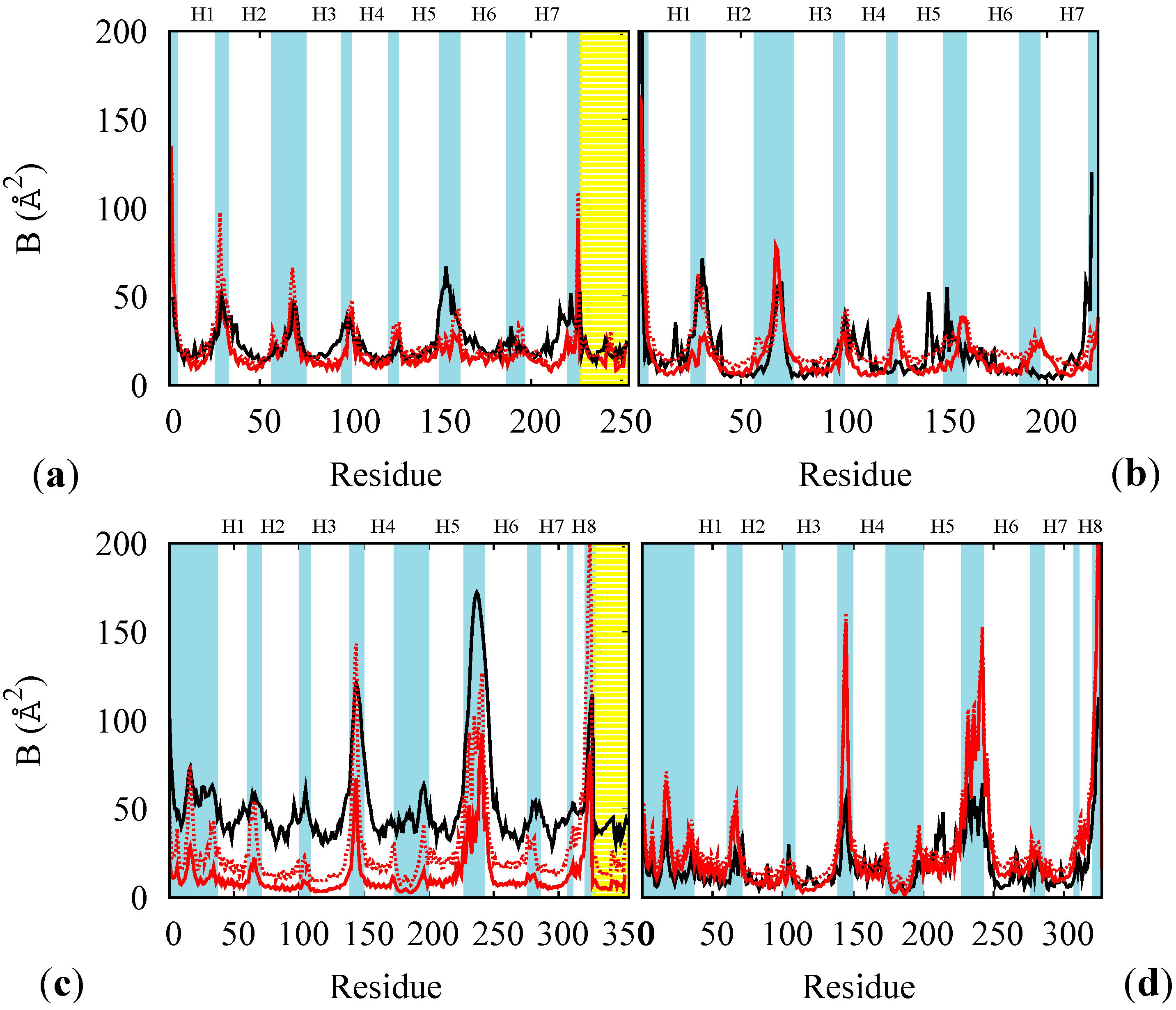
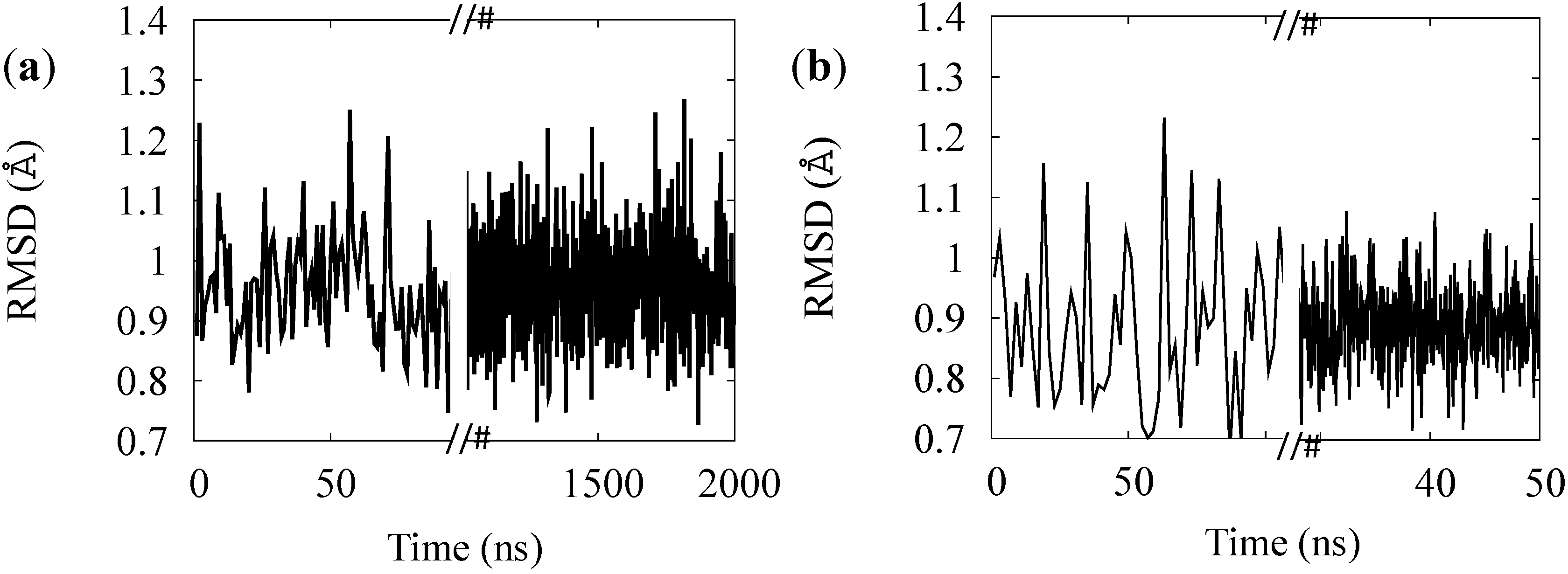
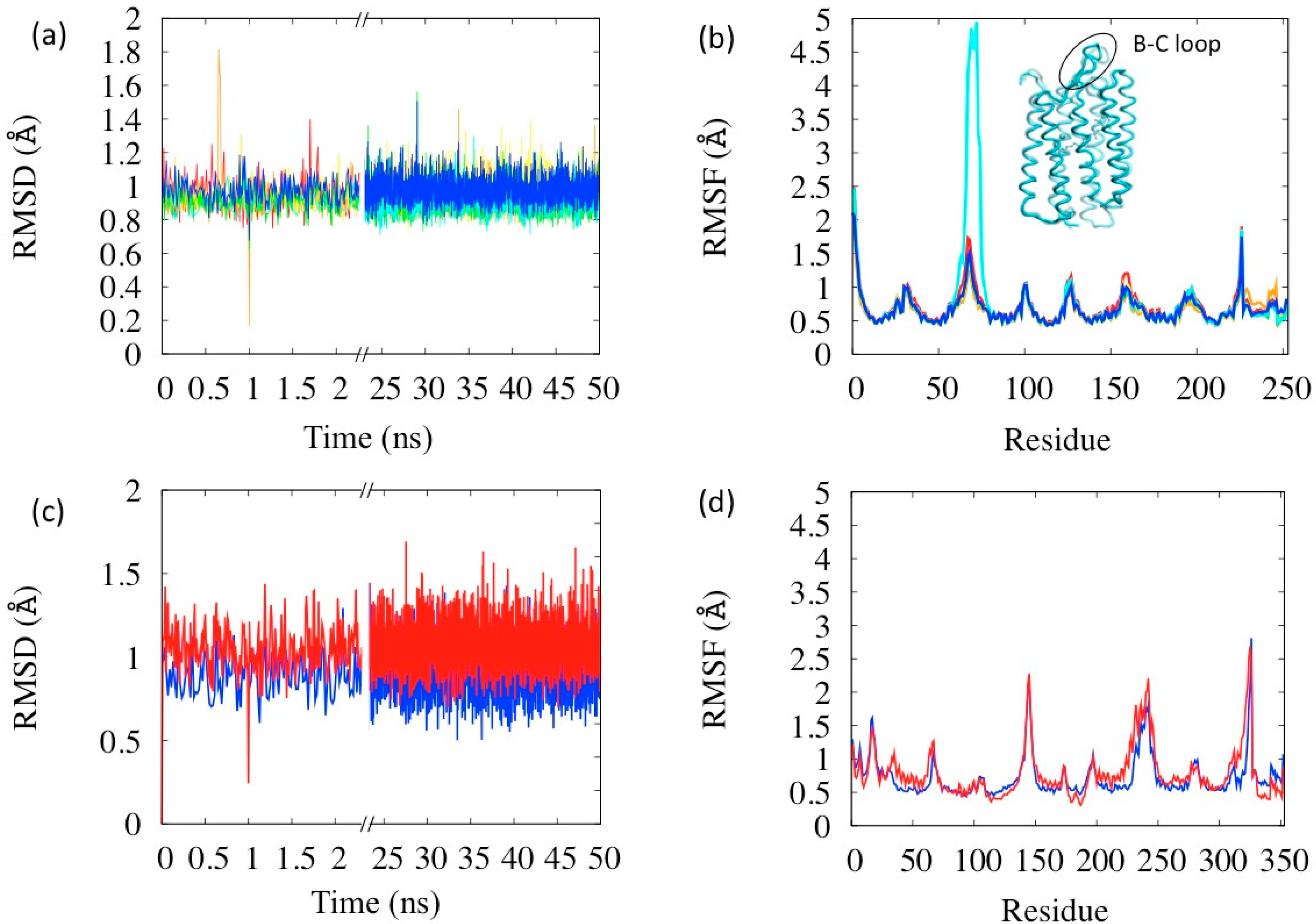
3.2. Simulation of the Photo-Cycle Dynamics
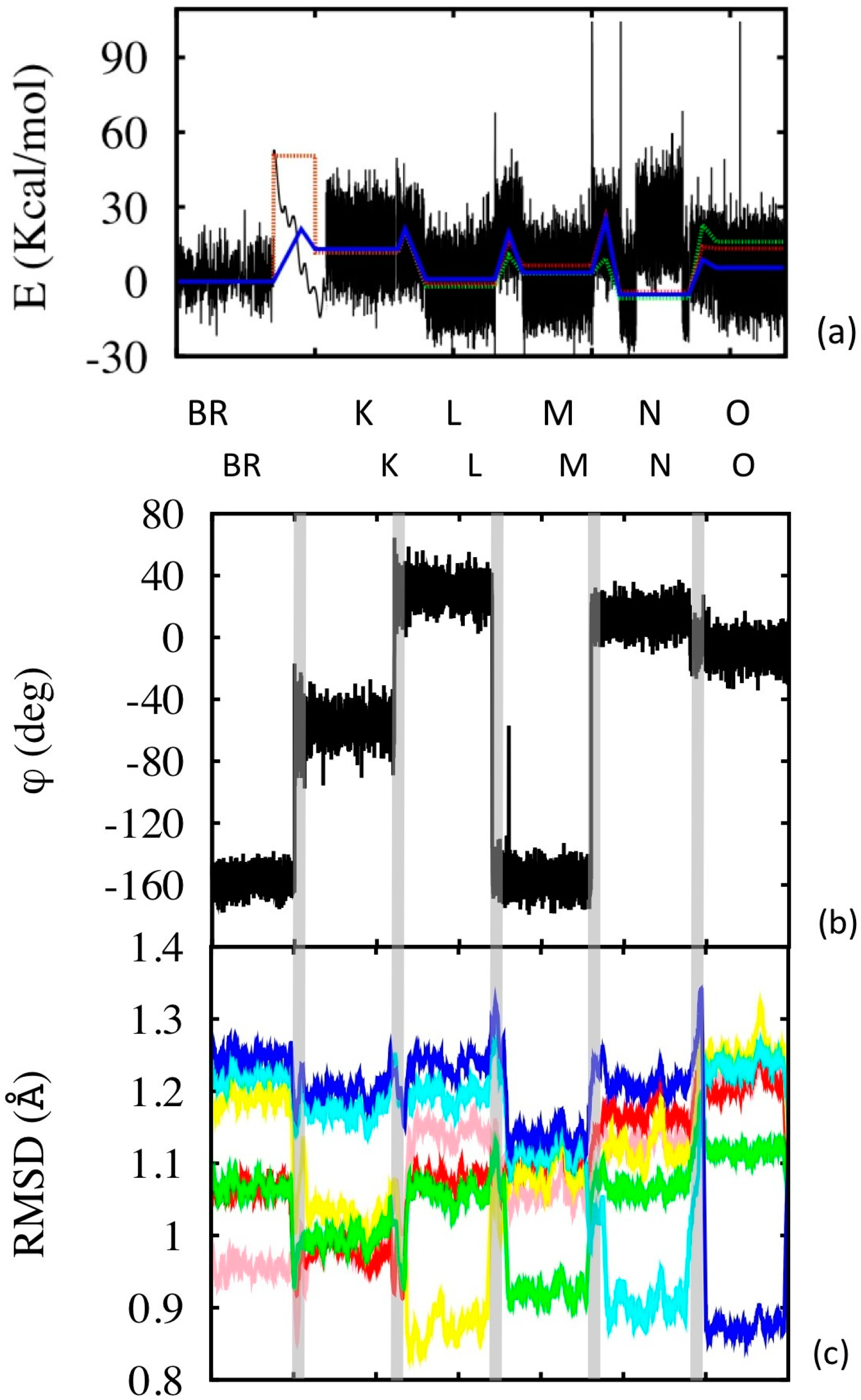
3.3. Structural Features of the Transitions
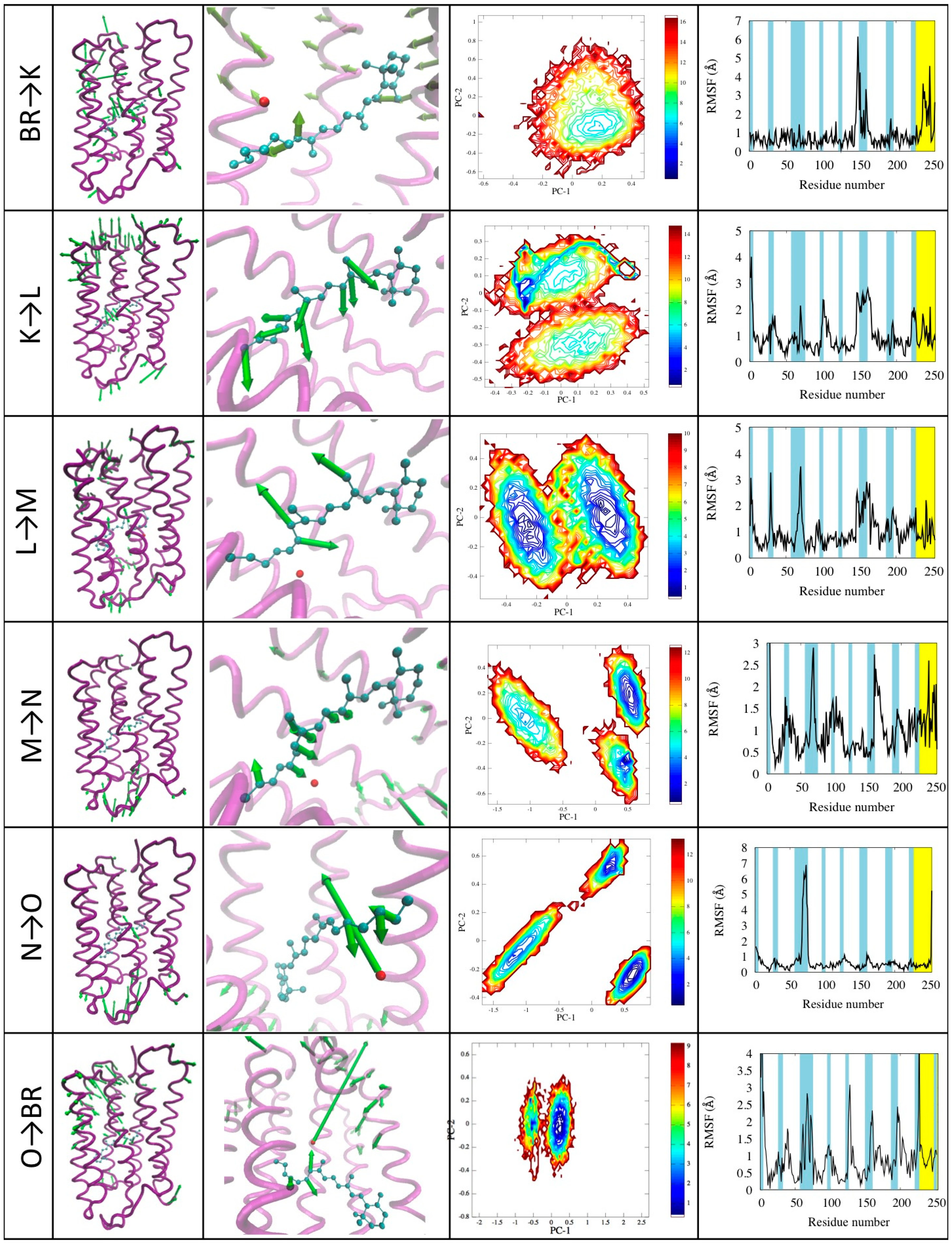
4. Experimental Section
4.1. Simulation Protocols and Tools
4.2. Simulation Analysis
5. Conclusions and Perspectives
Supplementary Materials
Supplementary Files
Supplementary File 1Acknowledgments
Author Contributions
Conflicts of Interest
References
- Lanyi, J.K. Bacteriorhodopsin. Annu. Rev. Physiol. 2004, 66, 665–688. [Google Scholar] [CrossRef] [PubMed]
- Kolbe, M.; Besir, H.; Essen, L.O.; Oesterhelt, D. Structure of the Light-Driven Chloride Pump Halorhodopsin at 1.8 Å Resolution. Science 2000, 288, 1390–1396. [Google Scholar]
- Lanyi, J.K.; Schobert, B. Mechanism of Proton Transport in Bacteriorhodopsin from Crystallographic Structures of the K, L, M1, M2, and M2' Intermediates of the Photocycle. J. Mol. Biol. 2003, 328, 439–450. [Google Scholar] [CrossRef] [PubMed]
- Teller, D.C.; Okada, T.; Behnke, C.A.; Palczewski, K.; Stenkamp, R.E. Advances in Determination of a High-Resolution Three- Dimensional Structure of Rhodopsin, a Model of G-Protein- Coupled Receptors (GPCRs). Biochemistry 2006, 40, 7761–7772. [Google Scholar] [CrossRef]
- Lozier, R.H.; Bogomolni, R.A.; Stoeckenius, W. Bacteriorhodopsin: A light-driven proton pump of Halobacterium Halobium. Biophys. J. 1975, 15, 955–962. [Google Scholar]
- Subramaniam, S.; Henderson, R. Molecular mechanism of vectorial proton translocation by bacteriorhodopsin. Nature 2000, 406, 653–657. [Google Scholar] [CrossRef] [PubMed]
- Royant, A.; Edman, K.; Ursby, T.; Pebay-Peyroula, E.; Landau, E.M.; Neutze, R. Helix deformation is coupled to vectorial proton transport in the photocycle of bacteriorhodopsin. Nature 2000, 406, 645–648. [Google Scholar]
- Piechnick, R.; Ritter, E.; Hildebrand, P.W.; Ernst, O.P.; Scheerer, P.; Hofmann, K.P.; Heck, M. Effect of channel mutations on the uptake and release of the retinal ligand in opsin. Proc. Natl. Acad. Sci. USA 2012, 109, 5247–5252. [Google Scholar] [CrossRef] [PubMed]
- Braun-Sand, S.; Sharma, P.K.; Chu, Z.T.; Pisliakov, A.V.; Warshel, A. The energetics of the primary proton transfer in bacteriorhodopsin revisited: It is a sequential light-induced charge separation after all. Biochim. Biophys. Acta 2008, 1777, 441–452. [Google Scholar] [CrossRef] [PubMed]
- Birge, R.R.; Cooper, T.M. Energy storage in the primary step of the photocycle of bacteriorhodopsin. Biophys. J. 1983, 42, 61–69. [Google Scholar] [CrossRef] [PubMed]
- Karplus, M.; McCammon, J.A. Molecular Dynamics Simulations of Biomolecules. Nat. Struct. Biol. 2002, 9, 646–652. [Google Scholar] [CrossRef] [PubMed]
- Rajamani, R.; Gao, J. Combined QM/MM Study of the Opsin Shift in Bacteriorhodopsin. J. Comput. Chem. 2002, 23, 96–105. [Google Scholar] [CrossRef] [PubMed]
- Valsson, O.; Campomanes, P.; Tavernelli, I.; Rothlisberger, U.; Filippi, C. Rhodopsin Absorption from First Principles: Bypassing Common Pitfalls. J. Chem. Theory Comput. 2013, 9, 2441–2454. [Google Scholar] [CrossRef]
- Weiner, S.J.; Kollman, P.A.; Case, D.A.; Singh, C.; Ghio, C.; Alagona, G.; Weiner, P. A New Force Field for Molecular Mechanical Simulation of Nucleic Acids and Proteins. J. Am. Chem. Soc. 1984, 106, 765–784. [Google Scholar] [CrossRef]
- Trovato, F.; Tozzini, V. Minimalist Models for Biopolymers: Open Problems, Latest Advances and Perspectives. AIP Conf. Proc. 2012, 1456, 187–200. [Google Scholar]
- Capece, L.; Nguyen, H.H.C.; Leguebe, M.; Giorgetti, A.; Carloni, P. Hybrid Molecular Mechanics/Coarse-Grained Calculations Applied to GPCR Receptors. Biophys. J. 2012, 102, 63a. [Google Scholar]
- Weiner, S.J.; Kollman, P.A; Nguyen, D.T.; Case, D.A. An all atom force field for simulations of proteins and nucleic acids. J. Comput. Chem. 1986, 7, 230–252. [Google Scholar]
- Tozzini, V.; McCammon, J.A. A coarse grained model for the dynamics of flap opening in HIV-1 Protease. Chem. Phys. Lett. 2005, 413, 123–128. [Google Scholar] [CrossRef]
- Trylska, J.; Tozzini, V.; McCammon, J.A. Exploring global motions and correlations in the ribosome. Biophys. J. 2005, 89, 1455–1463. [Google Scholar]
- Trovato, F.; Nifosì, R.; di Fenza, A.; Tozzini, V. A Minimalist Model of Protein Diffusion and Interactions: The Green Fluorescent Protein within the Cytoplasm. Macromolecules 2013, 46, 8311–8322. [Google Scholar] [CrossRef]
- Voltz, K.; Trylska, J.; Tozzini, V.; Kurkal-Siebert, V.; Langowski, J.; Smith, J. Coarse-grained force field for the nucleosome from self-consistent multiscaling. J. Comput. Chem. 2008, 29, 1429–1439. [Google Scholar] [CrossRef] [PubMed]
- Di Fenza, A.; Rocchia, W.; Tozzini, V. Complexes of HIV-1 integrase with HAT proteins: Multiscale models, dynamics, and hypotheses on allosteric sites of inhibition. Proteins 2009, 76, 946–958. [Google Scholar] [CrossRef] [PubMed]
- Edman, K.; Nollert, P.; Royant, A.; Belrhali, H.; Pebay-Peyroula, E.; Hajdu, J.; Neutze, R.; Landau, E.M. High-resolution X-ray structure of an early intermediate in the bacteriorhodopsin photocycle. Nature 1999, 401, 822–826. [Google Scholar] [CrossRef] [PubMed]
- Lanyi, J.K.; Schobert, B. Crystallographic Structure of the Retinal and the Protein after Deprotonation of the Schiff Base: The Switch in the Bacteriorhodopsin Photocycle. J. Mol. Biol. 2002, 2836, 727–737. [Google Scholar] [CrossRef]
- Chen, D.; Lanyi, J.K. Structural changes in the N and N' states of the bacteriorhodopsin photocycle. Biophys. J. 2009, 96, 2779–2788. [Google Scholar] [CrossRef] [PubMed]
- Zhang, J.; Yamazaki, Y.; Hikake, M.; Murakami, M.; Ihara, K.; Kouyama, T. Crystal structure of the O intermediate of the Leu93→Ala mutant of bacteriorhodopsin. Proteins 2012, 80, 2384–2396. [Google Scholar] [CrossRef] [PubMed]
- Berman, H.M.; Westbrook, J.; Feng, Z.; Gilliland, G.; Bhat, T.N.; Weissig, H.; Shindyalov, I.N.; Bourne, P.E. The Protein Data Bank. Nucleic Acids Res. 2000, 28, 235–242. [Google Scholar] [CrossRef] [PubMed]
- Jorgensen, W.L.; Tirado-Rives, J. The OPLS Potential Functions for Protins. Energy Minimizations for Crystals of Cyclic Peptides and Crambin. J. Am. Chem. Soc. 1998, 110, 1666–1671. [Google Scholar]
- Chu, J.-W.; Voth, G.A. Coarse-grained free energy functions for studying protein conformational changes: A double-well network model. Biophys. J. 2007, 93, 3860–3871. [Google Scholar] [CrossRef] [PubMed]
- Tekpinar, M.; Zheng, W. Predicting order of conformational changes during protein conformational transitions using an interpolated elastic network model. Proteins 2010, 78, 2469–2481. [Google Scholar] [PubMed]
- Maragakis, P.; Karplus, M.; Pasteur, L. Large Amplitude Conformational Change in Proteins Explored with a Plastic Network Model : Adenylate Kinase. J. Mol. Biol. 2005, 352, 807–822. [Google Scholar] [CrossRef] [PubMed]
- Trovato, F.; Tozzini, V. Diffusion within the cytoplasm: A meso-scale model of interacting macromolecules. 2014. submitted. [Google Scholar]
- Rueda, M.; Ferrer-Costa, C.; Meyer, T.; Pérez, A.; Camps, J.; Hospital, A.; Gelpì, J.L.; Orozco, M. A consensus of protein dynamics. Proc. Natl. Acad. Sci. USA 2007, 104, 796–801. [Google Scholar] [CrossRef] [PubMed]
- Poland, H.J.; Franz, M.A.; Zinth, W.; Kaiser, W.; Kölling, E.; Oesterhelt, D. Early picoseconds events in the photocycle of bacteriorhodopsin. Biophys. J. 1986, 49, 651–662. [Google Scholar]
- Takeda, K.; Matsui, Y.; Kamiya, N.; Adachi, S.; Okumura, H.; Kouyama, T. Crystal Structure of the M Intermediate of Bacteriorhodopsin: Allosteric Structural Changes Mediated by Sliding Movement of a Transmembrane Helix. J. Mol. Biol. 2004, 341, 1023–1037. [Google Scholar] [CrossRef] [PubMed]
- Verlet, L. Computer “Experiments” on Classical Fluids. I. Thermodynamical Properties of Lennad-Jones Molecules. Phys. Rev. 1967, 159, 98–103. [Google Scholar]
- Berendsen, H.J.C.; Postma, J.P.M.; van Gusteren, W.F.; DiNola, A.; Haak, J.R. Molecular dynamics with coupling to an external bath. J. Chem Phys. 1984, 81, 3684–3690. [Google Scholar] [CrossRef]
- Smith, W.; Forester, T.R. DL_POLY_2.0: A general-purpose parallel molecular dynamics simulation package Overall design. J. Mol. Graph. 1996, 7855, 136–141. [Google Scholar] [CrossRef]
- Yong, C.W. DL_FIELD-A Force Field and Model Development Tool for DL_POLY; Blake, R., Ed.; CSE Frontier, STFC Computational Science and Engineering, Daresbury Laboratory: Daresbury, UK, 2010; pp. 38–40. [Google Scholar]
- Jorgensen, W.L.; Maxwell, D.S.; Tirado-Rives, J. Development and testing of the OPLS all-atom force field on conformational energetics and properties of organic liquids. J. Am. Chem. Soc. 1996, 118, 11225–11236. [Google Scholar] [CrossRef]
- Lindahl, E.; Hess, B.; van der Spoel, D. GROMACS 3.0: A package for molecular simulation and trajectory analysis. J. Mol. Mod. 2001, 7, 306–317. [Google Scholar]
- Lindorff-Larsen, K.; Maragakis, P.; Piana, S.; Eastwood, M.P.; Dror, R.O.; Shaw, D.E. Systematic validation of protein force fields against experimental data. PLoS One 2012, 7, 1–6. [Google Scholar] [CrossRef]
- Garcia, A.E. Large-Amplitude Nonlinear Motions in Proteins. Phys. Rev. Lett. 1992, 68, 2696–2699. [Google Scholar] [CrossRef] [PubMed]
- Amadei, A.; Linssen, A.B.M.; Berendsen, H.J.C. Essential Dynamics of Proteins. Proteins 1993, 17, 412–425. [Google Scholar] [CrossRef] [PubMed]
- Tavernelli, I.; Cotesta, S.; di Iorio, E.E. Protein dynamics, thermal stability, and free-energy landscapes: A molecular dynamics investigation. Biophys. J. 2003, 85, 2641–2649. [Google Scholar] [CrossRef] [PubMed]
- Papaleo, E.; Mereghetti, P.; Fantucci, P.; Grandori, R.; de Gioia, L. Free-energy landscape, principal component analysis, and structural clustering to identify representative conformations from molecular dynamics simulations : The myoglobin case. J. Mol. Graph. Model. 2009, 27, 889–899. [Google Scholar] [CrossRef] [PubMed]
- Bakan, A.; Meireles, L.M.; Bahar, I. ProDy: Protein Dynamics Inferred from Theory and Experiments. Bioinformatics 2011, 27, 1575–1577. [Google Scholar] [CrossRef] [PubMed]
- Humphrey, W.; Dalke, A.; Schulten, K. VMD-Visual Molecular Dynamics. J. Mol. Graph. 1996, 14, 33–38. [Google Scholar] [CrossRef] [PubMed]
- Sample Availability: Not available.
© 2014 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Tavanti, F.; Tozzini, V. A Multi-Scale–Multi-Stable Model for the Rhodopsin Photocycle. Molecules 2014, 19, 14961-14978. https://doi.org/10.3390/molecules190914961
Tavanti F, Tozzini V. A Multi-Scale–Multi-Stable Model for the Rhodopsin Photocycle. Molecules. 2014; 19(9):14961-14978. https://doi.org/10.3390/molecules190914961
Chicago/Turabian StyleTavanti, Francesco, and Valentina Tozzini. 2014. "A Multi-Scale–Multi-Stable Model for the Rhodopsin Photocycle" Molecules 19, no. 9: 14961-14978. https://doi.org/10.3390/molecules190914961
APA StyleTavanti, F., & Tozzini, V. (2014). A Multi-Scale–Multi-Stable Model for the Rhodopsin Photocycle. Molecules, 19(9), 14961-14978. https://doi.org/10.3390/molecules190914961






