Studies on Cytotoxic Constituents from the Leaves of Elaeagnus oldhamii Maxim. in Non-Small Cell Lung Cancer A549 Cells
Abstract
:1. Introduction
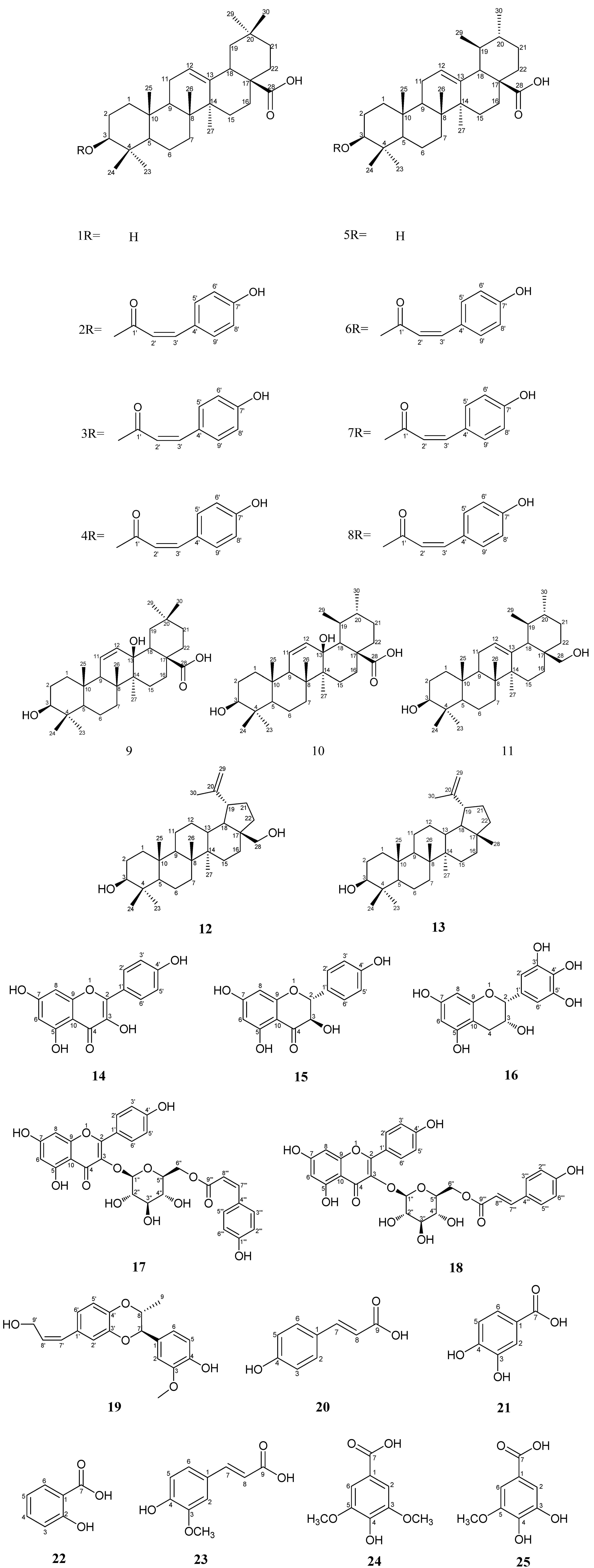
2. Results and Discussion
| Position | 1 d | 2 b | 3 c | 4 d | 5 d | 6 b | |||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 39.4 | 37.2 | 39.3 | 38.6 | 39.4 | 38.0 | |||||||
| 2 | 28.5 | 23.1 | 24.1 | 24.1 | 28.4 | 28.2 | |||||||
| 3 | 78.5 | 81.4 | 85.7 | 80.9 | 78.5 | 81.4 | |||||||
| 4 | 39.8 | 38.0 | 39.4 | 38.7 | 39.8 | 38.5 | |||||||
| 5 | 56.2 | 55.5 | 56.6 | 56.1 | 56.2 | 55.5 | |||||||
| 6 | 19.2 | 18.4 | 19.6 | 19.0 | 19.1 | 18.4 | |||||||
| 7 | 33.7 | 32.7 | 33.7 | 33.5 | 33.9 | 33.0 | |||||||
| 8 | 40.2 | 39.5 | 40.8 | 40.2 | 40.3 | 39.7 | |||||||
| 9 | 48.6 | 47.7 | 49.1 | 48.4 | 48.4 | 48.2 | |||||||
| 10 | 37.8 | 38.3 | 39.4 | 37.7 | 37.8 | 37.1 | |||||||
| 11 | 24.2 | 23.7 | 24.8 | 24.6 | 34.0 | 23.5 | |||||||
| 12 | 123.0 | 122.8 | 123.5 | 122.8 | 126.0 | 126.0 | |||||||
| 13 | 145.2 | 143.8 | 146.4 | 144.4 | 139.6 | 138.2 | |||||||
| 14 | 42.6 | 41.8 | 43.1 | 42.6 | 42.9 | 42.1 | |||||||
| 15 | 28.7 | 27.9 | 28.9 | 28.8 | 29.0 | 28.3 | |||||||
| 16 | 24.2 | 22.9 | 24.1 | 24.2 | 25.3 | 24.2 | |||||||
| 17 | 47.1 | 46.8 | 47.8 | 47.2 | 48.4 | 47.6 | |||||||
| 18 | 42.4 | 41.1 | 42.9 | 42.5 | 53.9 | 52.7 | |||||||
| 19 | 46.9 | 46.1 | 47.4 | 47.0 | 39.7 | 39.2 | |||||||
| 20 | 31.4 | 30.9 | 29.4 | 31.5 | 39.7 | 39.0 | |||||||
| 21 | 34.7 | 34.0 | 35.0 | 34.8 | 31.4 | 30.8 | |||||||
| 22 | 33.6 | 32.7 | 31.8 | 32.6 | 37.6 | 37.0 | |||||||
| 23 | 29.2 | 28.3 | 28.9 | 28.7 | 29.2 | 29.9 | |||||||
| 24 | 17.0 | 15.6 | 17.9 | 17.9 | 16.9 | 17.0 | |||||||
| 25 | 16.0 | 16.9 | 17.2 | 15.9 | 16.0 | 15.8 | |||||||
| 26 | 17.9 | 17.4 | 18.4 | 17.6 | 17.8 | 17.2 | |||||||
| 27 | 26.6 | 26.1 | 26.6 | 26.7 | 24.2 | 23.8 | |||||||
| 28 | 180.6 | 184.3 | 182.0 | 180.9 | 180.2 | 184.1 | |||||||
| 29 | 33.7 | 33.3 | 33.9 | 33.8 | 17.8 | 17.3 | |||||||
| 30 | 24.1 | 23.6 | 24.2 | 23.8 | 21.8 | 21.4 | |||||||
| 1' | - | 167.0 | 169.8 | 167.8 | - | 167.0 | |||||||
| 2' | - | 117.7 | 116.0 | 116.4 | - | 117.7 | |||||||
| 3' | - | 143.7 | 145.5 | 145.4 | - | 143.8 | |||||||
| 4' | - | 127.5 | 127.5 | 127.4 | - | 127.5 | |||||||
| 5' | - | 132.4 | 131.2 | 116.0 | - | 132.4 | |||||||
| 6' | - | 115.3 | 117.0 | 146.2 | - | 115.3 | |||||||
| 7' | - | 157.1 | 161.3 | 149.9 | - | 157.2 | |||||||
| 8' | - | 115.3 | 117.0 | 117.4 | - | 115.3 | |||||||
| 9' | - | 132.4 | 131.2 | 122.5 | - | 132.4 | |||||||
| Position | 7 c | 8 c | 9 b | 10 b | 11 b | 12 b | 13 b | ||||||
| 1 | 39.3 | 37.9 | 37.5 | 38.4 | 39.0 | 38.9 | 38.9 | ||||||
| 2 | 29.3 | 24.5 | 23.8 | 23.0 | 27.5 | 27.6 | 29.9 | ||||||
| 3 | 85.7 | 81.0 | 79.0 | 79.1 | 79.2 | 79.2 | 79.2 | ||||||
| 4 | 40.6 | 37.6 | 38.5 | 39.1 | 38.2 | 38.9 | 39.1 | ||||||
| 5 | 56.6 | 56.2 | 53.4 | 55.0 | 55.4 | 55.5 | 55.5 | ||||||
| 6 | 19.6 | 19.8 | 18.2 | 17.9 | 18.5 | 18.5 | 18.6 | ||||||
| 7 | 34.3 | 33.5 | 29.9 | 29.9 | 33.0 | 34.5 | 34.5 | ||||||
| 8 | 41.0 | 40.3 | 41.6 | 41.9 | 40.2 | 39.8 | 41.1 | ||||||
| 9 | 50.0 | 47.6 | 50.7 | 53.3 | 47.9 | 50.6 | 50.7 | ||||||
| 10 | 38.2 | 36.6 | 34.6 | 36.6 | 37.1 | 37.5 | 37.4 | ||||||
| 11 | 24.3 | 24.2 | 136.1 | 133.7 | 23.6 | 21.0 | 21.2 | ||||||
| 12 | 126.8 | 123.0 | 127.1 | 129.1 | 125.3 | 25.4 | 23.6 | ||||||
| 13 | 139.9 | 145.1 | 90.1 | 89.9 | 138.9 | 37.4 | 38.3 | ||||||
| 14 | 43.5 | 42.7 | 41.8 | 42.2 | 42.3 | 41.1 | 43.1 | ||||||
| 15 | 29.4 | 28.6 | 27.4 | 27.2 | 26.2 | 27.3 | 27.7 | ||||||
| 16 | 25.4 | 26.4 | 25.6 | 25.8 | 23.6 | 29.4 | 35.8 | ||||||
| 17 | 49.3 | 46.9 | 44.2 | 45.3 | 37.1 | 48.0 | 42.2 | ||||||
| 18 | 54.4 | 48.5 | 55.0 | 60.8 | 54.2 | 42.9 | 48.5 | ||||||
| 19 | 40.8 | 39.0 | 36.6 | 38.5 | 39.6 | 49.0 | 48.2 | ||||||
| 20 | 40.6 | 38.9 | 39.2 | 40.5 | 39.6 | 150.7 | 151.2 | ||||||
| 21 | 31.9 | 31.4 | 31.4 | 31.5 | 30.8 | 30.0 | 30.1 | ||||||
| 22 | 38.2 | 36.9 | 31.7 | 31.6 | 35.4 | 34.2 | 40.2 | ||||||
| 23 | 31.0 | 28.5 | 28.0 | 28.0 | 28.3 | 28.2 | 28.2 | ||||||
| 24 | 17.8 | 17.1 | 15.1 | 15.1 | 15.9 | 15.6 | 14.8 | ||||||
| 25 | 17.4 | 15.9 | 17.9 | 16.3 | 15.8 | 16.3 | 16.3 | ||||||
| 26 | 17.9 | 17.7 | 19.2 | 19.1 | 17.0 | 16.2 | 16.2 | ||||||
| 27 | 24.6 | 24.0 | 18.5 | 18.1 | 23.5 | 14.7 | 15.6 | ||||||
| 28 | 181.8 | 179.1 | 180.2 | 180.1 | 70.1 | 60.8 | 18.2 | ||||||
| 29 | 18.5 | 17.7 | 33.5 | 18.0 | 17.6 | 109.9 | 109.5 | ||||||
| 30 | 21.7 | 23.9 | 27.2 | 19.4 | 21.5 | 19.3 | 19.5 | ||||||
| 1' | 169.8 | 167.3 | - | - | - | - | - | ||||||
| 2' | 116.0 | 116.2 | - | - | - | - | - | ||||||
| 3' | 146.4 | 145.5 | - | - | - | - | - | ||||||
| 4' | 127.5 | 127.2 | - | - | - | - | - | ||||||
| 5' | 131.2 | 115.2 | - | - | - | - | - | ||||||
| 6' | 117.0 | 146.5 | - | - | - | - | - | ||||||
| 7' | 161.3 | 148.9 | - | - | - | - | - | ||||||
| 8' | 117.0 | 116.4 | - | - | - | - | - | ||||||
| 9' | 131.2 | 122.5 | - | - | - | - | - | ||||||
| Position | 14 c | 15 c | 16 c | 17 a | 18 b | ||||||
|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | - | - | - | - | - | ||||||
| 2 | 148.2 | 85.1 | 79.5 | 158.8 | 159.5 | ||||||
| 3 | 137.3 | 73.8 | 67.1 | 135.3 | 135.4 | ||||||
| 4 | 177.5 | 198.5 | 30.7 | 179.1 | 179.4 | ||||||
| 5 | 162.7 | 165.5 | 157.6 | 158.0 | 158.5 | ||||||
| 6 | 99.4 | 97.6 | 96.2 | 99.8 | 100.1 | ||||||
| 7 | 165.8 | 169.4 | 157.6 | 165.4 | 166.0 | ||||||
| 8 | 94.6 | 96.6 | 95.8 | 94.8 | 95.0 | ||||||
| 9 | 158.4 | 164.7 | 157.2 | 162.9 | 163.1 | ||||||
| 10 | 104.7 | 101.9 | 99.9 | 105.5 | 105.7 | ||||||
| 1' | 123.9 | 129.5 | 131.6 | 122.4 | 122.9 | ||||||
| 2' | 130.8 | 130.5 | 107.0 | 132.4 | 132.4 | ||||||
| 3' | 116.5 | 116.3 | 146.2 | 116.1 | 116.2 | ||||||
| 4' | 140.7 | 159.4 | 133.0 | 161.0 | 161.5 | ||||||
| 5' | 116.5 | 116.3 | 146.2 | 116.1 | 116.2 | ||||||
| 6' | 130.8 | 130.5 | 107.0 | 132.4 | 132.4 | ||||||
| 1'' | - | - | - | 104.5 | 104.2 | ||||||
| 2'' | - | - | - | 75.5 | 75.9 | ||||||
| 3'' | - | - | - | 71.1 | 71.9 | ||||||
| 4'' | - | - | - | 75.3 | 75.9 | ||||||
| 5'' | - | - | - | 78.1 | 78.1 | ||||||
| 6'' | - | - | - | 63.9 | 64.5 | ||||||
| 1''' | - | - | - | 127.3 | 127.2 | ||||||
| 2''' | - | - | - | 133.8 | 133.9 | ||||||
| 3''' | - | - | - | 115.7 | 115.9 | ||||||
| 4''' | - | - | - | 159.7 | 161.3 | ||||||
| 5''' | - | - | - | 115.7 | 115.9 | ||||||
| 6''' | - | - | - | 133.8 | 133.9 | ||||||
| 7''' | - | - | - | 144.5 | 146.7 | ||||||
| 8''' | - | - | - | 116.3 | 116.9 | ||||||
| 9''' | - | - | - | 166.3 | 169.0 | ||||||
| Position | 19 a | 20 c | 21 c | 22 a | 23 c | 24 c | 25 c | ||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|---|
| 1 | 129.4 | 127.4 | 123.2 | 114.3 | 128.1 | 120.6 | 122.4 | ||||||||
| 2 | 111.3 | 131.2 | 117.5 | 163.3 | 111.8 | 106.8 | 106.5 | ||||||||
| 3 | 148.5 | 116.9 | 145.7 | 118.3 | 150.6 | 147.4 | 146.3 | ||||||||
| 4 | 147.6 | 161.3 | 150.8 | 136.6 | 149.5 | 140.1 | 140.5 | ||||||||
| 5 | 117.9 | 116.9 | 115.8 | 120.2 | 116.6 | 147.4 | 149.2 | ||||||||
| 6 | 120.3 | 131.2 | 123.7 | 131.7 | 124.0 | 106.8 | 112.5 | ||||||||
| 7 | 78.1 | 146.7 | 167.8 | 173.8 | 146.7 | 167.2 | 170.5 | ||||||||
| 8 | 73.8 | 116.0 | - | - | 116.6 | - | - | ||||||||
| 9 | 13.7 | 171.3 | - | - | 171.5 | - | - | ||||||||
| 1' | 56.5 | - | - | - | - | - | - | ||||||||
| 2' | 132.8 | - | - | - | - | - | - | ||||||||
| 3' | 118.3 | - | - | - | - | - | - | ||||||||
| 4' | 143.9 | - | - | - | - | - | - | ||||||||
| 5' | 142.6 | - | - | - | - | - | - | ||||||||
| 6' | 115.9 | - | - | - | - | - | - | ||||||||
| 7' | 123.5 | - | - | - | - | - | - | ||||||||
| 8' | 129.6 | - | - | - | - | - | - | ||||||||
| 9' | 131.5 | - | - | - | - | - | - | ||||||||
| OCH3 | 59.8 | 56.6 | 55.9 | 56.8 | |||||||||||
| Sample Code | Compound Name | CC50 (μg/mL) | |
|---|---|---|---|
| 24 h | 48 h | ||
| Positive control | Cisplatin | 132.05 ± 1.46 | 14.87 ± 1.94 |
| 1 | Oleanolic acid | 64.86 ± 3.90 | 52.97 ± 1.22 |
| 2 | 3-O-(Z)-coumaroyl oleanolic acid | 9.23 ± 1.73 | 9.46 ± 1.50 |
| 3 | 3-O-(E)-coumaroyl oleanolic acid | 135.12 ± 3.08 | 65.21 ± 7.15 |
| 4 | 3-O-caffeoyl oleanolic acid | 12.35 ± 1.52 | 13.76 ± 3.45 |
| 5 | Ursolic acid | 153.37 ± 3.64 | 154.73 ± 7.86 |
| 6 | 3-O-(Z)-coumaroyl ursolic acid | 13.06 ± 2.26 | 8.56 ± 0.57 |
| 7 | 3-O-(E)-coumaroyl ursolic acid | 121.53 ± 9.68 | 79.59 ± 5.77 |
| 8 | 3-O-caffeoyl ursolic acid | 157.98 ± 5.02 | 15.70 ± 1.24 |
| 9 | 3β, 13β-Dihydroxyolean-11-en-28-oic acid | 92.71 ± 3.89 | 70.61 ± 8.27 |
| 10 | 3β, 13β-Dihydroxyurs-11-en-28-oic acid | 176.81 ± 6.33 | 156.20 ± 8.10 |
| 11 | Uvaol | 169.45 ± 9.47 | 163.36 ± 7.57 |
| 12 | Betulin | >500 | >500 |
| 13 | Lupeol | 142.70 ± 4.99 | 119.30 ± 5.28 |
| 14 | Kaempferol | 129.15 ± 5.16 | 113.48 ± 5.32 |
| 15 | Aromadendrin | 76.66 ± 4.35 | 55.67 ± 2.37 |
| 16 | Epigallocatechin | 193.77 ± 3.98 | 118.92 ± 4.25 |
| 17 | cis-Tiliroside | 68.05 ± 1.46 | 18.82 ± 3.64 |
| 18 | trans-Tiliroside | 149.90 ± 4.14 | 144.74 ± 5.37 |
| 19 | Isoamericanol B | >500 | 126.97 ± 4.83 |
| 20 | trans-p-Coumaric acid | >500 | >500 |
| 21 | Protocatechuic acid | >500 | >500 |
| 22 | Salicylic acid | >500 | 103.64 ± 4.59 |
| 23 | trans-Ferulic acid | 78.09 ± 5.78 | 60.42 ± 5.00 |
| 24 | Syringic acid | 73.41 ± 2.92 | 77.66 ± 4.04 |
| 25 | 3-O-methylgallic acid | >500 | 180.60 ± 8.58 |
| MOM | 84.67 ± 0.37 | 89.98 ± 7.03 | |
| MOE | 52.78 ± 0.78 | 45.15 ± 1.10 | |
| MOB | 192.46 ± 6.54 | 166.38 ± 12.65 | |
| MOW | 100.91 ± 1.01 | 136.47 ± 3.96 | |
3. Experimental
3.1. General
3.2. Plant Material
3.3. Extraction and Isolation
3.4. Spectroscopic Data
3.5. Cytotoxicity Assays
3.5.1. Cell Culture
3.5.2. Evaluation of Cell Proliferation by MTT Assay
4. Conclusions
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Huang, T.S. Elaeagnaceae. In Flora of Taiwan, 2nd ed.; Editorial Committee of the Flora of Taiwan: Taipei, Taiwan, 1998; Volume 3, pp. 753–759. [Google Scholar]
- Ahmad, S.D.; Sabir, M.S.; Juma, M.; Asad, H.S. Morphological and biochemical variations in Elaeagnus umbellata Thunb. From mountains of Pakistan. Acta Bot. Croat. 2005, 64, 121–128. [Google Scholar]
- Yuebin, G.; Liu, J.; Su, D. In vivo evaluation of the anti-asthmatic, antitussive and expectorant activities of extract and fractions from Elaeagnus pungens leaf. J. Ethnopharmacol. 2009, 126, 538–542. [Google Scholar] [CrossRef]
- Ahmadiani, A.; Hosseiny, J.; Semnanian, S.; Javan, M.; Saeedi, F.; Kamalinejad, M.; Saremi, S. Antinociceptive and anti-inflammatory effects of Elaeagnus angustifolia fruit extract. J. Ethnopharmacol. 2000, 72, 287–292. [Google Scholar]
- Mehrabani Natanzi, M.; Pasalar, P.; Kamalinejad, M.; Dehpour, A.R.; Tavangar, S.M.; Sharifi, R.; Ghanadian, N.; Rahimi-Balaei, M.; Gerayesh-Nejad, S. Effect of aqueous extract of Elaeagnus angustifolia fruit on experimental cutaneous wound healing in rats. Acta Med. Iran. 2012, 50, 589–596. [Google Scholar]
- Lee, Y.S.; Chang, Z.Q.; Oh, B.C.; Park, S.C.; Shin, S.R.; Kim, N.W. Antioxidant activity, anti-inflammatory activity, and whitening effects of extracts of Elaeagnus multiflora Thunb. J. Med. Food 2007, 10, 126–133. [Google Scholar]
- Talaei-Khozani, T.; Vojdani, Z.; Dehghani, F.; Heidari, E.; Kharazinejad, E.; Panjehshahin, M.R. Toxic effects of Elaeagnus angustifolia fruit extract on chondrogenesis and osteogenesis in mouse limb buds. Tokai J. Exp. Clin. Med. 2011, 36, 63–70. [Google Scholar]
- Wang, S.Y.; Bowman, L.; Ding, M. Variations in free radical scavenging capacity and antiproliferative activity among different genotypes of autumn olive (Elaeagnus umbellata). Planta Med. 2007, 73, 468–477. [Google Scholar]
- Wu, Y.B.; Gu, Y.; Ouyang, M.A. Water-soluble constituents from the bark of Elaeagnus pungens and their cytotoxic activities. J. Asian Nat.Prod. Res. 2010, 12, 278–285. [Google Scholar]
- Li, L.H.; Baek, I.K.; Kim, J.H.; Kang, K.H.; Koh, Y.S.; Jung, Y.D.; Cho, C.K.; Choi, S.Y.; Shin, B.A. Methanol extract of Elaeagnus glabra, a Korean medicinal plant, inhibits HT1080 tumor cell invasion. Oncol. Rep. 2009, 21, 559–563. [Google Scholar]
- Yan, M.C.; Liu, Y.; Chen, H.; Ke, Y.; Xu, Q.C.; Cheng, M.S. Synthesis and antitumor activity of two natural N-acetylglucosamine-bearing triterpenoid saponins: Lotoidoside D and E. Bioorg. Med. Chem. Lett. 2006, 16, 4200–4204. [Google Scholar]
- Liu, Q.; Liu, H.; Zhang, L.; Guo, T.; Wang, P.; Geng, M.; Li, Y. Synthesis and antitumor activities of naturally occurring oleanolic acid triterpenoid saponins and their derivatives. Eur. J. Med.Chem. 2013, 64, 1–15. [Google Scholar]
- Tu, J.; Sun, H.X.; Ye, Y.P. Immunomodulatory and antitumor activity of triterpenoid fractions from the rhizomes of Astilbe chinensis. J. Ethnopharmacol. 2008, 119, 266–271. [Google Scholar]
- Park, K.I.; Park, H.S.; Nagappan, A.; Hong, G.E.; Lee do, H.; Kang, S.R.; Kim, J.A.; Zhang, J.; Kim, E.H.; Lee, W.S.; et al. Induction of the cell cycle arrest and apoptosis by flavonoids isolated from Korean Citrus aurantium L. in non-small cell lung cancer cells. Food Chem. 2012, 135, 2728–2735. [Google Scholar]
- Hung, J.Y.; Hsu, Y.L.; Ko, Y.C.; Tsai, Y.M.; Yang, C.J.; Huang, M.S.; Kuo, P.L. Didymin, a dietary flavonoid glycoside from citrus fruits, induces Fas-mediated apoptotic pathway in human non-small cell lung cancer cells in vitro and in vivo. Lung Cancer 2010, 68, 366–374. [Google Scholar]
- Leung, H.W.; Lin, C.J.; Hour, M.J.; Yang, W.H.; Wang, M.Y.; Lee, H.Z. Kaempferol induces apoptosis in human lung non-small carcinoma cells accompanied by an induction of antioxidant enzymes. Food Chem. Toxicol. 2007, 45, 2005–2013. [Google Scholar]
- Peuhu, E.; Paul, P.; Remes, M.; Holmbom, T.; Eklund, P.; Sjöholm, R.; Eriksson, J.E. The antitumor lignan Nortrachelogenin sensitizes prostate cancer cells to TRAIL-induced cell death by inhibition of the Akt pathway and growth factor signaling. Biochem. Pharmacol. 2013, 86, 571–583. [Google Scholar]
- Vijayakurup, V.; Spatafora, C.; Tringali, C.; Jayakrishnan, P.C.; Srinivas, P.; Gopala, S. Phenethyl caffeate benzoxanthene lignan is a derivative of caffeic acid phenethyl ester that induces bystander autophagy in WiDr cells. Mol. Biol. Rep. 2014, 41, 85–94. [Google Scholar]
- Gao, X.M.; Wang, R.R.; Niu, D.Y.; Meng, C.Y.; Yang, L.M.; Zheng, Y.T.; Yang, G.Y.; Hu, Q.F.; Sun, H.D.; Xiao, W.L. Bioactive dibenzocyclooctadiene lignans from the stems of Schisandra neglecta. J. Nat. Prod. 2013, 76, 1052–1057. [Google Scholar]
- Giraldi, T.; Guarino, A.M.; Nisi, C.; Sava, G. Antitumor and antimetastatic effects of benzenoid triazenes in mice bearing lewis lung carcinoma. Pharmacol. Res. Commun. 1980, 12, 1–11. [Google Scholar]
- Chang, K.C.; Duh, C.Y.; Chen, I.S.; Tsai, I.L. A cytotoxic butenolide, two new dolabellane diterpenoids, a chroman and a benzoquinol derivative formosan Casearia membranace. Planta Med. 2003, 69, 667–672. [Google Scholar]
- Chen, P.Y.; Wu, J.D.; Tang, K.Y.; Yu, C.C.; Kuo, Y.H.; Zhong, W.B.; Lee, C.K. Isolation and synthesis of a bioactive benzenoid derivative from the fruiting bodies of Antrodia camphorata. Molecules 2013, 18, 7600–7608. [Google Scholar]
- Ge, Y.B.; Li, M.S.; Mei, Z.N.; Yang, G.Z. Two new flavonol glycosides from the leaves of Elaeagnus pungens. J. Asian Nat. Prod. Res. 2013, 15, 1073–1079. [Google Scholar]
- Lou, F.M.; Yang, J.; Bai, Z.C.; Wu, B.F. Studies on chemical constituents in rhizome of Elaeagnus bockii. Zhongguo Zhong Yao Za Zhi 2006, 31, 988–989. [Google Scholar]
- Cao, S.G.; Tanaka, T.; Mizuno, M.; Inoue, K. Flavonol glycosides from Elaeagnus lanceollata (Elaeagnaceae). Nat. Prod. Lett. 2001, 15, 211–216. [Google Scholar]
- Song, W.W.; Li, B.; Liu, J.K. A new lignan from Elaeagnus lanceolata (Elaeagnaceae). Acta. Bot. Yunnanica 2010, 32, 455–462. [Google Scholar]
- Bendaikha, S.; Gadaut, M.; Harakat, D.; Magid, A. Acylated flavonol glycosides from the flower of Elaeagnus angustifolia L. Phytochemistry 2014, 103, 129–136. [Google Scholar]
- Siegel, R.; Naishadham, D.; Jemal, A. Cancer statistics: 2013. CA: A Cancer J. Clin. 2013, 63, 11–30. [Google Scholar]
- Bray, F.; Jemal, A.; Grey, N.; Ferlay, J.; Forman, D. Global cancer transitions according to the Human Development Index (2008–2030): A population-based study. Lancet. Oncol. 2012, 13, 790–801. [Google Scholar]
- Tran, Q.N. A novel method for finding non-small cell lung cancer diagnosis biomarkers. BMC Med. Genomics 2013, 6, 11–20. [Google Scholar]
- Meoni, G.; Cecere, F.L.; Lucherini, E.; di Costanzo, F. Medical treatment of advanced non-small cell lung cancer in elderly patients: A review of the role of chemotherapy and targeted agents. J. Geriatr. Oncol. 2013, 4, 282–290. [Google Scholar]
- Seebacher, W.; Simic, N.; Weis, R.; Saf., R.; Kunert, O. Complete assignments of 1H and 13C NMR resonances of oleanolic acid, 18α-oleanolic acid, ursolic acid and their 11-oxo derivatives. Magn. Reson. Chem. 2003, 41, 636–638. [Google Scholar]
- Takahashi, H.; Iuchi, M.; Fujita, Y.; Minami, H.; Fukuyama, Y. Coumaroyl triterpenes from Casuarina equisetifolia. Phytochemistry 1999, 51, 543–550. [Google Scholar]
- Chen, B.; Duan, H.; Takaishi, Y. Triterpene caffeoyl esters and diterpenes from Celastrus stephanotifolius. Phytochemistry 1999, 51, 683–687. [Google Scholar]
- Murphy, B.T.; MacKinnon, S.L.; Yan, X.; Hammond, G.B.; Vaisberg, A.J.; Neto, C.C. Identification of triterpene hydroxycinnamates with in vitro antitumor activity from whole cranberry fruit (Vaccinium macrocarpon). J. Agric. Food Chem. 2003, 51, 3541–3545. [Google Scholar]
- Jeong, W.; Hong, S.S.; Kim, N.; Yang, Y.T.; Shin, Y.S.; Lee, C.; Hwang, B.Y.; Lee, D. Bioactive triterpenoids from Callistemon lanceolatus. Arch. Pharm. Res. 2009, 32, 845–849. [Google Scholar]
- Katai, M.; Terai, T.; Meguri, H. Triterpenoids of the bark of Pieris japonica D. DON (Japanese Name: Asebi). II. 13C nuclear magnetic resonance of the γ-lactones of ursane and oleanane-type triterpenes. Chem. Pharm. Bull. 1983, 31, 1567–1571. [Google Scholar]
- Huang, H.; Sun, H.; Zhao, S. Triterpenoids of Isodon loxothyrsus. Phytochemistry 1996, 42, 1665–1666. [Google Scholar]
- Collins, D.O.; Ruddock, P.L.D.; de Grasse, J.C.; Reynolds, W.F.; Reese, P.B. Microbial transformation of cadina-4,10(15)-dien-3-one, aromadendr-1(10)-en-9-one and methyl ursolate by Mucor plumbeus ATCC 4740. Phytochemistry 2002, 59, 479–488. [Google Scholar]
- Tijjani, A.; Ndukwe, I.G.; Ayo, R.G. Isolation and characterization of Lup-20(29)-ene-3, 28-diol (Betulin) from the stem-bark of Adenium obesum (Apocynaceae). Trop. J. Pharm. Res. 2012, 11, 259–262. [Google Scholar]
- Baek, M.Y.; Cho, J.G.; Lee, D.Y.; Ahn, E.M.; Jeong, T.S.; Baek, N.I. Isolation of triterpenoids from the stem bark of Albizia julibrissin and their inhibition activity on ACAT-1 and ACAT-2. J. Korean Soc. Appl. Biol. Chem. 2010, 53, 310–315. [Google Scholar]
- Liu, M.; Yang, S.; Jin, L.; Hu, D.; Wu, Z.; Yang, S. Chemical constituents of the ethyl acetate extract of Belamcanda chinensis (L.) DC roots and their antitumoractivities. Molecules 2012, 17, 6156–6169. [Google Scholar]
- Xu, S.; Shang, M.Y.; Liu, G.X.; Xu, F.; Wang, X.; Shou, C.C.; Cai, S.Q. Chemical constituents from the rhizomes of Smilax glabra and their antimicrobial activity. Molecules 2013, 18, 5265–5287. [Google Scholar]
- Wei, Y.; Xie, Q.; Dong, W.; Ito, Y. Separation of epigallocatechin and flavonoids from Hypericum perforatum L. by high-speed counter-current chromatography and preparative high-performance liquid chromatography. J. Chromatogr. A 2009, 1216, 4313–4318. [Google Scholar]
- Timmers, M.; Urban, S. On-line (HPLC-NMR) and off-line phytochemical profiling of the Australian plant, Lasiopetalum macrophyllum. Nat. Prod. Commun. 2012, 7, 551–560. [Google Scholar]
- Liao, C.R.; Ho, Y.L.; Huang, G.J.; Yang, C.S.; Chao, C.Y.; Chang, Y.S.; Kuo, Y.H. One lignanoid compound and four triterpenoid compounds with anti-inflammatory activity from the leaves of Elaeagnus oldhamii maxim. Molecules 2013, 18, 13218–13227. [Google Scholar]
- Świsłocka, R.; Kowczyk-Sadowy, M.; Kalinowska, M.; Lewandowski, W. Spectroscopic (FT-IR, FT-Raman, 1H and 13C NMR) and theoretical studies of p-coumaric acid and alkali metal p-coumarate. Spectroscopy 2012, 27, 35–48. [Google Scholar]
- Al-Musayeib, N.; Perveen, S.; Fatima, I.; Nasir, M.; Hussain, A. Antioxidant, anti-glycation and anti-inflammatory activities of phenolic constituents from Cordia sinensis. Molecules 2011, 16, 10214–10216. [Google Scholar]
- Jadrijević-Mladar Takac, M.; Vikić Topić, D. FT-IR and NMR spectroscopic studies of salicylic acid derivatives. II. Comparison of 2-hydroxy- and 2,4- and 2,5-dihydroxy derivatives. Acta Pharm. 2004, 54, 177–191. [Google Scholar]
- Yoshioka, T.; Inokuchi, T.; Fujioka, S.; Kimura, Y. Phenolic compounds and flavonoids as plant growth regulators from fruit and leaf of Vitex rotundifolia. Z. Naturforsch. C 2004, 59, 509–514. [Google Scholar]
- Ngan, L.T.; Moon, J.K.; Shibamoto, T.; Ahn, Y.J. Growth-inhibiting, bactericidal, and urease inhibitory effects of Paeonia lactiflora root constituents and related compounds on antibiotic-susceptible and resistant strains of Helicobacter pylori. J. Agric. Food Chem. 2012, 60, 9062–9073. [Google Scholar]
- Jeon, K.S.; Na, H.J.; Kim, Y.M.; Kwon, H.J. Antiangiogenic activity of 4-O-methylgallic acid from Canavalia gladiata, a dietary legume. Biochem. Biophys. Res. Commun. 2005, 330, 1268–1274. [Google Scholar]
- Liu, K.; Guo, L.; Miao, L.; Bao, W.; Yang, J.; Li, X.; Xi, T.; Zhao, W. Ursolic acid inhibits epithelial-mesenchymal transition by suppressing the expression of astrocyte-elevated gene-1 in human nonsmall cell lung cancer A549 cells. Anticancer Drugs 2013, 24, 494–503. [Google Scholar]
- Hao, J.; Liu, J.; Wen, X.; Sun, H. Synthesis and cytotoxicity evaluation of oleanolic acid derivatives. Bioorg. Med. Chem. Lett. 2013, 23, 2074–2077. [Google Scholar]
- Sun, H.; Fang, W.S.; Wang, W.Z.; Hu, C. Structure-activity relationships of Oleanane- and ursane- type triterpenoids. Bot. Stud. 2006, 47, 339–368. [Google Scholar]
- Min, B.S.; Kim, Y.H.; Lee, S.M.; Jung, H.J.; Lee, J.S.; Na, M.K.; Lee, C.O.; Lee, J.P.; Bae, K. Kaempferol-induced growth inhibition and apoptosis in A549 lung cancer cells is mediated by activation of MEK-MAPK. J. Cell Physiol. 2003, 197, 110–121. [Google Scholar]
- Maurya, D.K.; Nandakumar, N.; Devasagayam, T.P. Anticancer property of gallic acid in A549, a human lung adenocarcinoma cell line, and possible mechanisms. J. Clin. Biochem. Nutr. 2011, 48, 85–90. [Google Scholar]
- Sample Availability: Samples of the compounds are available from the authors.
© 2014 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Liao, C.-R.; Kuo, Y.-H.; Ho, Y.-L.; Wang, C.-Y.; Yang, C.-S.; Lin, C.-W.; Chang, Y.-S. Studies on Cytotoxic Constituents from the Leaves of Elaeagnus oldhamii Maxim. in Non-Small Cell Lung Cancer A549 Cells. Molecules 2014, 19, 9515-9534. https://doi.org/10.3390/molecules19079515
Liao C-R, Kuo Y-H, Ho Y-L, Wang C-Y, Yang C-S, Lin C-W, Chang Y-S. Studies on Cytotoxic Constituents from the Leaves of Elaeagnus oldhamii Maxim. in Non-Small Cell Lung Cancer A549 Cells. Molecules. 2014; 19(7):9515-9534. https://doi.org/10.3390/molecules19079515
Chicago/Turabian StyleLiao, Chi-Ren, Yueh-Hsiung Kuo, Yu-Ling Ho, Ching-Ying Wang, Chang -Syun Yang, Cheng-Wen Lin, and Yuan-Shiun Chang. 2014. "Studies on Cytotoxic Constituents from the Leaves of Elaeagnus oldhamii Maxim. in Non-Small Cell Lung Cancer A549 Cells" Molecules 19, no. 7: 9515-9534. https://doi.org/10.3390/molecules19079515
APA StyleLiao, C.-R., Kuo, Y.-H., Ho, Y.-L., Wang, C.-Y., Yang, C.-S., Lin, C.-W., & Chang, Y.-S. (2014). Studies on Cytotoxic Constituents from the Leaves of Elaeagnus oldhamii Maxim. in Non-Small Cell Lung Cancer A549 Cells. Molecules, 19(7), 9515-9534. https://doi.org/10.3390/molecules19079515





