A Benzoic Acid Derivative and Flavokawains from Piper species as Schistosomiasis Vector Controls
Abstract
:1. Introduction
2. Results and Discussion
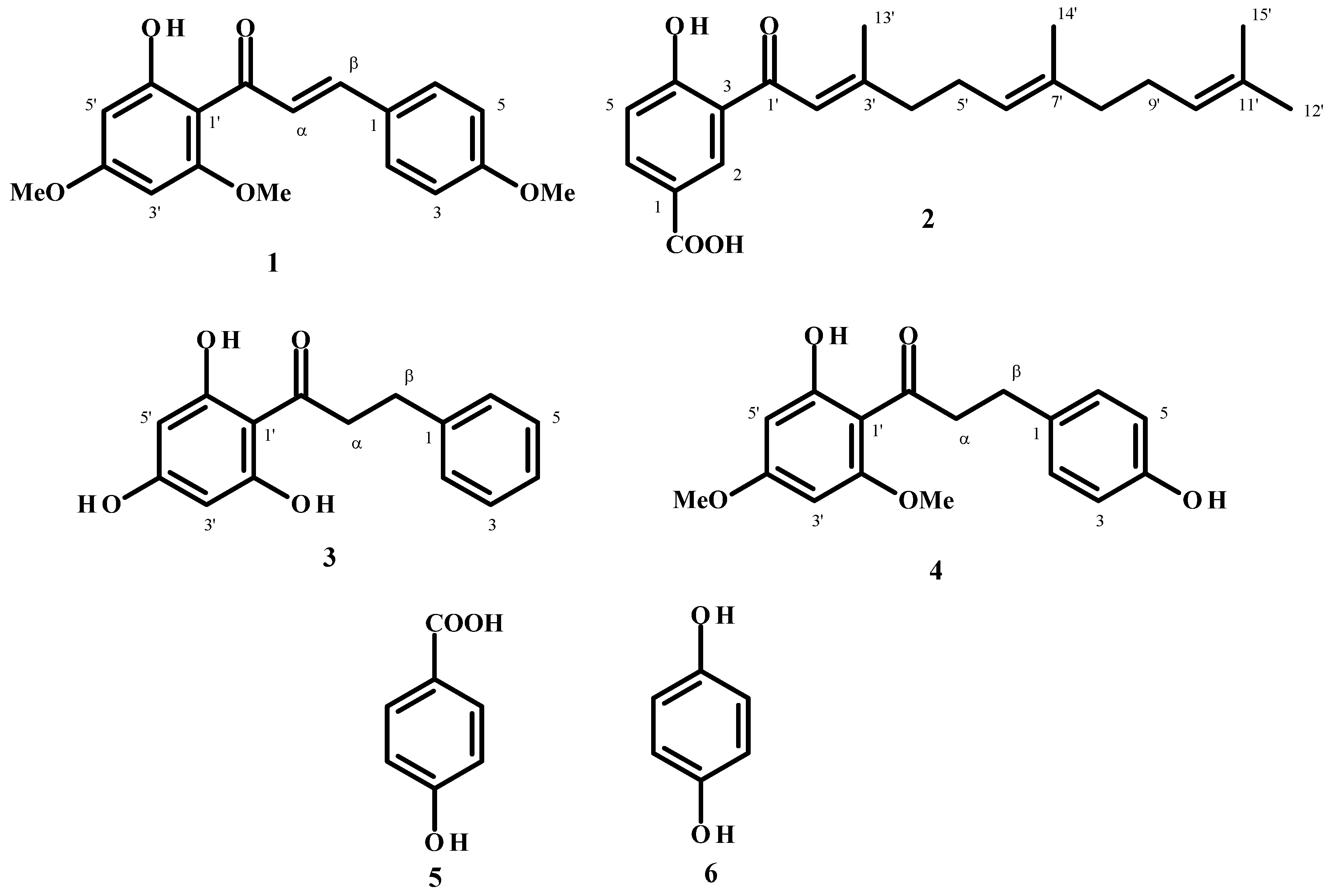
2.1. Compound Purification
| Compound 2 | ||
|---|---|---|
| Position | 1H (J Hz) | 13C |
| 1 | 120.31 | |
| 2 | 8.59 (1H, d, 2.0) | 133.16 |
| 3 | 119.70 | |
| 4 | 167.90 | |
| 5 | 7.03 (1H, d, 10.0) | 119.01 |
| 6 | 8.16 (1H, dd, 10.0, 2.0) | 137.17 |
| 1' | 195.63 | |
| 2' | 6.87 (1H, s) | 118.80 |
| 3' | 164.10 | |
| 4' | 2.37 (2H, m) | 42.11 |
| 5' | 2.04 (2H, m) | 26.81 |
| 6' | 5.17 (1H, m) | 127.70 |
| 7' | 136.82 | |
| 8' | 2.37 (2H, m) | 39.79 |
| 9' | 2.04 (2H, m) | 26.30 |
| 10' | 5.09 (1H, m) | 124.30 |
| 11' | 131.57 | |
| 12' | 1.65 (3H, s) | 25.91 |
| 13' | 2.26 (3H, s) | 20.57 |
| 14' | 1.65 (3H, s) | 16.28 |
| 15' | 1.57 (3H, s) | 17.80 |
| COOH | 13.49 (1H, s) | 170.90 |
2.2. Molluscicidal and Ovicidal Activities
| Compounds | Adults | Developmental stages | ||||
|---|---|---|---|---|---|---|
| Blastula | Gastula | Trocophore | Veliger | |||
| Flavokawain A (1) | LC50 | 21.85 [19.22–24.21] | nc | nc | nc | nc |
| LC90 | 27.97 [27.97–33.95] | nc | nc | nc | nc | |
| 4-Hydroxy-3-(3,7,11-trimethyldodeca-2,5,10-trienyl) benzoic acid (2) | LC50 | 7.28 [6.54–7.96] | nc | nc | nc | nc |
| LC90 | 10.04 [10.04–11.76] | nc | nc | nc | nc | |
| 2',4',6'-Trihydroxy-dihydrochalcone (3) | LC50 | 5.35 [4.28–6.30] | 10.18 [9.68–10.69] | 10.31 [9.86–10.79] | 10.71 [10.24–11.16] | 11.83 [11.35–12.31] |
| LC90 | 6.47 [6.47–9.06] | 14.12 [14.12–15.90] | 13.54 [13.54–15.28] | 14.44 [14.44–16.23] | 14.87 [14.87–16.19] | |
| Dihydroflavokawain C (4) | LC50 | nc | nc | nc | nc | nc |
| LC90 | nc | nc | nc | nc | nc | |
| p-Hydroxybenzoic acid (5) | LC50 | nc | nc | nc | nc | nc |
| LC90 | nc | nc | nc | nc | nc | |
| Hydroquinone (6) | LC50 | 3.12 [2.70–3.59] | 1.17 [1.14–1.20] | 1.98 [1.92–2.04] | 2.74 [2.60–2.89] | 4.26 [4.12–4.41] |
| LC90 | 5.27 [5.27–6.97] | 1.51 [1.51–1.59] | 2.86 [2.86–3.05] | 5.83 [5.83–6.45] | 6.17 [6.17–6.54] | |


3. Experimental Section
3.1. General
3.2. Plant Material
3.3. Extraction and Isolation
3.4. Spectral Data
3.5. Biological Assays
3.6. Molluscicidal Activity
3.7. Ovicidal Activity
3.8. Docking
3.9. Molecular Interaction Fields
4. Conclusions
Supplementary Materials
Acknowledgments
Author Contributions
Conflicts of Interest
References
- Prevention and Control of Schistosomiasis and Soil-transmitted Helminthiasis: Report of a WHO Expert Committee; Technical report series, 912; World Health Organization: Geneva, Switzerland, 2002.
- Chitsulo, L.; Engels, D.; Montresor, A.; Savioli, L. The global status of schistosomiasis and its control. Acta Trop. 2000, 77, 41–51. [Google Scholar] [CrossRef]
- World Health Organization. Weekly epidemiological record. WHO 2011, 9, 73–80. [Google Scholar]
- Lardans, V.; Dissous, C. Snail control strategies for reduction of schistosomiasis transmission. Parasitol. Today 1998, 14, 413–417. [Google Scholar] [CrossRef]
- Second Report of the WHO Expert Committee; 830; World Health Organization: Geneva, Switzerland, 1993; pp. 1–86.
- Pointier, J.P.; Giboda, M. The case for biological control of snail intermediate hosts of Schistosoma mansoni. Parasitol. Today 1999, 15, 395–397. [Google Scholar] [CrossRef]
- Hostettmann, K.; Potterat, O. Strategy for the isolation and analysis of antifungal, molluscicidal, and larvicidal agents from tropical plants. In Phytochemicals for Pest Control; ACS Symposium Series 658; American Chemical Society: Washington, DC, USA, 1997; pp. 14–26. [Google Scholar]
- Rapado, L.N.; Nakano, E.; Ohlweiler, F.P.; Kato, M.J.; Yamaguchi, L.F.; Pereira, C.A.B.; Kawano, T. Molluscicidal and ovicidal activities of plant extracts of the Piperaceae on Biomphalaria glabrata (Say, 1818). J. Helminthol. 2010, 85, 66–72. [Google Scholar]
- De Groot, A.; de Rosny, E.; Juillan-Binard, C.; Ferrer, J.L.; Laudet, V.; Pierce, R.J.; Pebay-Peyroula, E.; Fontecilla-Camps, J.C.; Borel, F. Crystal structure of a novel tetrameric complex of agonist-bound ligand-binding domain of Biomphalaria glabrata retinoid X receptor. J. Mol. Biol. 2005, 354, 841–853. [Google Scholar] [CrossRef]
- Robinson-Rechavi, M.; Garcia, H.E.; Laudet, V. The nuclear receptor superfamily. J. Cell Sci. 2003, 116, 585–586. [Google Scholar]
- Fang, H.; Tong, W.D.; Welsh, W.J.; Sheehan, D.M. QSAR models in receptor-mediated effects: The nuclear receptor superfamily. J. Mol. Struc-Theochem. 2003, 622, 113–125. [Google Scholar] [CrossRef]
- Cruciani, G.; Crivori, P.; Carrupt, P.A.; Testa, B. Molecular fields in quantitative structure-permeation relationships: The VolSurf approach. J. Mol. Struc-Theochem. 2000, 503, 17–30. [Google Scholar] [CrossRef]
- Dharmaratne, H.R.W.; Nanayakkara, N.P.; Khan, I.A. Kavalactones from Piper methysticum, and their 13C NMR spectroscopic analyses. Phytochemistry 2002, 59, 429–433. [Google Scholar] [CrossRef]
- Seeram, N.P.; Jacobs, H.; McLean, S.; Reynolds, W.F. Prenylated hydroxybenzoic acid derivatives from Piper murrayanum. Phytochemistry 1996, 43, 863–865. [Google Scholar] [CrossRef]
- Mustafa, K.; Kjaergaard, H.G.; Perry, N.B.; Weavers, R.T. Hydrogen-bonded rotamers of 2',4',6'-trihydroxy-3-formyldihydrochalcone, an intermediate in the synthesis of a dihydrochalcone from Leptospermum recurvum. Tetrahedron 2003, 59, 6113–6120. [Google Scholar] [CrossRef]
- Dutta, C.P.; Roy, L.P.K.; Chatterjee, A. Studies on the genus Piper. 5. Chemical investigation of Piper methysticum Forst (Piperaceae). Structure and synthesis of flavokawain C. J. Indian Chem. Soc. 1976, 53, 1194–1197. [Google Scholar]
- World Health Organization. Molluscicide screening and evaluation. Bull. World Health Organ. 1965, 33, 567–581. [Google Scholar]
- Second Report of the WHO Expert Committee; 745; World Health Organization: Geneva, Switzerland, 1983; p. 11.
- Rapado, L.N.; Pinheiro, A.S.; Lopes, P.O.M.V.; Fokoue, H.H.; Scotti, M.T.; Marques, J.V.; Ohlweiler, F.P.; Borrely, S.I.; Pereira, C.A.B.; Kato, M.J.; et al. Schistosomiasis control using piplartine against Biomphalaria glabrata at different developmental stages. PLoS Negl. Trop. Dis. 2013, 7, e2251. [Google Scholar] [CrossRef]
- Camey, T.; Verdonk, N.H. The early development of the snail Biomphalaria glabrata (Say) and the origin of the head organs. Neth. J. Zool. 1970, 20, 93–121. [Google Scholar] [CrossRef]
- Protein Data Bank. Available online: http://www.rcsb.org/pdb/home/home.do/ (accessed on 25 January 2014).
- Marvin Sketch, v. 6.1.4.; Chemaxon: Budapest, Hungary, 2014.
- Standardizer, v. 6.1.4.; Chemaxon: Budapest, Hungary, 2014.
- Thomsen, R.; Christensen, M.H. MolDock: A new technique for high-accuracy molecular docking. J. Med. Chem. 2006, 49, 3315–3321. [Google Scholar] [CrossRef]
- Docker, v. 6.0.1.; CLC Bio: Cambridge, MA, USA, 2014.
- Wold, P.; Esbensen, K.; Geladi, P. Principal Component Analysis. Chemometr. Intell. Lab. 1987, 2, 37–52. [Google Scholar] [CrossRef]
- VolSurf+, v. 1.0.7.; Molecular Discovery: Middlesex, UK, 2014.
- Sample Availability: Not available.
© 2014 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Rapado, L.N.; Freitas, G.C.; Polpo, A.; Rojas-Cardozo, M.; Rincón, J.V.; Scotti, M.T.; Kato, M.J.; Nakano, E.; Yamaguchi, L.F. A Benzoic Acid Derivative and Flavokawains from Piper species as Schistosomiasis Vector Controls. Molecules 2014, 19, 5205-5218. https://doi.org/10.3390/molecules19045205
Rapado LN, Freitas GC, Polpo A, Rojas-Cardozo M, Rincón JV, Scotti MT, Kato MJ, Nakano E, Yamaguchi LF. A Benzoic Acid Derivative and Flavokawains from Piper species as Schistosomiasis Vector Controls. Molecules. 2014; 19(4):5205-5218. https://doi.org/10.3390/molecules19045205
Chicago/Turabian StyleRapado, Ludmila N., Giovana C. Freitas, Adriano Polpo, Maritza Rojas-Cardozo, Javier V. Rincón, Marcus T. Scotti, Massuo J. Kato, Eliana Nakano, and Lydia F. Yamaguchi. 2014. "A Benzoic Acid Derivative and Flavokawains from Piper species as Schistosomiasis Vector Controls" Molecules 19, no. 4: 5205-5218. https://doi.org/10.3390/molecules19045205
APA StyleRapado, L. N., Freitas, G. C., Polpo, A., Rojas-Cardozo, M., Rincón, J. V., Scotti, M. T., Kato, M. J., Nakano, E., & Yamaguchi, L. F. (2014). A Benzoic Acid Derivative and Flavokawains from Piper species as Schistosomiasis Vector Controls. Molecules, 19(4), 5205-5218. https://doi.org/10.3390/molecules19045205







