Synthesis, DNA Binding and Topoisomerase I Inhibition Activity of Thiazacridine and Imidazacridine Derivatives
Abstract
:1. Introduction
2. Results and Discussion
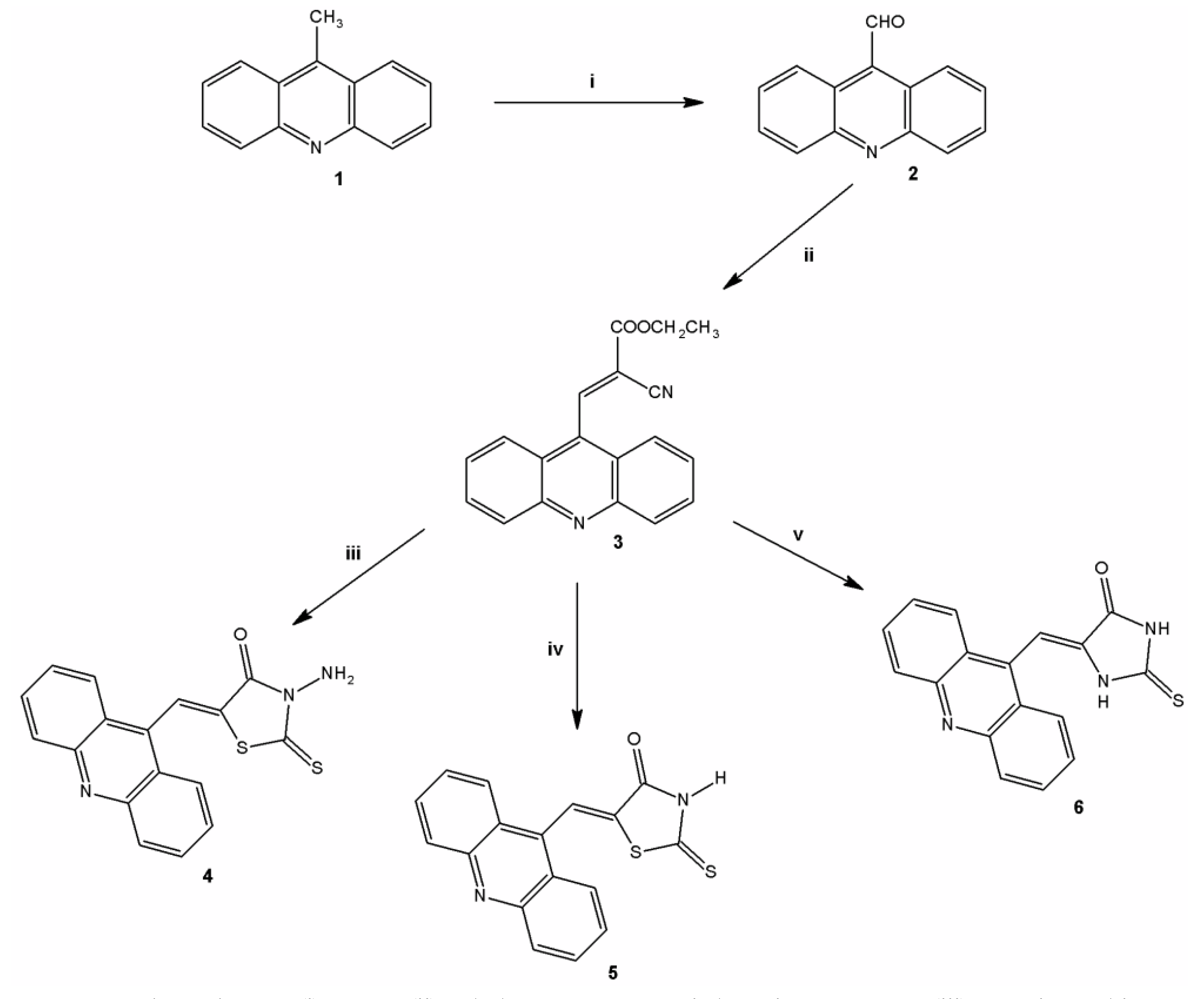
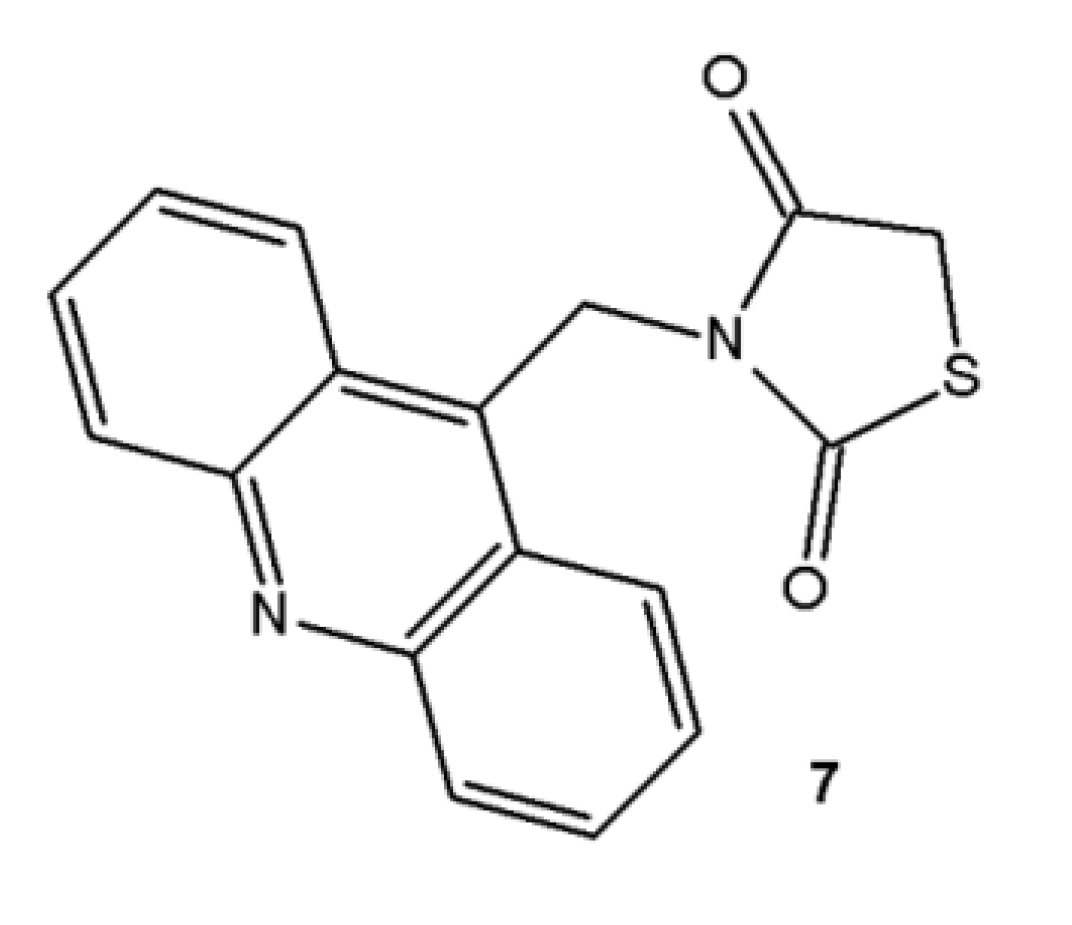
2.1. Chemistry
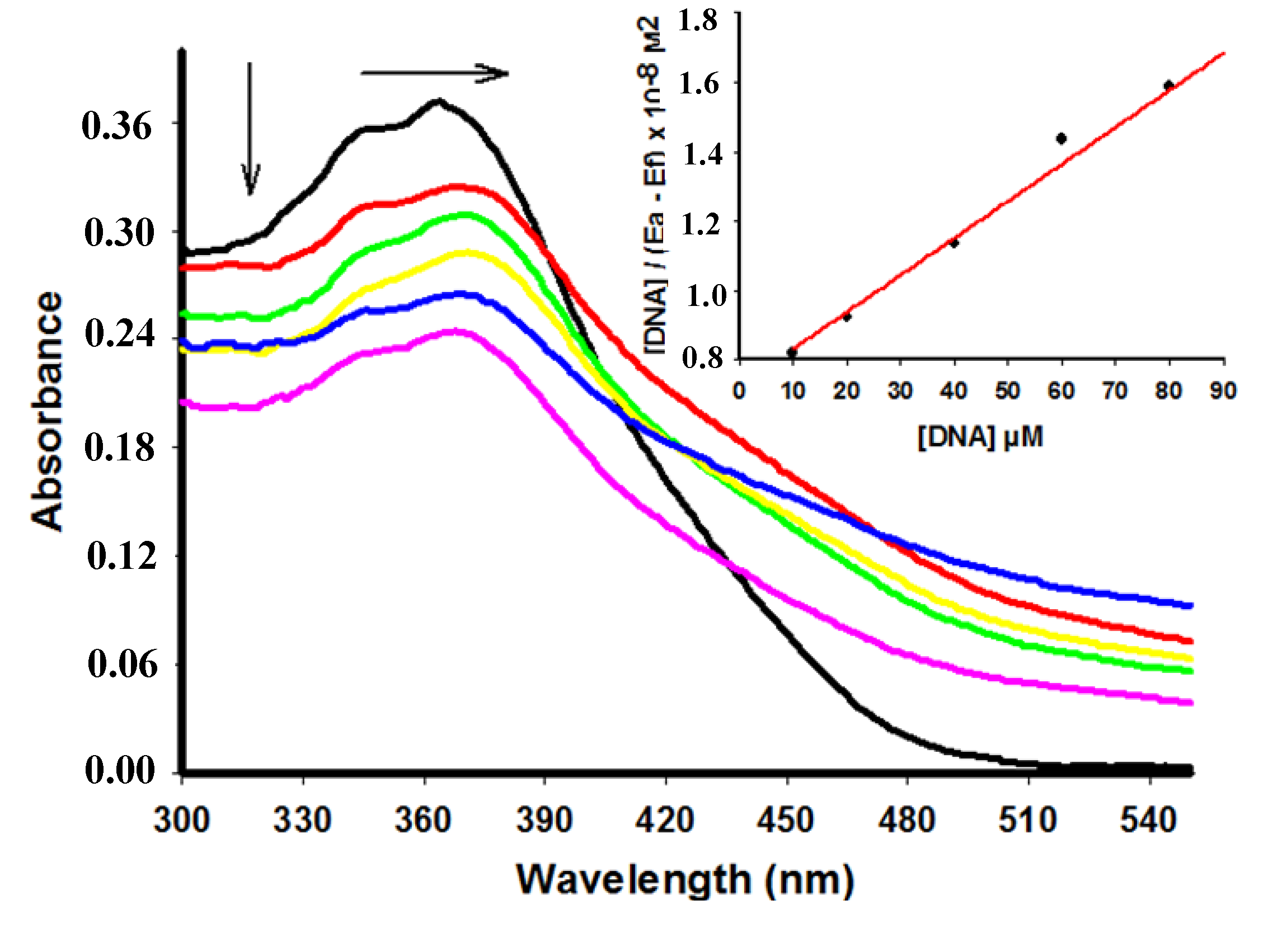
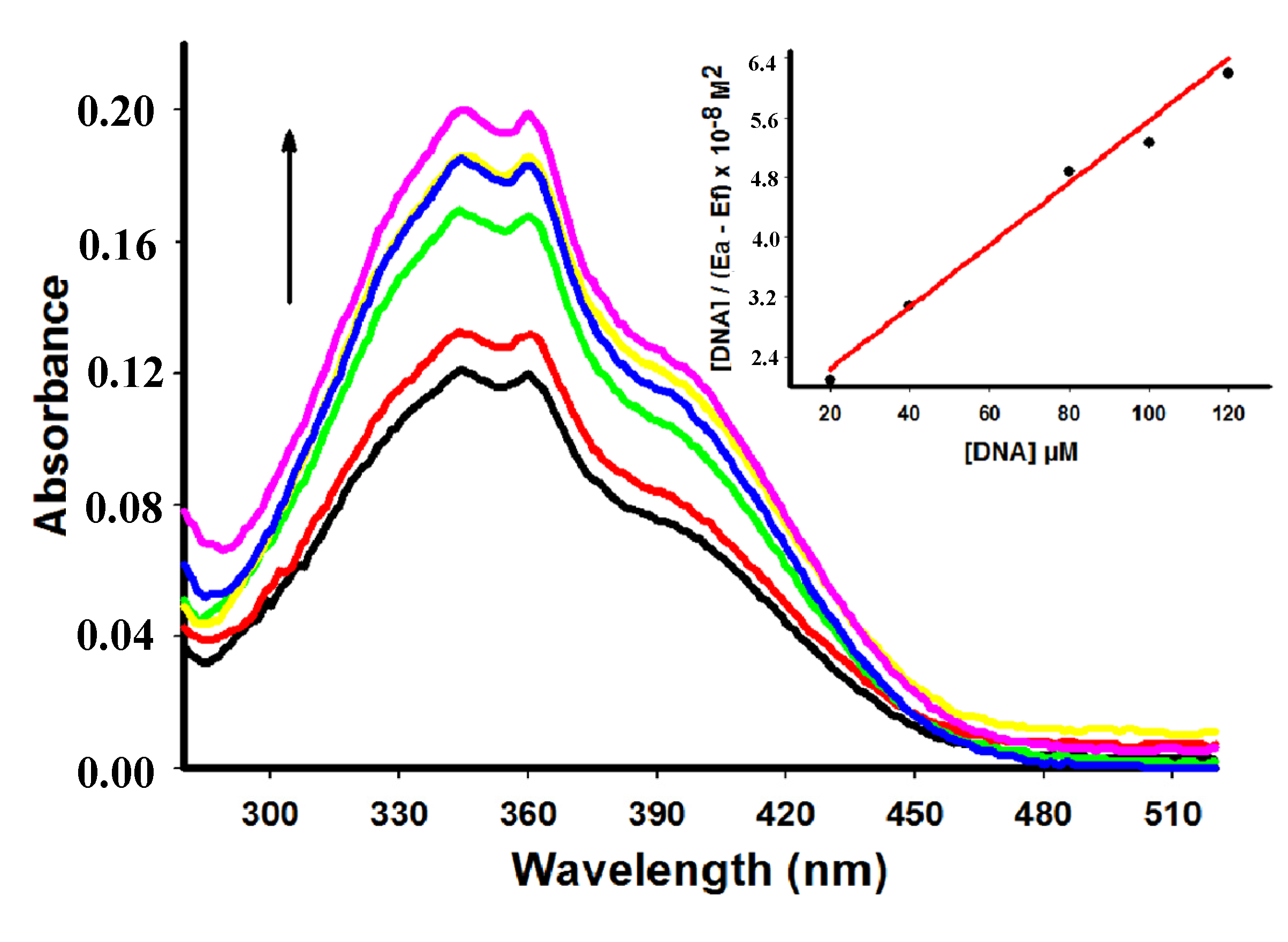
2.2. UV-Vis Spectral Absorbance
| Compound | λmax free (nm) | λmax bound (nm) | Extinction coefficient (ε) M−1 | Hypochromicity (%) | Kb M−1 | Log p |
|---|---|---|---|---|---|---|
| 4 | 346 | 346 | 10.340 | 40.43 | 1.46 × 104 | 3.37 |
| 5 | 345 | 345 | 2.420 | 0 | 2.37 × 104 | 2.93 |
| 6 | 364 | 368 | 14.840 | 28.85 | 3.25 × 104 | 3.44 |
| 7 | 361 | 361 | 8.000 | 15.50 | 6.01 × 104 | 2.24 |
2.3. Fluorescence Emission Spectra
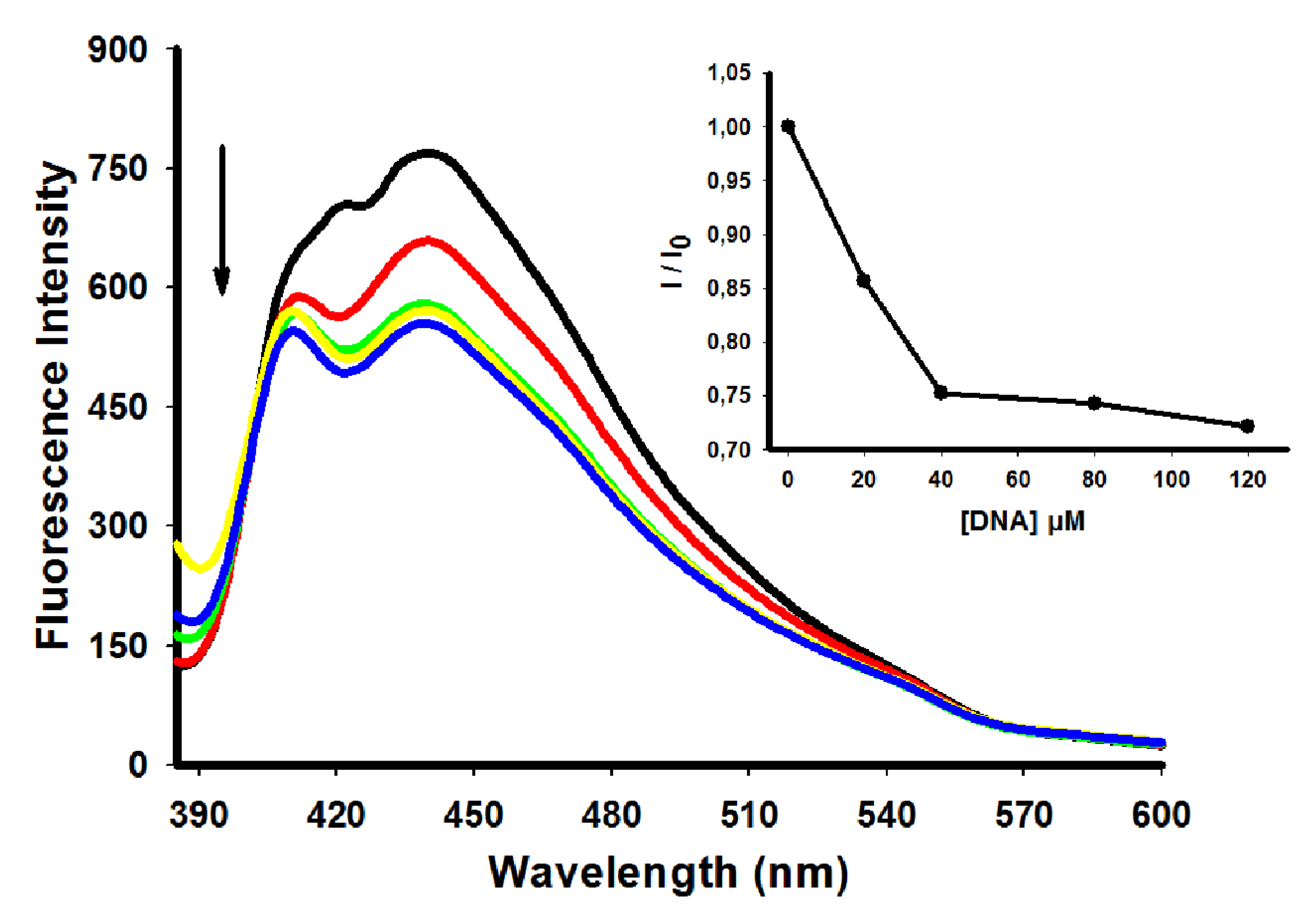
| Compoud | λ excitation (nm) | λ emission (nm) | I/I0 |
|---|---|---|---|
| 4 | 360 | 415 | 1.67 |
| 5 | 356 | 440 | 2.27 |
| 6 | 364 | 418 | 1.11 |
| 7 | 360 | 435 | 1.0 |
2.4. CD Spectroscopic Analysis
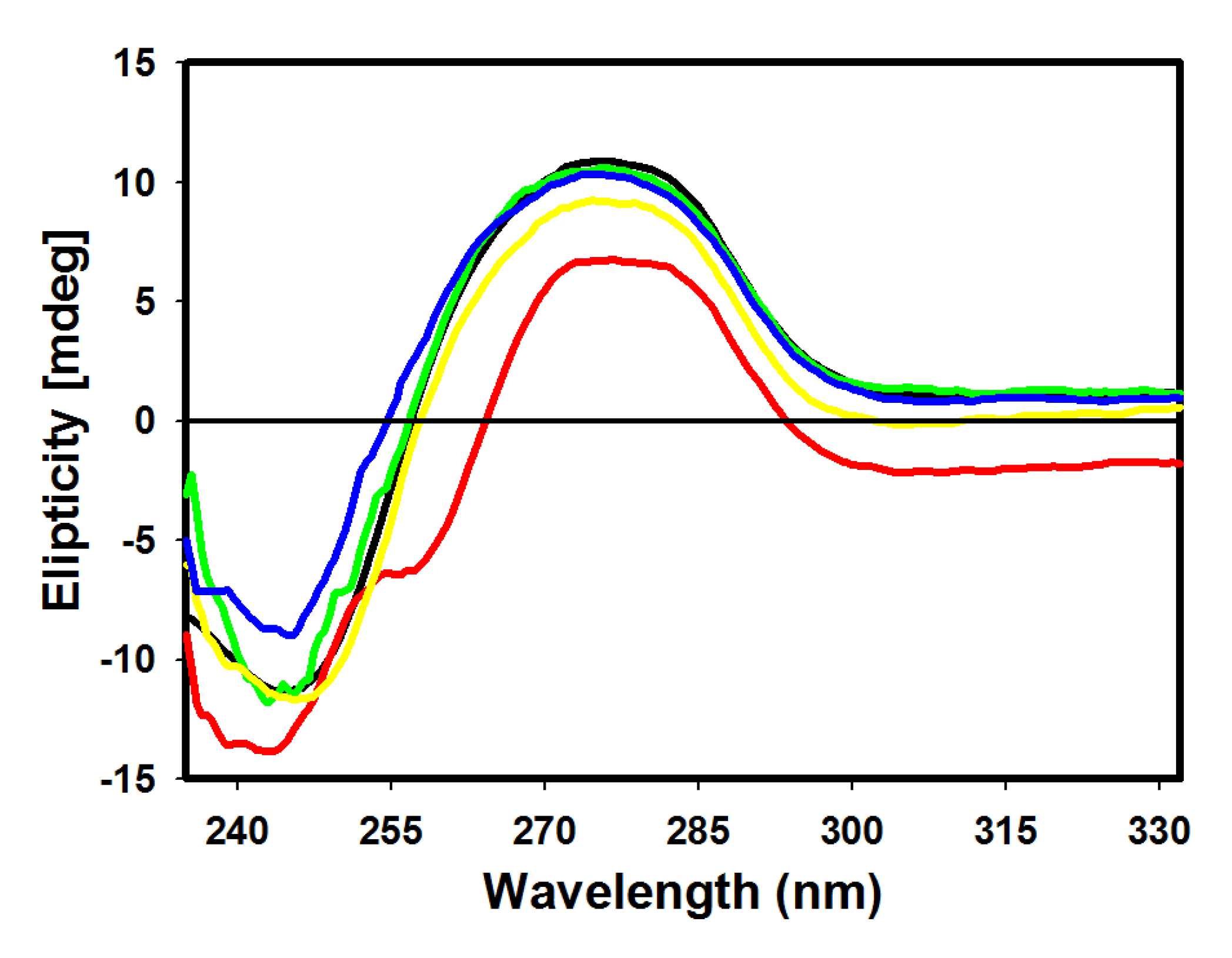
2.5. DNA Topoisomerase I Inhibition Assay
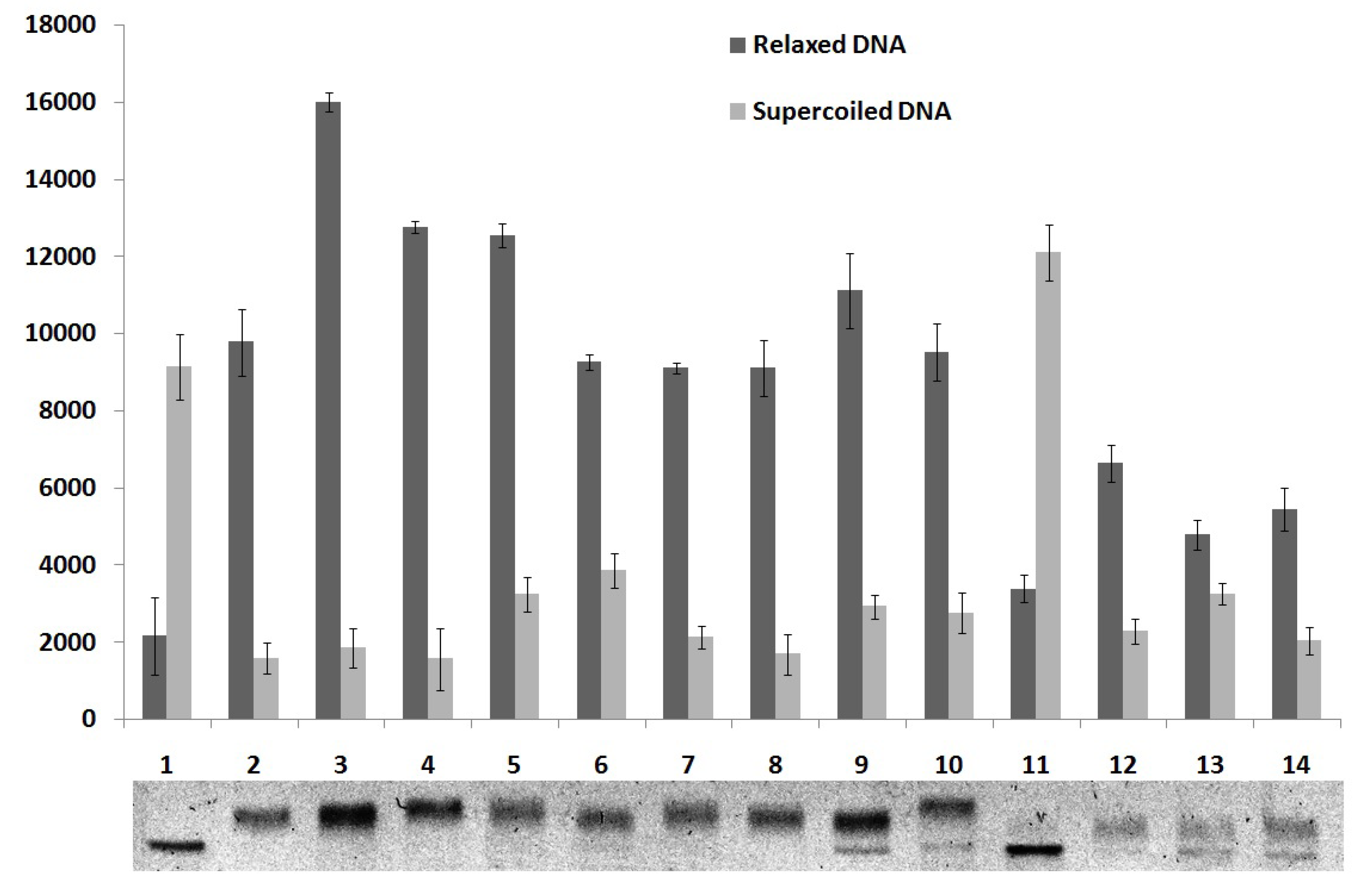
3. Experimental
3.1. Materials and Instrumentation
3.2. Preparation of the Acridine-Thiazolidines Derivatives 4–6
3.4. Fluorescence Measurements
3.5. Circular Dichroism
3.6. DNA Topoisomerase I Inhibition Assay
3.7. Determination of log p Values
4. Conclusions
Supplementary Materials
Acknowledgments
Conflicts of Interest
References
- Neto, B.A.D.; Lapis, A.A.M. Recent developments in the chemistry of desoxyribonucleic acid (DNA) intercalators: Principles, design, synthesis, applications and trends. Molecules 2009, 14, 1725–1746. [Google Scholar]
- Mukherjee, A.; Lavery, R.; Bagchi, B.; Hynes, J.T. On the molecular mechanism of drug intercalation into DNA: A simulation study of the intercalation pathway, free energy, and DNA structural changes. J. Am. Chem. Soc. 2008, 130, 9747–9755. [Google Scholar]
- Hurley, H.L. DNA and its associated processes as targets for cancer therapy. Nat. Rev. Cancer 2002, 2, 188–200. [Google Scholar]
- Ihmels, H.; Otto, D. Intercalation of organic dye molecules into double-stranded DNA general principles and recent developments. Top. Curr. Chem. 2005, 258, 161–204. [Google Scholar]
- Barros, F.W.A.; Silva, T.G.; Pitta, M.G.R.; Bezerra, D.P.; Costa-Lotufo, L.V.; Moraes, M.O.; Pessoa, C.; Moura, M.A.F.B.; Abreu, F.C.; Lima, M.C.A.; et al. Synthesis and cytotoxic activity of new acridine-thiazolidine derivatives. Bioorg. Med. Chem. 2012, 20, 3533–3539. [Google Scholar]
- Ghosh, R.; Bhowik, S.; Bagchi, A.; Das, D.; Ghosh, S. Chemotherapeutic potential of 9-phenyl acridine: Biophysical studies on its binding to DNA. Eur. Biophys. J. 2010, 39, 1243–1249. [Google Scholar]
- Gao, C.; Liu, F.; Luan, X.; Tan, C.; Liu, H.; Xie, Y.; Jin, Y.; Jiang, Y. Novel synthetic 2-amino-10-(3,5-dimethoxy)benzyl-9(10H)-acridinone derivatives as potent DNA-binding antiproliferative agents. Bioorg. Med. Chem. 2010, 18, 7507–7514. [Google Scholar]
- Janovec, L.; Kozurkova, M.; Sabolova, D.; Ungvarsky, J.; Paulikova, H.; Plsikova, J.; Vantosa, Z.; Imrich, J. Cytotoxic 3,6-bis((imidazolidionone)imino)acridines: Synthesis, DNA binding and molecular modeling. Bioorg. Med. Chem. 2011, 19, 1790–1801. [Google Scholar]
- Liu, Q.; Zhang, J.; Wang, M.; Zhang, D.; Lu, Q.; Huang, Y.; Lin, H.; Yu, X. Synthesis, DNA binding and cleavage activity of macrocyclic polyamines bearing mono- or bis-acridine moieties. Eur. J. Med. Chem. 2010, 45, 5302–5308. [Google Scholar]
- Plsikova, J.; Janovec, L.; Koval, J.; Ungvarsky, J.; Mikes, J.; Jendzelovsky, R.; Fedorocko, P.; Imrich, J.; Kristian, P.; Kasparkova, J.; et al. 3,6-Bis(3-alkyl-guanidino)acridines as DNA-intercalating antitumor agents. Eur. J. Med. Chem. 2012, 57, 283–295. [Google Scholar]
- Pommier, Y. Drugging Topoisomerases: Lessons and Challenges. ACS Chem. Biol. 2013, 8, 82–95. [Google Scholar] [CrossRef]
- Pommier, Y. DNA topoisomerase I inhibitors: Chemistry, biology and interfacial inhibition. Chem. Rev. 2009, 109, 2894–2902. [Google Scholar] [CrossRef]
- Wang, Y.-C.; Qiana, C.; Penga, Z.-L.; Hou, X.-J.; Wang, L.-L.; Chaoa, H.; Ji, L.-N. Dual topoisomerase I and II poisoning by chiral Ru(II) complexes containing 2-thiophenylimidazo[4,5-f][1,10]phenanthroline derivatives. J. Inorg. Biochem. 2014, 130, 15–27. [Google Scholar] [CrossRef]
- Jangir, D.K.; Dey, S.K.; Kundu, S.; Mehrotra, R. Assessment of amsacrine binding with DNA using UV-visible, circular dichroism and Raman spectroscopic techniques. J. Photochem. Photobiol. B 2012, 114, 38–43. [Google Scholar]
- Sanchez, I.; Reches, R.; Caignard, D.H.; Renard, P.; Pujol, M.D. Synthesis and biological evaluation of modified acridines: The effect of N- and O-substituent in the nitrogenated ring on antitumor activity. Eur. J. Med. Chem. 2006, 41, 340–352. [Google Scholar]
- Pitta, M.G.R.; Souza, E.S.; Barros, F.W.A.; Moraes, M.O.; Pessoa, C.O.; Hernandes, M.Z.; Lima, M.C.A.; Galdino, S.L.; Pitta, I.R. Synthesis and in vitro anticancer activity of novel thiazacridine derivatives. Med. Chem. Res 2012, 21, 3326–3334. [Google Scholar]
- Pigatto, M.C.; Lima, M.C.A.; Galdino, S.L.; Pitta, I.R.; Vessecchi, R.; Assis, M.D.; Santos, J.S.; Costa, T.D.; Lopes, N.P. Metabolism evaluation of the anticancer candidate AC04 by biomimetic oxidative model and rat liver microsomes. Eur. J. Med. Chem. 2011, 46, 4245–4251. [Google Scholar]
- Pigatto, M.C.; Uchoa, F.T.; Torres, B.; Haas, S.; Lima, M.C.A.; Galdino, S.L.; Lopes, N.P.; Costa, T.D. Pre-clinical pharmacokinetics of the acridine antitumour candidate AC04 and its 1-oxo-metabolite plasma profile. Xenobiotica 2012, 42, 1–7. [Google Scholar]
- Barros, F.W.A.; Bezerra, D.P.; Ferreira, P.M.P.; Cavalcanti, B.C.; Silva, T.G.; Pitta, M.G.R.; Lima, M.C.A.; Galdino, S.L.; Pitta, I.R.; Costa-Lotufo, L.V.; et al. Inhibition of DNA topoisomerase I activity and induction of apoptosis by thiazacridine derivatives. Toxicol. Appl. Pharmacol. 2013, 268, 37–46. [Google Scholar]
- Tsuge, O.; Nishinohara, M.; Tashiro, M.B. Compounds related to acridine. I. Condensation of acridine derivatives having active methyl group and aromatic nitroso compounds. Bull. Chem. Soc. 1963, 36, 1477–1485. [Google Scholar]
- Mosher, M.D.; Natale, N.R. The preparation of intercalating isoxazoles via a nitrile oxide cycloaddition. J. Heterocycl. Chem. 1995, 32, 779–781. [Google Scholar]
- Sabolova, D.; Kozurkova, M.; Kristian, P.; Danihel, I.; Podhradsk, D.; Imrich, J. Determination of the binding affinities of plasmid DNA using fluorescent intercalators possessing an acridine skeleton. Int. J. Biol. Macromol. 2006, 38, 94–98. [Google Scholar]
- David-Cordonnier, M.H.; Hildebrand, M.P.; Baldeyrou, B.; Lansiaux, A.; Keuser, C.; Benzchawel, K.; Lemster, T.; Pindur, U. Design, synthesis and biological evaluation of new oligopyrrole carboxamides linked with tricyclic DNA-intercalators as potential DNA ligands or topoisomerase inhibitors. Eur. J. Med. Chem. 2007, 42, 752–771. [Google Scholar]
- Kuruvilla, E.; Nandajan, P.; Schuster, G.B.; Ramaiah, D. Acridine-viologen dyads: Selective recognition of single-strand DNA through fluorescence enhancement. Org. Lett. 2008, 10, 4295–4298. [Google Scholar]
- Shahabadi, N.; Heidari, L. Binding studies of the antidiabetic drug, metformin to calf thymus DNA using multispectroscopic methods. Spectrochim. Acta A 2012, 97, 406–410. [Google Scholar]
- Busto, N.; Garcia, B.; Leal, J.M.; Gaspar, J.F.; Martins, C.; Boggioni, A.; Secco, F. ACMA (9-amino-6-chloro-2-methoxyacridine) forms three complexes in the presence of DNA. Phys. Chem. Chem. Phys. 2011, 13, 19534–19545. [Google Scholar]
- Zhou, C.Y.; Xi, X.L.; Yang, P. Studies on DNA binding to metal complexes of sal2trien. Biochemistry 2007, 72, 37–43. [Google Scholar]
- Kashanian, S.; Dolatabadi, E.N. DNA binding studies of 2-tert-butylhydroquinone (TBHQ) food additive. Food Chem. 2009, 116, 743–747. [Google Scholar]
- McGhee, J.D.; von Hippel, P.H. Theoretical aspects of DNA-Protein interactions: Co-operative and non-co-operative binding of larg ligands to a one-dimensional homogeneous lattice. J. Mol. Biol. 1974, 86, 469–489. [Google Scholar]
- Strekowski, L.; Wilson, B. Noncovalent interactions with DNA: An overview. Mutat. Res. 2007, 623, 3–13. [Google Scholar]
- Mahmood, I.; Paul, A.; Ladame, S. Synthesis and spectroscopic and DNA-binding properties of fluorogenic acridine-containing cyanine dyes. J. Org. Chem. 2010, 75, 204–207. [Google Scholar]
- Kypr, J.; Kejnovska, I.; Renciuk, D.; Vorlickova, M. Circular dichroism and conformational polymorphism of DNA. Nucleic Acids Res. 2009, 37, 1713–1725. [Google Scholar]
- Shahabadi, N.; Moghadam, N.H. Determining the mode of interaction of calf thymus DNA with the drug sumatriptan using voltammetric and spectroscopic techniques. Spectrochim. Acta A 2012, 99, 18–22. [Google Scholar]
- Shahabadi, N.; Kashanian, S.; Darabi, F. DNA binding and DNA cleavage studies of a water soluble cobalt(II) complex containing dinitrogen Schiff base ligands: The effect of metal on the mode of binding. Eur. J. Med. Chem. 2010, 45, 4239–4295. [Google Scholar]
- Bhowmik, S.; Bagchi, A.; Ghosh, R. Molecular modeling studies of some 9-arylacridines to elucidate their possible roles in topoisomerase I inhibition. Int. J. Integr. Biol. 2008, 2, 8–14. [Google Scholar]
- Drwal, M.N.; Agama, K.; Wakelin, L.P.G.; Pommier, Y.; Griffith, R. Exploring DNA topoisomerase I ligand space in search of novel anticancer agents. PloS One 2011, 6, e25150. [Google Scholar]
- Wolfe, A.; Shimer, G.H., Jr.; Meehan, T. Polycyclic aromatic hydrocarbons physically intercalate into duplex regions of denatured DNA. Biochemistry 1987, 26, 6392–6396. [Google Scholar]
- ACDLabs/ChemSketch. Available online: http://www.acdlabs.com/ (accessed on 2 December 2013).
- Sample Availability: Samples of the compounds 5-acridin-9-ylmethylidene-3-amino-2-thioxo-thiazolidin-4-one; 5-acridin-9-ylmethylidene-2-thioxo-thiazolidin-4-one; 5-acridin-9-ylmethylidene-2-thioxo-imidazolidin-4-one and 3-acridin-9-ylmethyl-thiazolidine-2,4-dione are available from the authors.
© 2013 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license (http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Lafayette, E.A.; Vitalino de Almeida, S.M.; Da Rocha Pitta, M.G.; Carneiro Beltrão, E.I.; Gonçalves da Silva, T.; Olímpio de Moura, R.; Da Rocha Pitta, I.; De Carvalho, L.B., Júnior; Do Carmo Alves de Lima, M. Synthesis, DNA Binding and Topoisomerase I Inhibition Activity of Thiazacridine and Imidazacridine Derivatives. Molecules 2013, 18, 15035-15050. https://doi.org/10.3390/molecules181215035
Lafayette EA, Vitalino de Almeida SM, Da Rocha Pitta MG, Carneiro Beltrão EI, Gonçalves da Silva T, Olímpio de Moura R, Da Rocha Pitta I, De Carvalho LB Júnior, Do Carmo Alves de Lima M. Synthesis, DNA Binding and Topoisomerase I Inhibition Activity of Thiazacridine and Imidazacridine Derivatives. Molecules. 2013; 18(12):15035-15050. https://doi.org/10.3390/molecules181215035
Chicago/Turabian StyleLafayette, Elizabeth Almeida, Sinara Mônica Vitalino de Almeida, Marina Galdino Da Rocha Pitta, Eduardo Isidoro Carneiro Beltrão, Teresinha Gonçalves da Silva, Ricardo Olímpio de Moura, Ivan Da Rocha Pitta, Luiz Bezerra De Carvalho, Júnior, and Maria Do Carmo Alves de Lima. 2013. "Synthesis, DNA Binding and Topoisomerase I Inhibition Activity of Thiazacridine and Imidazacridine Derivatives" Molecules 18, no. 12: 15035-15050. https://doi.org/10.3390/molecules181215035
APA StyleLafayette, E. A., Vitalino de Almeida, S. M., Da Rocha Pitta, M. G., Carneiro Beltrão, E. I., Gonçalves da Silva, T., Olímpio de Moura, R., Da Rocha Pitta, I., De Carvalho, L. B., Júnior, & Do Carmo Alves de Lima, M. (2013). Synthesis, DNA Binding and Topoisomerase I Inhibition Activity of Thiazacridine and Imidazacridine Derivatives. Molecules, 18(12), 15035-15050. https://doi.org/10.3390/molecules181215035