On the Reducible Character of Haldane-Radić Enzyme Kinetics to Conventional and Logistic Michaelis-Menten Models
Abstract
:1. Introduction
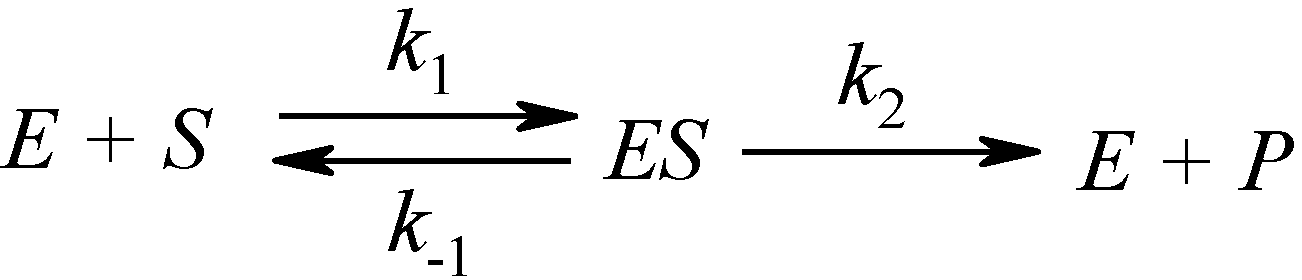
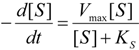
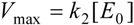

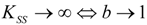
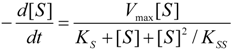
2. Background Theories
2.1. Haldane-Radić Equation
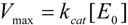
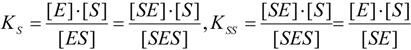
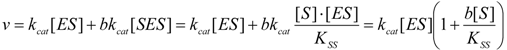
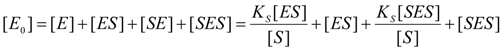
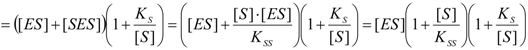
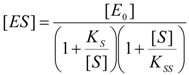
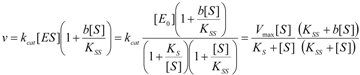
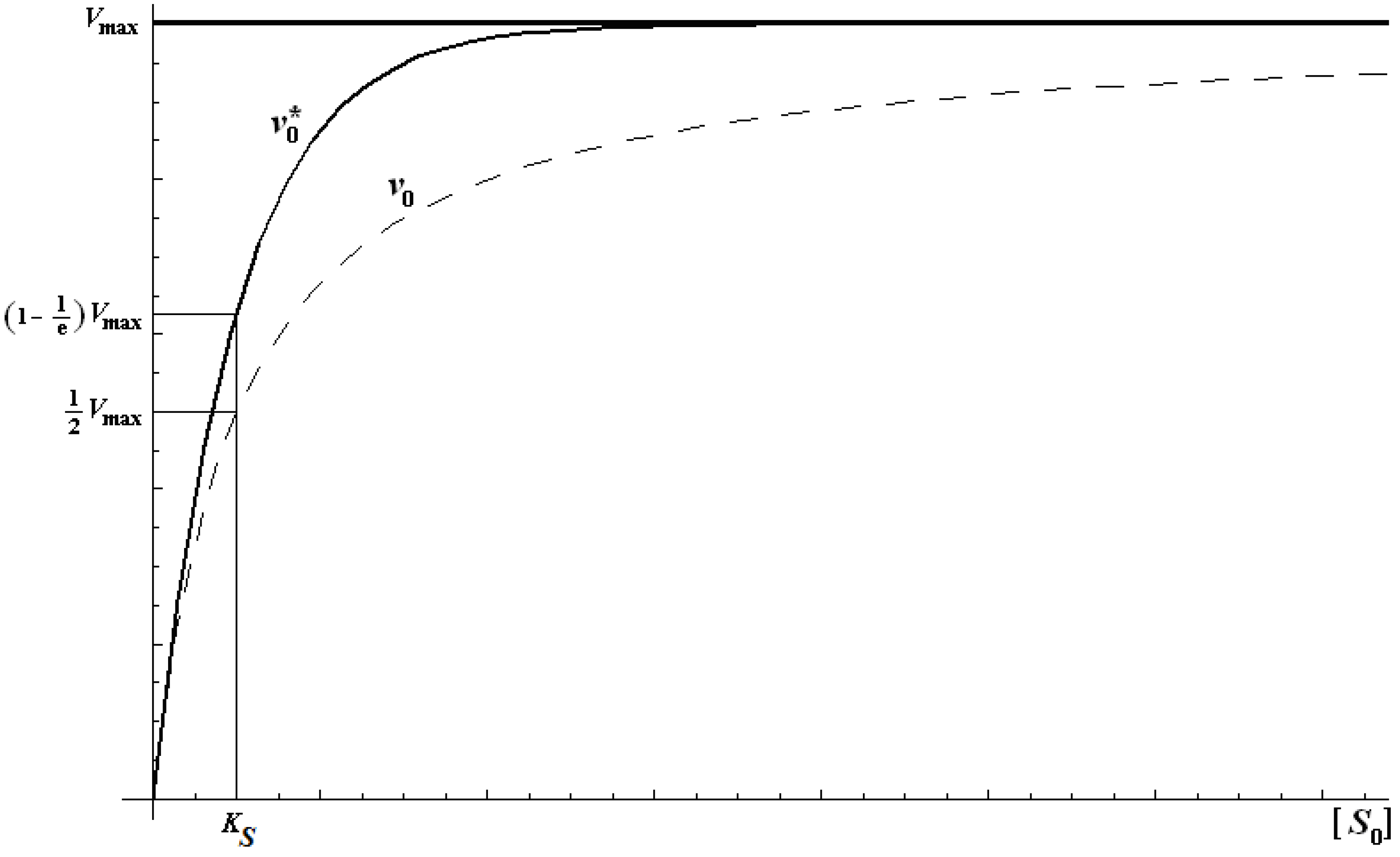
2.2. Logistic Enzyme Kinetics
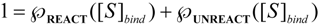
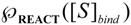 is the probability that the enzymatic reaction of Equation (1), for instance, proceeds at a certain concentration of substrate binding
is the probability that the enzymatic reaction of Equation (1), for instance, proceeds at a certain concentration of substrate binding 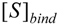 to the enzyme; it features the limits of the occurrence of products in E-S reactions:
to the enzyme; it features the limits of the occurrence of products in E-S reactions:
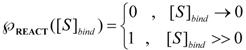
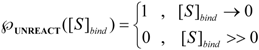
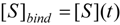 , followed by identification of the reactive term (16) as having the general form:
, followed by identification of the reactive term (16) as having the general form:
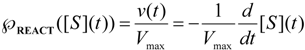
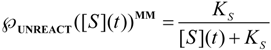
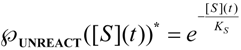
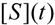 first order expansion for the latter case where the bound substrate approaches zero limits, i.e. within the low substrate concentrations.
first order expansion for the latter case where the bound substrate approaches zero limits, i.e. within the low substrate concentrations.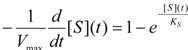
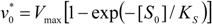 .
.
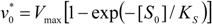 .
.
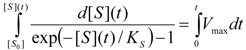
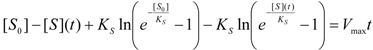
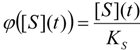
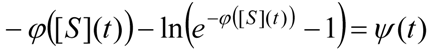
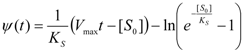
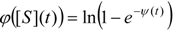
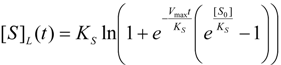
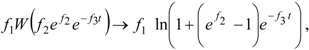
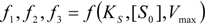
3. Results and Discussion
3.1. Probabilistic form of the Haldane-Radić Equation
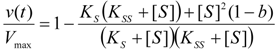
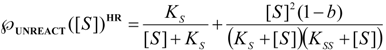
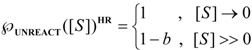

3.2. Temporal Solution of Haldane-Radić Equation by W-Lambert Functional
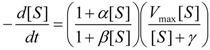
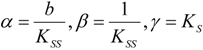
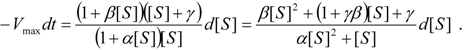
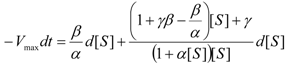
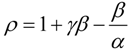
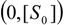 an the current one
an the current one 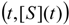 , it firstly yields:
, it firstly yields:
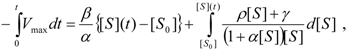
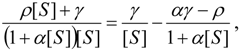
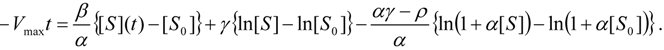
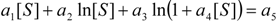
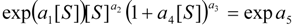
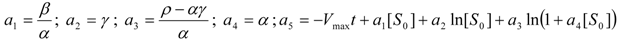
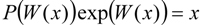

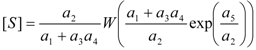
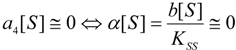
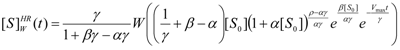
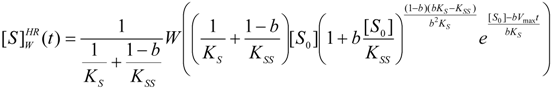
3.3. Temporal Solution of Haldane-Radić Equation by Analytic Logistic Transformation
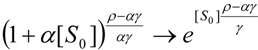
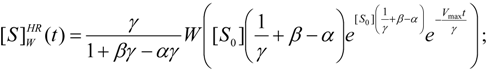
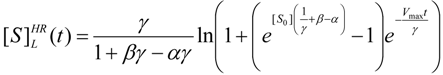
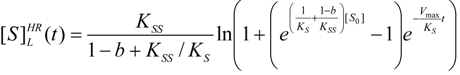
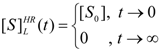
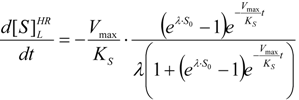
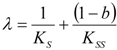
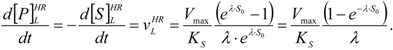
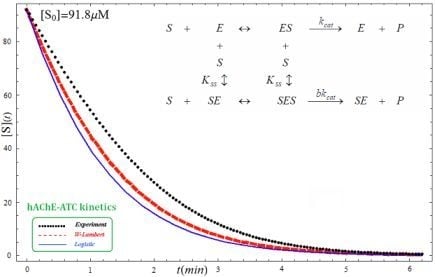
- The hAChE-ATC kinetics (Figure 3) differs from hBChE-ATC kinetics (Figure 4) essentially only in the lowering the Vmax and increasing b parameters for the last case, in accordance with the prescription associated with activation mechanism; moreover, the W-Lambert and logistic curves depart clock-wise from experimental record and more quickly for logistic case;
- The hBChE-ATC kinetics (Figure 4) differs from hBChE-BTC kinetics (Figure 5) essentially by further lowering the Vmax accompanied by decrease of KS parameter for the BTC kinetics, while the W-Lambert and logistic computationally fitting curves show in Figure 5 a departure tendency in anti-clock-wise respecting the experimental evidence; here is also recorded the clear failure of the numerical W-Lambert progress curve to reach the initial substrate concentration, a matter fully satisfied by the logistic counterpart instead;
- Comparison between hBChE-BTC kinetics (Figure 5) and BSCh-BTC hydrolysis (Figure 6) reveals that by maintaining the same kinetic parameters between these two cases, in the latter, the computational fitting with respect the experimental data oscillate from clock-wise to anti-clock-wise departure of the logistic model as the initial substrate concentration goes from lower (<100 µM) to higher (>100 µM) values, respectively; here, again, the initial time discrepancy between W-Lambert and logistic kinetics is obviously in the favor of the latter approach.
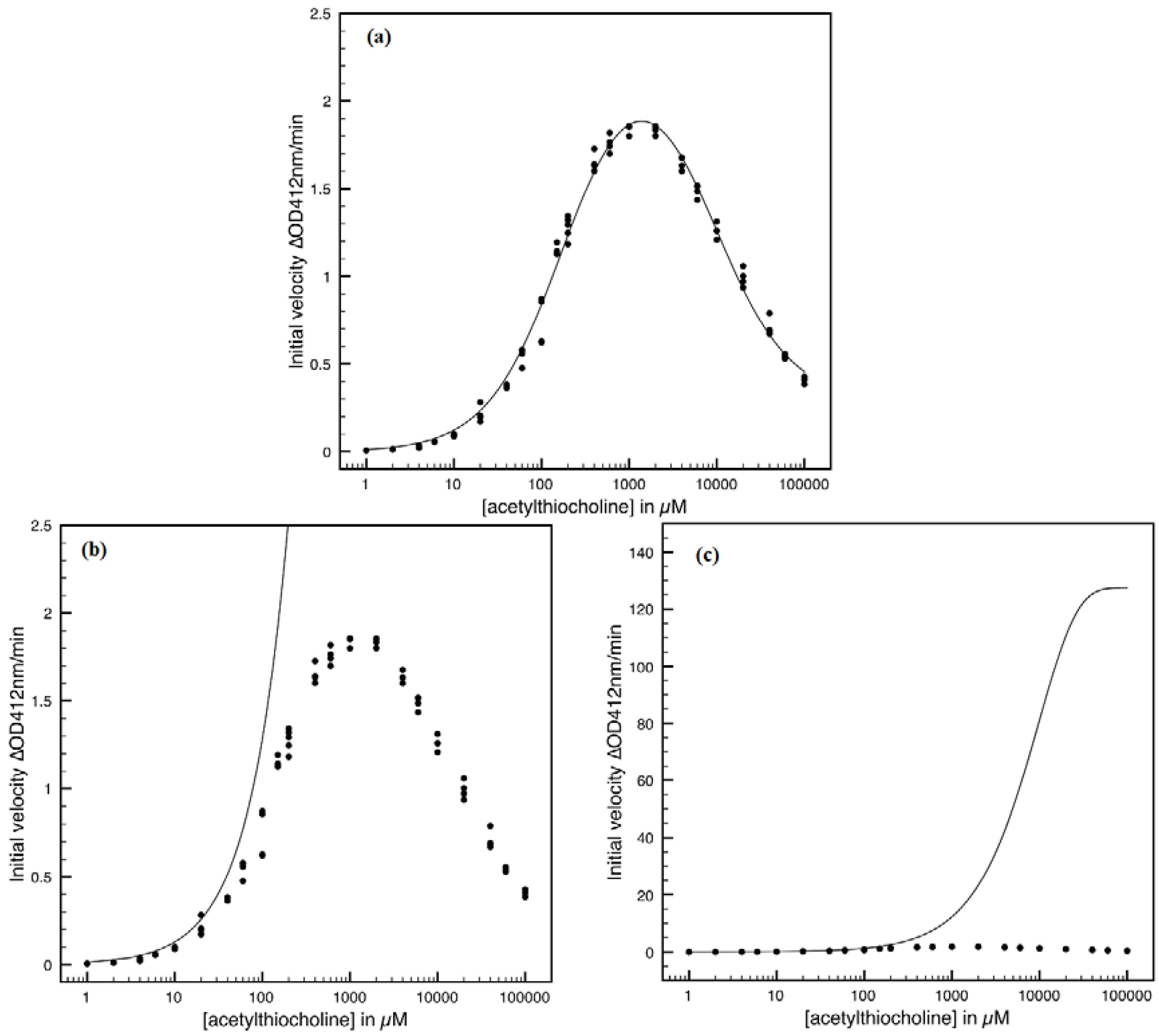
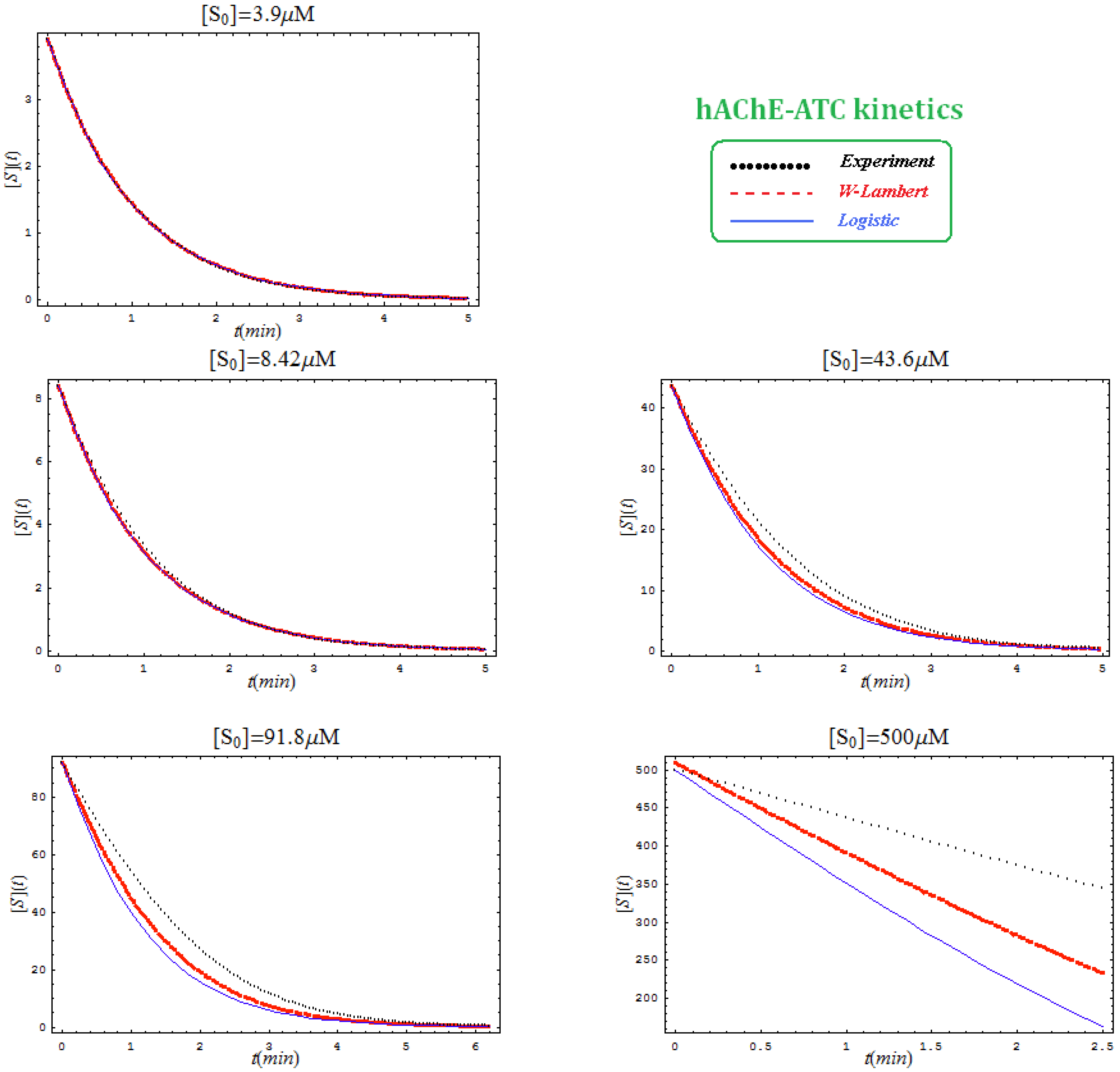
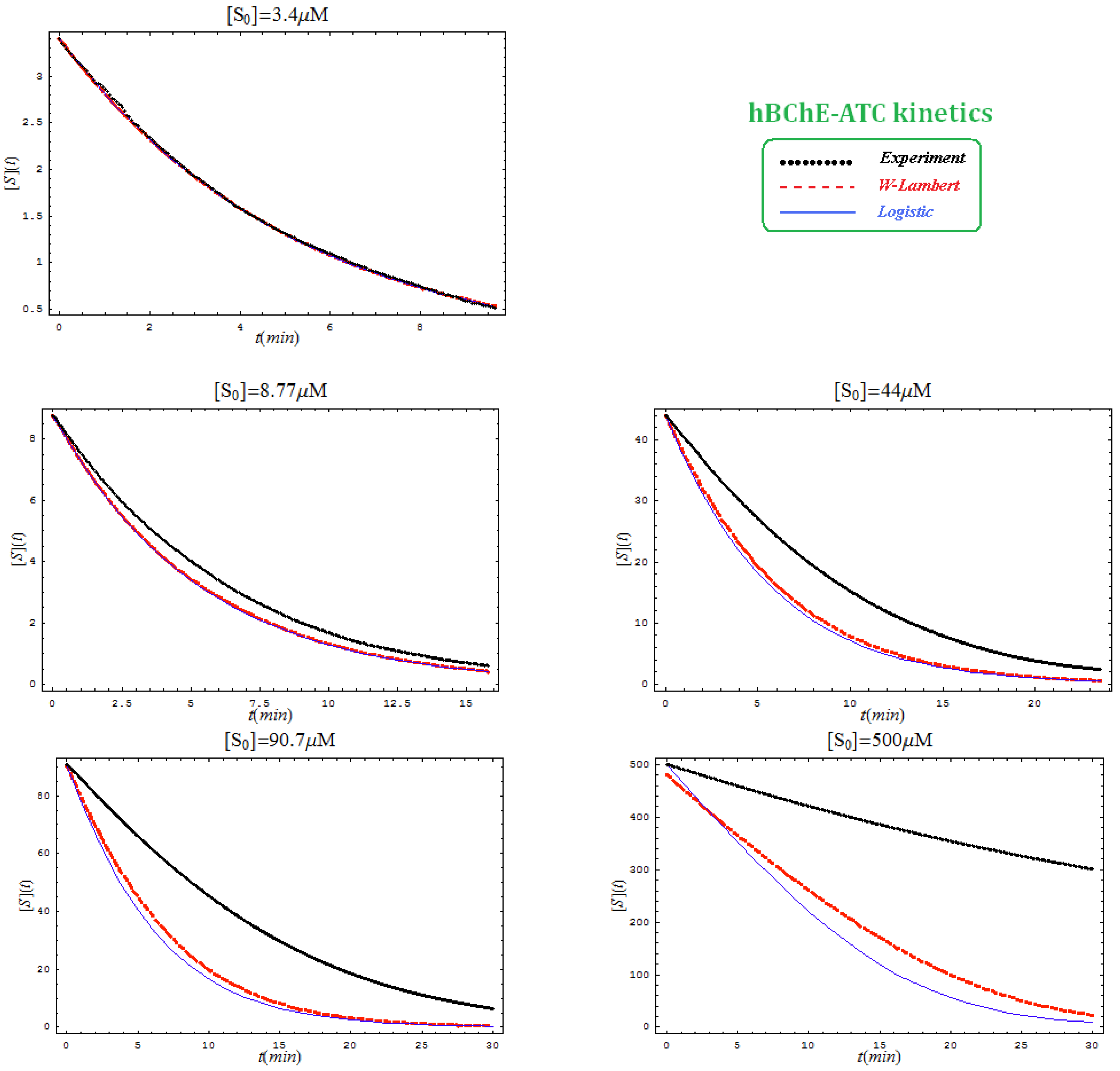

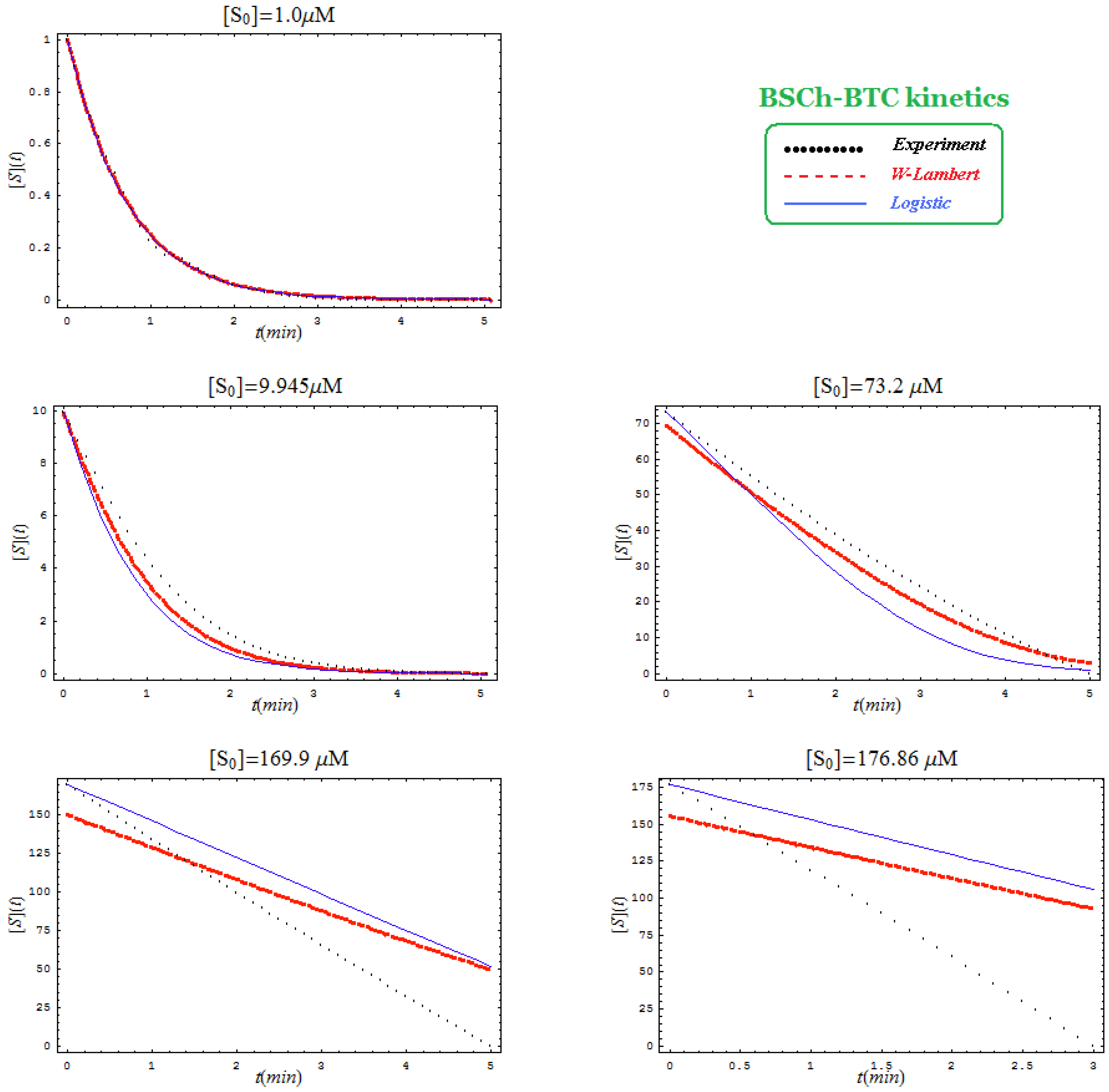
- The Haldane-Radić kinetics may be quite well modeled by its Michaelis-Menten counterpart progress curves for substrate kinetics below 100 µM in all studied cases, being this condition susceptible to be a general fact that is independent of ideal approach for the Haldane-Radić kinetic parameters KSS and b as prescribed in Equation (7);
- Haldane-Radić kinetics display full specificity in looping S-E mechanisms of inhibition/activation for higher concentration of the substrate, i.e. within the mili-molar range;
4. Conclusions
- Higher dissociation constant for inhibition/activation substrate site interaction to enzyme;
- Equal catalytic efficiency of inhibition/activation substrate-enzyme loop as provided by E-S hydrolysis;
- Lower substrate concentration, typically in the range up to the 100 µM.
Acknowledgements
References
- Henri, V. Über das gesetz der wirkung des invertins. Z. Phys. Chem. 1901, 39, 194–216. [Google Scholar]
- Michaelis, L.; Menten, M.L. Die kinetik der invertinwirkung. Biochem. Z. 1913, 49, 333–369. [Google Scholar]
- Rubinow, S.I. Introduction to Mathematical Biology; Wiley: New York, NY, USA, 1975. [Google Scholar]
- Cantor, C.R.; Schimmel, P.R. Biophysical Chemistry. Part III: The Behavior of Biological Macromolecules; Freeman & Co.: San Francisco, CA, USA, 1980. [Google Scholar]
- Murray, J.D. Mathematical Biology; Springer Verlag: Berlin-Heidelberg: Germany, 1990. [Google Scholar]
- Voet, D.; Voet, J.G. Biochemistry, 2nd ed; John Wiley & Sons, Inc.: New York, NY, USA, 1995. [Google Scholar]
- Copeland, R.A. Enzymes; Wiley-VCH: New York, NY, USA, 2000. [Google Scholar]
- Ross, J.; Schreiber, I.; Vlad, M.O. Determination of Complex Reaction Mechanisms: Analysis of Chemical, Biological and Genetic Networks; Oxford University Press: Oxford, UK, 2006. [Google Scholar]
- Schnell, S.; Maini, P.K. A Century of enzyme kinetics: Reliability of the KM and vmax estimates. Comm. Theor. Biol. 2003, 8, 169–187. [Google Scholar] [CrossRef]
- Goudar, C.T.; Sonnad, J.R.; Duggleby, R.G. Parameter estimation using a direct solution of the integrated Michaelis-Menten equation. Biochim. Biophys. Acta 1999, 1429, 377–383. [Google Scholar] [CrossRef]
- Schnell, S.; Mendoza, C. Closed form solution for time-dependent enzyme kinetics. J. Theor. Biol. 1997, 187, 207–212. [Google Scholar] [CrossRef]
- Schnell, S.; Mendoza, C. Enzyme kinetics of multiple alternative substrates. J. Math. Chem. 2000, 27, 155–170. [Google Scholar] [CrossRef]
- Schnell, S.; Mendoza, C. Time-dependent closed form solution for fully competitive enzyme reactions. Bull. Math. Biol. 2000, 62, 321–336. [Google Scholar] [CrossRef]
- Goudar, C.T.; Harris, S.K.; McInerney, M.J.; Suflita, J.M. Progress curve analysis for enzyme and microbial kinetic reactions using explicit solutions based on the Lambert W function. J. Microbiol. Methods 2004, 59, 317–326. [Google Scholar] [CrossRef]
- Euler, L. De serie Lambertina plurimisque eius insignibus proprietatibus (1779), reprinted at B.G. Teubner, Leipzig and Berlin. Opera Omnia 1921, 6, 350–369. [Google Scholar]
- Hayes, B. Why W? Amer. Sci. 2005, 93, 104–108. [Google Scholar]
- Hirs, C.H.W. Methods Enzymol. 11 (Enzyme Structure); Academic Press: New York, NY, USA, 1967. [Google Scholar]
- Rigby, P.W.J.; Burleigh, B.D.; Hartley, B.S. Gene duplication in experimental enzyme evolution. Nature 1974, 251, 200–204. [Google Scholar]
- Graham, L.D.; Haggett, K.D.; Hayes, P.J.; Schober, P.A.; Jennings, P.A.; Whittaker, R.G. A new library of alpha-lytic protease S1 mutants generated by combinatorial random substitution. Biochem. Mol. Biol. Int. 1994, 32, 831–839. [Google Scholar]
- Tyagi, R.; Lee, Y.-T.; Guddat, L.W.; Duggleby, R.G. Probing the mechanism of the bifunctional enzyme ketol-acid reductoisomerase by site-directed mutagenesis of the active site. FEBS J. 2005, 272, 593–602. [Google Scholar]
- Stemmer, W.P.C. Rapid evolution of a protein in vitro by DNA shuffling. Nature 1994, 370, 389–391. [Google Scholar] [CrossRef]
- Radić, Z.; Gibney, G.; Kawamoto, S.; MacPhee-Quigley, K.; Bongiorno, C.; Taylor, P. Expression of recombinant acetylcholinesterase in a baculovirus system: Kinetic properties of glutamate 199 mutants. Biochemistry 1992, 31, 9760–9767. [Google Scholar] [CrossRef]
- Radić, Z.; Pickering, N.A.; Vellom, D.C.; Camp, S. Palmer Taylor Three distinct domains in the cholinesterase molecule confer selectivity for acetyl- and butyrylcholinesterase inhibitors. Biochemistry 1993, 32, 12074–12084. [Google Scholar]
- Stojan, J.; Marcel, V.; Estrada-Mondaca, S.; Klaebe, A.; Masson, P.; Fournier, D. A putative kinetic model for substrate metabolisation by Drosophila acetylcholinesterase. FEBS Lett. 1998, 440, 85–88. [Google Scholar] [CrossRef]
- Stojan, J.; Brochier, L.; Aliès, C.; Colletier, J.P.; Fournier, D. Inhibition of Drosophila melanogaster acetylcholinesterase by high concentrations of substrate. Eur. J. Biochem. 2004, 271, 1364–1371. [Google Scholar]
- Nachon, F.; Stojan, J.; Fournier, D. Insights into substrate and product traffic in the Drosophila melanogaster acetylcholinesterase active site gorge by enlarging a back channel. FEBS J. 2008, 275, 2659–2664. [Google Scholar] [CrossRef]
- Haldane, J.B.S. The Enzymes; Longmans-Green: London, UK, 1930. [Google Scholar]
- Grady, C.P.L., Jr.; Daigger, G.T.; Lim, H.C. Biological Wastewater Treatment, 2nd ed; Marcel Dekker: New York, NY, USA, 1999. [Google Scholar]
- Ellis, T.G.; Barbeau, D.S.; Smets, B.F.; Grady, C.P.L., Jr. Respirometric technique for determination of extant kinetic parameters describing biodegradation. Wat. Env. Res. 1996, 68, 917–926. [Google Scholar] [CrossRef]
- Sonad, J.R.; Goudar, C.T. Solution of the Haldane equation for substrate inhibition enzyme kinetics using the decomposition method. Math. Comput. Model. 2004, 40, 573–582. [Google Scholar] [CrossRef]
- Putz, M.V.; Lacrămă, A.-M.; Ostafe, V. Introducing logistic enzyme kinetics. J. Optoelectron. Adv. Mat. 2007, 9, 2910–2916. [Google Scholar]
- Putz, M.V.; Lacrămă, A.-M.; Ostafe, V. Full Analytic Progress Curves of the Enzymic Reactions in Vitro. Int. J. Mol. Sci. 2006, 7, 469–484. [Google Scholar] [CrossRef]
- Putz, M.V.; Lacrămă, A.-M. Enzymatic control of the bio-inspired nanomaterials at the spectroscopic level. J. Optoelectron. Adv. Mat. 2007, 9, 2529–2534. [Google Scholar]
- Putz, M.V.; Putz, A.M. Logistic vs. W-Lambert information in quantum modeling of enzyme kinetics. Int. J. Chemoinf. Chem. Eng. 2011, 1, 42–60. [Google Scholar]
- Laidler, K.J. Theory of transient phase in kinetics, with special reference to enzyme systems. Can. J. Chem. 1955, 33, 1614–1624. [Google Scholar] [CrossRef]
- Duggleby, R.G.; Morrison, J.F. The use of steady-state rate equations to analyse progress curve data. Biochim. Biophys. Acta 1979, 568, 357–362. [Google Scholar]
- Sample Availability: Not available.
© 2011 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Putz, M.V. On the Reducible Character of Haldane-Radić Enzyme Kinetics to Conventional and Logistic Michaelis-Menten Models. Molecules 2011, 16, 3128-3145. https://doi.org/10.3390/molecules16043128
Putz MV. On the Reducible Character of Haldane-Radić Enzyme Kinetics to Conventional and Logistic Michaelis-Menten Models. Molecules. 2011; 16(4):3128-3145. https://doi.org/10.3390/molecules16043128
Chicago/Turabian StylePutz, Mihai V. 2011. "On the Reducible Character of Haldane-Radić Enzyme Kinetics to Conventional and Logistic Michaelis-Menten Models" Molecules 16, no. 4: 3128-3145. https://doi.org/10.3390/molecules16043128
APA StylePutz, M. V. (2011). On the Reducible Character of Haldane-Radić Enzyme Kinetics to Conventional and Logistic Michaelis-Menten Models. Molecules, 16(4), 3128-3145. https://doi.org/10.3390/molecules16043128