Identification of Calpain Substrates by ORF Phage Display
Abstract
:1. Introduction
2. Results and Discussion
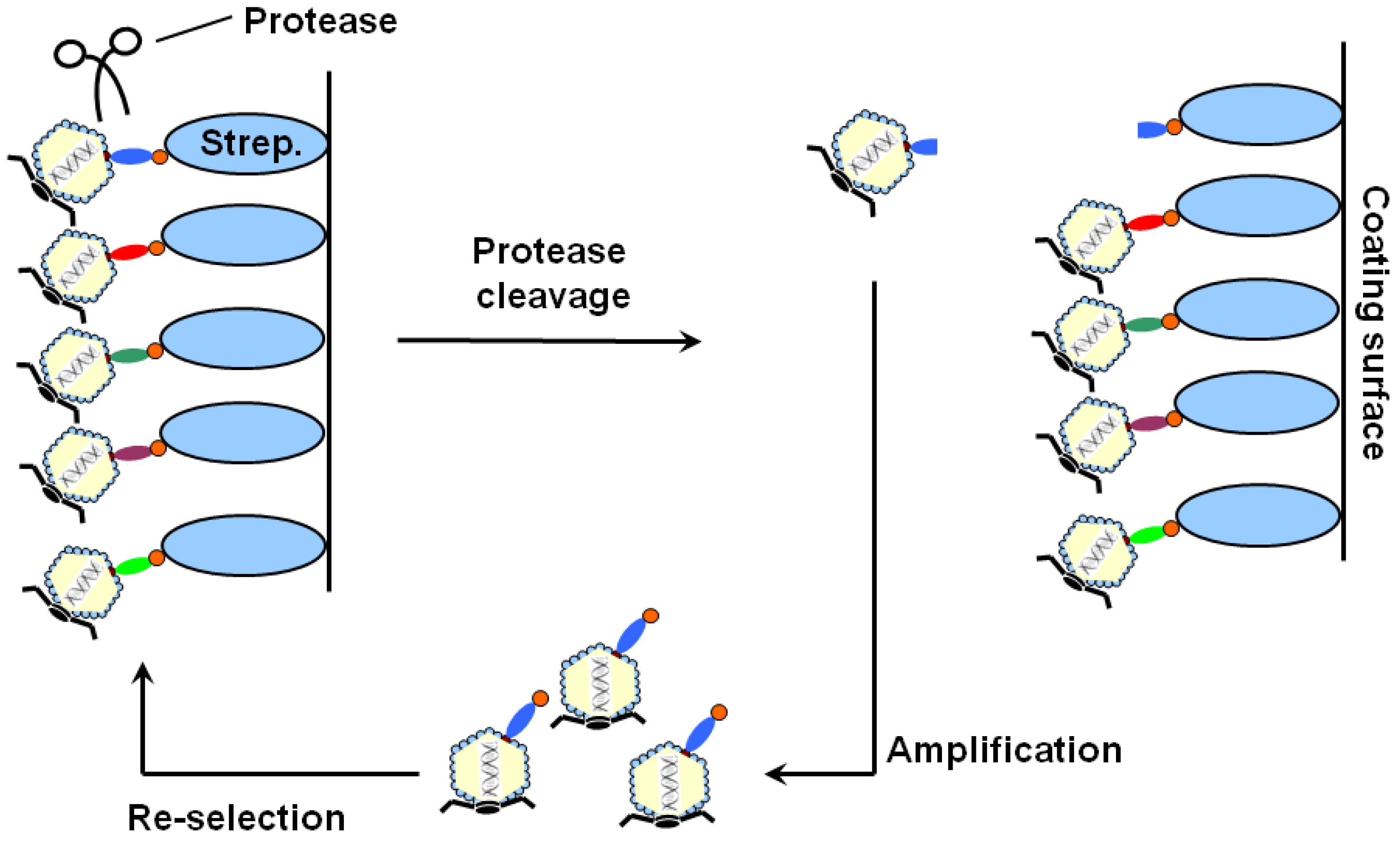
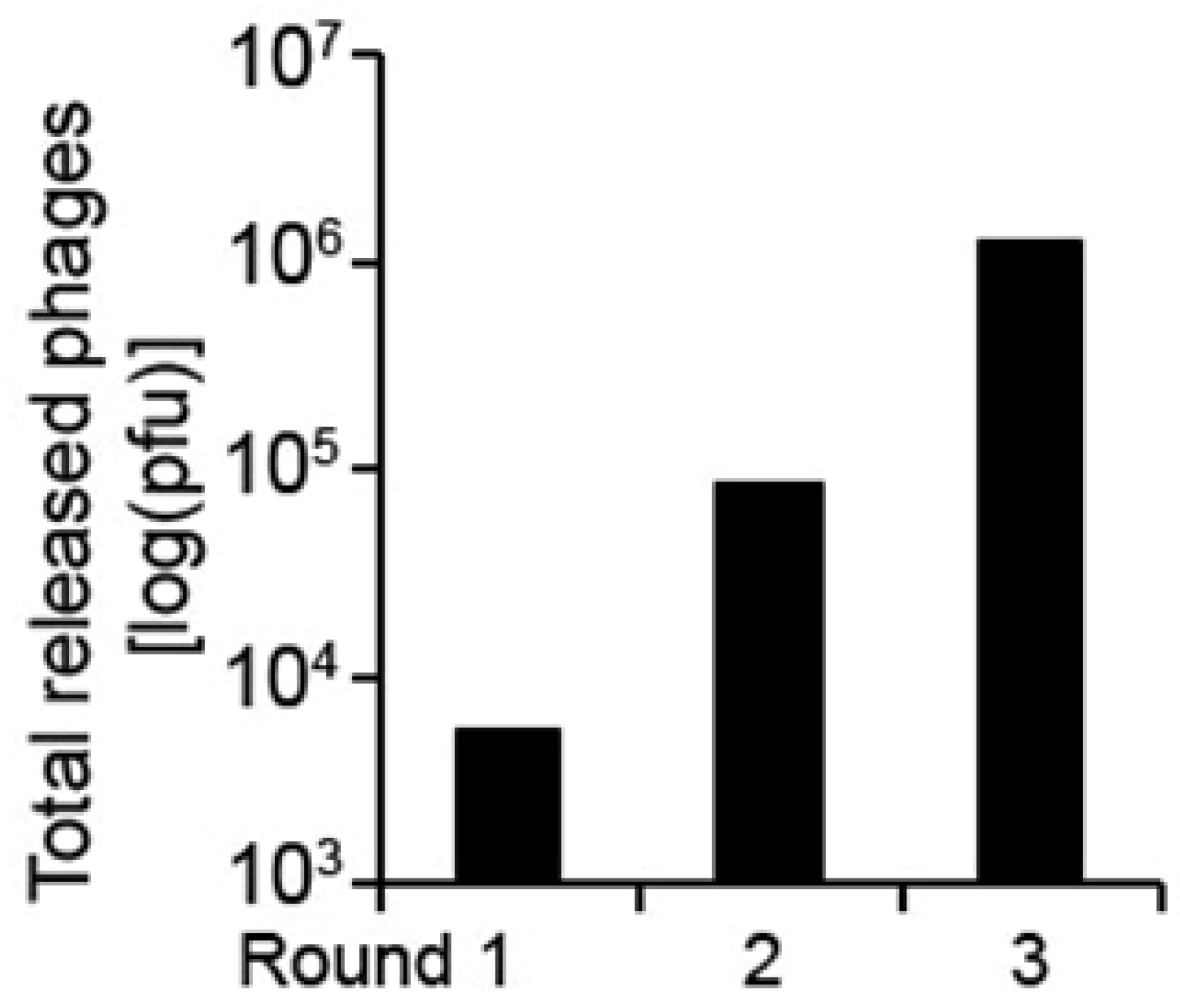
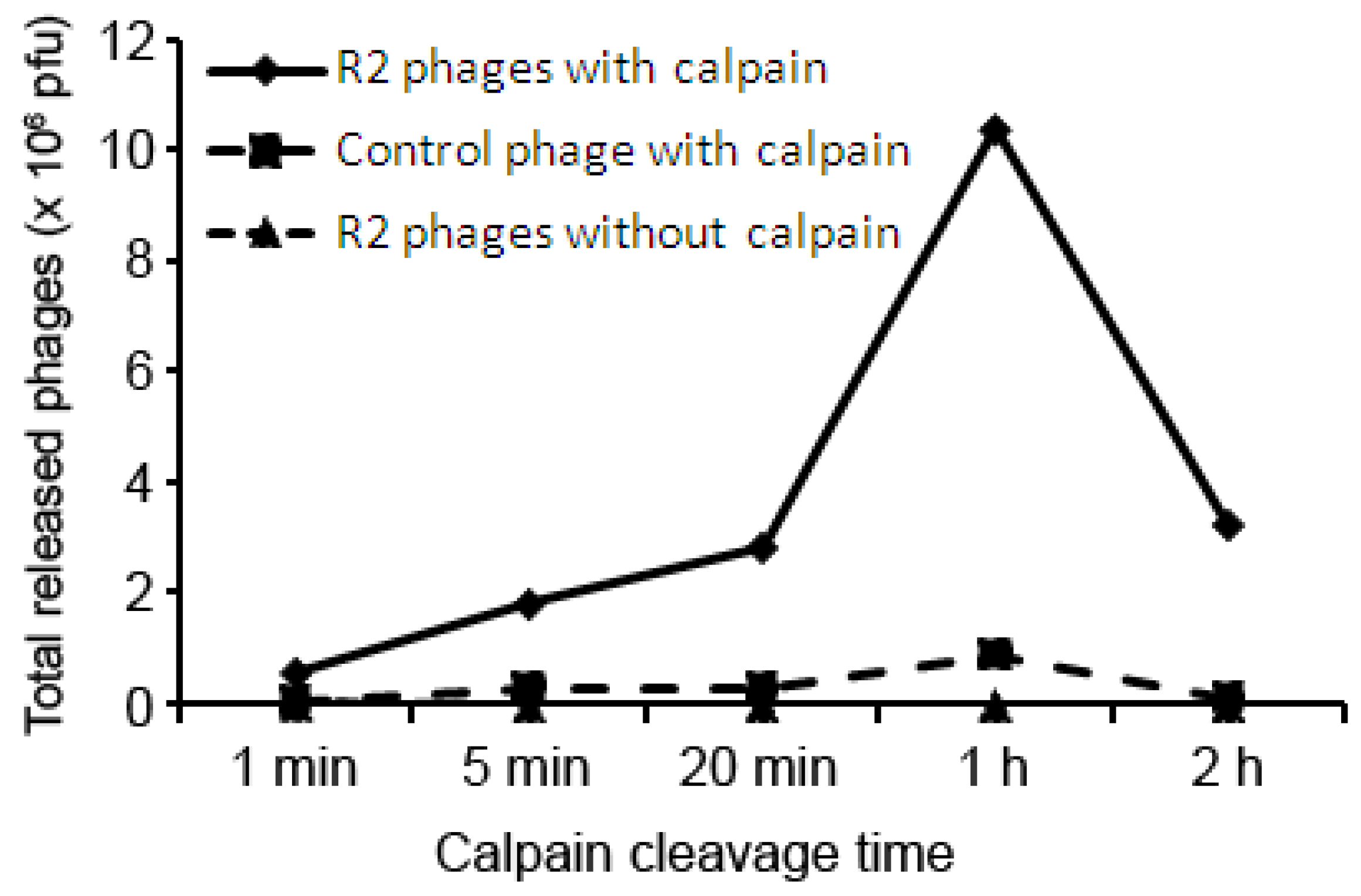
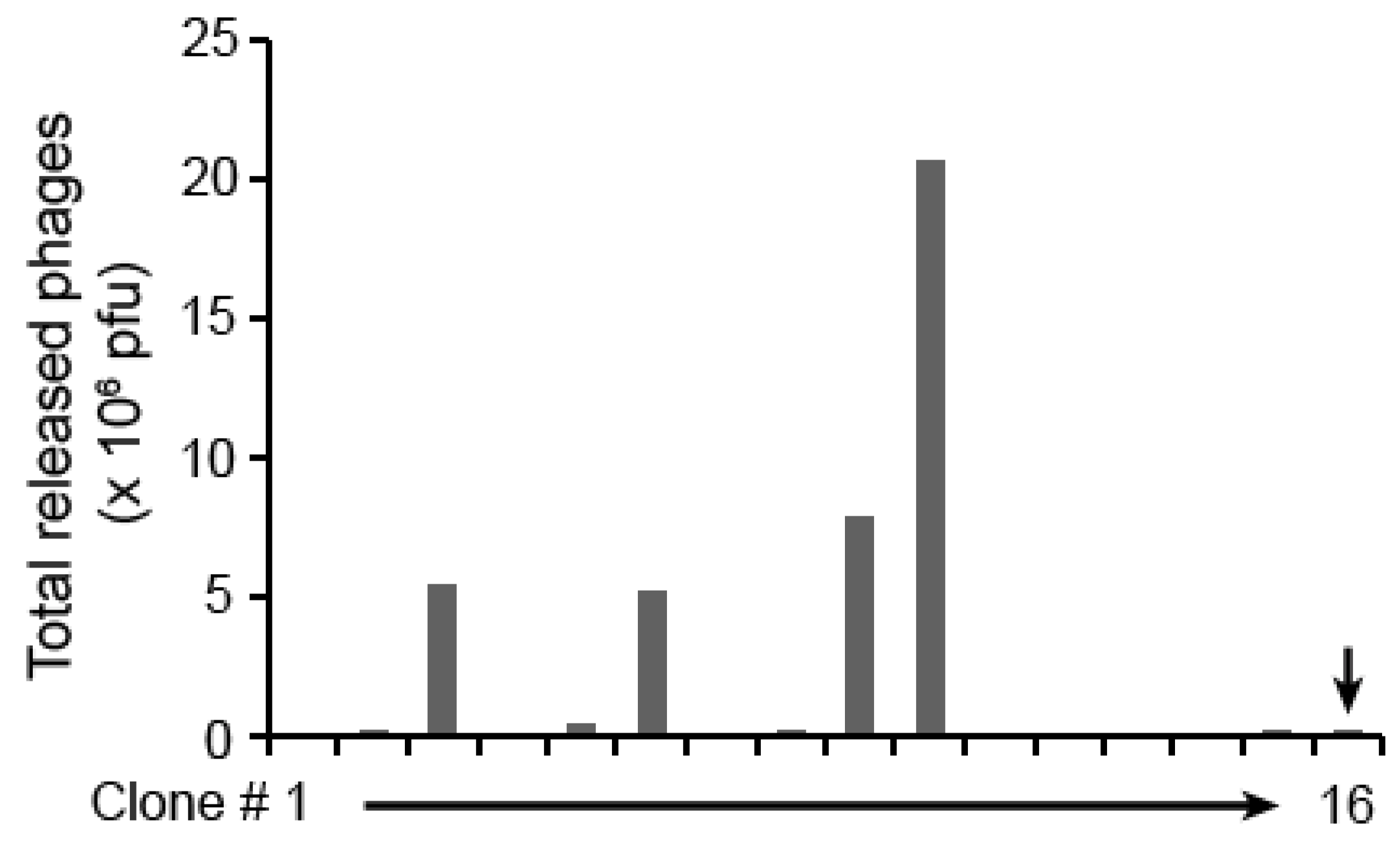
| Protein name | Accession number | Matched aa residues | Substrate relative cleavage activity | |
|---|---|---|---|---|
| Calpain vs. controla | Ca2+ vs. EDTAb | |||
| Son cell proliferation protein (Son) | NM_178880 | 690L-833T | ~90.6 | ~28.4 |
| CCR4-NOT transcription complex, subunit 3 (Cnot3) | NM_146176 | 64T-232E | ~16.7 | ~7.9 |
| Skeletal muscle heavy polypeptides | Multiplec | Multiple | ~35.1 | ~117.0 |
| Ring finger protein 146 (Rnf146) | NM_026518 | 147V-338G | ~62.4 | ~24.9 |
| Catenin (cadherin associated protein), delta 2 (Ctnnd2) | NM_008729 | 1081S-11969H | ~16.5 | ~16.8 |
| Caldesmon 1 (Cald1) | NM_145575 | 121D-255R | ~27.9 | ~14.1 |
| Thymopoietin (Tmpo) | NM_001080132 | 89A-266E | ~60.3 | ~43.7 |
| Golgin subfamily a, 4 (Golga4) | NM_018748 | 1553K-1631K | ~16.2 | ~9.1 |
| Caldesmon 1 (Cald1) | NM_145575 | 131M-255R | ~30.3 | ~11.1 |
| Specc1 | NM_001029936 | 163E-197Q | ~36.4 | ~11.2 |
| Calpastatin (Cast) | NM_009817 | 523P-623G | ~436.1 | ~54.5 |
3. Experimental
3.1. General
3.2. Phage selection
3.3. Clone screening
3.4. Cleavage efficiency
4. Conclusions
Acknowledgements
References
- Suzuki, K.; Hata, S.; Kawabata, Y.; Sorimachi, H. Structure, activation, and biology of calpain. Diabetes 2004, 53 Suppl. 1, S12–S18. [Google Scholar]
- Paquet-Durand, F.; Johnson, L.; Ekstrom, P. Calpain activity in retinal degeneration. J. Neurosci. Res. 2007, 85, 693–702. [Google Scholar] [CrossRef]
- Huang, W.; Fileta, J.; Rawe, I.; Qu, J.; Grosskreutz, C.L. Calpain activation in experimental glaucoma. Invest. Ophthalmol. Vis. Sci. 2010, 51, 3049–3054. [Google Scholar] [CrossRef]
- Hood, J.L.; Brooks, W.H.; Roszman, T.L. Subcellular mobility of the calpain/calpastatin network: an organelle transient. Bioessays 2006, 28, 850–859. [Google Scholar] [CrossRef]
- Diamond, S.L. Methods for mapping protease specificity. Curr. Opin. Chem. Biol. 2007, 11, 46–51. [Google Scholar] [CrossRef]
- Deperthes, D. Phage display substrate: A blind method for determining protease specificity. Biol. Chem. 2002, 383, 1107–1112. [Google Scholar]
- Lien, S.; Francis, G.L.; Graham, L.D.; Wallace, J.C. Isolating substrates for an engineered alpha-lytic protease by phage display. J. Protein. Chem. 2003, 22, 155–166. [Google Scholar] [CrossRef]
- McCarter, J.D.; Stephens, D.; Shoemaker, K.; Rosenberg, S.; Kirsch, J.F.; Georgiou, G. Substrate specificity of the Escherichia coli outer membrane protease OmpT. J. Bacteriol. 2004, 186, 5919–5925. [Google Scholar] [CrossRef]
- Kerr, F.K.; O'Brien, G.; Quinsey, N.S.; Whisstock, J.C.; Boyd, S.; de la Banda, M.G.; Kaiserman, D.; Matthews, A.Y.; Bird, P.I.; Pike, R.N. Elucidation of the substrate specificity of the C1s protease of the classical complement pathway. J. Biol. Chem. 2005, 280, 39510–39514. [Google Scholar]
- Lin, H.S.; Talwar, H.S.; Tarca, A.L.; Ionan, A.; Chatterjee, M.; Ye, B.; Wojciechowski, J.; Mohapatra, S.; Basson, M.D.; Yoo, G.H.; Peshek, B; Lonardo, F.; Pan, C.J.; Folbe, A.J.; Draghici, S.; Abrams, J.; Tainsky, M.A. Autoantibody approach for serum-based detection of head and neck cancer. Cancer Epidemiol. Biomarkers Prev. 2007, 16, 2396–2405. [Google Scholar]
- Kalnina, Z.; Silina, K.; Meistere, I.; Zayakin, P.; Rivosh, A.; Abols, A.; Leja, M.; Minenkova, O.; Schadendorf, D.; Line, A. Evaluation of T7 and lambda phage display systems for survey of autoantibody profiles in cancer patients. J. Immunol. Methods 2008, 334, 37–50. [Google Scholar] [CrossRef]
- Li, W.; Caberoy, N.B. New perspective for phage display as an efficient and versatile technology of functional proteomics. Appl. Microbiol. Biotechnol. 2010, 85, 909–919. [Google Scholar] [CrossRef]
- Caberoy, N.B.; Zhou, Y.; Jiang, X.; Alvarado, G.; Li, W. Efficient identification of tubby-binding proteins by an improved system of T7 phage display. J. Mol. Recognit. 2010, 23, 74–83. [Google Scholar]
- Caberoy, N.B.; Zhou, Y.; Alvarado, G.; Fan, X.; Li, W. Efficient identification of phosphatidylserine-binding proteins by ORF phage display. Biochem. Biophys. Res. Commun. 2009, 386, 197–201. [Google Scholar] [CrossRef]
- Caberoy, N.B.; Maiguel, D.; Kim, Y.; Li, W. Identification of tubby and tubby-like protein 1 as eat-me signals by phage display. Exp. Cell Res. 2010, 316, 245–257. [Google Scholar] [CrossRef]
- Kim, Y.; Caberoy, N.B.; Alvarado, G.; Davis, J.L.; Feuer, W.J.; Li, W. Identification of Hnrph3 as an autoantigen for acute anterior uveitis. Clin. Immunol. 2011, 138, 60–66. [Google Scholar] [CrossRef]
- Rosenberg, A.; Griffin, K.; Studier, F.W.; McCormick, M.; Berg, J.; Novy, R.; Mierendorf, R. T7Select phage display system: A powerful new protein display system based on bacteriophage T7. InNovations 1996, 6, 1–6. [Google Scholar]
- Schatz, P.J. Use of peptide libraries to map the substrate specificity of a peptide-modifying enzyme: A 13 residue consensus peptide specifies biotinylation in Escherichia coli. Biotechnology (N Y) 1993, 11, 1138–1143. [Google Scholar] [CrossRef]
- Ansuini, H.; Cicchini, C.; Nicosia, A.; Tripodi, M.; Cortese, R.; Luzzago, A. Biotin-tagged cDNA expression libraries displayed on lambda phage: A new tool for the selection of natural protein ligands. Nucl. Acids Res. 2002, 30, e78. [Google Scholar]
- Caberoy, N.B.; Zhou, Y.; Li, W. Can phage display be used as a tool to functionally identify endogenous eat-me signals in phagocytosis? J. Biomol. Screen. 2009, 14, 653–661. [Google Scholar] [CrossRef]
- Wendt, A.; Thompson, V.F.; Goll, D.E. Interaction of calpastatin with calpain: A review. Biol. Chem. 2004, 385, 465–472. [Google Scholar]
- Goll, D.E.; Thompson, V.F.; Li, H.; Wei, W.; Cong, J. The calpain system. Physiol. Rev. 2003, 83, 731–801. [Google Scholar]
- Schilling, O.; Overall, C.M. Proteomic discovery of protease substrates. Curr. Opin. Chem. Biol. 2007, 11, 36–45. [Google Scholar] [CrossRef]
- Agard, N.J.; Wells, J.A. Methods for the proteomic identification of protease substrates. Curr. Opin. Chem. Biol. 2009, 13, 503–509. [Google Scholar] [CrossRef]
- Bozoky, Z.; Alexa, A.; Dancsok, J.; Gogl, G.; Klement, E.; Medzihradszky, K.F.; Friedrich, P. Identifying calpain substrates in intact S2 cells of Drosophila. Arch. Biochem. Biophys. 2009, 481, 219–225. [Google Scholar] [CrossRef]
- Cuerrier, D.; Moldoveanu, T.; Davies, P.L. Determination of peptide substrate specificity for mu-calpain by a peptide library-based approach: The importance of primed side interactions. J. Biol. Chem. 2005, 280, 40632–40641. [Google Scholar] [CrossRef]
- Dias-Neto, E.; Nunes, D.N.; Giordano, R.J.; Sun, J.; Botz, G.H.; Yang, K.; Setubal, J.C.; Pasqualini, R.; Arap, W. Next-generation phage display: Integrating and comparing available molecular tools to enable cost-effective high-throughput analysis. PLoS One 2009, 4, e8338. [Google Scholar]
- Di Niro, R.; Sulic, A.M.; Mignone, F.; D'Angelo, S.; Bordoni, R.; Iacono, M.; Marzari, R.; Gaiotto, T.; Lavric, M.; Bradbury, A.R.; Biancone, L.; Zevin-Sonkin, D.; De Bellis, G.; Santoro, C.; Sblattero, D. Rapid interactome profiling by massive sequencing. Nucl. Acids Res. 2010, 38, e110. [Google Scholar]
- Fowler, D.M.; Araya, C.L.; Fleishman, S.J.; Kellogg, E.H.; Stephany, J.J.; Baker, D.; Fields, S. High-resolution mapping of protein sequence-function relationships. Nat. Methods 2010, 7, 741–746. [Google Scholar] [CrossRef]
- Caberoy, N.B.; Zhou, Y.; Li, W. Tubby and tubby-like protein 1 are new MerTK ligands for phagocytosis. EMBO J. 2010, 29, 3898–3910. [Google Scholar] [CrossRef]
- Paschke, M. Phage display systems and their applications. Appl. Microbiol. Biotechnol. 2006, 70, 2–11. [Google Scholar] [CrossRef]
- Krumpe, L.R.; Atkinson, A.J.; Smythers, G.W.; Kandel, A.; Schumacher, K.M.; McMahon, J.B.; Makowski, L.; Mori, T. T7 lytic phage-displayed peptide libraries exhibit less sequence bias than M13 filamentous phage-displayed peptide libraries. Proteomics 2006, 6, 4210–4222. [Google Scholar]
- Bratkovic, T. Progress in phage display: Evolution of the technique and its application. Cell. Mol. Life Sci. 2010, 67, 749–767. [Google Scholar] [CrossRef]
- Zhang, Y.; Davis, J.L.; Li, W. Identification of tribbles homolog 2 as an autoantigen in autoimmune uveitis by phage display. Mol. Immunol. 2005, 42, 1275–1281. [Google Scholar] [CrossRef]
- Sample Availability: Samples of the compounds in Table 1are available from the authors.
© 2011 by the authors; licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution license ( http://creativecommons.org/licenses/by/3.0/).
Share and Cite
Caberoy, N.B.; Alvarado, G.; Li, W. Identification of Calpain Substrates by ORF Phage Display. Molecules 2011, 16, 1739-1748. https://doi.org/10.3390/molecules16021739
Caberoy NB, Alvarado G, Li W. Identification of Calpain Substrates by ORF Phage Display. Molecules. 2011; 16(2):1739-1748. https://doi.org/10.3390/molecules16021739
Chicago/Turabian StyleCaberoy, Nora B., Gabriela Alvarado, and Wei Li. 2011. "Identification of Calpain Substrates by ORF Phage Display" Molecules 16, no. 2: 1739-1748. https://doi.org/10.3390/molecules16021739
APA StyleCaberoy, N. B., Alvarado, G., & Li, W. (2011). Identification of Calpain Substrates by ORF Phage Display. Molecules, 16(2), 1739-1748. https://doi.org/10.3390/molecules16021739




