3. Plant Grinding
High throughput grinding and extraction was a requirement of the new Anticancer Drug Discovery Program that was established in 1987, as the extracts which were produced were to serve as the ‘feedstock’ for the new 60 human cancer cell line panel which was to serve as the primary discovery tool used for detection of anticancer lead compounds [
3,
4]. Processing protocols had to be developed which were capable of producing many thousands of extracts from thousands of dry plant collections per year. But the goal was not simply to produce thousands of extracts, or to maximize the weight of extract obtained in the extraction process, but rather to produce crude extracts containing a representation of all molecules found in the specimen in their unchanged, un-degraded, and hopefully, biologically active state. Certain factors are widely believed to be enemies of the preservation of biological activity, including heat, chemical reactivity, time in solution, light, oxygen,
etc. Consideration was therefore given to grinding methods, temperature exposure, solvents used for extraction and time in contact with solvent, solvent removal, and how materials would be stored when dry to achieve this goal. Following a period of methods development studies, which will be presented first, a generic processing protocol designed to achieve high throughput grinding and extraction with preservation of biological activity was developed, which is reported in detail here.
In grinding operations, care must always be taken to avoid plant dust cross-contamination of other specimens, as well as protecting the technician from possible irritants and other deleterious effects of plant dust. Gloves, lab coat or other protective clothing, earplugs, dust mask and safety glasses are worn. The grinding mill is placed inside a 4 foot deep chemical fume hood (
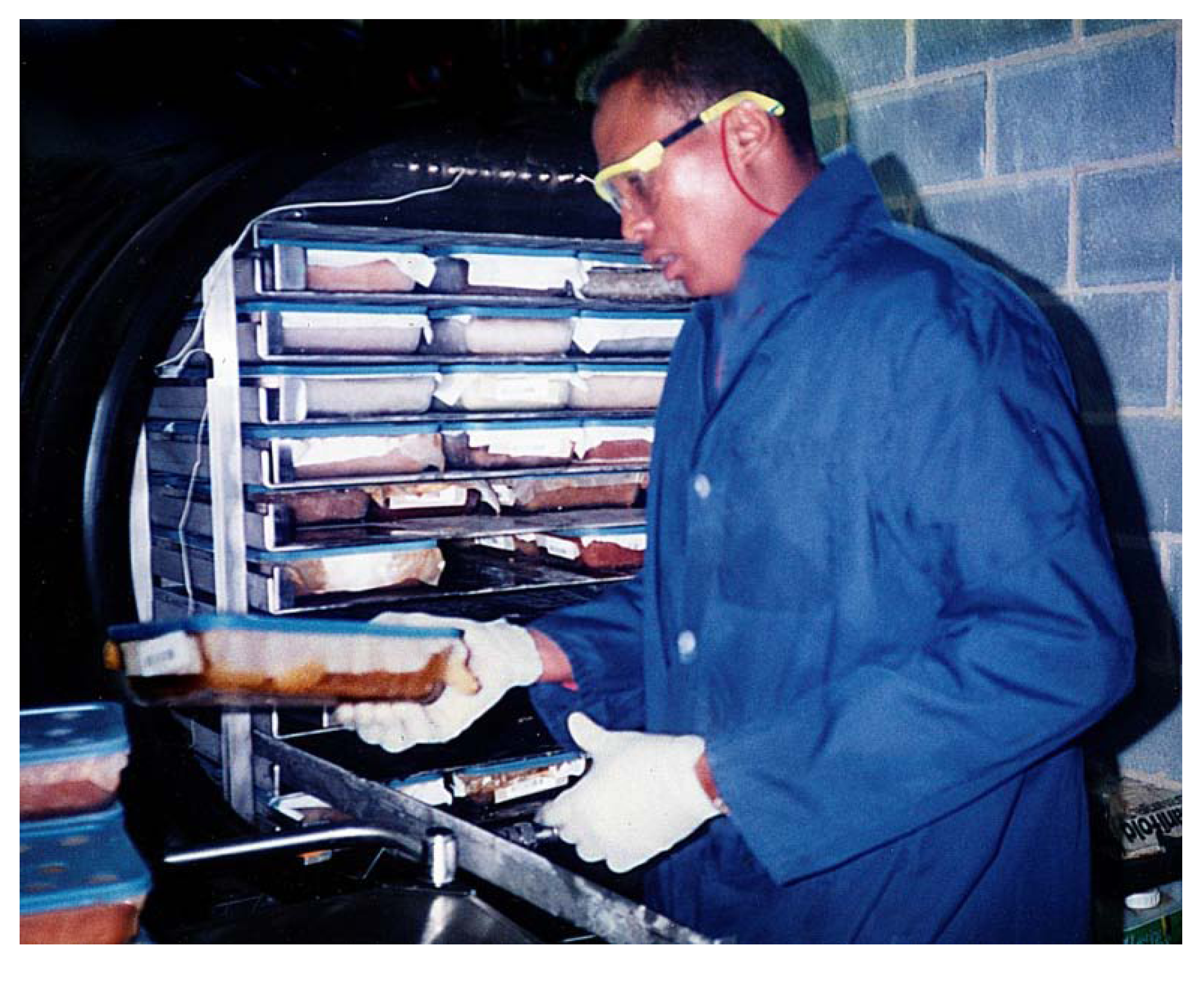
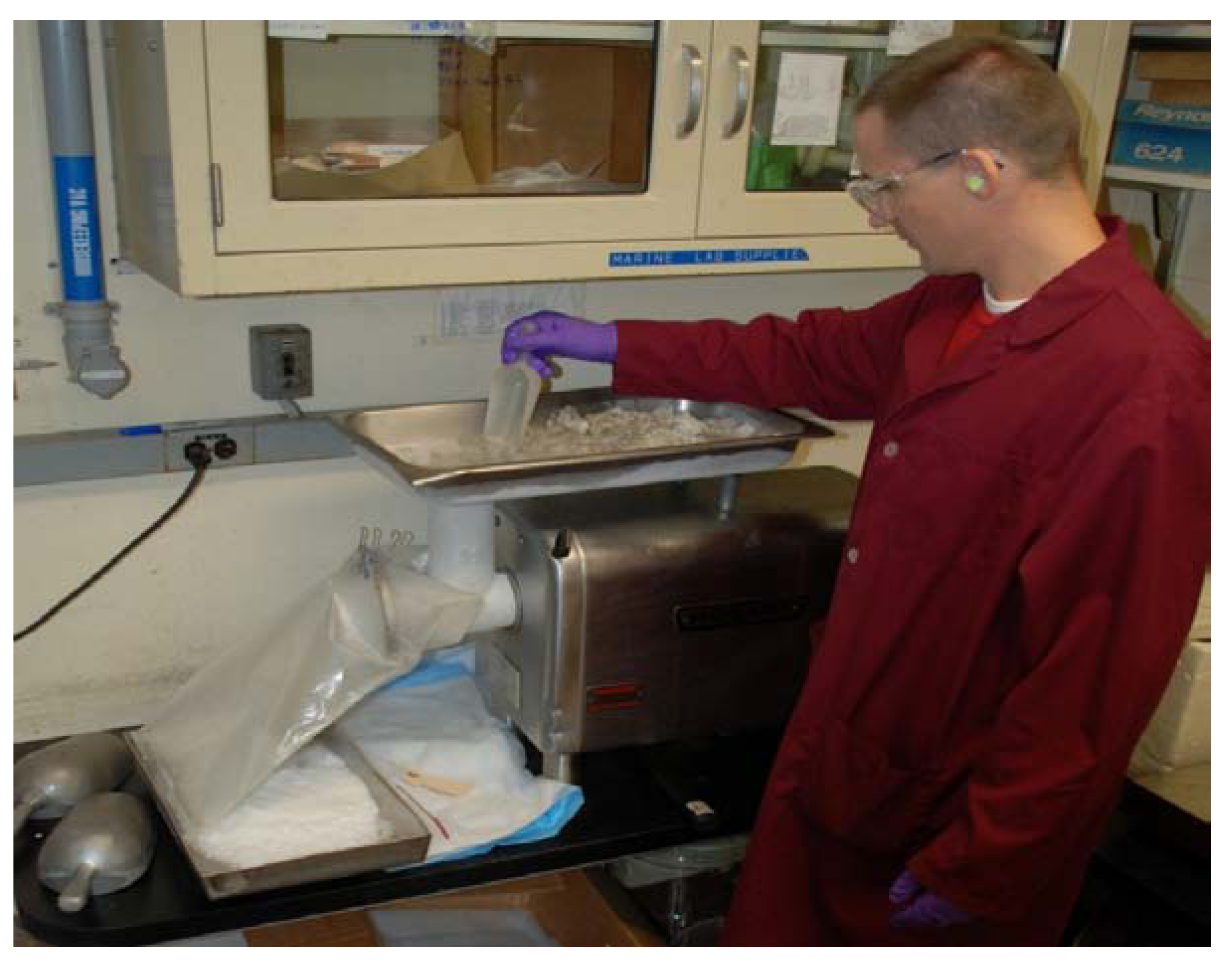
Figure 3) which has >1,000 linear feet per minute exhaust with the sashes pulled nearly closed, so that the dust which will be generated while grinding is taking place is vigorously pulled away from the technician and captured in a particulates filter. Specimen-contaminated materials including the bags which contained plants, contaminated protective equipment, floor sweepings, contents of vacuum cleaners and dirty particulate filters are incinerated on site.
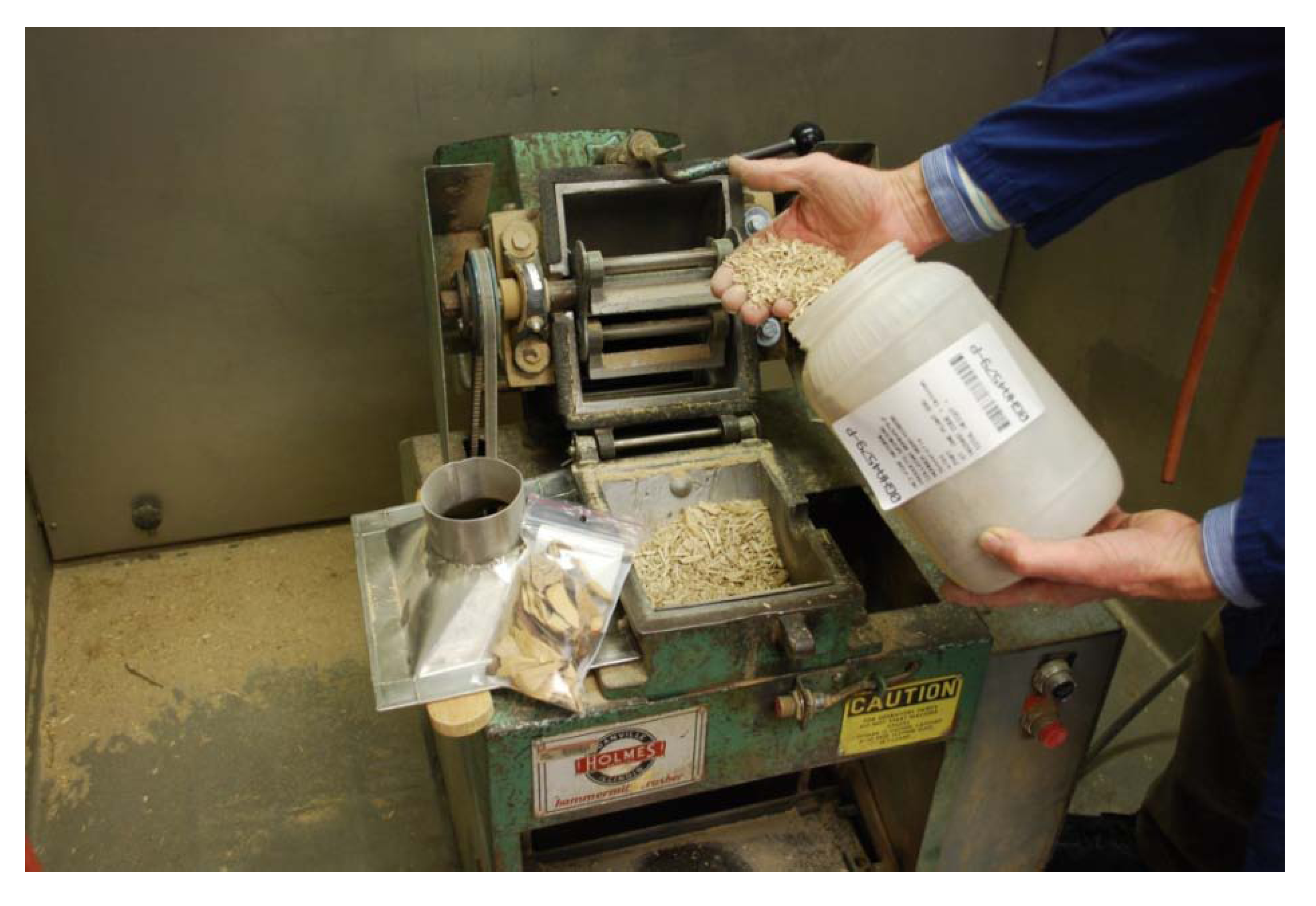
Figure 3.
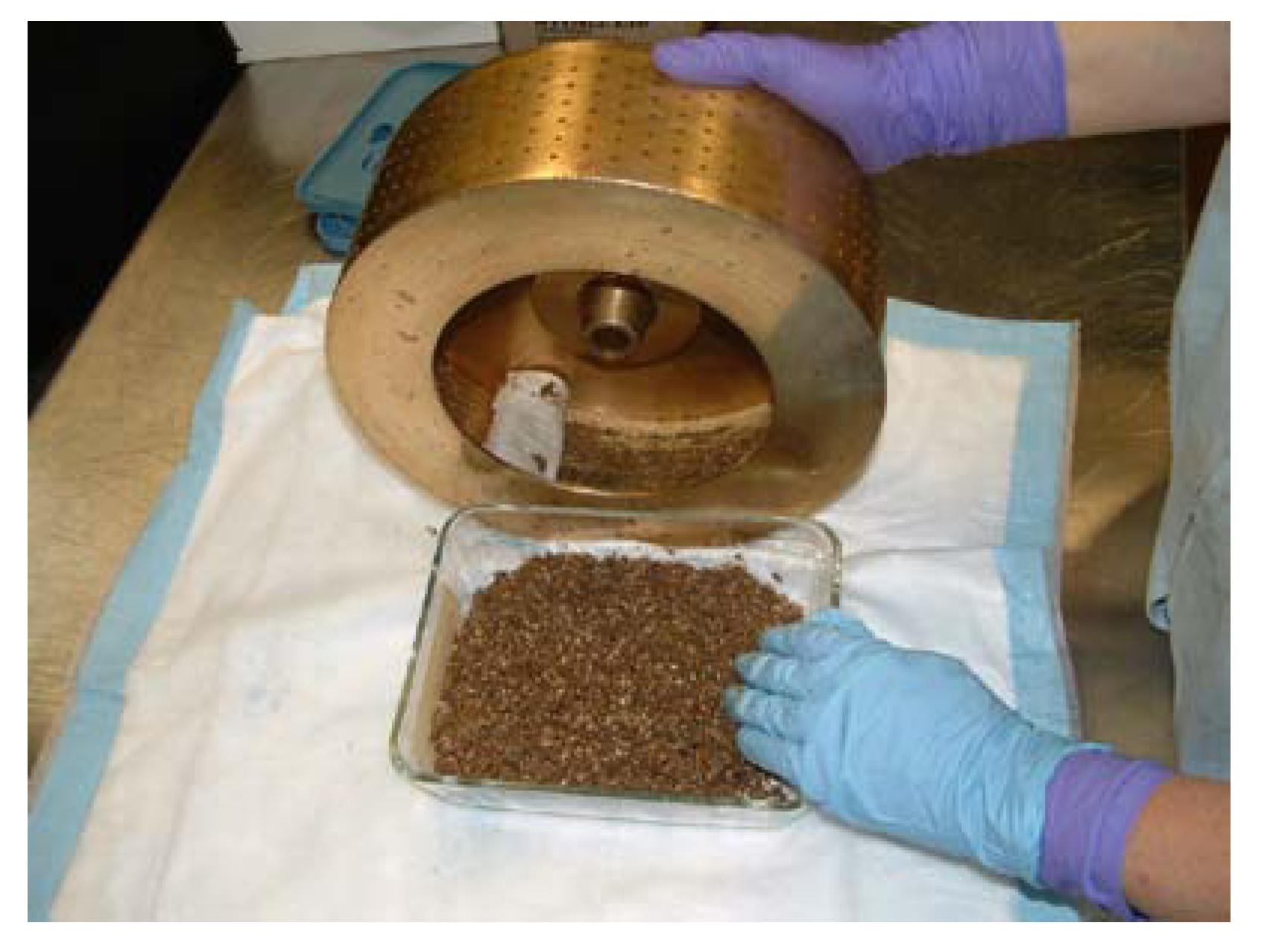
Dry Plant Grinding with a Holmes Hammermill. Note voucher, purpose-built stainless steel funnel, and barcode labeling.
Figure 3.
Dry Plant Grinding with a Holmes Hammermill. Note voucher, purpose-built stainless steel funnel, and barcode labeling.
Since most plants acquired for this screening program are imported, this required that the facility obtain a United States Department of Agriculture (USDA) foreign plant importation permit. Typically the USDA insists that dried, imported plants be fumigated: this cannot be done in a drug discovery situation as a toxic residue from pesticides would be present in the extracts which are produced, and will kill the cells used to test for anticancer activity. Therefore, sealed boxes of dried plants as prepared by the collectors are moved through the import stations and while still sealed, moved into storage in -20 ºC freezers at NCI-Frederick. Specimens are stored at -20 ºC often for months until processed. The un-opened cloth bags containing the dried plant specimens are typically be removed from freezers and lay on the bench in the grinding area for 1-2 days to come to room temperature prior to grinding. To assure that no foreign plant diseases or pests could escape into the local environment, this grinding facility has been inspected and approved by the USDA.
Exposure to heat is believed to be detrimental to preservation of biological activity. Heat is always generated in the grinding process. Milling dry plant specimens into extremely fine particles is detrimental in two respects: first - it takes extra time inside the mill, and as a consequence increases exposure to heat. And secondly, solvent extraction is a diffusion process, and is most effective and occurs most quickly when the particle size of the material being extracted is small. Yet if the particle size is extremely small, gravity flow of solvent through a glass column percolator is very slow, reducing the rate at which an extraction can be completed, clearly a negative factor to high throughput extraction. In order to achieve a high specimen grinding and extraction throughput, compromises must be made in grinding, particle size and completeness of extraction. Therefore, the protocol that was developed to achieve high throughput extraction is a compromise in which plant particle size distribution following grinding yields a powder with 80% of the specimen in the 0.5–2 mm particle size range. This compromise has benefit by limiting grinding time and exposure to heat, and in providing a material with good flow-through properties.
A number of different mills were tested for their ability to grind the wide variety of plant materials that exist, their ease of operation and cleaning, and to produce the desired particle size range. There are many types of grinding mills, (knife or Wiley-type, impact, pin,
etc.) which could be used to grind dried plants, but there is no single type which is ideal for all materials. Our experience has been that a hammer mill (model 201XL, Holmes Bros., Danville, IL, USA,
http://www.holmesbrosinc.com) has been the most generally useful for the sort of work described here, being capable of pulverizing 3-centimeter-cubes of the hardest heartwood, nut or root, in addition to powdering leaves, twigs, and bark. As is shown in the photograph, (
Figure 3), this mill is easily opened for cleaning. An infrequent drawback is that fibrous material will wrap around the rotor. A custom-fabricated stainless steel funnel (shown on the left side in
Figure 3) directs the comminuted plant directly into a 4,000 mL plastic wide-mouth bottle, thus saving a transfer step from the metal ‘bin’ that comes with the mill. The choice of a suitable grinding mill is also dependent on the size of the specimens being ground, with micro-mills unsuitable for kilogram specimens, and larger mills unsuitable for specimens of a few grams. An unacceptable amount of sample loss will occur when small specimens are ground in unsuitably large mills. Other mills, used when needed, include the Wiley knife mill, Fitzpatrick high speed impact mill, Hobart hamburger grinder and Glen Mills hammer-cutter type V with top feed. An example of the appropriate use of this last item was the grinding of ~1,400 individual leaf collections for quantification of an alkaloid [
5]. Mills are equipped with the largest motor available, so they can run longer without overheating, and pulverize larger pieces of plant without jamming.
Before grinding, the specimen is examined to determine if it is dry enough to be ground, if it is properly labeled, and if there is anything unusual about it (such as fungus growing on it) that requires special treatment. If drying is needed the specimen goes into a vacuum chamber without heating. Attention is given to any HAZARD information that might have been included by the collector, such as THORNS, IRRITANT or ALLERGEN, with appropriate precautions taken (i.e. leather gloves, sleeve protectors, respirator) when handling the specimen.
Only non-toxic lubricant such as cooking oil is used on mills to avoid contamination of a specimen with potentially toxic hydrocarbon grease. During grinding in any mill, care must be taken to avoid the introduction of other potentially contaminating materials such as paper (inks, glues, sulfites), cloth or non-plant foreign materials, which will later get into an extract.
Prior to grinding, the identifying number placed on the specimen by the collector is read into the Natural Products Support Group (NPSG) local area computer network, thus creating an ‘inventory’ of what is in this lab, and which documents the processing steps. This software, which is based on dBase III+, has been made available to NCI collaborators as ‘The NCI Field Taxonomy Program’, is user configurable, and can be run on a PC. Plant taxonomy, grinding and extraction steps and barcoding are supported in this software.
A small sample of each plant specimen is not ground, but is saved as a ‘grind voucher’, which is selected to contain all the different components seen in the specimen (i.e. leaf, twig, bark), even obvious ‘contaminants’ (i.e. moss growing on bark) in approximately the same ratio as is seen in the bulk of the specimen. These vouchers, have been useful for comparison with the conventional taxonomic voucher or for chemotaxonomy, and are stored at room temperature in the NCI-Frederick Natural Products Repository.
Reducing the size of plant pieces as provided by the collectors may be needed before they can easily be fed into the mill. This may include breaking twigs by hand, using a limb lopper to cut branches into smaller lengths, or use of a band saw to reduce hard woods and roots to sizes appropriate for the mill, typically under 3 cm. Cutting fibrous stems and bark across the fiber with the band saw is done to forestall a ‘ball of long fibers’ from accumulating around the armature of the hammer mill. The band saw is the most dangerous piece of apparatus in routine laboratory use, made more so by extremely hard and oddly-shaped roots which must be cut, and which sometimes contain embedded pebbles and even bits of metal.
The particle size obtained in grinding, and the residence time inside the hammer mill, is determined by the size of the holes in the outlet screen. Trials determined that for high throughput, minimum exposure to heating, and good rate of solvent flow-through during extraction, a 3/16" (4.7 mm hole diameter) mesh size installed in a Holmes 201XL hammer mill provided relatively small particles, most less than 2 mm, but not so fine that the flow rate of solvent through the sample, when packed in a columnar percolator, was unacceptably slow. A random sampling of plants and individual plant parts were ground with the 4.7 mm screen and other sizes. Then, utilizing a Tyler Industrial Products model RX24 portable shaker, the ground plant sample was sieved through a series of mesh sizes to obtain information about particle size distribution. Following vigorous agitation, the contents of each sieve was weighed.
Table 1 shows an example representative of results obtained with a 4.7 mm screen, expressed as the percent falling into each size range.
Table 1.
Particle size distribution for nine plant samples, expressed as a percentage.
Table 1.
Particle size distribution for nine plant samples, expressed as a percentage.
| Plant Part | Size (mm) |
| 10 – 2.8 | 2.8 - 1 | 1 – 0.5 | 0.5 – 0.18 | < 0.18 |
| leaf/stem | 11 | 57 | 21 | 7 | 2 |
| root | 12 | 38 | 24 | 14 | 9 |
| root | 44 | 30 | 13 | 6 | 4 |
| bark | 24 | 39 | 20 | 10 | 6 |
| bark | 28 | 37 | 15 | 14 | 6 |
| wood | 38 | 46 | 8 | 5 | 2 |
| wood | 50 | 30 | 9 | 7 | 3 |
| fruit | 43 | 20 | 16 | 12 | 8 |
| fruit | 30 | 37 | 17 | 12 | 5 |
These data document that when the 4.7 mm screen is used in the hammer mill, ~70% of the resultant ground product is smaller than 2.8 mm diameter, while an excessive quantity of fines <0.18 mm is not produced. Grinding time for a typical 1 kg plant specimen is 5 minutes, with roughly twice that spent in cleaning the mill prior to the next specimen. Between each grinding, the mill is opened, cleaned first with a vacuum cleaner, followed by a cloth towel moistened with ethanol or isopropanol, and finally with a blast of compressed air to remove any residual fine dust or solvent. An occasional specimen will leave a ‘waxy’ residue in the mill, which is most easily removed by passing corn cobs through the operating grinder. The ground weight of each specimen is recorded to the database. This weight is important to know so that the percent yield of any molecule of interest that is obtained from this specimen can be calculated. Should gram quantities of a biologically active molecule be required in the future for more comprehensive biological evaluation, it is important to be able to inform the botanist whether recollection of one or one hundred kilos of the plant is needed to obtain the required quantity of compound. Yield of a compound from its source should always be given in any publication, and recording a ground weight, which in our situation is also the weight extracted, allows this calculation to be made.
7. Miscellaneous Extractions and Processes
Several hundreds of specimens do not neatly fit into any of the three above delineated major categories. Dried lichens have been extracted as if they are dry plants. Mature fungal basidiocarps, collected fresh and delivered wet frozen to the lab, have sometimes been freeze dried, then extracted with organic solvent much like a dried plant. Alternatively, these high water content specimens can be placed into a large spark-free Waring blender with 75% MeOH/25% DCM and thoroughly minced. The organic solvent extract is clarified by passing through filter paper, and the marc re-extracted in the same way with water. Extracts were made of several Caenorhabditis elegans strains by pelleting the nematodes in a centrifuge tube then homogenizing and adding organic solvent to create a two phase system. “Traditional medicines” of various sorts, often poorly-described mixtures of components, are frequently sent to the NCI, and forwarded to this program, by private individuals as well as organizations doing research on Oriental and African traditional medicines, often with a protocol for preparation and administration. These submissions have generally been split into two parts with half being processed as faithfully to the instructions provided as possible, while the other half is processed by NPSG standard methods. In these cases, four or more extracts are submitted to anticancer testing per specimen, with testing results provided to the submitter. Semi-hardened plant saps and exudates are not amenable to standard processes, and have essentially no water soluble components (excepting simple sugars) in them, but sometimes an organic solvent extract can be made by mechanical stirring in an appropriate organic solvent, often methyl t-butyl ether being the best. There continue to be new challenges even after 20+ years, and improvisation, based on experience, is frequently required.
During the mid-1990’s when extraction of specimens was a high priority of the NCI, seven technicians working in ~3,800 sq. ft. of laboratory space were routinely producing > 10,000 crude natural products extracts each year for biological evaluation.
Removal of dried organic solvent extract from lab glassware, even utilizing hot detergent water under pressure, has always been problematic. After each use, laboratory glassware is passed through an industrial glassware washer which begins its cycle with an alkaline detergent, then an acid detergent, then a rinse and a de-ionized water rinse. A universally satisfactory detergent has not been found, so inspection and hand-washing of glassware is a continuing requirement. Long-term use of a single detergent was observed to lead to a residue in the washer and presumably on glassware as well, so the best solution to the problem is to change brands and types of detergent every few months. Each detergent has seemed to remove the residue left by its predecessor. There have been instances of stainless steel and borosilicate ware being permanently stained by substances present in an extract. Borosilicate dishes used to freeze dry aqueous extracts were treated with a silanizing solution, which makes a noticeable difference in the ease of which dried extract can be scraped from the dish.
With >230,000 extracts produced there has not been one totally lost because of a spill, breakage or accident (
Figure 12). About a dozen 3L flasks with flattened bottoms have imploded while rotovaping, and in all cases the plastic safety coating contained both the glass and the extract. There have been several instances when biological test and retest did not correspond, so an examination of extract in comparison to grind voucher content was done by HPLC fingerprinting. In general, three extracts prior to and three past the extract in question were examined. There has been no occasion in which the composition of the extract differed significantly from the composition of the voucher. In laboratory operations as labor intensive as grinding and extraction are, there have unquestionably been some mistakes, but very few. Tracking through barcoding has helped greatly in minimizing mistakes through transcriptional error or mixing, keeping the identity of a specimen or extract at each step during processing beyond question.
The size of the specimens processed for this program has been relatively larger than those processed in most screening operations. The philosophy of this program from the start is that the initial specimen is large enough so that sufficient extract will be obtained to allow for the active substance to be isolated and its structure determined without a recollection of the source material. This has almost always been possible. The extraction protocols presented above were designed to be generic, high throughput, rapid, and to preserve those biologically active substances that may be contained, but these methods may not produce the highest yield of some substances. Once the structure of a compound of interest has been determined, customized extraction and processing methods have often been developed to optimize its yield. The same extraction laboratory personnel and equipment are used to perform up-scaled grinding, extraction by modified methods and large column chromatography (i.e. several kg of silica) to produce a semi-purified extract for delivery to the isolations chemist.
Figure 12.
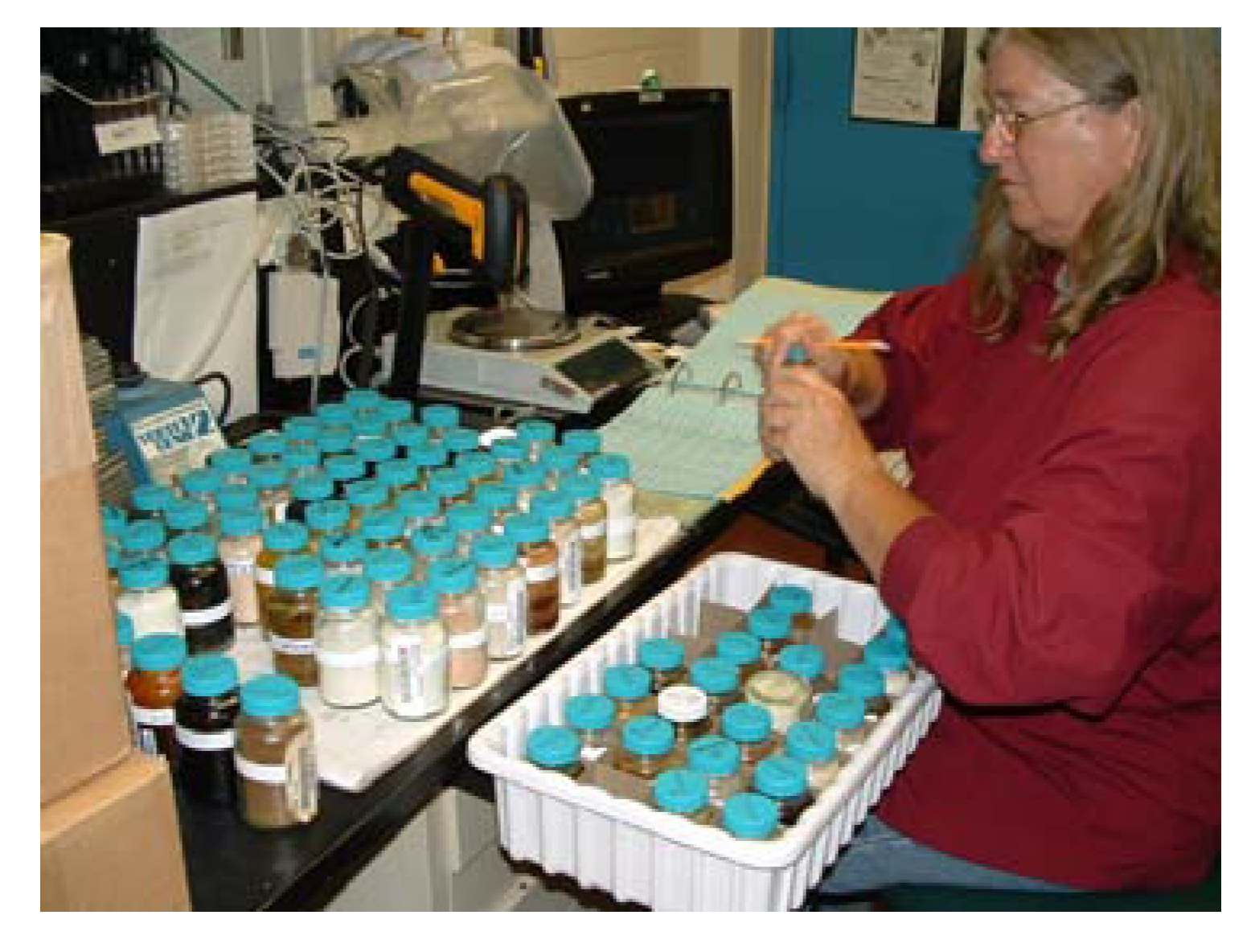
Recording weights of ‘bulk bottles’ containing vacuum dried extracts. Note multiple labels, non-contact laser barcode reader, and pan balance connected to computer work station.
Figure 12.
Recording weights of ‘bulk bottles’ containing vacuum dried extracts. Note multiple labels, non-contact laser barcode reader, and pan balance connected to computer work station.
9. Bioactive Compounds from the NCI Natural Products Screening Library: “The Proof’s in the Pudding”
The creation of a large and diverse screening library is not an end in itself: the discovery of new, useful pharmaceuticals is the goal. At NCI-Frederick the primary anticancer screen is the 60 human tumor cell line panel, (
http://dtp.nci.nih.gov/branches/btb/ivclsp.html), which detects cytotoxic and growth-inhibitory substances but additionally, provides information on selective cytotoxicity that, with the aid of the COMputerized PAttern REcognition algorithm (COMPARE), (
http://dtp.nci.nih.gov/timeline/noflash/milestones/M7_COMPARE.htm), can be useful in predicting a mechanism of action for a compound, and in prioritizing those extracts for fractionation which exhibit a particularly unusual cytotoxicity pattern, perhaps indicative of a previously unexploited anticancer target. Nearly all of the 230,000 extracts produced in this program have been through this preliminary tissue-culture-based anticancer screening. A frequently asked, but ultimately meaningless, question has been: ‘What is your hit rate?’ The answer is entirely dependent on how you define a ‘hit’ and will be different for each individual assay. There are commonly interfering substances present in crude extracts, but these do not adversely affect all bioassays. Some bioassays are sensitive to tannins, others to lipids, others to highly colored substances. When 634 preferentially selected Euphorbiaceae crude organic solvent extracts were screened for PDBu displacement activity, 153 exhibited this property, so the hit rate was 24%, but from a very select ‘feed’ into the bioassay [
14]. After >140,000 extracts had been screened for toxicity toward azole resistant
Candida albicans, a ‘hit list’ of only 140 extracts was selected for dereplication, a 0.1% hit rate. But to make it onto the hit list, the extract had to show significantly lower toxicity toward bacteria and a human cell line being screened concomitantly. Thus, the emphasis is on selective toxicity toward fungi [
21]. Simple toxicity in the 60 cell primary anticancer screen is not sufficient to make an extract interesting. From a subset of >19,000 crude plants extracts tested at 5 × 1 log doses in the 60 cell screen, 452 showed LC50’s below 50 µg/mL. But a highly toxic compound does not necessarily make a good clinically-useful chemotherapeutic. Only a few hundred extracts have shown patterns in 60 cell screening indicative of selective toxicity toward a tissue type. Fewer still have yielded novel bioactive molecules.
Even the finding of the same or similar compounds can be useful, for example in identifying an abundant source of a previously scarce substance, in establishing chemotaxonomy, or in the identification of new homologues of a bioactive molecule helpful in the pursuit of Structure Activity Relationship (SAR) information. Examples from each of these categories follow:
Jaspamide, (jasplakinolide, CAS 219774-75-1) was such a scarce resource, obtainable in only mg amounts following tedious purification from a sponge. The compound could not be fully investigated as an anticancer lead drug for lack of a supply. The NPSG lab was requested to explore our existing NCI-Frederick Natural Products Repository as a source. Approx. 85 sponge extracts obtained from
Jaspis,
Auletta,
Dorypleres, and other genera from a broad region of the South Pacific where jaspamide had been reported were examined by HPLC. Jaspamide was detectable in a number of extracts which had been obtained from New Guinea, Fiji, the Philippines, and elsewhere, but a single collection identified as a
Dorypleres sp. which had come from Palau, was very rich in the desired molecule. From the 845 gms of organic solvent soluble extract, 1.3 gms of jaspamide was purified for NCI research [
9].
Anticancer screening identified the known fungal-produced compound brefeldin (CAS 20350-15-6) as of interest. Brefeldin A has, of course, been detected from many sources, not all of them fungal [
10,
11,
12]. Fermentation of a number of fungal strains lead to a
Eupenicillium brefeldianum culture as a moderate producer
, which was adapted for large-scale liquid fermentation. At the NCI-Frederick fermentation facility, four fermentations at the 12,000 L scale, with purification of the compound by the NPSG, resulted in the production of 4 kg of crystalline brefeldin A for NCI research [
13].
Camptothecin (CAS 7689-03-4), a natural product originally isolated from the Asian tree
Camptotheca acuminata [
14], which has been semisynthetically modified into the clinically used anticancer drug topotecan (CAS 123948-87-8), shows bioactivity in many test systems, including a high throughput screen to detect inhibitors of the promising anticancer target HIF-1α. In screening extracts for inhibitors of HIF-1α, an extract obtained from an
Ophiorrhiza trichocarpon specimen was identified as of interest. Upon dereplication, the extract was found to contain not only camptothecin but additionally, 9-methoxy-, 10-methoxy- and 9,10-methylenedioxy camptothecins [
15]. This genus had not previously been reported to contain camptothecin.
A large crude extract library can have value in chemotaxonomic investigations. The marine natural product bryostatin 1 (CAS 83314-01-6), obtained from the bryozoan
Bugula neritina, is of interest as an anticancer compound having activity through interaction with the diacylglycerol binding site on the cell membrane, the same binding site affected by the irritant and co-carcinogenic phorbol esters which are present in some plants. Bryostatin is present in the marine organism at only a few thousandths of a percent by weight, so tracking its presence during extraction and initial purification via a sensitive bioassay was preferable to chromatographic analysis. At this facility a project to isolate a multi-gram quantity of bryostatin A for research use was initiated [
16], and a bioassay measuring displacement of radioactive phorbol dibutyrate (PDBu, CAS 37558-16-0) from rat brain homogenate was developed. Though some members of the plant family Euphorbiaceae are known to produce phorbol esters, there had been no chemotaxonomic survey of the family to determine how widely spread these substances were. To demonstrate the robustness of the PDBu displacement assay when applied to a chemically complex crude fraction, several hundred organic solvent extracts from the Euphorbiaceae, obtained from the NCI-Frederick repository, were run in the displacement assay. When PDBu displacement results were organized according to traditional taxonomy of the Euphorbiaceae, the correspondence was found to be quite close [
17,
18,
19], adding to our knowledge of the distribution of these potentially hazardous natural substances in the environment, and to the chemotaxonomy of the family Euphorbiaceae. Additional investigation would be required to determine the chemotype responsible for the positive PDBu displacement result, or whether a displacement result was produced by a non-phorbol molecule. And at the conclusion of the bryostatin scale-up project, several grams of highly purified bryostatin 1 were produced for drug development studies.
Serendipitous discoveries are made only because both HTS and a large natural products screening library are available. An example of this came with the observation of an anomalous response in the HTS screen designed to detect compounds which inhibit the HIF-1α protein. An extract demonstrating HIF-1α inhibition would give a decrease in signal from the luciferase reporter engineered into the U251-HRE human glioma cell line, but a small subset of extracts gave a great increase in signal. Upon database mining, this small group of ‘enhancer’ extracts was found to have come from Rubiaceae which had been collected in east Africa and Madagascar. Dereplication of several extracts revealed a common constituent: a known substance, FK228, which had only been reported previously as bacterially-produced [
20]. FK228, or FR901228, also known as romidepsin, is a potent histone deacetylase inhibitor (HDAC). Thus the unexpected response was attributable to hyperacetylation, leading to over-expression of the luciferase reporter. By HPLC/MS, a survey of ‘enhancer’ extracts showed the presence of FK228 in all those Rubiaceae extracts giving the high luciferase signal [
21].
Some particularly interesting stories have developed from the screening program, one of which involved the discovery of a new compound, a new chemotype and description of a new species of plant. A routine specimen of leaves received from Cameroon, West Africa, was identified as
Ancistrocladus abbreviatus. It was ground, extracted, and the extract made a part of the screening library. In HTS for substances active against the Human Immunodeficiency Virus (HIV), the virus responsible for causing Acquired Immunodeficiency Syndrome (AIDS) in humans, this
Ancistrocladus extract inhibited cell death of HIV-infected human CEM-SS cells. The compound responsible for this desirable biological response was identified and found to be of a new chemotype of alkaloid, an unusual ‘dimer of dimers’, and given the trivial name of michellamine B, (CAS 137793-81-8), [
22], but several subsequent recollections of
A. abbreviatus obtained from Cameroon failed to contain the same compound. Upon detailed examination of the genus
Ancistrocladus by plant taxonomists, it was determined that the biologically active molecule was contained in a new species, subsequently described [
23]. This rare species of the genus
Ancistrocladus is localized to a small region of western Cameroon and eastern Nigeria, a significant portion of its range protected in Korup National Park - thus the new species name it received:
A. korupensis. It is probable, even likely, that this species would never have been described, nor the unusual alkaloid discovered, except for a random collection of the plant and screening of its extract for anti-HIV activity. The NPSG ground and extracted several hundred kg of
Ancistrocladus korupensis leaves for large scale isolation of michellamine B. But because unacceptable toxicity was observed when michellamine B was administered to dogs, no further development was done.
A collection of
Calophyllum lanigerum from the Malaysian State of Sarawak, likewise showed activity in the anti-HIV screen, with the bioactive new substance identified and named calanolide A, (CAS 142632-32-4), [
24]. When collectors went back to the site to obtain additional plant material, the original source trees had disappeared, a sad example of loss of tropical biodiversity. A chemo-taxonomic survey of a great many trees from the surrounding area failed to find another containing calanolide A. Subsequently, a structurally related molecule having similar biological activity, costatolide, (CAS 63023-58-5) was found which is obtainable in quantity from the latex of a
Calophyllum spp., [
25]. The NPSG developed a procedure for extraction and purification of costatolide which lead to several hundred grams being produced for research. These discoveries contributed to moves by the Sarawak Government to promote conservation measures and the establishment of the Sarawak Biodiversity Center (
http://www.sbc.org.my/) which is now conducting in-state investigation of the State’s valuable and diverse biological resources.
HTS in a whole-organism screen designed to uncover compounds useful in the treatment of drug-resistant microbial infections identified many extracts toxic to the test microorganisms, of which two examples deserve mention in this context. An extract from
Aniba panurensis collected in Guyana showed potent and selective killing of an azole-resistant strain of
Candida albicans. Following an activity-directed isolation procedure, the toxic substance was isolated, and its structure determined to be a new indolizinium alkaloid (CAS 725265-00-9), [
26]. Unfortunately, this molecule does not possess drug-like properties and has low water solubility, so despite its anti-Candida activity, there was no interest in development of this substance as a new drug.
The same HTS identified an extract from fermentation of an at-that-time unidentified microorganism, later to be determined as a strain of
Aspergillus flavus, as highly cytotoxic to azole-resistant
C. albicans. Following activity-guided fractionation, the most fungitoxic substance in the extract was identified as an already known molecule, aspirochlorine (CAS 59978-04-0), but additionally in this extract were trace amounts of two previously unknown aspirochlorine homologues wherein the sulfur bridge contained three or four sulfur atoms, [
27].
During the late 1960’s and early 1970’s the DTP supported an anticancer drug discovery program looking for new chemotherapeutic agents in plants. A success from this previous screening program was discovery of the highly effective anticancer drug taxol [
28]. Remaining from this program were approximately 6,000 plant collections that had not been extracted. These were ground and extracted by the new protocol given here, and these extracts added into the present screening library. It has not been an infrequent question and implied criticism: “What do you expect to find in 20 year old plants?” In addition to detection of known compounds with new screens, and the resupply of scarce known compounds for research use, new bioactive substances continue to be detected even from old plant specimens. An example is found from an extract prepared from a
Crossosoma bigelovii specimen collected in Mexico in 1990, and during the intervening years stored dry but not in a temperature-controlled location, which showed activity in an HTS HIF-1α inhibition assay. Activity directed isolation lead to the identification of a new strophanthidin glycoside as being the molecule responsible for HIF-1α inhibitory activity [
29].
More importantly than the discovery and re-discovery of bioactive compounds in organic solvent extracts is the exploration of water extracts for new anticancer drugs. Twenty years ago the chromatographic techniques available for the purification of water soluble substances were quite limited, and the spectroscopic equipment and techniques available for solving the larger molecular structures found in water extracts were primitive at best, so most drug discovery organizations made no effort at looking for new drugs in water extracts. Over the intervening years new chromatographic supports (such as C18 phase bonded materials suitable for use in 100% water, hydrophilic interaction columns, etc.) applicable to resolution of water soluble compounds and new analytical equipment and methods (countercurrent and pH electrophoresis, TOF mass spectrometry, automated Edman degradation sequencing or sequencing by MS, etc.) have been developed, overcoming many of the previous deficiencies, and making the search for new drug substances contained in highly polar water soluble mixtures possible.
With twenty years of experience we can now say that quite a wide range of biologically-active substances have been identified in water extracts. Those chemotypes that would be anticipated, such as sulfated sterols in marine invertebrate extracts,
i.e. the anti-HIV substance ibisterol sulfate (CAS 148101-50-2), [
31], and relatively small macrolide lactones,
i.e. a compound discovered as a result of anticancer activity in the NCI 60 cell line screen: lobatamide, (CAS 200563-47-9), [
32] have been found. But most prominent among the compounds discovered from water-soluble extracts are cyanovirin, [
33] produced by a blue-green alga, and griffithsin, [
34] produced by a red alga. Both are small proteins, 101 residues for cyanovirin, and 121 for griffithsin, and both are currently being pursued as potential new anti-HIV chemotherapeutics with different modes of action.
An aqueous extract from the sponge
Pipestella candelabra sp. nov. displayed a cytotoxicity pattern in the 60 cell primary screen which resulted in its selection for testing in the
in vivo hollow fiber mouse anticancer model, [
35]. Interesting
in vivo biological activity was detected, which resulted in dereplication of this extract. Activity directed isolation lead to the discovery that this sponge extract contains three known bioactive substances, jaspamide (jasplakinolide), hemiasterlin, and brefeldin A, none of which had displayed a comparable 60 cell activity pattern when tested as pure compounds, but in this combination, showed an as-yet unexplained synergy [
36].