Hyper-Null Models and Their Applications
Abstract
:1. Introduction
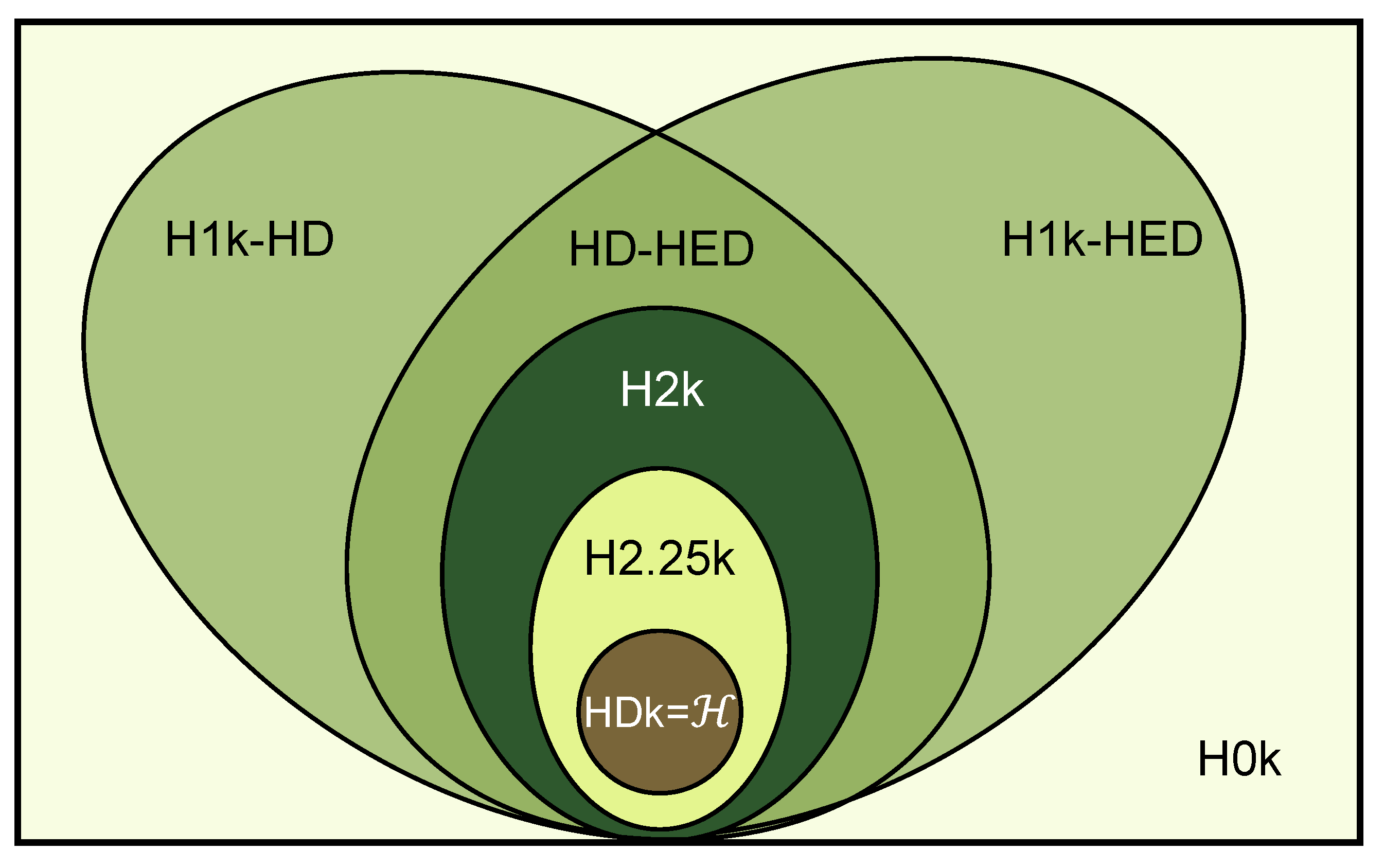
- We propose a new method to construct null models for hypergraphs and summarize the relationship between our proposed hypergraph models.
- We introduce the concept of entropy to quantify the randomness of networks, retaining different statistical properties, and explore the relationship between randomness and the network structure.
- We utilize topological statistics analysis, network dismantling, and epidemic contagion to showcase the universality of the framework employed in the original network and its hyper-null models.
2. Methods
2.1. Hypergraphs
2.2. Hyperedges and Hypertriangles
2.3. Statistics of Hypergraphs
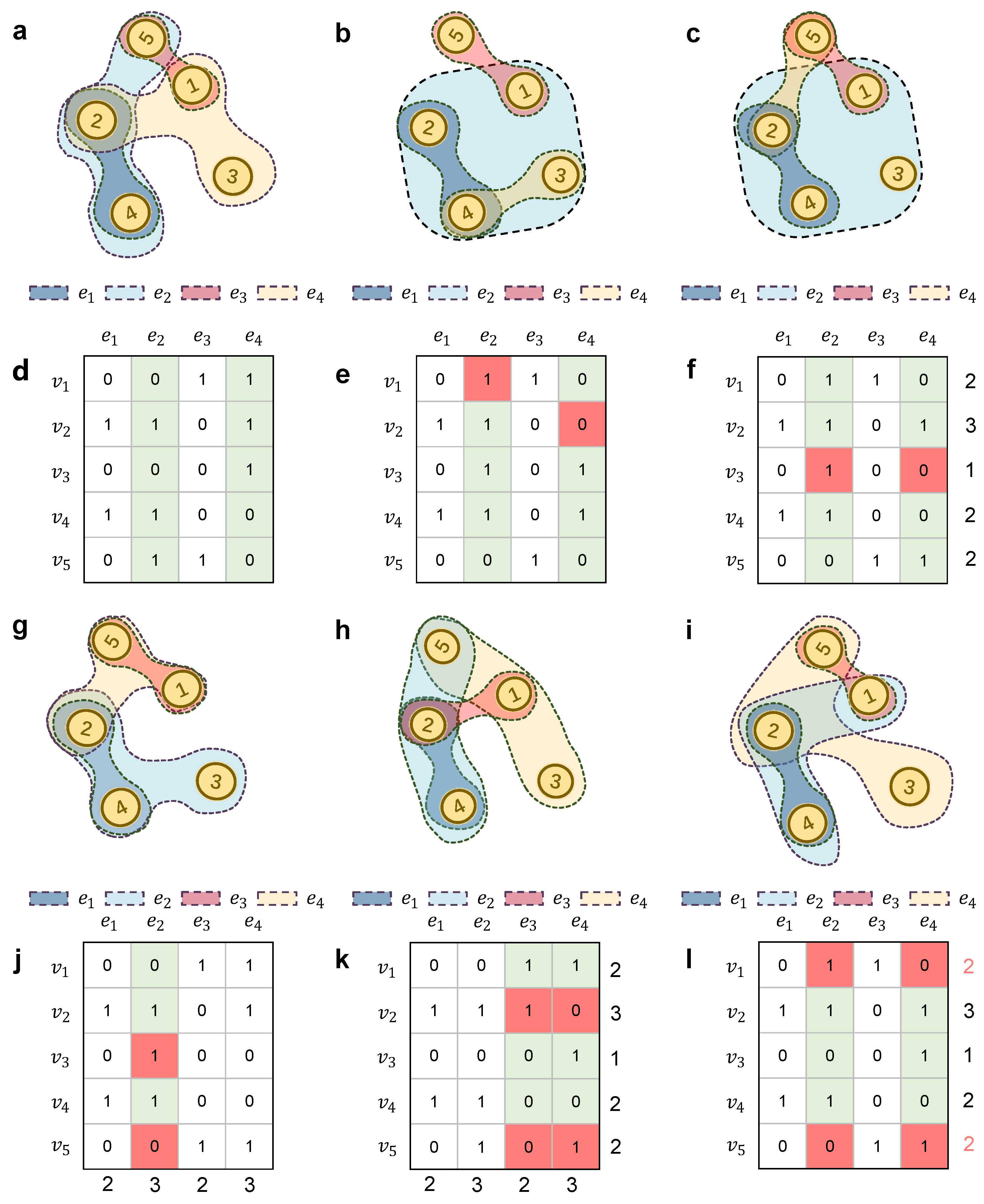
2.4. Matrices of Hypergraphs
2.4.1. Hyperdegree Matrices
2.4.2. Hyperedge degree Matrices
2.4.3. Incidence Matrices
2.4.4. Adjacency Matrices
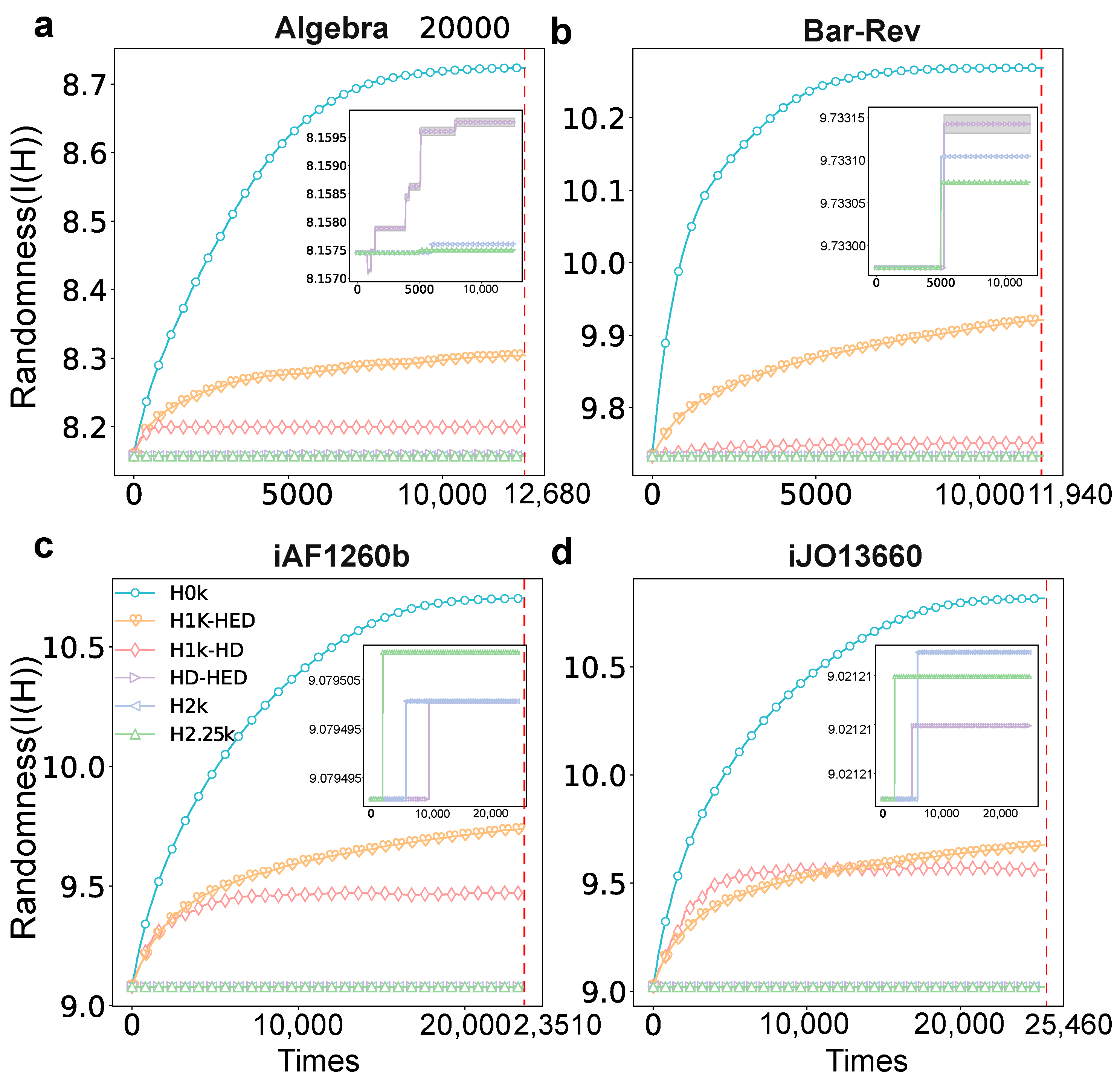
2.5. Randomness
2.6. Hyper-Null Models Based on Hyperedge Swapping
2.6.1. Hyper-0k Null Model
2.6.2. Hyper-1k Null Model with Hyperdegree Constant
2.6.3. Hyper-1k Null Model with Hyperedge Degree Constant
2.6.4. Hyperdegree–Hyperedge Degree Null Model
2.6.5. Hyper-2k Null Model
2.6.6. Hyper-2.25k Null Model
3. Results
3.1. Data Description
- Algebra: A question–answer network, which is collected from MathOverflow.net, where the nodes denote users, and the users who answered the same question are enclosed in a hyperedge.
- Bars-Rev: A review hypergraph collected from Yelp.com, where a hyperedge consists of the users who reviewed the same bars.
- iAF1260b and iJO1366: The metabolic hypergraph where nodes denote metabolites and the hyperedges represent the reaction that is involved in the same metabolic.
3.2. Statistical Analysis Based on Hyper-Null Models
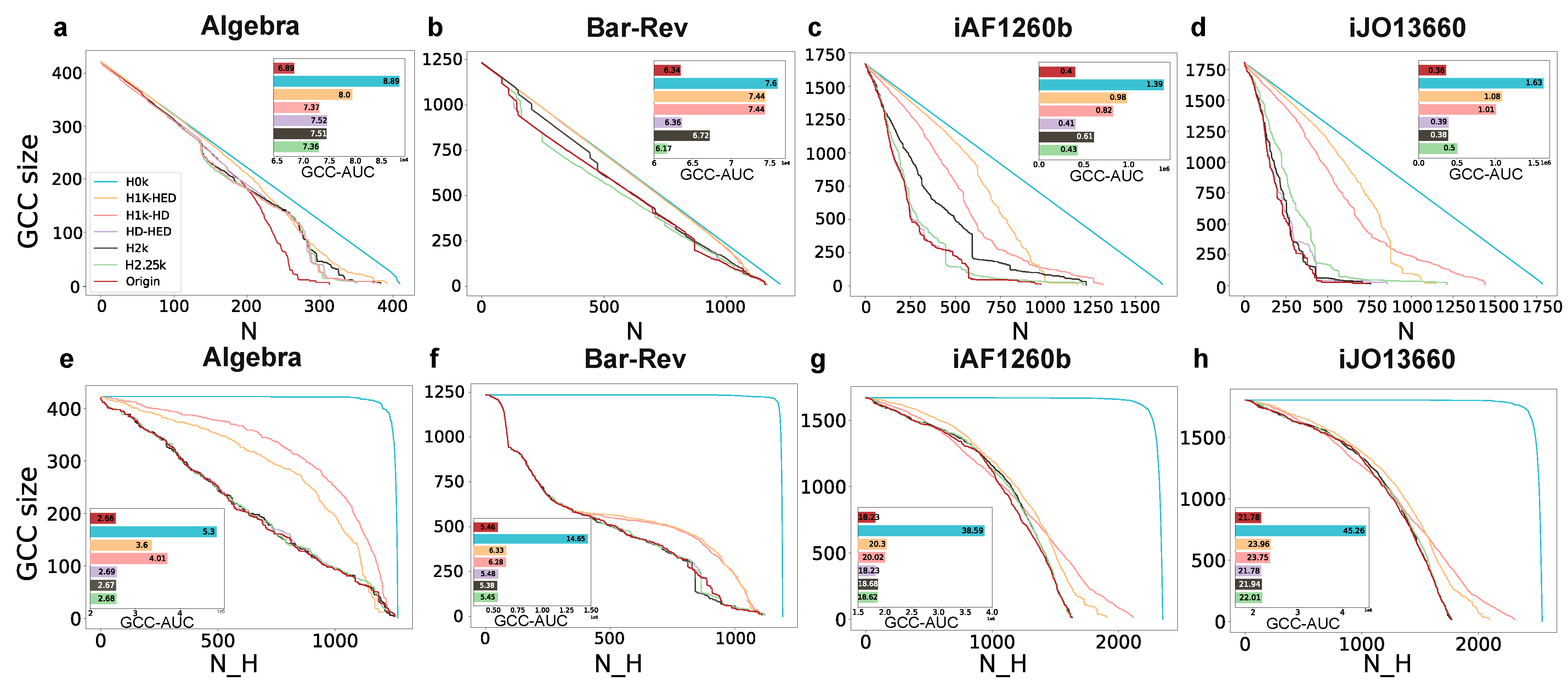
3.3. Hypergraph Dismantling
3.3.1. Hypergraph Dismantling by Removing Nodes
3.3.2. Hypergraph Dismantling by Removing Hyperedges
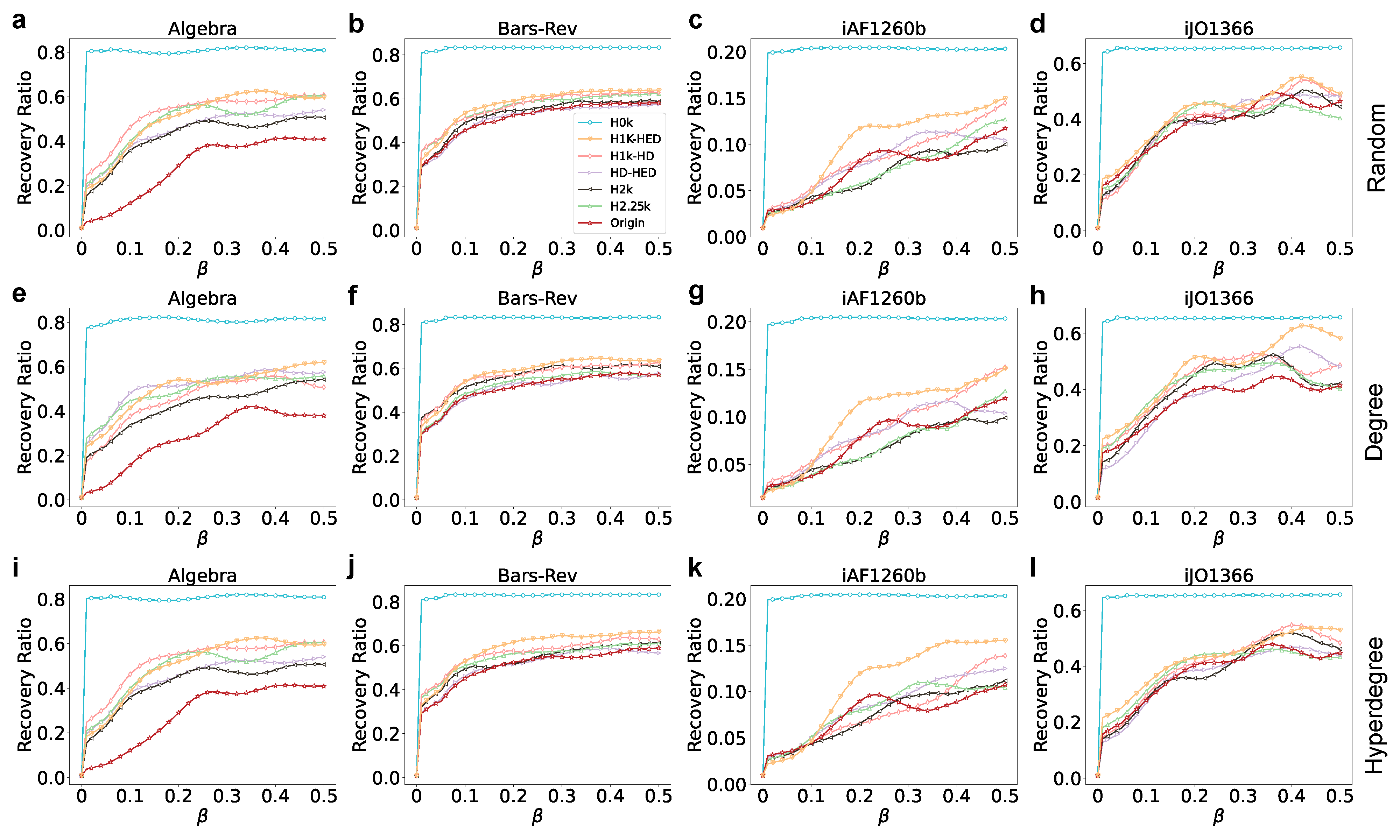
3.4. Hypergraph Epidemic Contagion
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Conflicts of Interest
References
- Barabási, A.L. Network Science; Cambridge University Press: Cambridge, UK, 2016. [Google Scholar]
- Newman, M. Networks; Oxford University Press: Oxford, UK, 2018. [Google Scholar]
- Majhi, S.; Perc, M.; Ghosh, D. Dynamics on higher-order networks: A review. J. R. Soc. Interface 2022, 19, 20220043. [Google Scholar] [CrossRef] [PubMed]
- Battiston, F.; Cencetti, G.; Iacopini, I.; Latora, V.; Lucas, M.; Patania, A.; Young, J.G.; Petri, G. Networks beyond pairwise interactions: Structure and dynamics. Phys. Rep. 2020, 874, 1–92. [Google Scholar]
- Battiston, F.; Amico, E.; Barrat, A.; Bianconi, G.; Ferraz de Arruda, G.; Franceschiello, B.; Iacopini, I.; Kéfi, S.; Latora, V.; Moreno, Y. The physics of higher-order interactions in complex systems. Nat. Phys. 2021, 17, 1093–1098. [Google Scholar] [CrossRef]
- Zeng, Y.; Huang, Y.; Ren, X.L.; Lü, L. Identifying vital nodes through augmented random walks on higher-order networks. arXiv 2023, arXiv:2305.06898. [Google Scholar]
- Bianconi, G. Higher-Order Networks; Cambridge University Press: Cambridge, UK, 2021. [Google Scholar]
- Berge, C. Hypergraphs: Combinatorics of Finite Sets; Elsevier: Amsterdam, The Netherlands, 1984; Volume 45. [Google Scholar]
- LaRock, T.; Lambiotte, R. Encapsulation Structure and Dynamics in Hypergraphs. arXiv 2023, arXiv:2307.04613. [Google Scholar]
- Feng, Y.; You, H.; Zhang, Z.; Ji, R.; Gao, Y. Hypergraph neural networks. In Proceedings of the AAAI Conference on Artificial Intelligence, Hilton, HI, USA, 27 January–1 February 2019; Volume 33, pp. 3558–3565. [Google Scholar]
- Liao, X.; Xu, Y.; Ling, H. Hypergraph neural networks for hypergraph matching. In Proceedings of the IEEE/CVF International Conference on Computer Vision, Montreal, BC, Canada, 11–17 October 2021; pp. 1266–1275. [Google Scholar]
- Milo, R.; Shen-Orr, S.; Itzkovitz, S.; Kashtan, N.; Chklovskii, D.; Alon, U. Network motifs: Simple building blocks of complex networks. Science 2002, 298, 824–827. [Google Scholar] [CrossRef]
- Milo, R.; Itzkovitz, S.; Kashtan, N.; Levitt, R.; Shen-Orr, S.; Ayzenshtat, I.; Sheffer, M.; Alon, U. Superfamilies of evolved and designed networks. Science 2004, 303, 1538–1542. [Google Scholar] [CrossRef] [PubMed]
- Liu, B.; Xu, S.; Li, T.; Xiao, J.; Xu, X.K. Quantifying the effects of topology and weight for link prediction in weighted complex networks. Entropy 2018, 20, 363. [Google Scholar] [CrossRef]
- Orsini, C.; Dankulov, M.M.; Colomer-de Simón, P.; Jamakovic, A.; Mahadevan, P.; Vahdat, A.; Bassler, K.E.; Toroczkai, Z.; Boguná, M.; Caldarelli, G.; et al. Quantifying randomness in real networks. Nat. Commun. 2015, 6, 8627. [Google Scholar] [CrossRef]
- Gjoka, M.; Kurant, M.; Markopoulou, A. 2.5K-graphs: From sampling to generation. In Proceedings of the 2013 IEEE INFOCOM, Turin, Italy, 14–19 April 2013; pp. 1968–1976. [Google Scholar]
- Mahadevan, P.; Hubble, C.; Krioukov, D.; Huffaker, B.; Vahdat, A. Orbis: Rescaling degree correlations to generate annotated internet topologies. In Proceedings of the 2007 Conference on Applications, Technologies, Architectures, and Protocols for Computer Communications, Kyoto, Japan, 27–31 August 2007; pp. 325–336. [Google Scholar]
- Zeng, Y.; Huang, Y.; Wu, Q.; Lü, L. Influential Simplices Mining via Simplicial Convolutional Network. arXiv 2023, arXiv:2307.05841. [Google Scholar]
- Mahadevan, P.; Krioukov, D.; Fall, K.; Vahdat, A. Systematic topology analysis and generation using degree correlations. ACM Sigcomm Comput. Commun. Rev. 2006, 36, 135–146. [Google Scholar] [CrossRef]
- Klimm, F.; Deane, C.M.; Reinert, G. Hypergraphs for predicting essential genes using multiprotein complex data. J. Complex Netw. 2021, 9, cnaa028. [Google Scholar] [CrossRef]
- Chodrow, P.S. Configuration models of random hypergraphs. J. Complex Netw. 2020, 8, cnaa018. [Google Scholar] [CrossRef]
- Miyashita, R.; Nakajima, K.; Fukuda, M.; Shudo, K. Randomizing Hypergraphs Preserving Two-mode Clustering Coefficient. In Proceedings of the 2023 IEEE International Conference on Big Data and Smart Computing (BigComp), Jeju, Republic of Korea, 13–16 February 2023; pp. 316–317. [Google Scholar]
- Nakajima, K.; Shudo, K.; Masuda, N. Randomizing hypergraphs preserving degree correlation and local clustering. IEEE Trans. Netw. Sci. Eng. 2021, 9, 1139–1153. [Google Scholar] [CrossRef]
- Omar, Y.M.; Plapper, P. A survey of information entropy metrics for complex networks. Entropy 2020, 22, 1417. [Google Scholar] [CrossRef]
- Shannon, C.E. A mathematical theory of communication. Bell Syst. Tech. J. 1948, 27, 379–423. [Google Scholar] [CrossRef]
- Fortunato, S.; Barthelemy, M. Resolution limit in community detection. Proc. Natl. Acad. Sci. USA 2007, 104, 36–41. [Google Scholar] [CrossRef] [PubMed]
- Newman, M.E.; Girvan, M. Finding and evaluating community structure in networks. Phys. Rev. E 2004, 69, 026113. [Google Scholar] [CrossRef] [PubMed]
- Hu, D.; Li, X.L.; Liu, X.G.; Zhang, S.G. Extremality of graph entropy based on degrees of uniform hypergraphs with few edges. Acta Math. Sin. Engl. Ser. 2019, 35, 1238–1250. [Google Scholar] [CrossRef]
- Berge, C. Graphs and Hypergraphs; North-Holland Publishing Company: Amsterdam, The Netherlands, 1973. [Google Scholar]
- Estrada, E.; Rodríguez-Velázquez, J.A. Subgraph centrality and clustering in complex hyper-networks. Phys. A Stat. Mech. Its Appl. 2006, 364, 581–594. [Google Scholar] [CrossRef]
- Newman, M.E. Assortative Mixing in Networks. Phys. Rev. Lett. 2002, 89, 208701. [Google Scholar] [CrossRef] [PubMed]
- Amburg, I.; Veldt, N.; Benson, A. Clustering in graphs and hypergraphs with categorical edge labels. In Proceedings of the Web Conference 2020, Taipei, Taiwan, 20–24 April 2020; pp. 706–717. [Google Scholar]
- King, Z.A.; Lu, J.; Dräger, A.; Miller, P.; Federowicz, S.; Lerman, J.A.; Ebrahim, A.; Palsson, B.O.; Lewis, N.E. BiGG Models: A platform for integrating, standardizing and sharing genome-scale models. Nucleic Acids Res. 2016, 44, D515–D522. [Google Scholar] [CrossRef] [PubMed]
- Peng, H.; Qian, C.; Zhao, D.; Zhong, M.; Ling, X.; Wang, W. Disintegrate hypergraph networks by attacking hyperedge. J. King Saud-Univ.-Comput. Inf. Sci. 2022, 34, 4679–4685. [Google Scholar] [CrossRef]
- Bodó, Á.; Katona, G.Y.; Simon, P.L. SIS epidemic propagation on hypergraphs. Bull. Math. Biol. 2016, 78, 713–735. [Google Scholar] [CrossRef] [PubMed]

| Networks | n | e | C | Node | Hyperedge | ||
|---|---|---|---|---|---|---|---|
| Algebra | 423 | 1268 | 78.90 | 19.53 | 0.79 | User | Question |
| Bars-Rev | 1234 | 1194 | 174.30 | 9.62 | 0.58 | User | Bar review |
| iAF1260b | 1668 | 2351 | 13.26 | 5.46 | 0.55 | Metabolite | Metabolic interaction |
| iJO1366 | 1805 | 2546 | 16.92 | 5.55 | 0.58 | Metabolite | Metabolic interaction |
| Statistics | Network | Original | H0k | H1k-HED | H1k-HD | HD-HED | H2k | H2.5k |
|---|---|---|---|---|---|---|---|---|
| Assortativity () | Algebra | −0.10 | −0.02 (0.20) | −0.03 (0.30) | −0.05 (0.50) | −0.08 (0.80) | −0.09(0.90) | −0.09(0.90) |
| Bars-Rev | 0.30 | −0.002 (−0.01) | 0.08 (0.27) | 0.26 (0.87) | 0.27 (0.90) | 0.29 (0.97) | 0.29 (0.97) | |
| iAF1260b | −0.30 | −0.03 (0.10) | −0.16 (0.53) | −0.22 (0.73) | −0.26 (0.87) | −0.28 (0.93) | −0.28 (0.93) | |
| iJO1366 | −0.29 | −0.03 (0.10) | −0.14 (0.48) | −0.23 (0.79) | −0.25 (0.86) | −0.27 (0.93) | −0.28 (0.97) | |
| Clustering coefficient () | Algebra | 0.80 | 0.66 (0.83) | 0.75 (0.93) | 0.76 (0.95) | 0.79 (0.99) | 0.80 (1.00) | 0.80 (1.00) |
| Bars-Rev | 0.58 | 0.28 (0.48) | 0.44 (0.76) | 0.49 (0.84) | 0.57 (0.98) | 0.58 (1.00) | 0.58 (1.00) | |
| iAF1260b | 0.55 | 0.46 (0.84) | 0.54 (0.98) | 0.55 (1.00) | 0.55 (1.00) | 0.55 (1.00) | 0.55 (1.00) | |
| iJO1366 | 0.58 | 0.45 (0.77) | 0.55 (0.95) | 0.57 (0.98) | 0.57 (0.98) | 0.57 (0.98) | 0.58 (0.98) | |
| Average Neighbor Degree () | Algebra | 44.35 | 22.88 (0.52) | 38.15 (0.86) | 42.12 (0.95) | 44.13 (0.99) | 44.25 (0.99) | 44.35 (1.00) |
| Bars-Rev | 13.28 | 8.94 (0.67) | 12.32 (0.93) | 11.95 (0.90) | 12.29 (0.93) | 13.28 (1.00) | 13.28 (1.00) | |
| iAF1260b | 141.425 | 25.66 (0.18) | 37.21 (0.26) | 135.27 (0.96) | 139.49 (0.99) | 141.395 (1.00) | 141.425 (1.00) | |
| iJO1366 | 159.46 | 30.85 (0.19) | 150.43 (0.94) | 42.79 (0.27) | 157.45 (0.99) | 157.79 (0.99) | 159.46 (1.00) |
| Origin | H0k | H1k-HED | H1k-HD | HD-HED | H2K | H2.25k | |
|---|---|---|---|---|---|---|---|
| Algebra | 0.15 | 0.81 | 0.42 | 0.45 | 0.41 | 0.36 | 0.39 |
| Bars-Rev | 0.43 | 0.83 | 0.55 | 0.54 | 0.49 | 0.49 | 0.51 |
| iAF1260b | 0.04 | 0.20 | 0.04 | 0.04 | 0.05 | 0.04 | 0.05 |
| iJO1366 | 0.29 | 0.65 | 0.31 | 0.28 | 0.27 | 0.29 | 0.30 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Zeng, Y.; Liu, B.; Zhou, F.; Lü, L. Hyper-Null Models and Their Applications. Entropy 2023, 25, 1390. https://doi.org/10.3390/e25101390
Zeng Y, Liu B, Zhou F, Lü L. Hyper-Null Models and Their Applications. Entropy. 2023; 25(10):1390. https://doi.org/10.3390/e25101390
Chicago/Turabian StyleZeng, Yujie, Bo Liu, Fang Zhou, and Linyuan Lü. 2023. "Hyper-Null Models and Their Applications" Entropy 25, no. 10: 1390. https://doi.org/10.3390/e25101390
APA StyleZeng, Y., Liu, B., Zhou, F., & Lü, L. (2023). Hyper-Null Models and Their Applications. Entropy, 25(10), 1390. https://doi.org/10.3390/e25101390







