Silhouette Analysis for Performance Evaluation in Machine Learning with Applications to Clustering
Abstract
1. Introduction
2. Background
2.1. K-Means
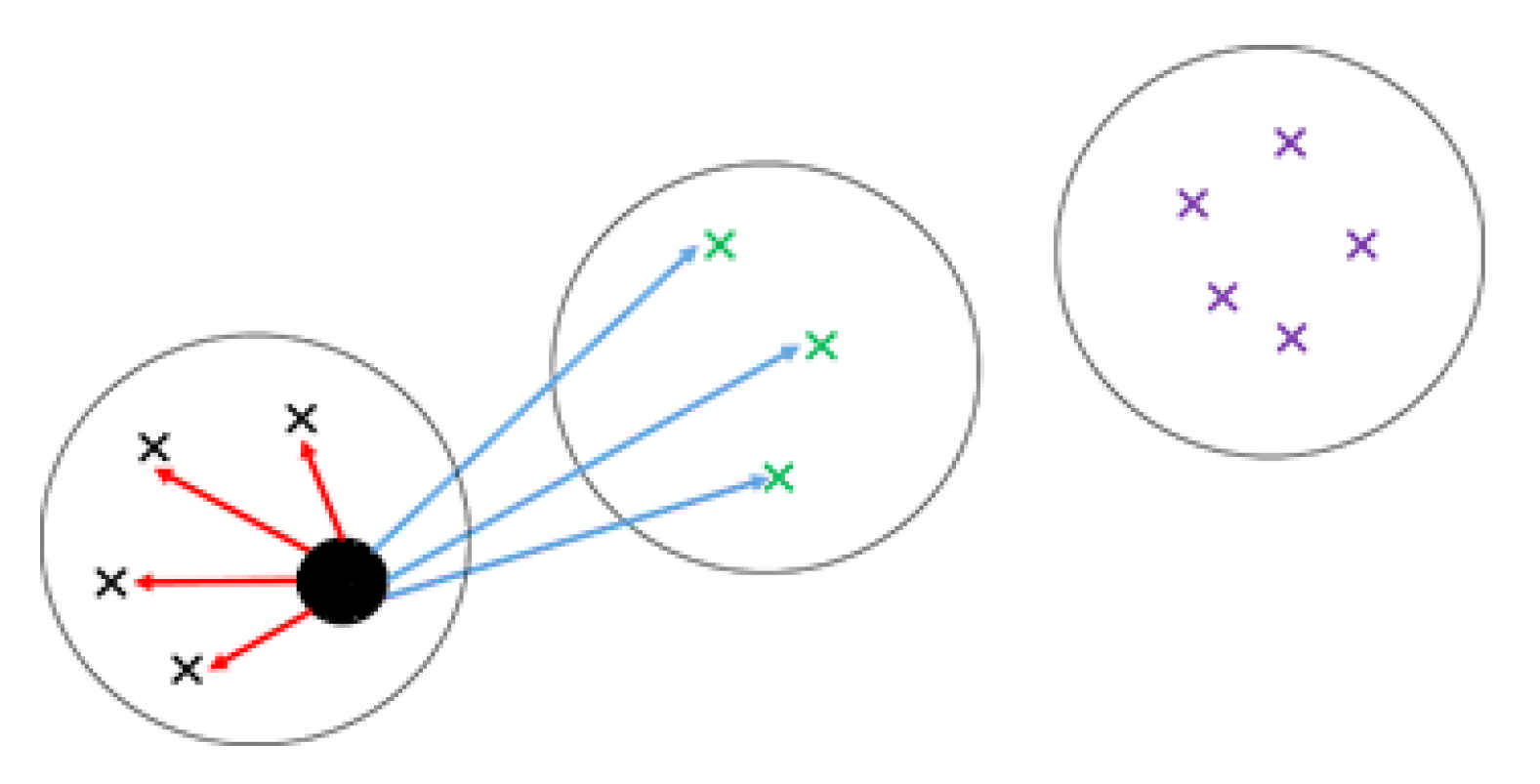
2.2. Kernel K-Means
2.3. Silhouette Index
3. Weighted Clustering Method Using Silhouette Index
- Let be a set of kernel functions to . Perform kernel k-means method using kernels in the kernel set and generate clustering results ;
- Compute average Silhouette width for clustering results obtained by kernel for all kernels ;
- Shift Silhouette values from {−1 to 1} to {0 to 2} to compute non-negative weights for each kernel;
- For each data point, use the computed weights in step (3) to combine the clustering results as follows:
- Sum up the weights corresponding to the kernels that assign the same cluster label to the data point;
- Compare the total weight of each cluster for the data point;
- Group the data point to the cluster with the highest total weight.
4. Simulation
- Bensaid data: it contains 49 two-dimensional elements grouped in three classes [16].
- Dunn data: it contains 90 two-dimensional elements grouped in two classes [17].
- Iris data: it contains 150 four-dimensional elements grouped in three classes [18].
- Seed data: it contains 210 seven-dimensional elements grouped in three classes [19].
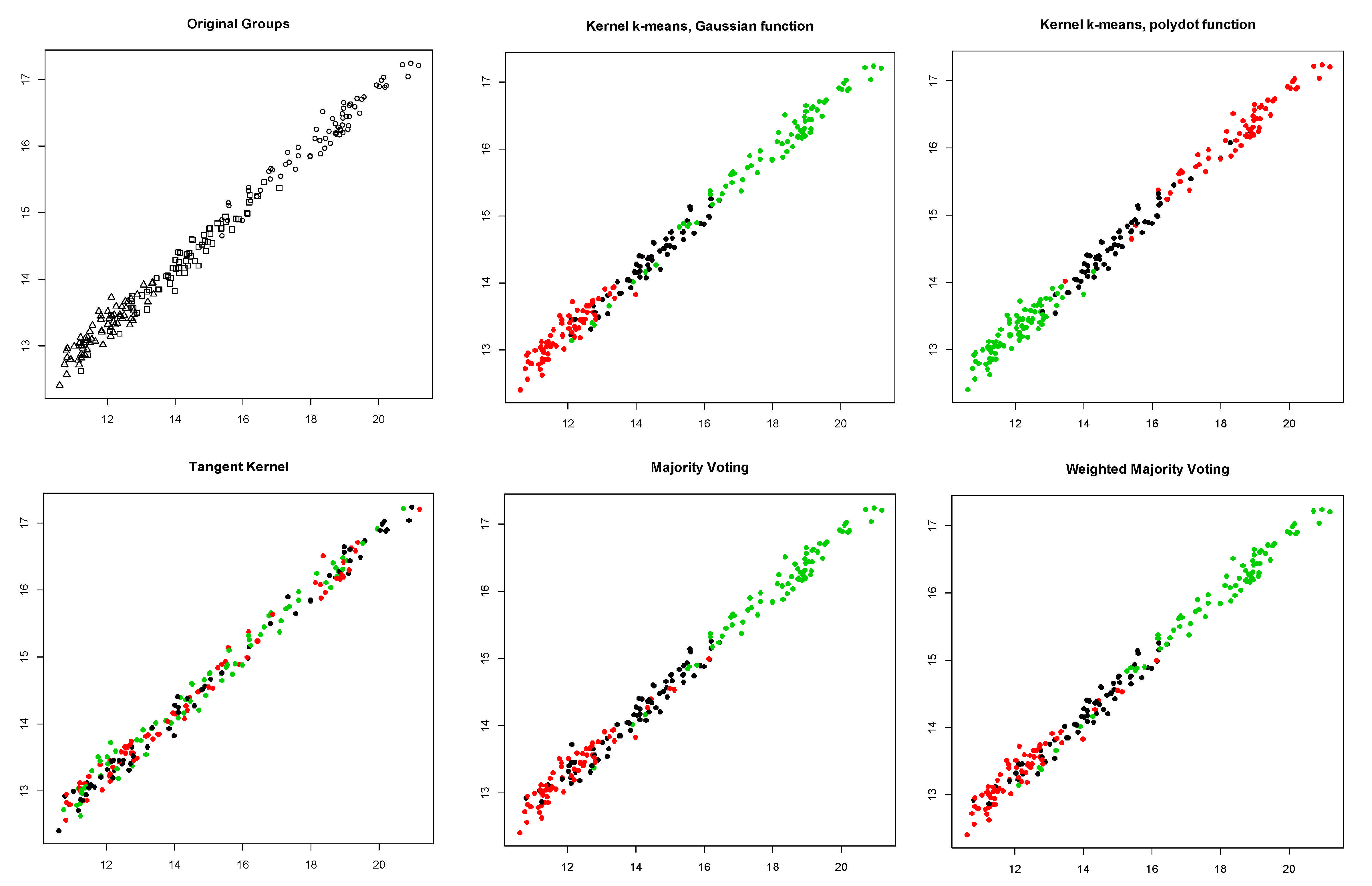
5. Results
6. Conclusions and Future Research
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Gentleman, R.; Carey, V.; Huber, W.; Irizarry, R.; Dudoit, S. (Eds.) Bioinformatics and Computational Biology Solutions Using R and Bioconductor; Springer Science & Business Media: New York, NY, USA, 2006. [Google Scholar]
- Monti, S.; Tamayo, P.; Mesirov, J.; Golub, T. Consensus clustering: A resampling-based method for class discovery and visualization of gene expression microarray data. Mach. Learn. 2003, 52, 91–118. [Google Scholar] [CrossRef]
- Wu, J. Advances in K-Means Clustering: A Data Mining Thinking; Springer Science & Business Media: New York, NY, USA, 2012. [Google Scholar]
- Kassambara, A. Practical Guide to Cluster Analysis in R: Unsupervised Machine Learning; CreateSpace Independent Publishing Platform: Scotts Valley, CA, USA, 2017; Volume 1. [Google Scholar]
- Brocard, D.; Gillet, F.; Legendre, P. Numerical Ecology with R (Use R!); Springer: New York, NY, USA, 2011; pp. 978–981. [Google Scholar] [CrossRef]
- Nguyen, N.; Caruana, R. Consensus clusterings. In Proceedings of the Seventh IEEE International Conference on Data Mining (ICDM 2007), Omaha, NE, USA, 28–31 October 2007; pp. 607–612. [Google Scholar]
- Jain, A.K.; Murty, M.N.; Flynn, P.J. Data clustering: A review. ACM Comput. Surv. (CSUR) 1999, 31, 264–323. [Google Scholar] [CrossRef]
- Arthur, D.; Vassilvitskii, S. Society for Industrial and Applied Mathematics. k-means++: The advantages of careful seeding. In Proceedings of the eighteenth annual ACM-SIAM symposium on Discrete algorithms, New Orleans, LA, USA, 7–9 January 2007; pp. 1027–1035. [Google Scholar]
- Kachouie, N.N.; Shutaywi, M. Weighted Mutual Information for Aggregated Kernel Clustering. Entropy 2020, 22, 351. [Google Scholar] [CrossRef] [PubMed]
- Dhillon, I.S.; Guan, Y.; Kulis, B. Kernel k-means: Spectral clustering and normalized cuts. In Proceedings of the Tenth ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, Sydney, Australia, 10–13 August 2015; ACM: New York, NY, USA, 2004; pp. 551–556. [Google Scholar]
- Camps-Valls, G. (Ed.) Kernel Methods in Bioengineering, Signal and Image Processing; Igi Global: Hershey, PA, USA, 2006. [Google Scholar]
- Campbell, C. An introduction to kernel methods. Stud. Fuzziness Soft Comput. 2001, 66, 155–192. [Google Scholar]
- Yeung, K.Y.; Haynor, D.R.; Ruzzo, W.L. Validating clustering for gene expression data. Bioinformatics 2001, 17, 309–318. [Google Scholar] [CrossRef] [PubMed]
- Hubert, L.; Arabie, P. Comparing partitions. J. Classif. 1985, 2, 193–218. [Google Scholar] [CrossRef]
- Rousseeuw, P.J. Silhouettes: A graphical aid to the interpretation and validation of cluster analysis. J. Comput. Appl. Math. 1987, 20, 53–65. [Google Scholar] [CrossRef]
- Bensaid, A.M.; Hall, L.O.; Bezdek, J.C.; Clarke, L.P.; Silbiger, M.L.; Arrington, J.A.; Murtagh, R.F. Validity-guided (re) clustering with applications to image segmentation. IEEE Trans. Fuzzy Syst. 1996, 4, 112–123. [Google Scholar] [CrossRef]
- Bezdek, J.C.; Keller, J.; Krisnapuram, R.; Pal, N. Fuzzy Models and Algorithms for Pattern Recognition and Image Processing; Springer Science & Business Media: New York, NY, USA, 1999; Volume 4. [Google Scholar]
- Kozak, M.; Łotocka, B. What should we know about the famous Iris data? Curr. Sci. 2013, 104, 579–580. [Google Scholar]
- Charytanowicz, M.; Niewczas, J.; Kulczycki, P.; Kowalski, P.A.; Łukasik, S.; Żak, S. Complete Gradient Clustering Algorithm for Features Analysis of X-Ray Images. In Information Technologies in Biomedicine; Springer: Berlin/Heidelberg, Germany, 2010; pp. 15–24. [Google Scholar]


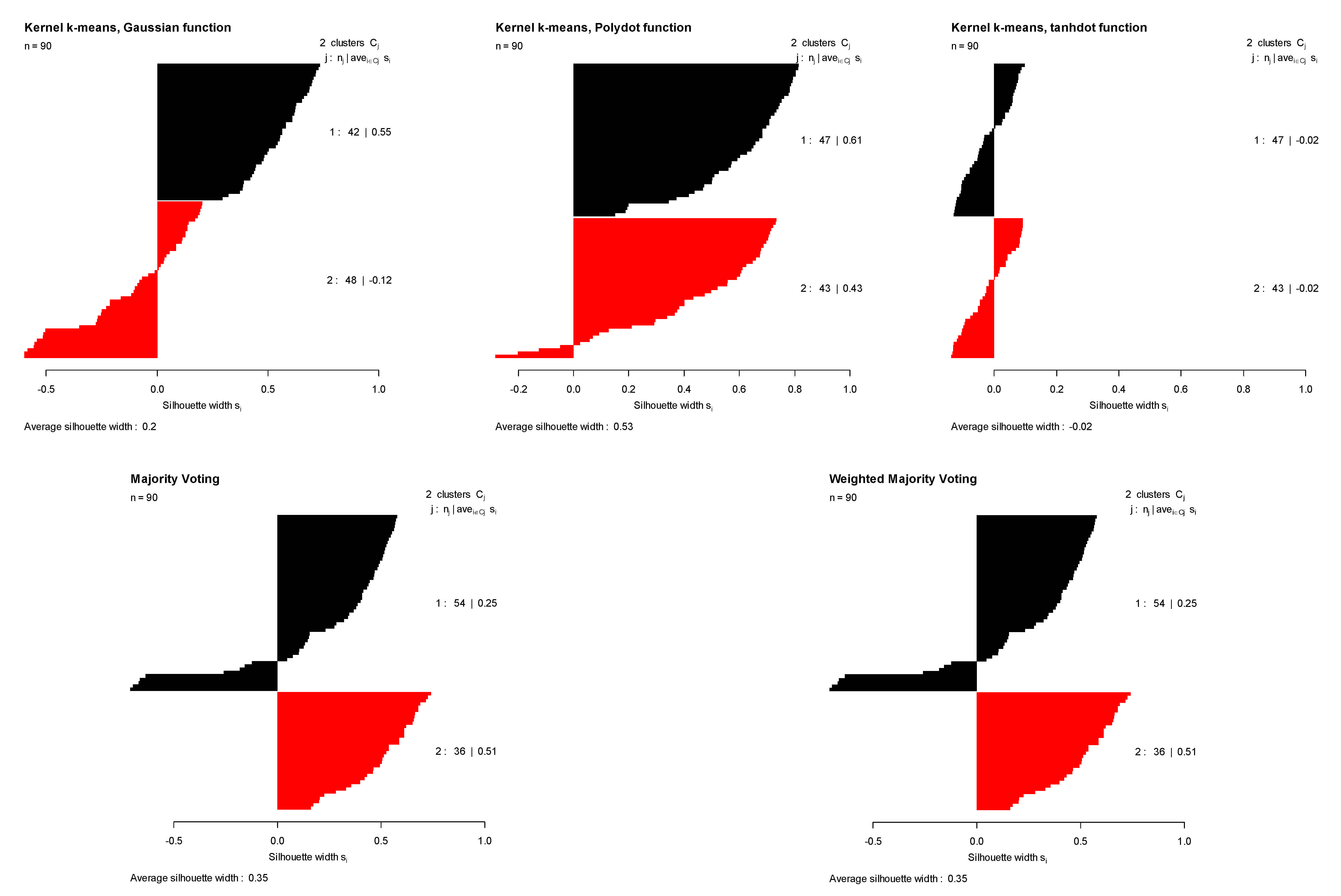
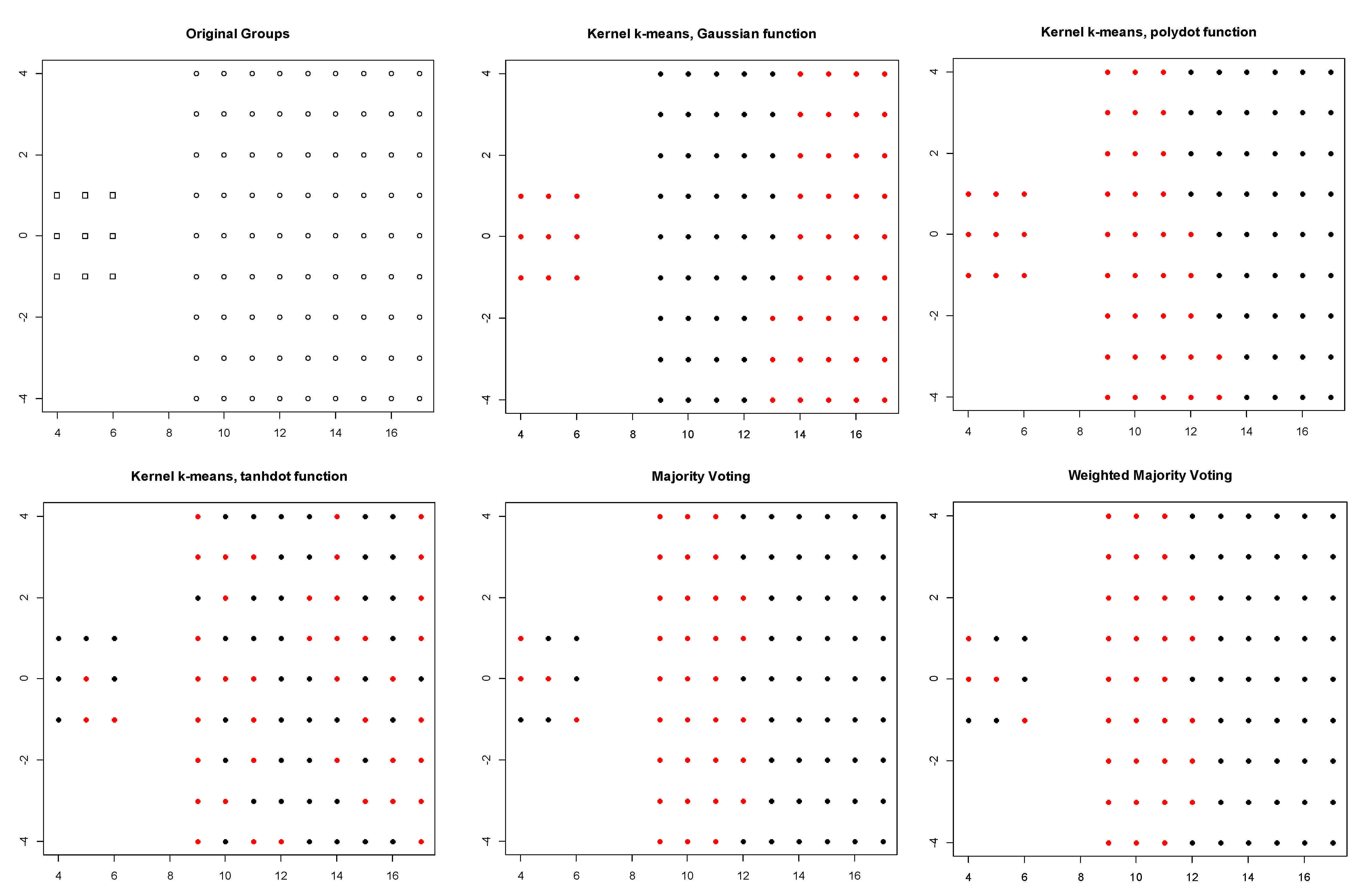
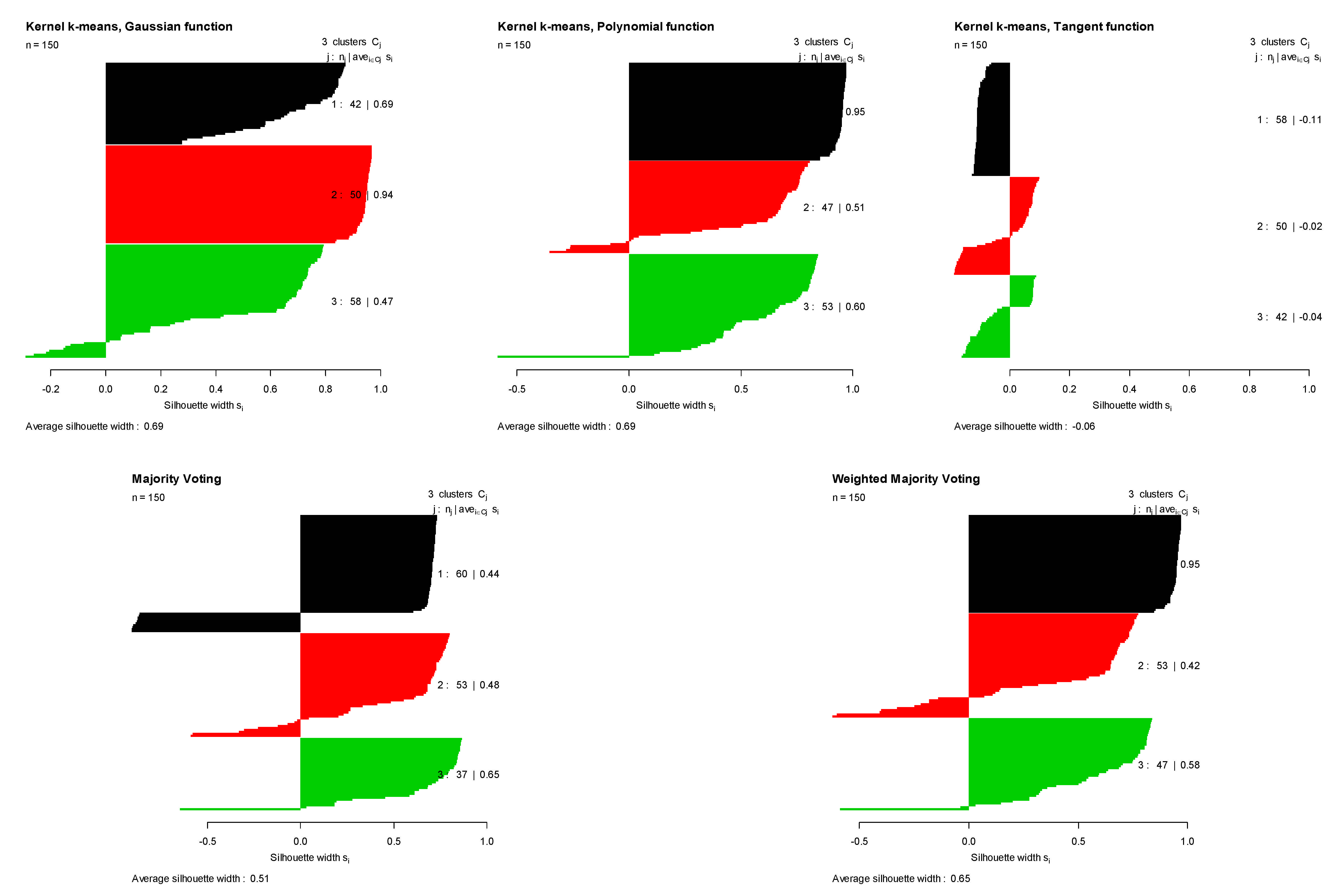


| DATA | Gaussian | Polynomial | Tangent | Majority Voting | Weighted Majority Voting | |
|---|---|---|---|---|---|---|
| Bensaid | ASI | 0.159 | 0.453 | −0.127 | 0.132 | 0.167 |
| TR | 0.528 | 0.641 | 0.413 | 0.592 | 0.611 | |
| DUNN | ASI | 0.269 | 0.418 | −0.003 | 0.293 | 0.293 |
| TR | 0.453 | 0.521 | 0.417 | 0.474 | 0.474 | |
| IRIS | ASI | 0.609 | 0.609 | −0.068 | 0.482 | 0.589 |
| TR | 0.856 | 0.850 | 0.381 | 0.833 | 0.873 | |
| SEED | ASI | 0.594 | 0.601 | −0.052 | 0.537 | 0.570 |
| TR | 0.869 | 0.842 | 0.376 | 0.852 | 0.862 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Shutaywi, M.; Kachouie, N.N. Silhouette Analysis for Performance Evaluation in Machine Learning with Applications to Clustering. Entropy 2021, 23, 759. https://doi.org/10.3390/e23060759
Shutaywi M, Kachouie NN. Silhouette Analysis for Performance Evaluation in Machine Learning with Applications to Clustering. Entropy. 2021; 23(6):759. https://doi.org/10.3390/e23060759
Chicago/Turabian StyleShutaywi, Meshal, and Nezamoddin N. Kachouie. 2021. "Silhouette Analysis for Performance Evaluation in Machine Learning with Applications to Clustering" Entropy 23, no. 6: 759. https://doi.org/10.3390/e23060759
APA StyleShutaywi, M., & Kachouie, N. N. (2021). Silhouette Analysis for Performance Evaluation in Machine Learning with Applications to Clustering. Entropy, 23(6), 759. https://doi.org/10.3390/e23060759






