Scikit-Dimension: A Python Package for Intrinsic Dimension Estimation
Abstract
:1. Introduction
2. Materials and Methods
2.1. Software Features
2.1.1. id Module
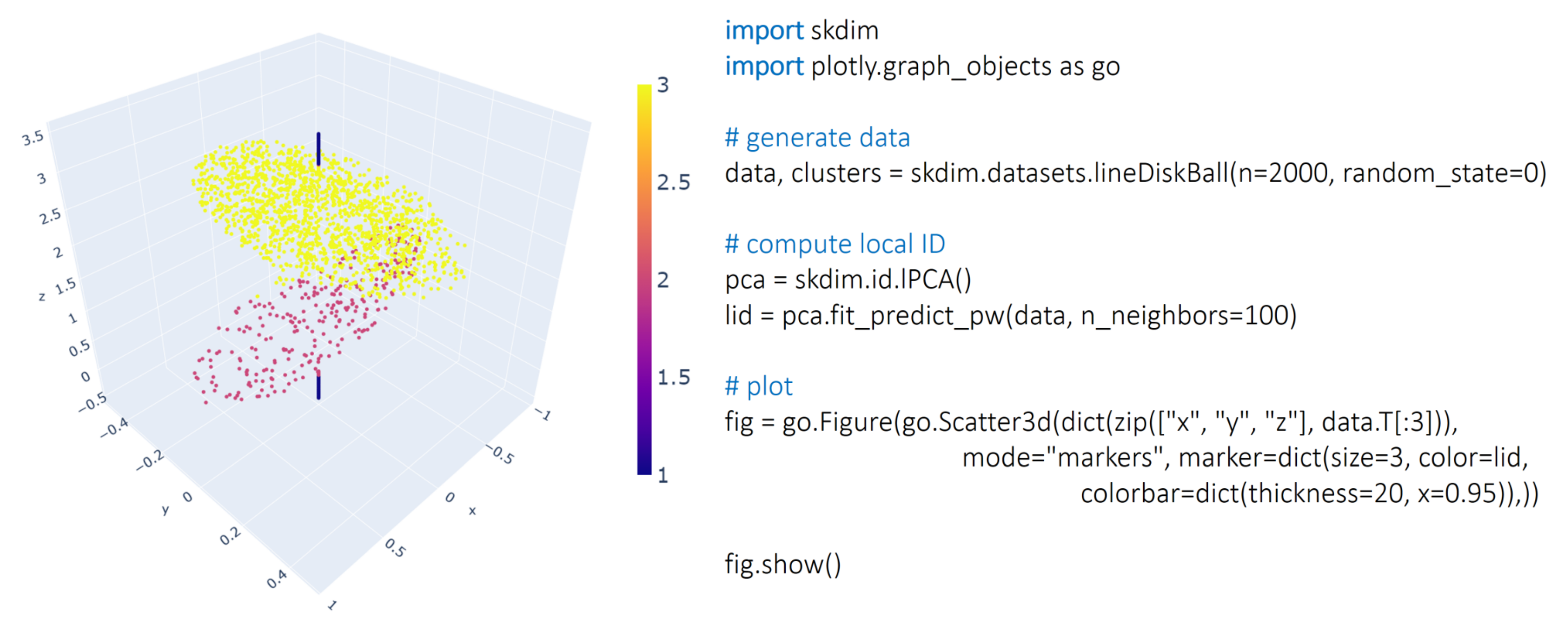
2.1.2. Datasets Module
2.2. Development
2.3. Dependencies
2.4. Related Software
3. Results
3.1. Benchmarking Scikit-Dimension on a Large Collection of Datasets
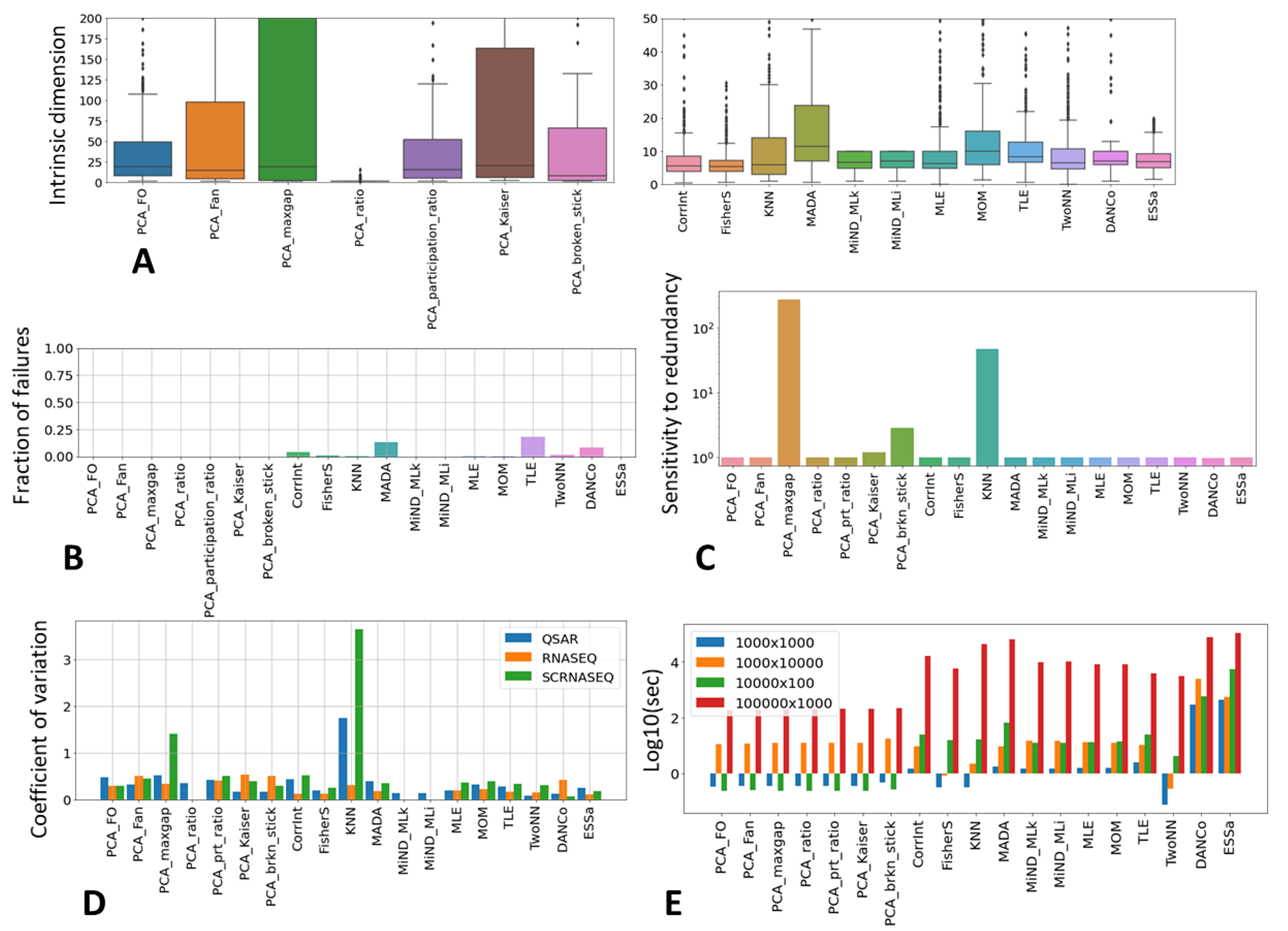
3.1.1. Scikit-Dimension ID Estimator Method Features
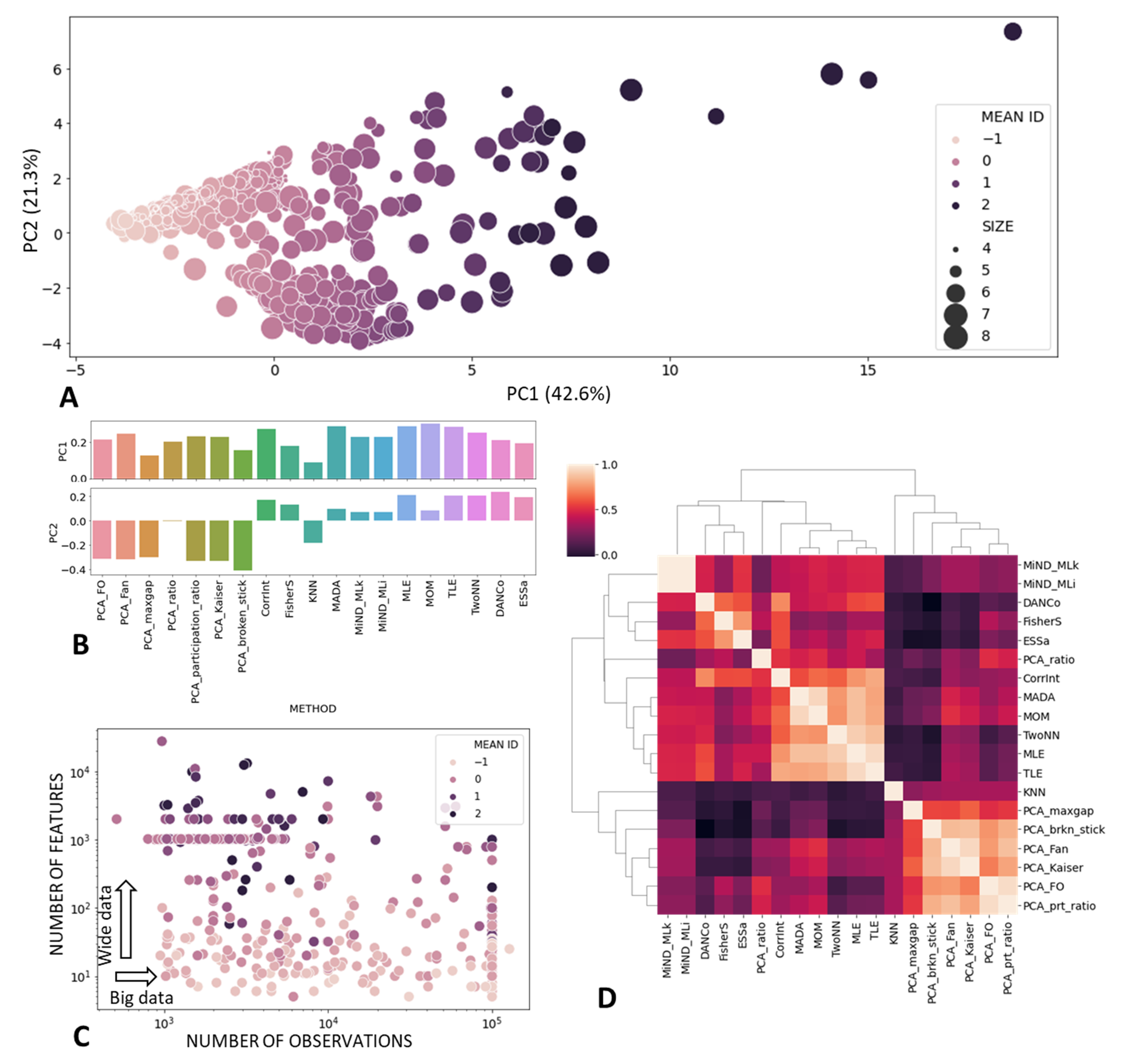
3.1.2. Metanalysis of Scikit-Dimension ID Estimates
4. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V.; et al. Scikit-learn: Machine learning in Python. J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Bishop, C.M. Neural Networks for Pattern Recognition; Oxford University Press: Oxford, UK, 1995. [Google Scholar] [CrossRef]
- Fukunaga, K. Intrinsic dimensionality extraction. In Pattern Recognition and Reduction of Dimensionality, Handbook of Statistics; Krishnaiah, P.R., Kanal, L.N., Eds.; North-Holland: Amsterdam, The Netherlands, 1982; Volume 2, pp. 347–362. [Google Scholar]
- Albergante, L.; Bac, J.; Zinovyev, A. Estimating the effective dimension of large biological datasets using Fisher separability analysis. In Proceedings of the 2019 International Joint Conference on Neural Networks (IJCNN), Budapest, Hungary, 14–19 July 2019; pp. 1–8. [Google Scholar]
- Giudice, M.D. Effective Dimensionality: A Tutorial. Multivar. Behav. Res. 2020, 1–16. [Google Scholar] [CrossRef]
- Palla, K.; Knowles, D.; Ghahramani, Z. A nonparametric variable clustering model. In Advances in Neural Information Processing Systems; MIT Press: Cambridge, MA, USA, 2012; Volume 4, pp. 2987–2995. [Google Scholar]
- Giuliani, A.; Benigni, R.; Sirabella, P.; Zbilut, J.P.; Colosimo, A. Nonlinear Methods in the Analysis of Protein Sequences: A Case Study in Rubredoxins. Biophys. J. 2000, 78, 136–149. [Google Scholar] [CrossRef] [Green Version]
- Jiang, H.; Kim, B.; Guan, M.Y.; Gupta, M.R. To Trust Or Not To Trust A Classifier. In NeurIPS; Montreal Convention Centre: Montreal, QC, Canada, 2018; pp. 5546–5557. [Google Scholar] [CrossRef]
- Bac, J.; Zinovyev, A. Lizard Brain: Tackling Locally Low-Dimensional Yet Globally Complex Organization of Multi-Dimensional Datasets. Front. Neurorobotics 2020, 13, 110. [Google Scholar] [CrossRef] [Green Version]
- Hino, H. ider: Intrinsic Dimension Estimation with R. R J. 2017, 9, 329–341. [Google Scholar] [CrossRef]
- Campadelli, P.; Casiraghi, E.; Ceruti, C.; Rozza, A. Intrinsic Dimension Estimation: Relevant Techniques and a Benchmark Framework. Math. Probl. Eng. 2015, 2015, 759567. [Google Scholar] [CrossRef] [Green Version]
- Camastra, F.; Staiano, A. Intrinsic dimension estimation: Advances and open problems. Inf. Sci. 2016, 328, 26–41. [Google Scholar] [CrossRef]
- Little, A.V.; Lee, J.; Jung, Y.; Maggioni, M. Estimation of intrinsic dimensionality of samples from noisy low-dimensional manifolds in high dimensions with multiscale SVD. In Proceedings of the 2009 IEEE/SP 15th Workshop on Statistical Signal Processing, Cardiff, UK, 31 August–3 September 2009; pp. 85–88. [Google Scholar] [CrossRef]
- Hein, M.; Audibert, J.Y. Intrinsic dimensionality estimation of submanifolds in Rd. In Proceedings of the 22nd International Conference on Machine Learning, Bonn, Germany, 7–11 August 2005; ACM: New York, NY, USA, 2005; pp. 289–296. [Google Scholar] [CrossRef] [Green Version]
- Mirkes, E.; Allohibi, J.; Gorban, A.N. Fractional Norms and Quasinorms Do Not Help to Overcome the Curse of Dimensionality. Entropy 2020, 22, 1105. [Google Scholar] [CrossRef] [PubMed]
- Golovenkin, S.E.; Bac, J.; Chervov, A.; Mirkes, E.M.; Orlova, Y.V.; Barillot, E.; Gorban, A.N.; Zinovyev, A. Trajectories, bifurcations, and pseudo-time in large clinical datasets: Applications to myocardial infarction and diabetes data. GigaScience 2020, 9, giaa128. [Google Scholar] [CrossRef] [PubMed]
- Zinovyev, A.; Sadovsky, M.; Calzone, L.; Fouché, A.; Groeneveld, C.S.; Chervov, A.; Barillot, E.; Gorban, A.N. Modeling Progression of Single Cell Populations Through the Cell Cycle as a Sequence of Switches. bioRxiv 2021. [Google Scholar] [CrossRef]
- Grassberger, P.; Procaccia, I. Measuring the strangeness of strange attractors. Phys. D Nonlinear Phenom. 1983, 9, 189–208. [Google Scholar] [CrossRef]
- Farahmand, A.M.; Szepesvári, C.; Audibert, J.Y. Manifold-adaptive dimension estimation. In Proceedings of the 24th International Conference on Machine Learning, Corvallis, OR, USA, 20–24 June 2007; pp. 265–272. [Google Scholar] [CrossRef]
- Amsaleg, L.; Chelly, O.; Furon, T.; Girard, S.; Houle, M.E.; Kawarabayashi, K.; Nett, M. Extreme-value-theoretic estimation of local intrinsic dimensionality. Data Min. Knowl. Discov. 2018, 32, 1768–1805. [Google Scholar] [CrossRef]
- Jackson, D.A. Stopping rules in principal components analysis: A comparison of heuristical and statistical approaches. Ecology 1993, 74, 2204–2214. [Google Scholar] [CrossRef]
- Fukunaga, K.; Olsen, D.R. An Algorithm for Finding Intrinsic Dimensionality of Data. IEEE Trans. Comput. 1971, C-20, 176–183. [Google Scholar] [CrossRef]
- Mingyu, F.; Gu, N.; Qiao, H.; Zhang, B. Intrinsic dimension estimation of data by principal component analysis. arXiv 2010, arXiv:1002.2050. [Google Scholar]
- Hill, B.M. A simple general approach to inference about the tail of a distribution. Ann. Stat. 1975, 1163–1174. [Google Scholar] [CrossRef]
- Levina, E.; Bickel, P.J. Maximum Likelihood estimation of intrinsic dimension. In Proceedings of the 17th International Conference on Neural Information Processing Systems, Vancouver, Canada, 1 December 2004; MIT Press: Cambridge, MA, USA, 2004; pp. 777–784. [Google Scholar] [CrossRef]
- Haro, G.; Randall, G.; Sapiro, G. Translated poisson mixture model for stratification learning. Int. J. Comput. Vis. 2008, 80, 358–374. [Google Scholar] [CrossRef]
- Carter, K.M.; Raich, R.; Hero, A.O. On Local Intrinsic Dimension Estimation and Its Applications. IEEE Trans. Signal Process. 2010, 58, 650–663. [Google Scholar] [CrossRef] [Green Version]
- Rozza, A.; Lombardi, G.; Ceruti, C.; Casiraghi, E.; Campadelli, P. Novel high intrinsic dimensionality estimators. Mach. Learn. 2012, 89, 37–65. [Google Scholar] [CrossRef]
- Ceruti, C.; Bassis, S.; Rozza, A.; Lombardi, G.; Casiraghi, E.; Campadelli, P. DANCo: An intrinsic dimensionality estimator exploiting angle and norm concentration. Pattern Recognit. 2014, 47, 2569–2581. [Google Scholar] [CrossRef]
- Johnsson, K. Structures in High-Dimensional Data: Intrinsic Dimension and Cluster Analysis. Ph.D. Thesis, Faculty of Engineering, LTH, Perth, Australia, 2016. [Google Scholar]
- Facco, E.; D’Errico, M.; Rodriguez, A.; Laio, A. Estimating the intrinsic dimension of datasets by a minimal neighborhood information. Sci. Rep. 2017, 7, 12140. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gorban, A.; Golubkov, A.; Grechuk, B.; Mirkes, E.; Tyukin, I. Correction of AI systems by linear discriminants: Probabilistic foundations. Inf. Sci. 2018, 466, 303–322. [Google Scholar] [CrossRef] [Green Version]
- Amsaleg, L.; Chelly, O.; Houle, M.E.; Kawarabayashi, K.; Radovanović, M.; Treeratanajaru, W. Intrinsic dimensionality estimation within tight localities. In Proceedings of the 2019 SIAM International Conference on Data Mining, Calgary, AB, Canada, 2–4 May 2019; SIAM: Philadelphia, PA, USA, 2019; pp. 181–189. [Google Scholar]
- Harris, C.R.; Millman, K.J.; van der Walt, S.J.; Gommers, R.; Virtanen, P.; Cournapeau, D.; Wieser, E.; Taylor, J.; Berg, S.; Smith, N.J.; et al. Array programming with NumPy. Nature 2020, 585, 357–362. [Google Scholar] [CrossRef]
- Hunter, J.D. Matplotlib: A 2D graphics environment. Comput. Sci. Eng. 2007, 9, 90–95. [Google Scholar] [CrossRef]
- The Pandas Development Team.Pandas-Dev/Pandas: Pandas 1.3.4, Zenodo. Available online: https://zenodo.org/record/5574486#.YW50jhpByUk (accessed on 18 October 2021). [CrossRef]
- Lam, S.K.; Pitrou, A.; Seibert, S. Numba: A llvm-based python jit compiler. In Proceedings of the Second Workshop on the LLVM Compiler Infrastructure in HPC, Austin, TX, USA, 15 November 2015; pp. 1–6. [Google Scholar] [CrossRef]
- Virtanen, P.; Gommers, R.; Oliphant, T.E.; Haberland, M.; Reddy, T.; Cournapeau, D.; Burovski, E.; Peterson, P.; Weckesser, W.; Bright, J.; et al. SciPy 1.0: Fundamental Algorithms for Scientific Computing in Python. Nat. Methods 2020, 17, 261–272. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Johnsson, K. intrinsicDimension: Intrinsic Dimension Estimation (R Package). 2019. Available online: https://rdrr.io/cran/intrinsicDimension/ (accessed on 6 September 2021).
- You, K. Rdimtools: An R package for Dimension Reduction and Intrinsic Dimension Estimation. arXiv 2020, arXiv:2005.11107. [Google Scholar]
- Denti, Francesco intRinsic: An R package for model-based estimation of the intrinsic dimension of a dataset. arXiv 2021, arXiv:2102.11425.
- Hein, M.J.Y.A. IntDim: Intrindic Dimensionality Estimation. 2016. Available online: https://www.ml.uni-saarland.de/code/IntDim/IntDim.htm (accessed on 6 September 2021).
- Lombardi, G. Intrinsic Dimensionality Estimation Techniques (MATLAB Package). 2013. Available online: https://fr.mathworks.com/matlabcentral/fileexchange/40112-intrinsic-dimensionality-estimation-techniques (accessed on 6 September 2021).
- Van der Maaten, L. Drtoolbox: Matlab Toolbox for Dimensionality Reduction. 2020. Available online: https://lvdmaaten.github.io/drtoolbox/ (accessed on 6 September 2021).
- Radovanović, M. Tight Local Intrinsic Dimensionality Estimator (TLE) (MATLAB Package). 2020. Available online: https://perun.pmf.uns.ac.rs/radovanovic/tle/ (accessed on 6 September 2021).
- Gomtsyan, M.; Mokrov, N.; Panov, M.; Yanovich, Y. Geometry-Aware Maximum Likelihood Estimation of Intrinsic Dimension (Python Package). 2019. Available online: https://github.com/stat-ml/GeoMLE (accessed on 6 September 2021).
- Gomtsyan, M.; Mokrov, N.; Panov, M.; Yanovich, Y. Geometry-Aware Maximum Likelihood Estimation of Intrinsic Dimension. In Proceedings of the Eleventh Asian Conference on Machine Learning, Nagoya, Japan, 17–19 November 2019; pp. 1126–1141. [Google Scholar]
- Erba, V. pyFCI: A Package for Multiscale-Full-Correlation-Integral Intrinsic Dimension Estimation. 2019. Available online: https://github.com/vittorioerba/pyFCI (accessed on 6 September 2021).
- Granata, D. Intrinsic-Dimension (Python Package). 2016. Available online: https://github.com/dgranata/Intrinsic-Dimension (accessed on 6 September 2021).
- Bac, J.; Zinovyev, A. Local intrinsic dimensionality estimators based on concentration of measure. In Proceedings of the International Joint Conference on Neural Networks (IJCNN), Glasgow, UK, 19–24 July 2020; pp. 1–8. [Google Scholar]
- Gorban, A.N.; Makarov, V.A.; Tyukin, I.Y. The unreasonable effectiveness of small neural ensembles in high-dimensional brain. Phys. Life Rev. 2019, 29, 55–88. [Google Scholar] [CrossRef]
- Vanschoren, J.; van Rijn, J.N.; Bischl, B.; Torgo, L. OpenML: Networked Science in Machine Learning. SIGKDD Explor. 2013, 15, 49–60. [Google Scholar] [CrossRef] [Green Version]
- Gulati, G.; Sikandar, S.; Wesche, D.; Manjunath, A.; Bharadwaj, A.; Berger, M.; Ilagan, F.; Kuo, A.; Hsieh, R.; Cai, S.; et al. Single-cell transcriptional diversity is a hallmark of developmental potential. Science 2020, 24, 405–411. [Google Scholar] [CrossRef]
- Giuliani, A. The application of principal component analysis to drug discovery and biomedical data. Drug Discov. Today 2017, 22, 1069–1076. [Google Scholar] [CrossRef] [PubMed]
- Cangelosi, R.; Goriely, A. Component retention in principal component analysis with application to cDNA microarray data. Biol. Direct 2007, 2, 2. [Google Scholar] [CrossRef] [Green Version]
- Johnsson, K.; Soneson, C.; Fontes, M. Low Bias Local Intrinsic Dimension Estimation from Expected Simplex Skewness. IEEE Trans. Pattern Anal. Mach. Intell. 2015, 37, 196–202. [Google Scholar] [CrossRef] [PubMed]
- Jolliffe, I.T. Principal Component Analysis; Springer Series in Statistics; Springer: Berlin/Heidelberg, Germany, 2002. [Google Scholar]
- Kaiser, H. The Application of Electronic Computers to Factor Analysis. Educ. Psychol. Meas. 1960, 20, 141–151. [Google Scholar] [CrossRef]
- Frontier, S. Étude de la décroissance des valeurs propres dans une analyse en composantes principales: Comparaison avec le modèle du bâton brisé. J. Exp. Mar. Biol. Ecol. 1976, 25, 67–75. [Google Scholar] [CrossRef]
- Gorban, A.N.; Sumner, N.R.; Zinovyev, A.Y. Topological grammars for data approximation. Appl. Math. Lett. 2007, 20, 382–386. [Google Scholar] [CrossRef] [Green Version]
- Albergante, L.; Mirkes, E.; Bac, J.; Chen, H.; Martin, A.; Faure, L.; Barillot, E.; Pinello, L.; Gorban, A.; Zinovyev, A. Robust and scalable learning of complex intrinsic dataset geometry via ElPiGraph. Entropy 2020, 22, 296. [Google Scholar] [CrossRef] [Green Version]
- Lähnemann, D.; Köster, J.; Szczurek, E.; McCarthy, D.J.; Hicks, S.C.; Robinson, M.D.; Vallejos, C.A.; Campbell, K.R.; Beerenwinkel, N.; Mahfouz, A.; et al. Eleven grand challenges in single-cell data science. Genome Biol. 2020, 21, 1–31. [Google Scholar] [CrossRef]
- Chen, H.; Albergante, L.; Hsu, J.Y.; Lareau, C.A.; Lo Bosco, G.; Guan, J.; Zhou, S.; Gorban, A.N.; Bauer, D.E.; Aryee, M.J.; et al. Single-cell trajectories reconstruction, exploration and mapping of omics data with STREAM. Nat. Commun. 2019, 10, 1–14. [Google Scholar] [CrossRef] [Green Version]
- Sritharan, D.; Wang, S.; Hormoz, S. Computing the Riemannian curvature of image patch and single-cell RNA sequencing data manifolds using extrinsic differential geometry. Proc. Natl. Acad. Sci. USA 2021, 118, e2100473118. [Google Scholar] [CrossRef]
- Radulescu, O.; Gorban, A.N.; Zinovyev, A.; Lilienbaum, A. Robust simplifications of multiscale biochemical networks. BMC Syst. Biol. 2008, 2, 86. [Google Scholar] [CrossRef] [Green Version]
- Gorban, A.N.; Zinovyev, A. Principal manifolds and graphs in practice: From molecular biology to dynamical systems. Int. J. Neural Syst. 2010, 20, 219–232. [Google Scholar] [CrossRef] [Green Version]
- Donoho, D.L. High-dimensional data analysis: The curses and blessings of dimensionality. AMS Math Challenges Lect. 2000, 1, 1–32. [Google Scholar]
- Gorban, A.N.; Tyukin, I.Y. Blessing of dimensionality: Mathematical foundations of the statistical physics of data. Philos. Trans. R. Soc. A Math. Phys. Eng. Sci. 2018, 376, 20170237. [Google Scholar] [CrossRef] [Green Version]
- Kainen, P.C.; Kůrková, V. Quasiorthogonal dimension of euclidean spaces. Appl. Math. Lett. 1993, 6, 7–10. [Google Scholar] [CrossRef] [Green Version]
- Tyukin, I.Y.; Higham, D.J.; Gorban, A.N. On Adversarial Examples and Stealth Attacks in Artificial Intelligence Systems. In Proceedings of the International Joint Conference on Neural Networks (IJCNN), Glasgow, UK, 19–24 July 2020; pp. 1–6. [Google Scholar]
- Gorban, A.N.; Grechuk, B.; Mirkes, E.M.; Stasenko, S.V.; Tyukin, I.Y. High-Dimensional Separability for One- and Few-Shot Learning. Entropy 2021, 23, 1090. [Google Scholar] [CrossRef] [PubMed]
- Amblard, E.; Bac, J.; Chervov, A.; Soumelis, V.; Zinovyev, A. Hubness reduction improves clustering and trajectory inference in single-cell transcriptomic data. bioRxiv 2021. [Google Scholar] [CrossRef]
- Gionis, A.; Hinneburg, A.; Papadimitriou, S.; Tsaparas, P. Dimension Induced Clustering. In KDD ’05: Proceedings of the Eleventh ACM SIGKDD International Conference on Knowledge Discovery in Data Mining; Association for Computing Machinery: New York, NY, USA, 2005; pp. 51–60. [Google Scholar] [CrossRef]
- Allegra, M.; Facco, E.; Denti, F.; Laio, A.; Mira, A. Data segmentation based on the local intrinsic dimension. Sci. Rep. 2020, 10, 1–12. [Google Scholar] [CrossRef] [PubMed]
- Grechuk, B.; Gorban, A.N.; Tyukin, I.Y. General stochastic separation theorems with optimal bounds. Neural Netw. 2021, 138, 33–56. [Google Scholar] [CrossRef]


| Method Name | Short Name(s) | Ref(s) | Valid Result | Insensitivity to Redundancy | Uniform ID Estimate in Similar Datasets | Performance with Many Observations | Performance with Many Features |
|---|---|---|---|---|---|---|---|
| PCA Fukunaga-Olsen | PCA FO, PFO | [15,22] | +++ | +++ | +++ | +++ | +++ |
| PCA Fan | PFN | [23] | +++ | +++ | +++ | +++ | +++ |
| PCA maxgap | PMG | [56] | +++ | + | +++ | +++ | |
| PCA ratio | PRT | [57] | +++ | +++ | + | +++ | +++ |
| PCA participation ratio | PPR | [57] | +++ | +++ | ++ | +++ | +++ |
| PCA Kaiser | PKS | [54,58] | +++ | − | +++ | +++ | +++ |
| PCA broken stick | PBS | [55,59] | +++ | +++ | +++ | +++ | |
| Correlation (fractal) dimensionality | CorrInt, CID | [18] | + | +++ | ++ | + | + |
| Fisher separability | FisherS, FSH | [4,32] | ++ | +++ | +++ | ++ | +++ |
| K-nearest neighbours | KNN | [27] | ++ | − | ++ | ||
| Manifold-adaptive fractal dimension | MADA, MDA | [19] | − | +++ | +++ | − | + |
| Minimum neighbor distance—ML | MIND_ML,MMk, MMi | [28] | +++ | +++ | ++ | ++ | + |
| Maximum likelihood | MLE | [25] | ++ | +++ | ++ | ++ | + |
| Methods of moments | MOM | [20] | +++ | +++ | +++ | ++ | + |
| Estimation within tight localities | TLE | [33] | +++ | +++ | ++ | + | |
| Minimal neighborhood information | TwoNN,TNN | [31] | ++ | +++ | +++ | ++ | +++ |
| Angle and norm concentration | DANCo,DNC | [29] | + | +++ | +++ | ||
| Expected simplex skewness | ESS | [56] | +++ | +++ | +++ |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Bac, J.; Mirkes, E.M.; Gorban, A.N.; Tyukin, I.; Zinovyev, A. Scikit-Dimension: A Python Package for Intrinsic Dimension Estimation. Entropy 2021, 23, 1368. https://doi.org/10.3390/e23101368
Bac J, Mirkes EM, Gorban AN, Tyukin I, Zinovyev A. Scikit-Dimension: A Python Package for Intrinsic Dimension Estimation. Entropy. 2021; 23(10):1368. https://doi.org/10.3390/e23101368
Chicago/Turabian StyleBac, Jonathan, Evgeny M. Mirkes, Alexander N. Gorban, Ivan Tyukin, and Andrei Zinovyev. 2021. "Scikit-Dimension: A Python Package for Intrinsic Dimension Estimation" Entropy 23, no. 10: 1368. https://doi.org/10.3390/e23101368
APA StyleBac, J., Mirkes, E. M., Gorban, A. N., Tyukin, I., & Zinovyev, A. (2021). Scikit-Dimension: A Python Package for Intrinsic Dimension Estimation. Entropy, 23(10), 1368. https://doi.org/10.3390/e23101368










