1. Introduction
String-matching processes are included in applications in many areas, like applications for information retrieval, information analysis, computational biology, multiple variations of practical software implementations in all operating systems, etc. String-matching forms the basis for other computer science fields, and it is one of the most researched areas in theory as well as in practice. An increasing amount and availability of textual data require the development of new approaches and tools to search useful information more effectively from such a large amount of data. Different string-matching algorithms perform better or worse, depending on the application domain, making it hard to choose the best one for any particular case [
1,
2,
3,
4,
5].
The main reason for analyzing an algorithm is to discover its features and compare them with other algorithms in a similar environment. When features are the focus, the mostly and primarily used parts are time and space resources and researchers want to know how long the implementation of a particular algorithm will run on a specific computer and how much space it will require. The implementation quality and compiler properties, computer architecture, etc., have huge effects on performance. Establishing differences between an algorithm and its implementation features can be challenging [
6,
7].
An algorithm is efficient if its resource consumption in the process of execution is below some acceptable or desirable level. The algorithm will end execution on an available computer in a reasonable amount of time, or space in efficiently acceptable contexts. Multiple factors can affect an algorithm’s efficiency, such as algorithm implementation, accuracy requirements, and lack of computational power. A few frameworks exist for testing string matching algorithms [
8]. Hume and Sunday presented a framework for testing string matching algorithms in 1991. It was developed in the C programming language, and it has been used in the 90′. [
9] Faro presented the String Matching Algorithm Research Tool (SMART) framework in 2010 and its improved version six years later. SMART is a framework designed to develop, test, compare, and evaluate string matching algorithms [
3].
This paper introduces a model-building methodology for selecting the most efficient string search algorithm, based on a pattern entropy while expressing the algorithm’s efficiency using platform independent metrics. According to their efficiency, the developed methodology for ranking algorithms considers properties of the searched string and properties of the texts that are being searched. This methodology does not depend on algorithm implementation, computer architecture, programming languages specifics, and it provides a way to investigate algorithm strengths and weaknesses. More information about the formal metric definition is described in
Section 3.1.3. The paper covers only the fundamental algorithms. Selected algorithms are the basis for the most state-of-the-art algorithms, which belong to the classical approach. More information is described in
Section 3.2.2. We also analyzed different approaches, metrics for measuring algorithms’ efficiency, and algorithm types.
The paper is organized as follows: The basic concepts of string-matching are described in
Section 2. Entropy, formal metrics, and related work are described in
Section 3 along with the proposed methodology. Experimental results with string search algorithms evaluation models developed according to the proposed methodology are presented in
Section 4.
Section 5 presents the validation results for the developed models and a discussion.
Section 6 concludes this paper.
2. String-Matching
String-matching consists of finding all occurrences of a given string in a text. String-matching algorithms are grouped into exact and approximate string-matching algorithms. Exact string-matching does not allow any tolerance, while approximate string-matching allows some tolerance. Further, exact string-matching algorithms are divided into two groups: single pattern matching and multiple pattern matching. These two categories are also divided into software and hardware-based methods. The software-based string- matching algorithms can be divided into character comparison, hashing, bit-parallel, and hybrid approaches. This research focuses on on-line software-based exact string-matching using a character comparison approach. On-line searching means that there is no built data structures in the text. The character-based approach is known as a classical approach that compares characters to solve string-matching problems [
10,
11].
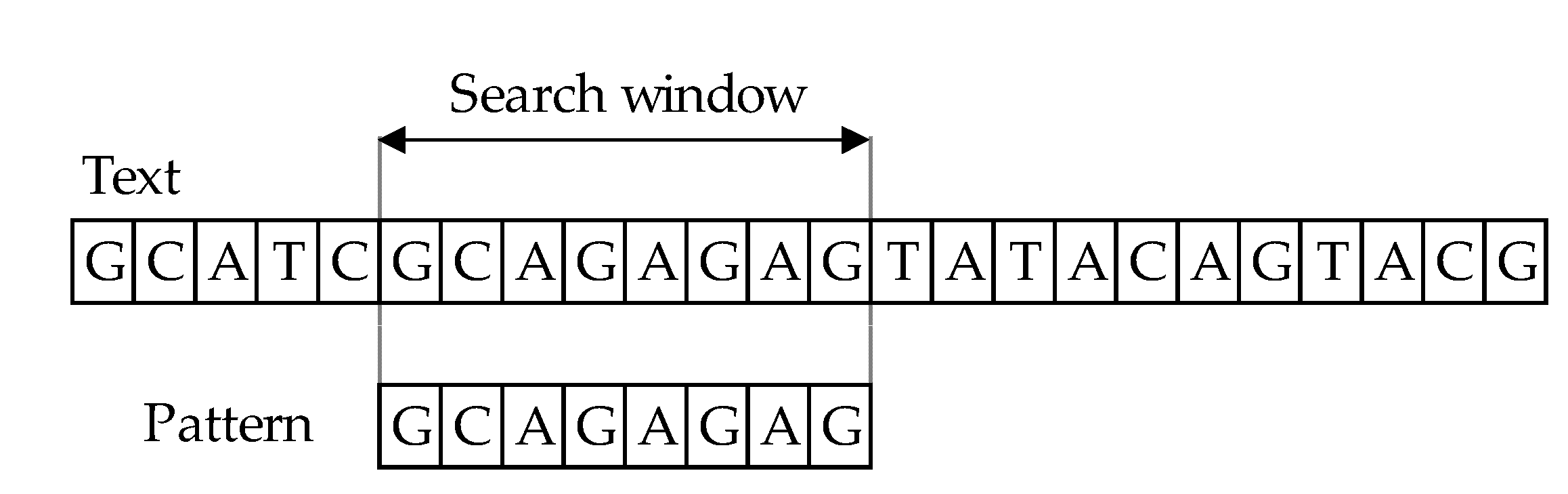
The character-based approach has two key stages: matching and shift phases. The principle behind algorithms for string comparison covers text scanning with the window of size 𝑚, commonly referred to as the sliding window mechanism (or search window). In the process of comparing the main text
T [1…n] and a pattern
P [1…m], where
m ≤ n, the aim is to find all occurrences, if any, of the exact pattern 𝑃 in the text 𝑇 (
Figure 1). The result of comparing patterns with text is information that they match if they are equal or they mismatch. The length of both windows must be of equal in length, during the comparison phase. First, one must align the window and the text’s left end and then compare the characters from the window with the pattern’s characters. After an exact matching (or mismatch) of pattern with the text, the window is moved to the right. The same procedure repeats until the right end of the window has reached the right end of the text [
11,
12,
13,
14,
15].
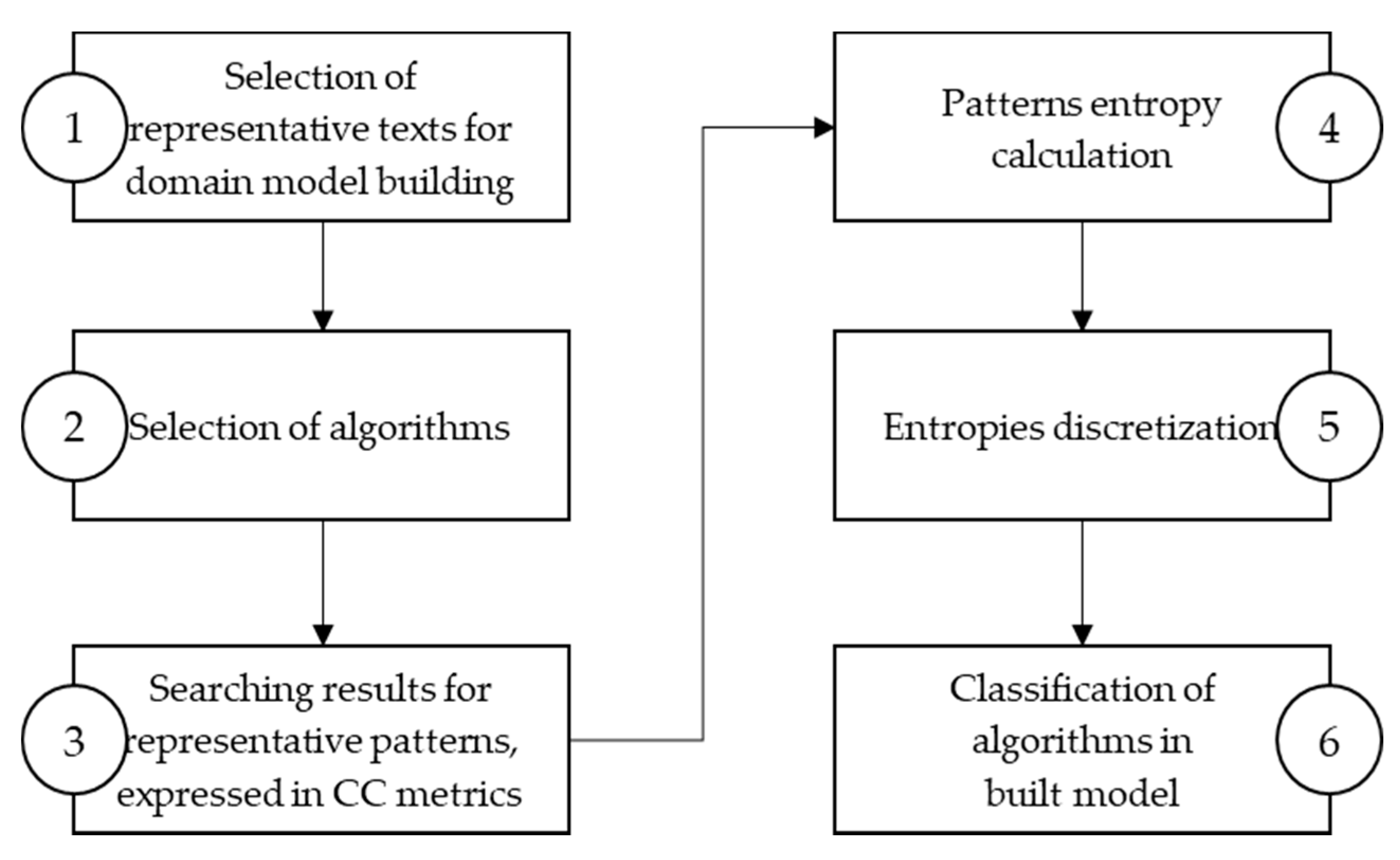
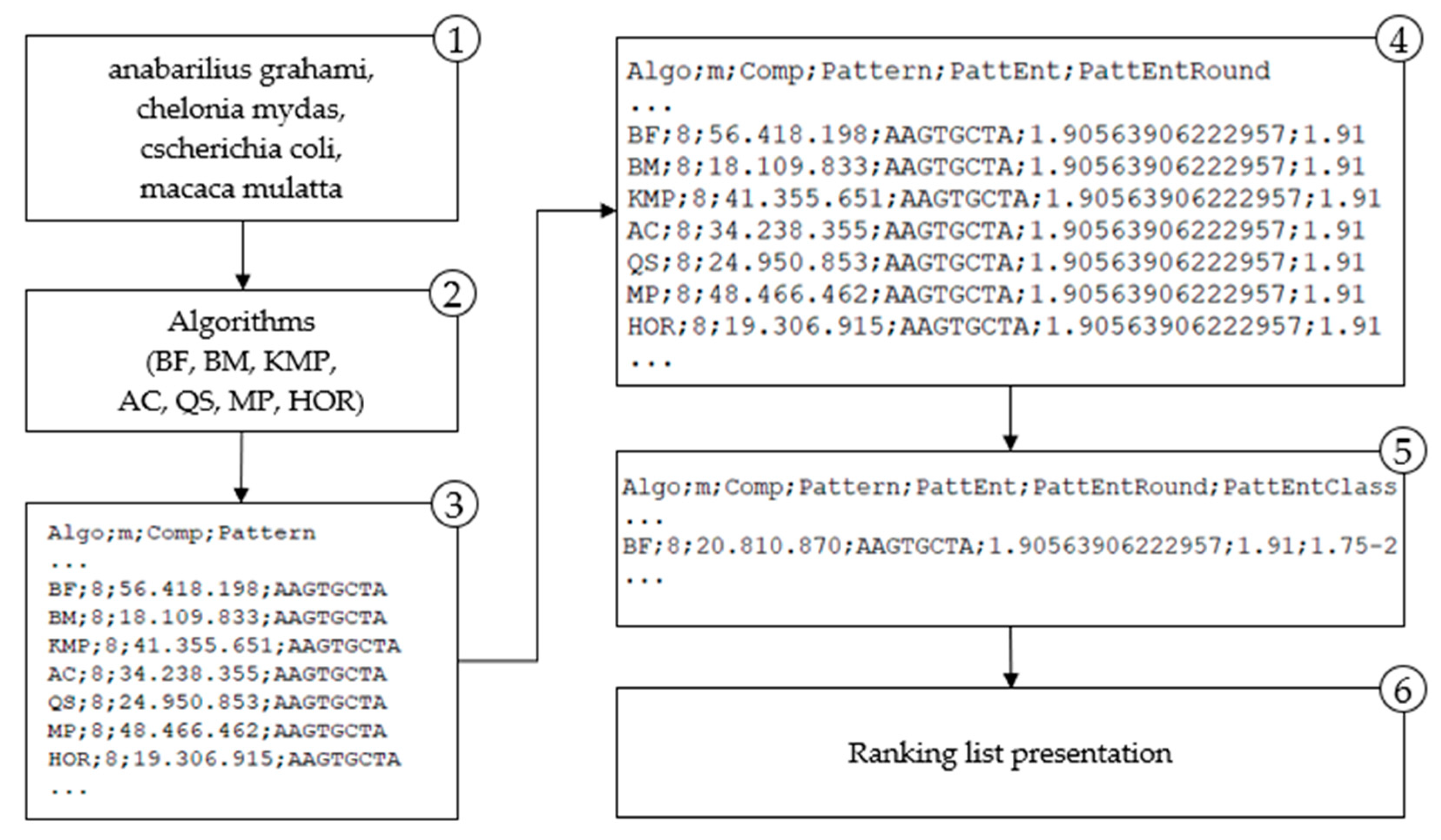
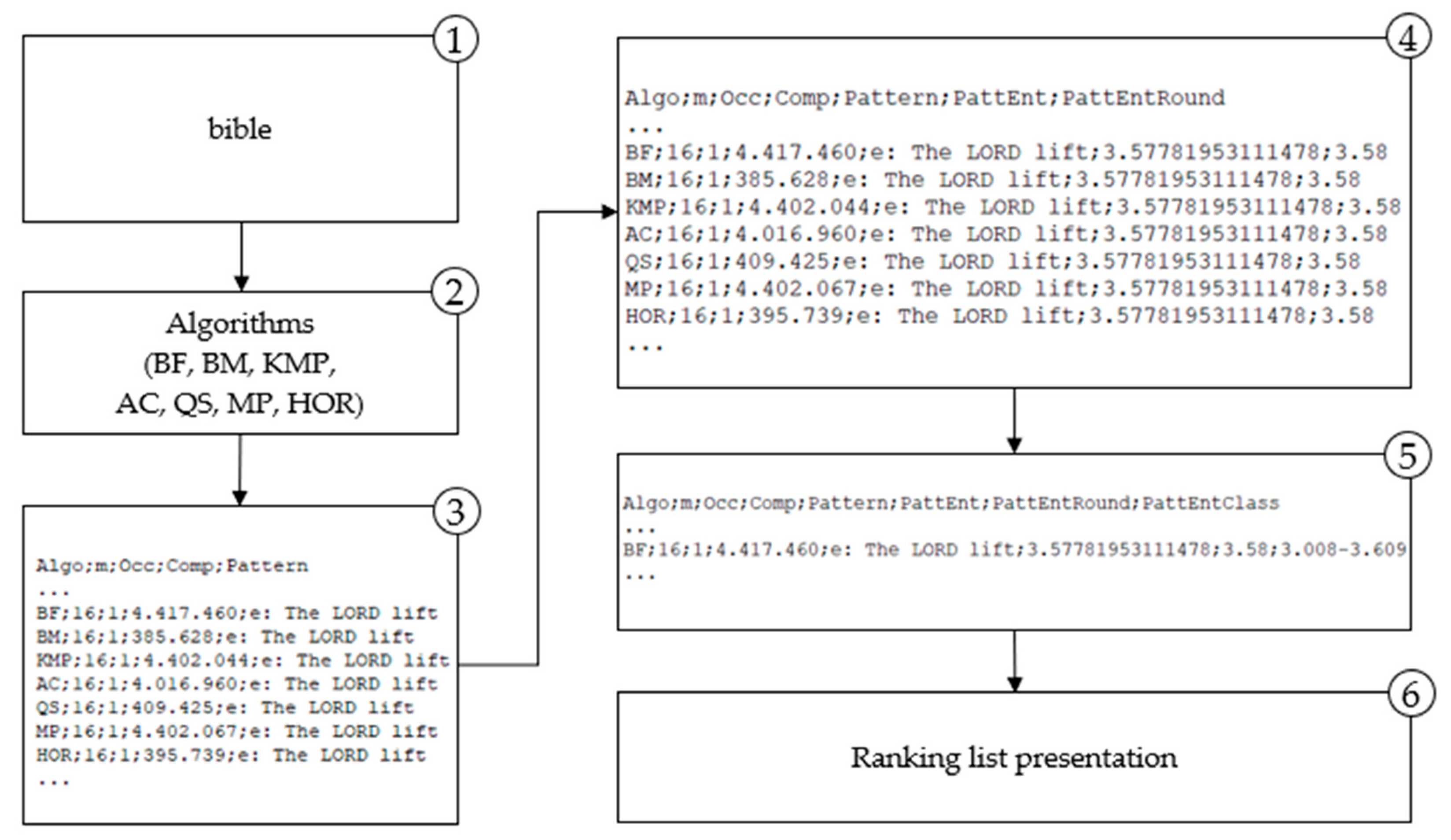
4. Classification of Algorithms in the Built Model
The algorithm analysis results integrated into a model, provide a ranking list of algorithms by their efficiency, measured with character comparison metrics, correlated with the searched pattern entropy. More efficient algorithms perform fewer characters comparison when finding a pattern in a text. The model proposes a more efficient algorithm for string matching for a given pattern based on the entropy class to which the observed pattern belongs.
The results presented in
Table 5 and
Table 6 give a ranking list of the selected algorithms grouped by the entropy class. The percentages shown in the result tables represent a proportion of pattern searching results for a particular algorithm, which might be smaller or greater than the remaining algorithms inside the quartile. For example, in
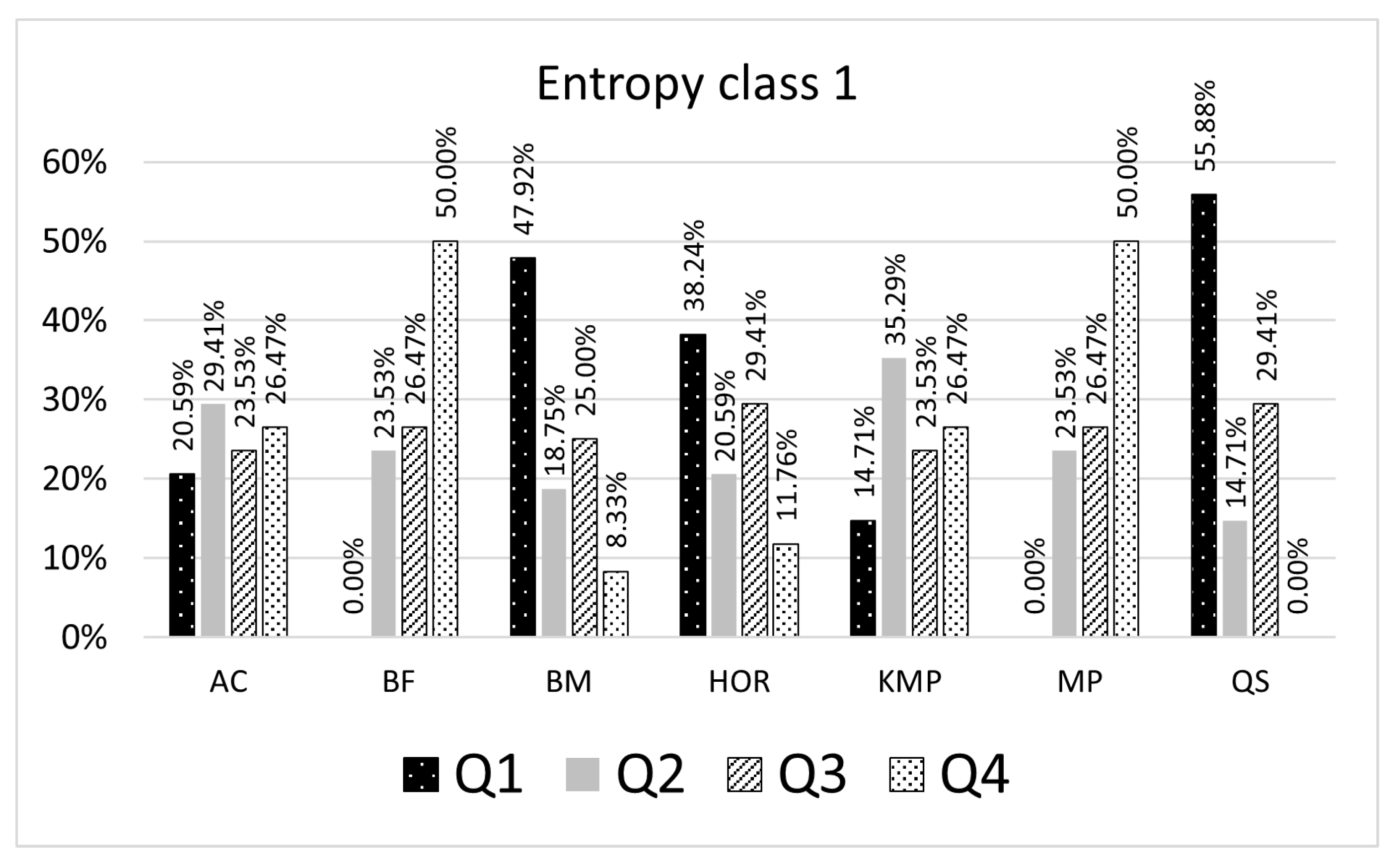
Table 5, if the searched pattern belongs to the entropy class 1 (number of representative patterns is 9), 55.88% of the searching results for a given entropy class with the QS algorithm are in the first quartile, 14.71% are in the second quartile, 29.41% are in the third quartile (
Figure 5). When patterns are searched with the BM algorithm, 47.92% of the searching results expressed as CC count is in the first quartile, 23.53% are in the second quartile, and 25% are in the third, and 8.33% are in the fourth quartile. In this case, for a given pattern, the built model suggests using the QS algorithm as the most efficient algorithm. The selected algorithm is considered an optimal algorithm that will make fewer character comparisons (CC) than others for most patterns being searched belonging to the entropy class 1.
In
Table 5, for the entropy class 8 (number of representative patterns searched is 1451), the model shows that the BM algorithm is the most efficient. In 61.95% of cases for patterns in the entropy class 8, the BM algorithm made the least characters comparison versus the other six algorithms evaluated with the model. In 24.38% cases, BM was second best; in 13.68% cases was the third and never was the worse.
In
Table 6, for example, for the entropy class 6 (number of representative patterns searched is 393), the model shows that the QS algorithm is the most efficient. In 70.13% of cases for patterns in the entropy class 6, the QS algorithm made the least characters comparison versus the other six algorithms evaluated with the model. In 29.87% of cases, QS was second best and never was the worse. For the entropy class 7 (number of representative patterns searched is 283), the model shows that the most efficient is the BM algorithm. In 65.02% of cases for patterns in the entropy class 7, the BM algorithm made the least characters comparison versus the other six algorithms evaluated with the model. In 34.98% of cases, BM was second best and never was the worse.
5. Methodology Validation and Discussion
For model validation, the seventh and ninth entropy classes (961 and 4692 patterns) were selected for the DNA domain, and the sixth (393 patterns) and ninth classes (221 patterns) were selected for the natural language domain. The model classes chosen for validation have the highest number of representative patterns and are characteristic for the specific domains.
The selected patterns for validation are not part of the patterns set with which the model was created. For the DNA domain model, also a different text is chosen for validation. The DNA domain model is validated with the DNA sequence
Homo sapiens isolate HG00514 chromosome 9 genomic scaffold HS_NIOH_CHR9_SCAFFOLD_1, whole genome shotgun sequence, 43.213.237 bp, 39 Mb as the text [
58]. The natural language domain is validated with the natural language text set from the Canterbury Corpus. [
43]
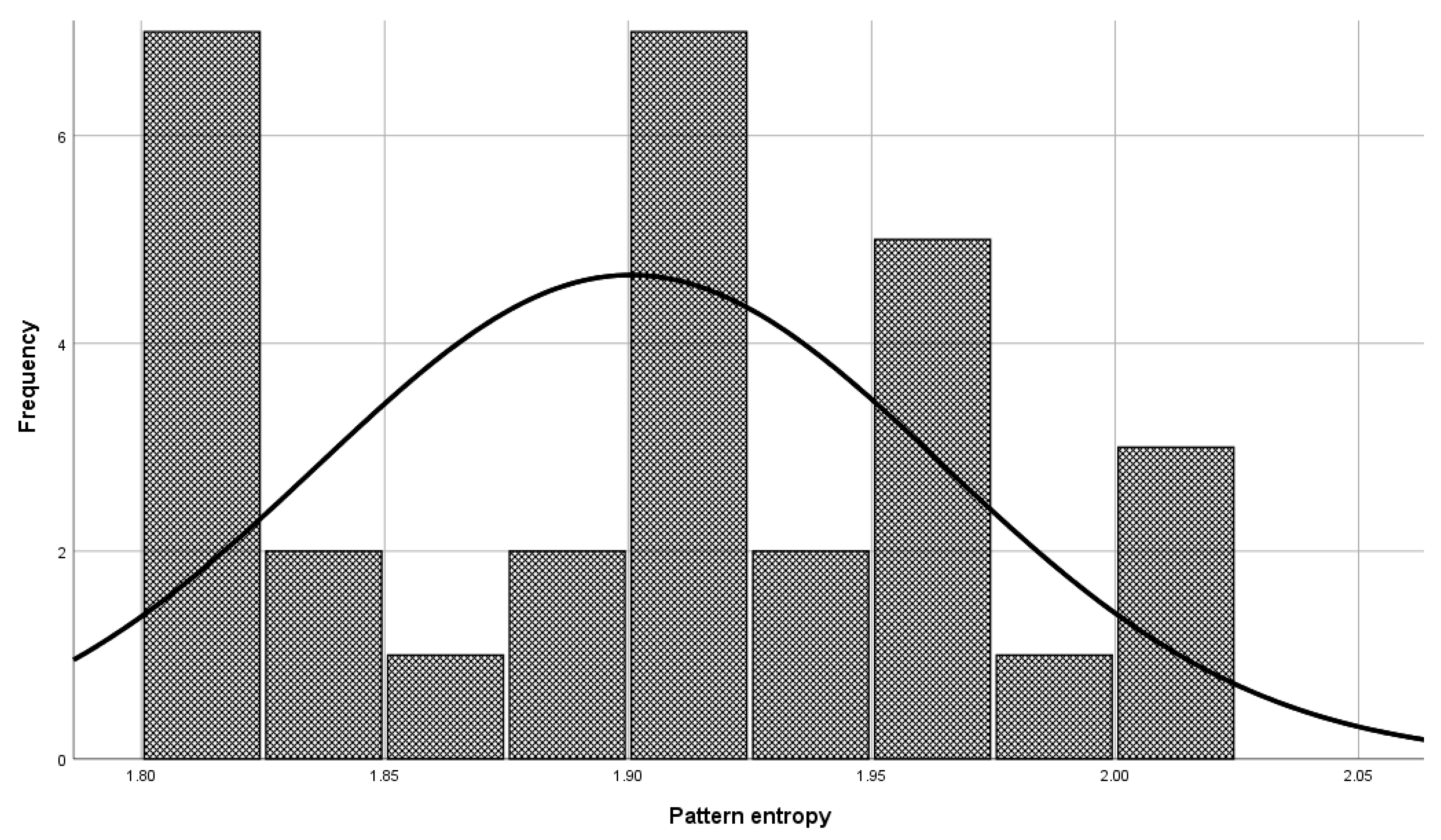
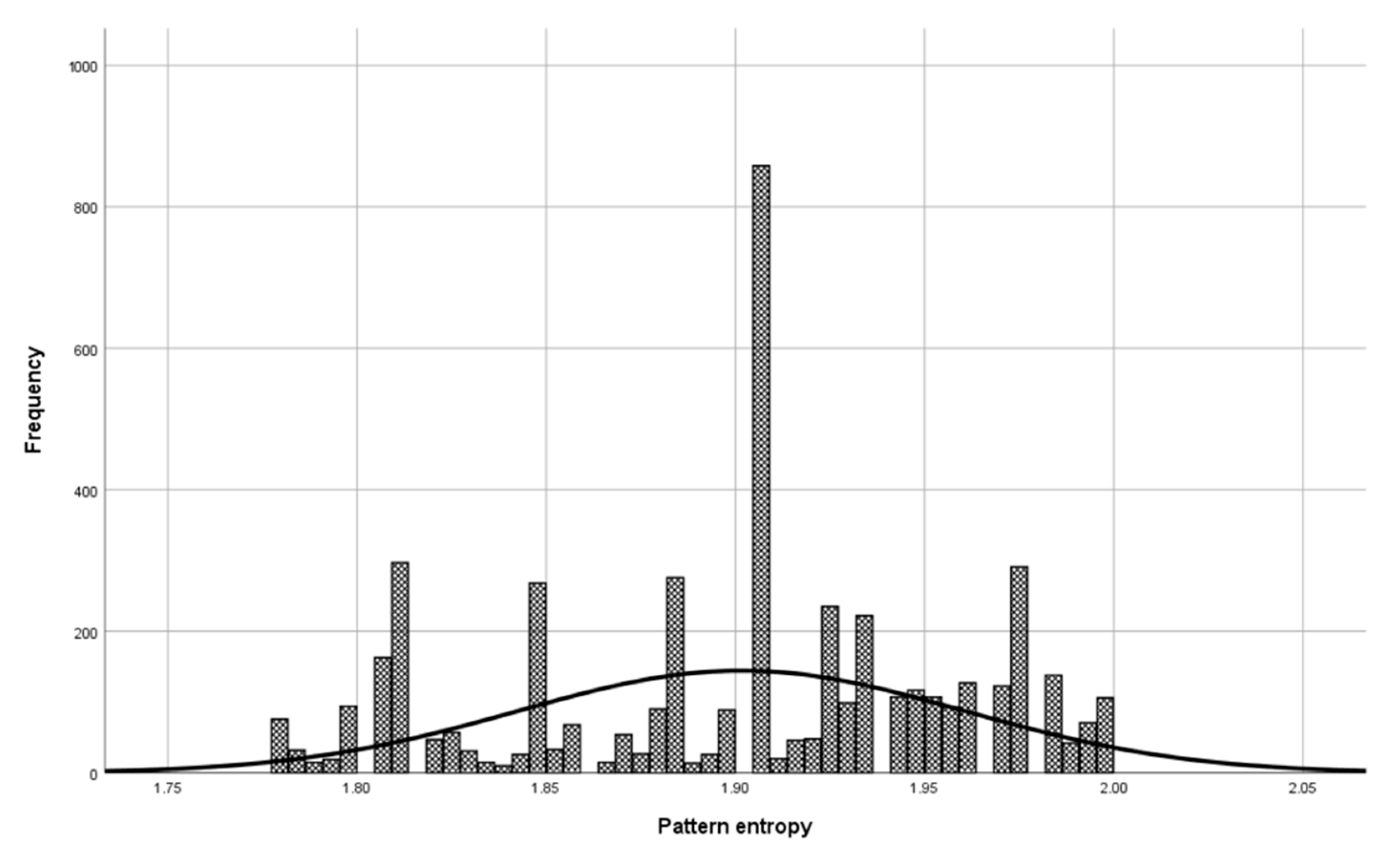
Before the model validation process, a check was made to see if the selected patterns were sufficiently representative for model validation. The check was done with the central limit theorem. The set of patterns used in the validation phase has a normal distribution (
Figure 6, Mean = 1.900, and Std. Dev = 0.064) as a set of patterns used in model building (
Figure 7, Mean = 1.901, and Std. Dev = 0.059), which means that patterns used to validate the model represent a domain.
Other entropy classes of patterns discretized character comparisons also follow the normal distribution. The basis in the model validation phase is to verify if the test results differ from the developed model results presented in
Table 5 and
Table 6.
For comparing the two data sets (model results and test results), double-scaled Euclid distance, and Pearson correlation coefficient were used.
Double-scaled Euclidian distance normalizes raw Euclidian distance into a range of 0–1, where 1 represents the maximum discrepancy between the two variables. The first step in comparing two datasets with double-scaled Euclidian methods is to compute the maximum possible squared discrepancy (
md) per variable
i of
v variables, where
v is the number of observed variables in the data set. The
mdi = (Maximum for variable i-Minimum for variable i)2, where 0 (0%) is used for minimum and 1 (100%) for maximum values for double-scaled Euclid distance. The second step’s goal is to produce the scaled variable Euclidean distance, where the sum of squared discrepancies per variable is divided by the maximum possible discrepancy for that variable, Equation (7):
The final step is dividing scaled Euclidian distance with the root of
v, where
v is the number of observed variables, Equation (8). Double-scaled Euclid distance easily turns into a measure of similarity by subtracting it from 1.0. [
16,
59,
60,
61,
62]:
Table 7 shows the usage of the double-scaled Euclidian distance method for entropy class 7 of DNA
Applying Equation (8) on
Table 7, column “Scaled Euclidean (
d1)”, gives a double-scaled Euclidian distance of 0.227. Subtracting double-scaled Euclidian Distance from 1 gives a similarity coefficient of 0.773 or 77%.
Table 8 shows the results of the calculated double-scaled Euclid distance and corresponding similarity coefficient.
Converting double-scaled Euclidian distance to a context of similarity, it is possible to conclude that the built model is similar to the validation results with a high degree of similarity. The seventh and ninth classes from the built model for the DNA domain have a similarity coefficient with their validation results of 77%. The high percentage of similarity also has the sixth and ninth classes from the built model for the natural language domain with their validation results of 80% and 86%. The results for validated classes obtained in the validation process are extremely similar to the results from the built model. A proportion of searched pattern character comparisons for a particular algorithm inside the quartile is similar to the built model.
Pearson’s correlation coefficient is used to check the correlation between data from the model and data from the validation phase. Pearson correlation coefficients per classes are shown in
Table 9.
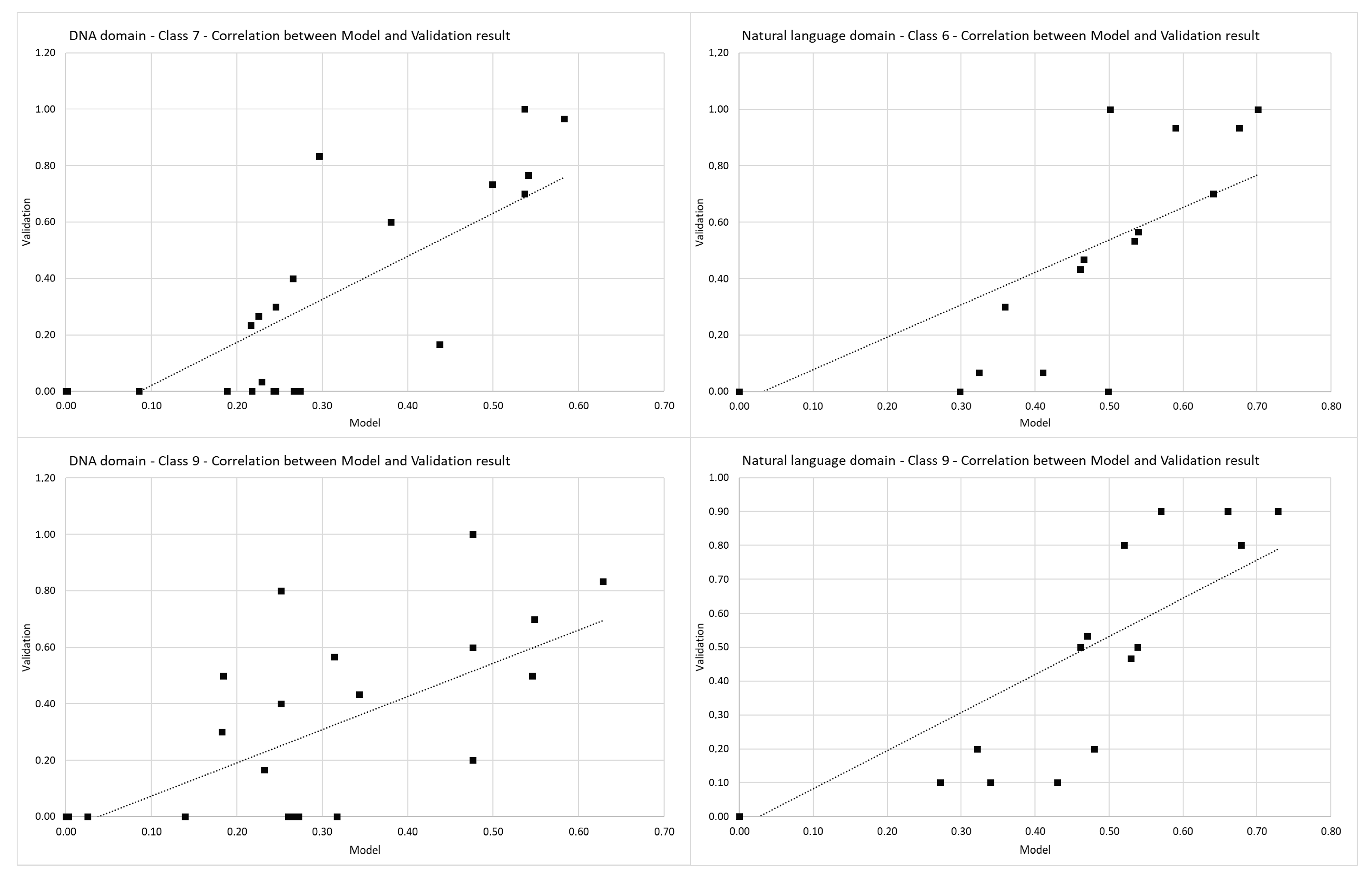
The seventh and ninth classes from the built model for the DNA domain have a linear Pearson’s correlation coefficient with their validation results. The sixth and ninth classes from the natural language domain’s built model have a linear Pearson’s correlation coefficient with their validation results. Pearson’s correlation coefficient shown in
Figure 8 indicate that the values from the built model (
x-axis, Model) and their corresponding validation result (
y-axis, validation) follow each other with a strong positive relationship.
Using the double-scaled Euclidean distance in the validation process shows a strong similarity between the built model and validation results. In addition to the similarity, a strong positive relationship exists between classes selected from the built model and validation results proven by Pearson’s correlation coefficient. Presented results show that it is possible to use the proposed methodology to build a domain model for selecting an optimal algorithm for the exact string matching. Except for optimal algorithm selection for a specific domain, this methodology can be used to improve the efficiency of string- matching algorithms in the context of performance, which is in correlation with empirical measurements.
The data used to build and validate the model can be downloaded from the website [
63].
6. Conclusions
Proposed methodology for ranking algorithms is based on properties of the searched string and properties of the texts being searched. Searched strings are classified according to the pattern entropy. This methodology is expressing algorithms efficiency using platform independent metrics thus not depending on algorithm implementation, computer architecture or programming languages characteristics. This work focuses on classical software-based algorithms that use exact string-matching techniques with a character comparison approach. For any other type of algorithms, this methodology cannot be used. The used character comparisons metrics is platform-independent in the context of formal approaches, but the number of comparisons directly affects the time needed for algorithm execution and usage of computational resource. Studying the methodology, complexity, and limitations of all available algorithms is a complicated and long-term task. The paper discusses, in detail, available metrics for string searching algorithms properties evaluation and proposing a methodology for building a domain model for selecting an optimal string searching algorithm. The methodology is based on presenting exact string-matching results to express algorithm efficiency regardless of query pattern length and dataset size. We considered the number of compared characters of each algorithm expressed by the searched string entropy for our baseline analysis. High degrees of similarity and a strong correlation between the validation results and the built model data have been proven, making this methodology a useful tool that can help researchers choose an efficient string- matching algorithm according to the needs and choose a suitable programming environment for developing new algorithms. Everything that is needed is a pattern from a specific domain by which the model is built, and the model will suggest using the most optimal algorithm for usage. The defined model finally selects the algorithm that will most likely run up the least character comparison count in pattern matching. This research does not intend to evaluate the algorithm logic and programming environment in any way; the main reason for comparing the results of algorithms is the construction of the algorithm selection model. The built model is straightforwardly extendable with other algorithms; all required is adequate training data sets. Further research is directed to find additional string characteristics, besides pattern entropy, that can enhance developed methodology precision for selecting more efficient string search algorithms.