Identifying Influencers in Social Networks
Abstract
1. Introduction
2. Related Works
3. Modeling and Methods
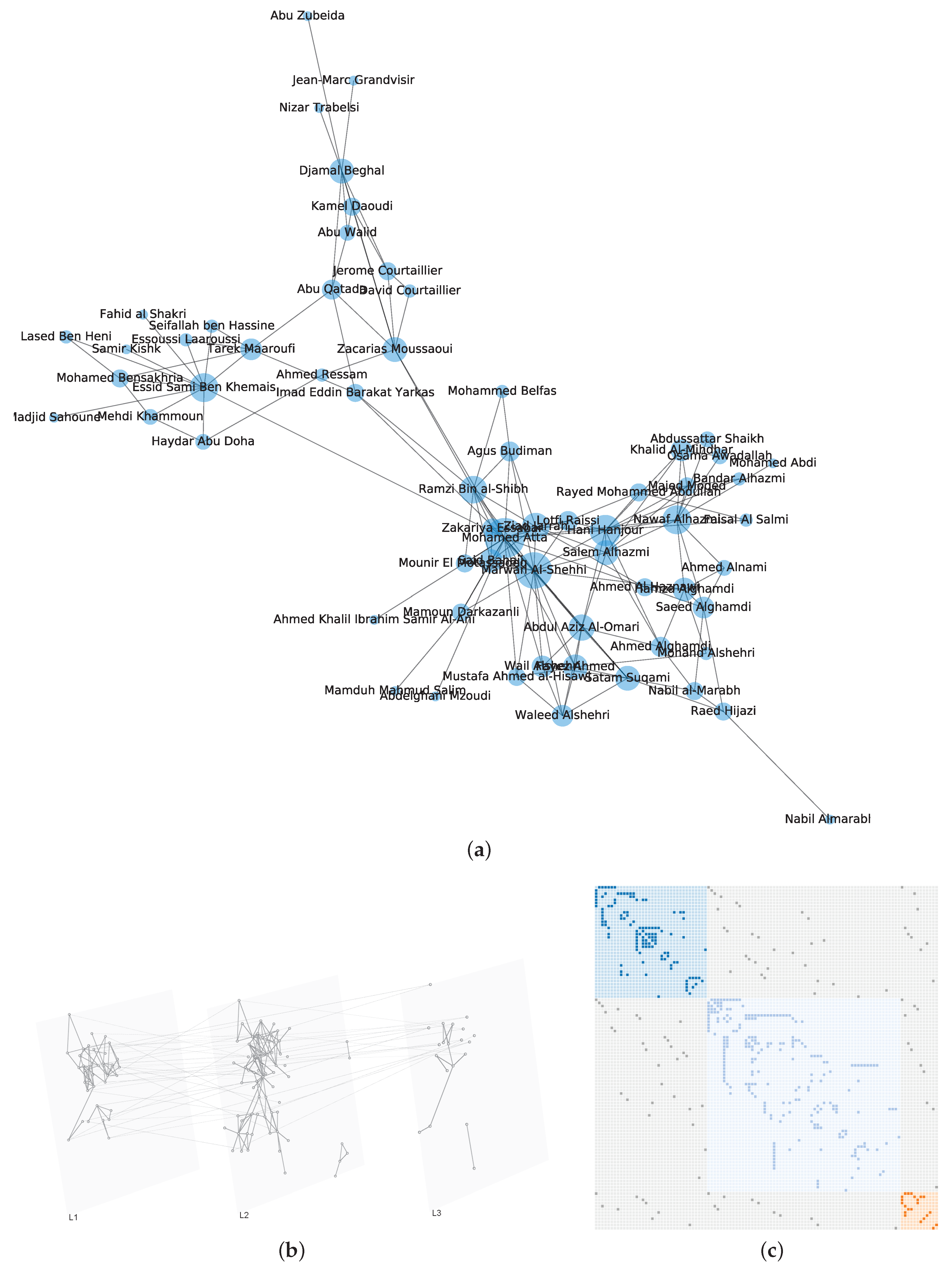
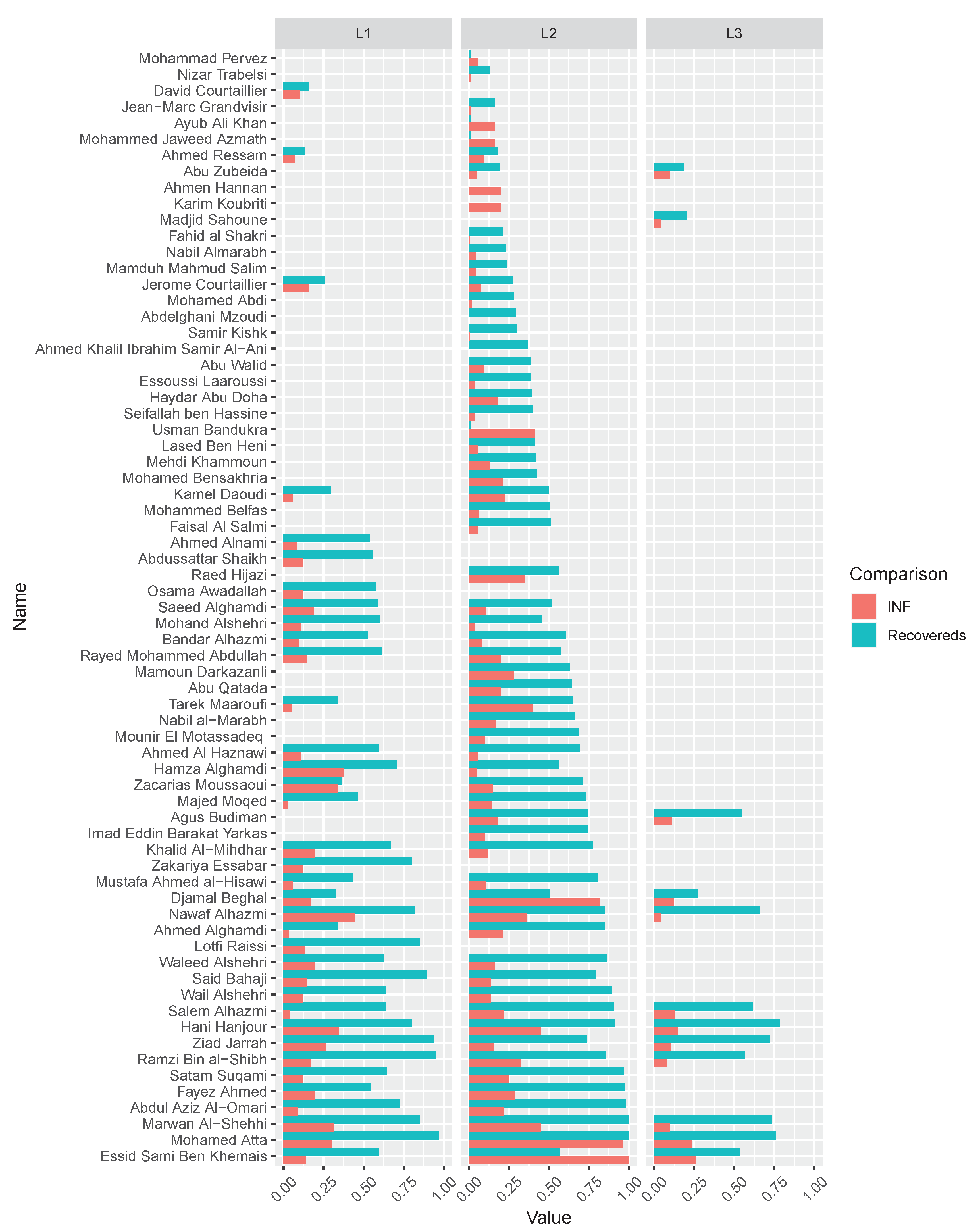
3.1. Network Modeling
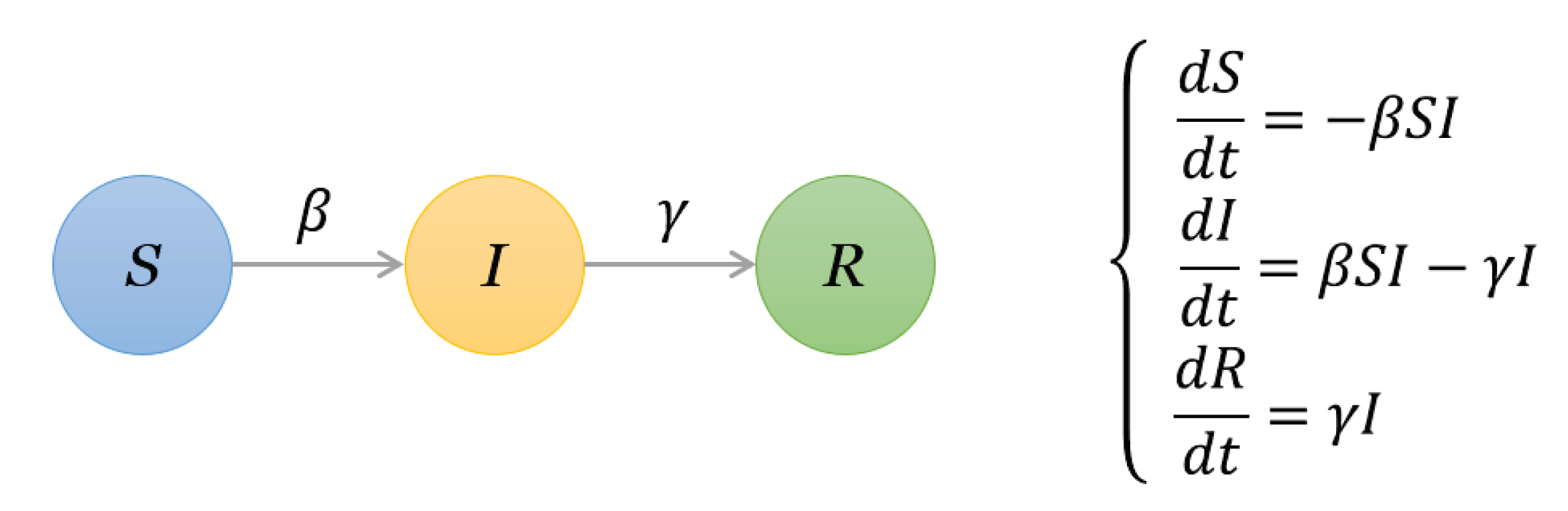
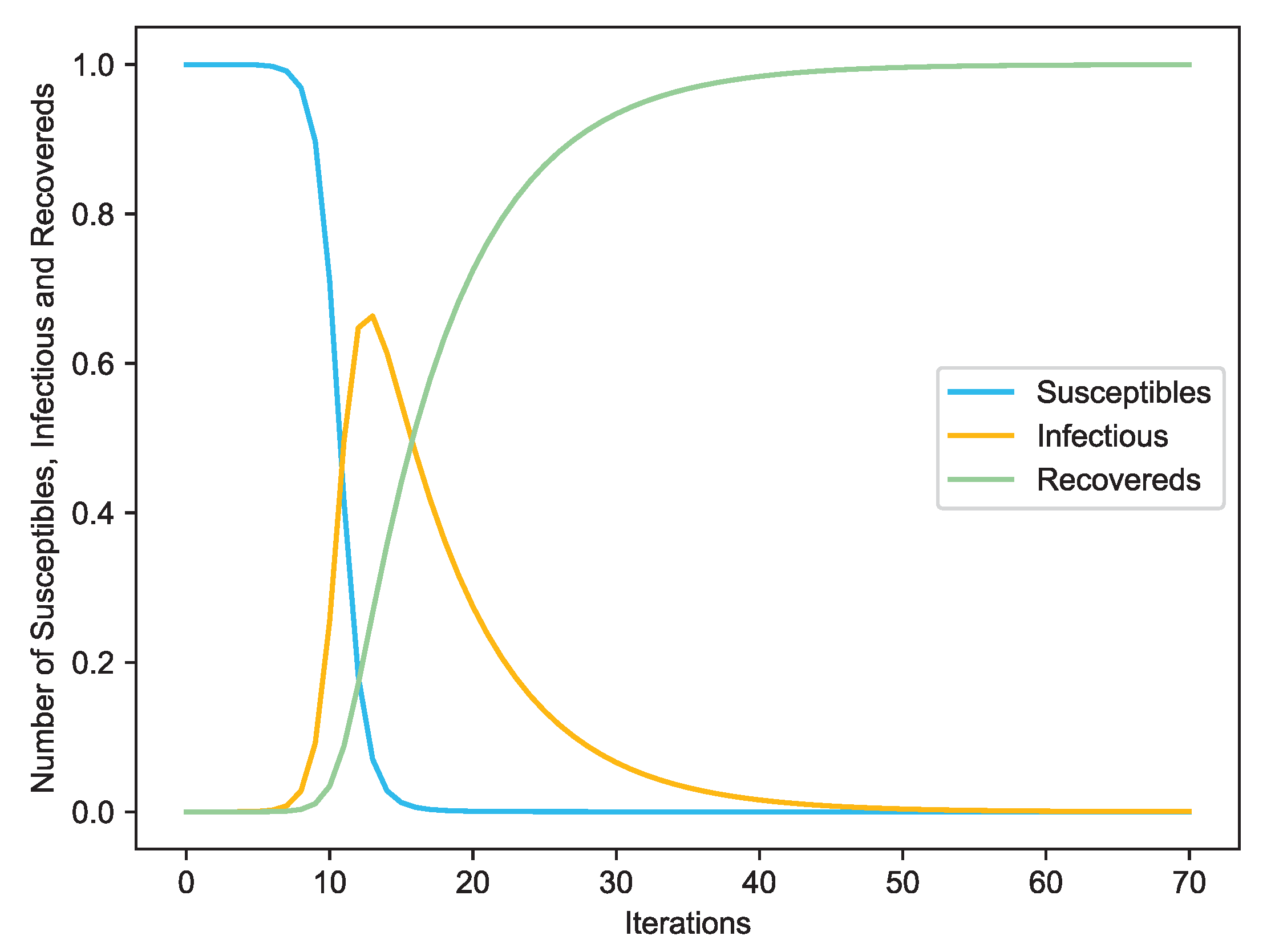
3.2. Methods
- Susceptible (S) state, where a node is vulnerable to infection.
- Infectious (I) state, where a node tries to infect its susceptible neighbors.
- Recovered (R) state, where a node has recovered (or isolated) and can no longer infect others.
3.3. Complexity Analysis
4. Experiments and Discussion
4.1. Experimental Datasets
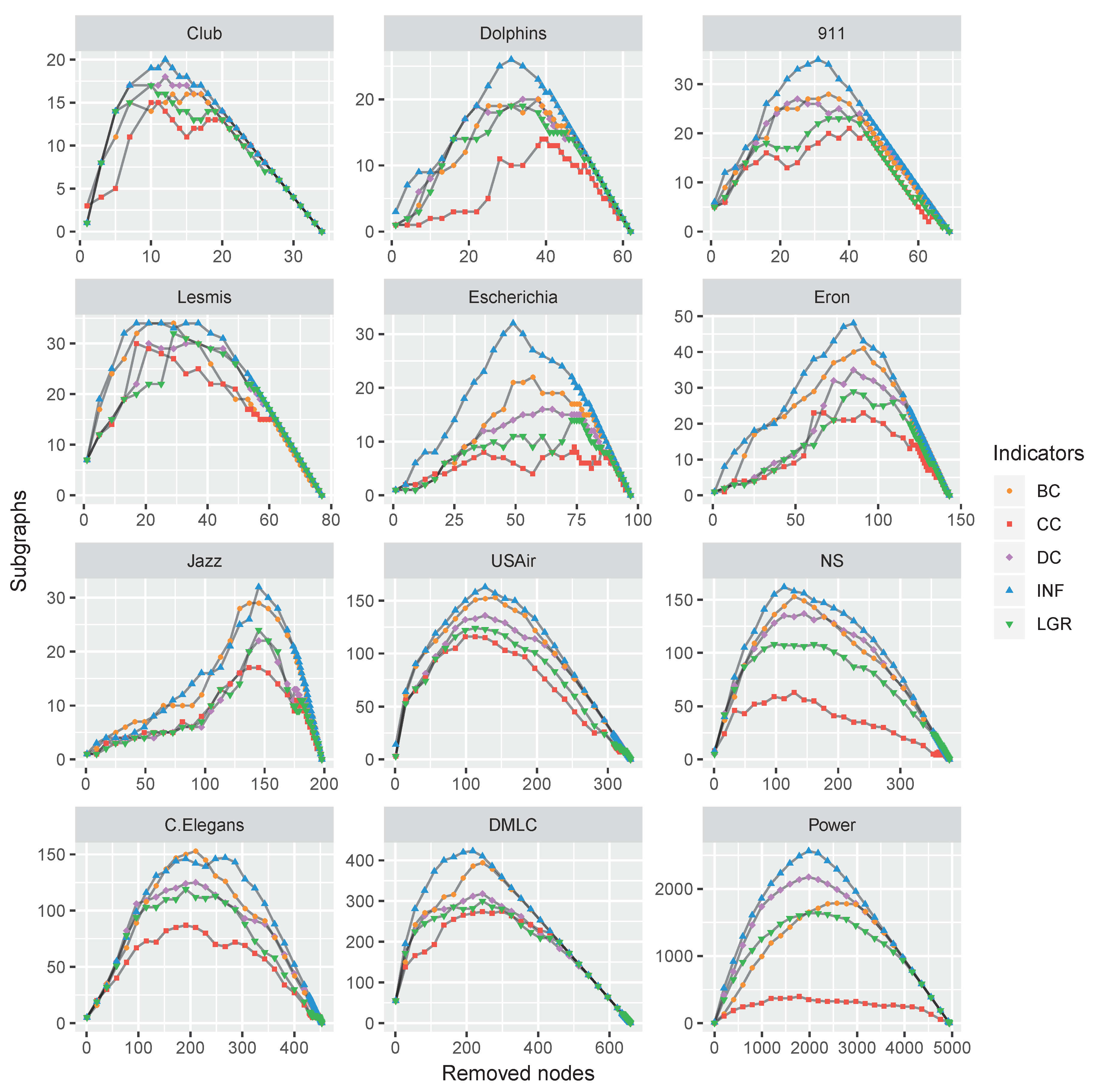
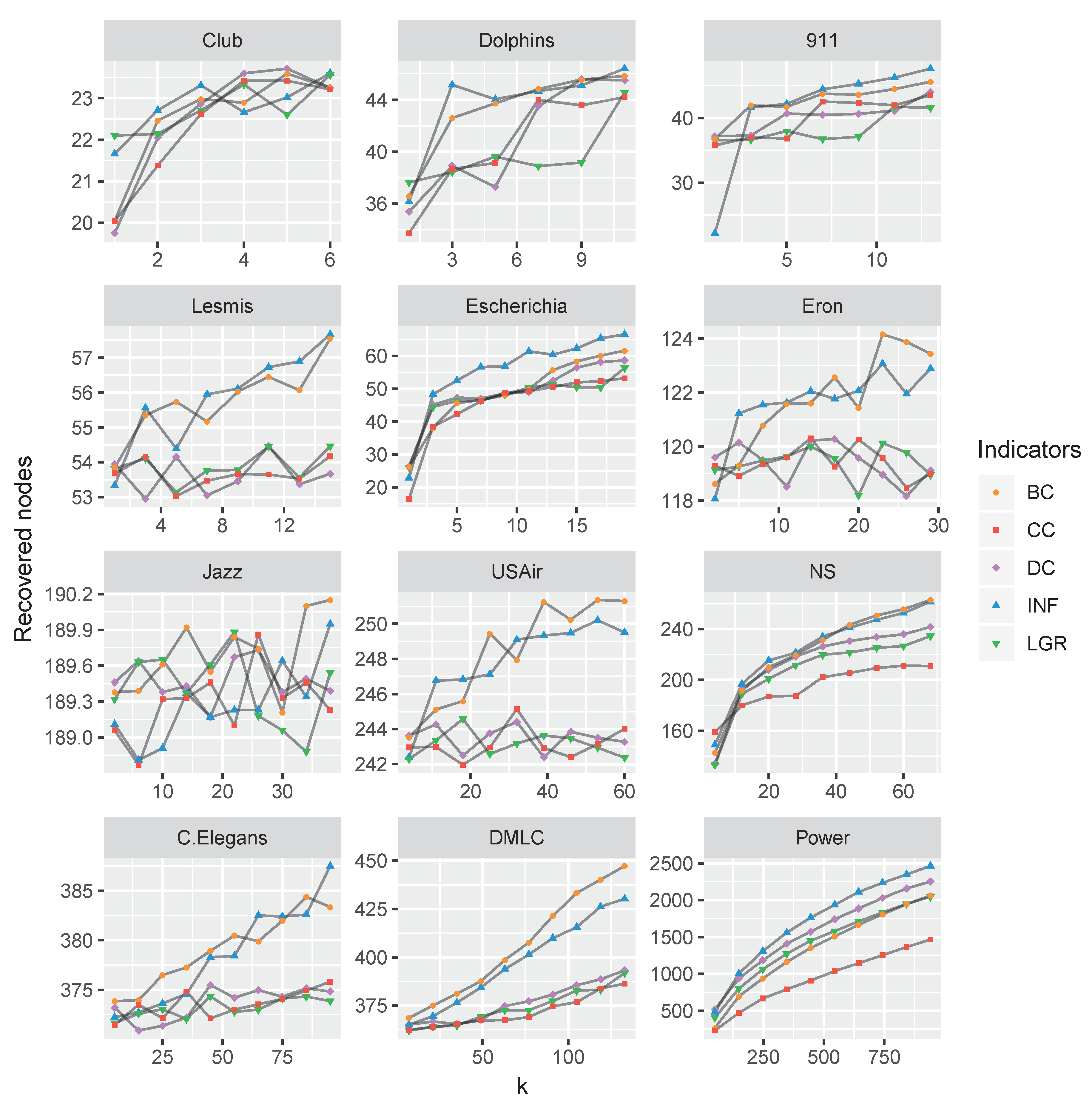
4.2. Performance Comparison
4.3. Discussion
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Aldecoa, R.; Marín, I. Surprise maximization reveals the community structure of complex networks. Sci. Rep. 2013, 3, 1–9. [Google Scholar] [CrossRef]
- Pei, S.; Muchnik, L.; Andrade, J.S., Jr.; Zheng, Z.; Makse, H.A. Searching for superspreaders of information in real-world social media. Sci. Rep. 2014, 4, 5547. [Google Scholar] [CrossRef] [PubMed]
- Xu, Z.; Rui, X.; He, J.; Wang, Z.; Hadzibeganovic, T. Superspreaders and superblockers based community evolution tracking in dynamic social networks. Knowl.-Based Syst. 2019, 105377. [Google Scholar] [CrossRef]
- Malliaros, F.D.; Rossi, M.E.G.; Vazirgiannis, M. Locating influential nodes in complex networks. Sci. Rep. 2016, 6, 19307. [Google Scholar] [CrossRef] [PubMed]
- Salavati, C.; Abdollahpouri, A.; Manbari, Z. BridgeRank: A novel fast centrality measure based on local structure of the network. Phys. A Stat. Mech. Appl. 2018, 496, 635–653. [Google Scholar] [CrossRef]
- Huang, X.; Chen, D.; Ren, T. Social network coalescence based on multilayer network model. J. Nonlinear Convex Anal. 2019, 20, 1465–1474. [Google Scholar]
- Wang, D.; Wang, H.; Zou, X. Identifying key nodes in multilayer networks based on tensor decomposition. Chaos Interdiscip. J. Nonlinear Sci. 2017, 27, 063108. [Google Scholar] [CrossRef]
- Kivelä, M.; Arenas, A.; Barthelemy, M.; Gleeson, J.P.; Moreno, Y.; Porter, M.A. Multilayer networks. J. Complex Netw. 2014, 2, 203–271. [Google Scholar] [CrossRef]
- Boccaletti, S.; Bianconi, G.; Criado, R.; Del Genio, C.I.; Gómez-Gardenes, J.; Romance, M.; Sendina-Nadal, I.; Wang, Z.; Zanin, M. The structure and dynamics of multilayer networks. Phys. Rep. 2014, 544, 1–122. [Google Scholar] [CrossRef]
- Rodrigues, F.A. Network centrality: an introduction. In A Mathematical Modeling Approach from Nonlinear Dynamics to Complex Systems; Springer: Berlin, Germany, 2019; pp. 177–196. [Google Scholar]
- Peng, S.; Zhou, Y.; Cao, L.; Yu, S.; Niu, J.; Jia, W. Influence analysis in social networks: A survey. J. Netw. Comput. Appl. 2018, 106, 17–32. [Google Scholar] [CrossRef]
- Grover, A.; Leskovec, J. node2vec: Scalable feature learning for networks. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco, CA, USA, 13–17 August 2016; pp. 855–864. [Google Scholar]
- Jeong, H.; Mason, S.P.; Barabási, A.L.; Oltvai, Z.N. Lethality and centrality in protein networks. Nature 2001, 411, 41. [Google Scholar] [CrossRef] [PubMed]
- Kitsak, M.; Gallos, L.K.; Havlin, S.; Liljeros, F.; Muchnik, L.; Stanley, H.E.; Makse, H.A. Identification of influential spreaders in complex networks. Nat. Phys. 2010, 6, 888. [Google Scholar] [CrossRef]
- Albert, R.; Albert, I.; Nakarado, G.L. Structural vulnerability of the North American power grid. Phys. Rev. E 2004, 69, 025103. [Google Scholar] [CrossRef] [PubMed]
- Liu, Y.Y.; Slotine, J.J.; Barabási, A.L. Controllability of complex networks. Nature 2011, 473, 167. [Google Scholar] [CrossRef]
- Yan, G.; Zhou, T.; Hu, B.; Fu, Z.Q.; Wang, B.H. Efficient routing on complex networks. Phys. Rev. E 2006, 73, 046108. [Google Scholar] [CrossRef]
- Xie, W.; Yu, W.; Zou, X. Diversity-maintained differential evolution embedded with gradient-based local search. Soft Comput. 2013, 17, 1511–1535. [Google Scholar] [CrossRef]
- Bonacich, P. Factoring and weighting approaches to status scores and clique identification. J. Math. Sociol. 1972, 2, 113–120. [Google Scholar] [CrossRef]
- McPherson, M.; Smith-Lovin, L.; Cook, J.M. Birds of a feather: Homophily in social networks. Annu. Rev. Sociol. 2001, 27, 415–444. [Google Scholar] [CrossRef]
- Borgatti, S.P. Centrality and network flow. Soc. Netw. 2005, 27, 55–71. [Google Scholar] [CrossRef]
- Freeman, L.C. A set of measures of centrality based on betweenness. Sociometry 1977, 40, 35–41. [Google Scholar] [CrossRef]
- Freeman, L.C. Centrality in social networks conceptual clarification. Soc. Netw. 1978, 1, 215–239. [Google Scholar] [CrossRef]
- Bonacich, P. Power and centrality: A family of measures. Am. J. Sociol. 1987, 92, 1170–1182. [Google Scholar] [CrossRef]
- Brin, S.; Page, L. The anatomy of a large-scale hypertextual web search engine. Comput. Netw. ISDN Syst. 1998, 30, 107–117. [Google Scholar] [CrossRef]
- Watts, D.J.; Strogatz, S.H. Collective dynamics of ‘small-world’networks. Nature 1998, 393, 440. [Google Scholar] [CrossRef] [PubMed]
- Barrat, A.; Barthelemy, M.; Pastor-Satorras, R.; Vespignani, A. The architecture of complex weighted networks. Proc. Natl. Acad. Sci. USA 2004, 101, 3747–3752. [Google Scholar] [CrossRef] [PubMed]
- Chen, D.B.; Gao, H.; Lü, L.; Zhou, T. Identifying influential nodes in large-scale directed networks: The role of clustering. PLoS ONE 2013, 8, e77455. [Google Scholar] [CrossRef] [PubMed]
- Ma, L.l.; Ma, C.; Zhang, H.F.; Wang, B.H. Identifying influential spreaders in complex networks based on gravity formula. Phys. A Stat. Mech. Appl. 2016, 451, 205–212. [Google Scholar] [CrossRef]
- Gleiser, P.M.; Danon, L. Community structure in jazz. Adv. Complex Syst. 2003, 6, 565–573. [Google Scholar] [CrossRef]
- Newman, M.E. Modularity and community structure in networks. Proc. Natl. Acad. Sci. USA 2006, 103, 8577–8582. [Google Scholar] [CrossRef]
- Colizza, V.; Pastor-Satorras, R.; Vespignani, A. Reaction-diffusion processes and metapopulation models in heterogeneous networks. Nat. Phys. 2007, 3, 276–282. [Google Scholar] [CrossRef]
- Li, Z.; Ren, T.; Ma, X.; Liu, S.; Zhang, Y.; Zhou, T. Identifying influential spreaders by gravity model. Sci. Rep. 2019, 9, 8387. [Google Scholar] [CrossRef] [PubMed]
- Hirsch, J.E. An index to quantify an individual’s scientific research output. Proc. Natl. Acad. Sci. USA 2005, 102, 16569–16572. [Google Scholar] [CrossRef] [PubMed]
- Lü, L.; Zhang, Y.C.; Yeung, C.H.; Zhou, T. Leaders in social networks, the delicious case. PLoS ONE 2011, 6, e21202. [Google Scholar] [CrossRef] [PubMed]
- Zhuang, Y.; Yağan, O. Information propagation in clustered multilayer networks. IEEE Trans. Netw. Sci. Eng. 2016, 3, 211–224. [Google Scholar] [CrossRef]
- Basaras, P.; Iosifidis, G.; Katsaros, D.; Tassiulas, L. Identifying influential spreaders in complex multilayer networks: A centrality perspective. IEEE Trans. Netw. Sci. Eng. 2017, 6, 31–45. [Google Scholar] [CrossRef]
- Cozzo, E.; Kivelä, M.; De Domenico, M.; Solé, A.; Arenas, A.; Gómez, S.; Porter, M.A.; Moreno, Y. Clustering coefficients in multiplex networks. arXiv 2013, arXiv:1307.6780. [Google Scholar]
- Rahmede, C.; Iacovacci, J.; Arenas, A.; Bianconi, G. Centralities of nodes and influences of layers in large multiplex networks. J. Complex Netw. 2018, 6, 733–752. [Google Scholar] [CrossRef]
- Paidar, M.; Chaharborj, S.S.; Harounabadi, A. Identifying Top-k Most Influential Nodes by using the Topological Diffusion Models in the Complex Networks. Network 2017, 4, 5. [Google Scholar] [CrossRef]
- Ohara, K.; Saito, K.; Kimura, M.; Motoda, H. Resampling-based predictive simulation framework of stochastic diffusion model for identifying top-K influential nodes. Int. J. Data Sci. Anal. 2020, 9, 175–195. [Google Scholar] [CrossRef]
- Tang, J.; Zhang, R.; Yao, Y.; Yang, F.; Zhao, Z.; Hu, R.; Yuan, Y. Identification of top-k influential nodes based on enhanced discrete particle swarm optimization for influence maximization. Phys. A Stat. Mech. Appl. 2019, 513, 477–496. [Google Scholar] [CrossRef]
- Silber, M.D. The Al Qaeda Factor: Plots against the West; University of Pennsylvania Press: Philadelphia, PA, USA, 2011. [Google Scholar]
- Hethcote, H.W. The mathematics of infectious diseases. SIAM Rev. 2000, 42, 599–653. [Google Scholar] [CrossRef]
- Guan-Rong, C.; Xiao-Fan, W.; Xiang, L. Introduction to Complex Networks: Models, Structures and Dynamics; Higher Education Press: Beijing, China, 2012. [Google Scholar]
- Krackhardt, D. Assessing the political landscape: Structure, cognition, and power in organizations. Adm. Sci. Q. 1990, 35, 342–369. [Google Scholar] [CrossRef]
- Zachary, W.W. An information flow model for conflict and fission in small groups. J. Anthropol. Res. 1977, 33, 452–473. [Google Scholar] [CrossRef]
- Lusseau, D.; Schneider, K.; Boisseau, O.J.; Haase, P.; Slooten, E.; Dawson, S.M. The bottlenose dolphin community of Doubtful Sound features a large proportion of long-lasting associations. Behav. Ecol. Sociobiol. 2003, 54, 396–405. [Google Scholar] [CrossRef]
- Tsvetovat, M.; Kouznetsov, A. Social Network Analysis for Startups: Finding Connections on the Social Web; OŔeilly Media, Inc.: Sebastopol, CA, USA, 2011. [Google Scholar]
- Knuth, D.E. The Stanford GraphBase: A Platform for Combinatorial Algorithms; SODA: Austin, TX, USA, 1993; Volume 93, pp. 41–43. [Google Scholar]
- Shen-Orr, S.S.; Milo, R.; Mangan, S.; Alon, U. Network motifs in the transcriptional regulation network of Escherichia coli. Nat. Genet. 2002, 31, 64–68. [Google Scholar] [CrossRef] [PubMed]
- Rossi, R.; Ahmed, N. The network data repository with interactive graph analytics and visualization. In Proceedings of the Twenty-Ninth AAAI Conference on Artificial Intelligence, Austin, TX, USA, 25–30 January 2015. [Google Scholar]
- Duch, J.; Arenas, A. Community detection in complex networks using extremal optimization. Phys. Rev. E 2005, 72, 027104. [Google Scholar] [CrossRef] [PubMed]
- Cho, A.; Shin, J.; Hwang, S.; Kim, C.; Shim, H.; Kim, H.; Kim, H.; Lee, I. WormNet v3: A network-assisted hypothesis-generating server for Caenorhabditis elegans. Nucleic Acids Res. 2014, 42, W76–W82. [Google Scholar] [CrossRef]
- Action, R. The Rise of the Medici. Am. J. Sociol. 1993, 98, 1259–1319. [Google Scholar]
- Krackhardt, D. Cognitive social structures. Soc. Netw. 1987, 9, 109–134. [Google Scholar] [CrossRef]
- Vickers, M.; Chan, S. Representing Classroom Social Structure; Victoria Institute of Secondary Education: Melbourne, Australia, 1981. [Google Scholar]
- Kapferer, B. Strategy and Transaction in an African Factory: African Workers and Indian Management in a Zambian Town; Manchester University Press: Manchester, UK, 1972. [Google Scholar]
- Lazega, E. The Collegial Phenomenon: The Social Mechanisms of Cooperation Among Peers in a Corporate Law Partnership; Oxford University Press on Demand: Oxford, UK, 2001. [Google Scholar]
- Snijders, T.A.; Pattison, P.E.; Robins, G.L.; Handcock, M.S. New specifications for exponential random graph models. Sociol. Methodol. 2006, 36, 99–153. [Google Scholar] [CrossRef]
- Stark, C.; Breitkreutz, B.J.; Reguly, T.; Boucher, L.; Breitkreutz, A.; Tyers, M. BioGRID: a general repository for interaction datasets. Nucleic Acids Res. 2006, 34, D535–D539. [Google Scholar] [CrossRef] [PubMed]
- Magnani, M.; Micenkova, B.; Rossi, L. Combinatorial analysis of multiple networks. arXiv 2013, arXiv:1303.4986. [Google Scholar]
- De Domenico, M.; Solé-Ribalta, A.; Gómez, S.; Arenas, A. Navigability of interconnected networks under random failures. Proc. Natl. Acad. Sci. USA 2014, 111, 8351–8356. [Google Scholar] [CrossRef] [PubMed]
- Cardillo, A.; Gómez-Gardenes, J.; Zanin, M.; Romance, M.; Papo, D.; Del Pozo, F.; Boccaletti, S. Emergence of network features from multiplexity. Sci. Rep. 2013, 3, 1344. [Google Scholar] [CrossRef] [PubMed]
- Kendall, M.G. A new measure of rank correlation. Biometrika 1938, 30, 81–93. [Google Scholar] [CrossRef]



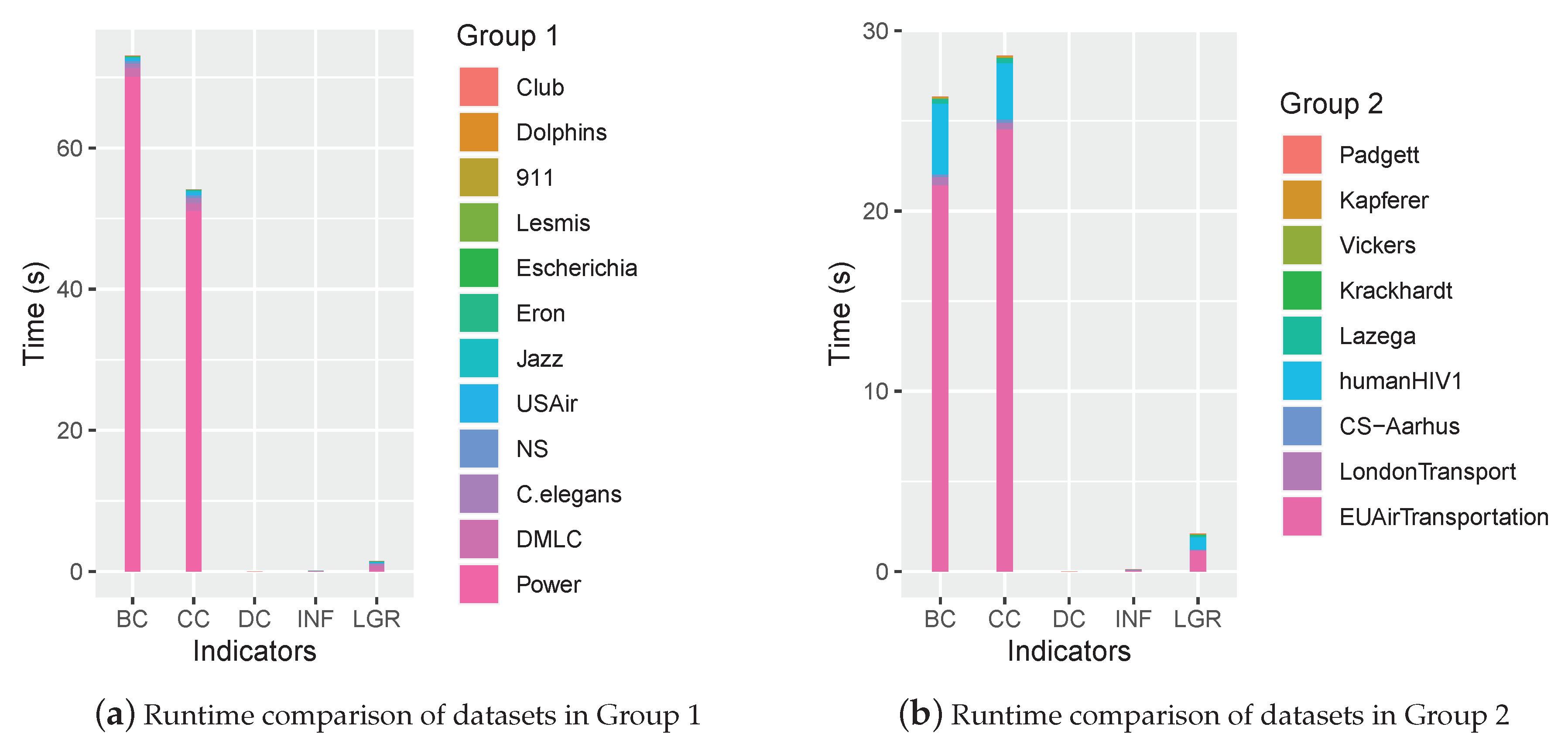
| Metric | Topology | Complexity | Advantages | Disadvantages |
|---|---|---|---|---|
| DC [19] | Local | simple | incapable of dealing with “bridge” nodes | |
| BC [22] | Global | finding “bridge-like” nodes | cannot differentiate most marginal nodes | |
| CC [23] | Global | finding “nearest” nodes | incapable in disconnected graphs | |
| EC [24] | Global | consider both of the quality and quantity of neighbors | may be non-convergent | |
| PR [25] | Global | efficient, widely applied in search engine | may be non-convergent | |
| H-index [34] | Semi-local | famous for academic evaluation | lack of global information | |
| k-shell [14] | Global | suitable for large-scale networks | indistinguishable | |
| LR [35] | Global | no parameters; robustness | may be non-convergent | |
| GR, GR+ [29] | Global | accuracy | high complexity in k-shell | |
| LGR [33] | Semi-local | simple and capable in most cases | additional parameters R determination |
| ID | Name | Degree | Betweenness | Closeness | Katz | Eigenvector | INF |
|---|---|---|---|---|---|---|---|
| 1 | Andre | 0.4444 | 0.0231 | 0.5294 | 0.3307 | 0.3522 | 0.7211 |
| 2 | Beverley | 0.4444 | 0.0231 | 0.5294 | 0.3307 | 0.3522 | 0.7211 |
| 3 | Carol | 0.3333 | 0.0000 | 0.5000 | 0.3006 | 0.2858 | 0.6495 |
| 4 | Diane | 0.6667 | 0.1019 | 0.6000 | 0.3907 | 0.4810 | 0.8273 |
| 5 | Ed | 0.3333 | 0.0000 | 0.5000 | 0.3006 | 0.2858 | 0.6495 |
| 6 | Fernando | 0.5556 | 0.2315 | 0.6429 | 0.3595 | 0.3977 | 0.7830 |
| 7 | Garth | 0.5556 | 0.2315 | 0.6429 | 0.3595 | 0.3977 | 0.7830 |
| 8 | Heather | 0.3333 | 0.3889 | 0.6000 | 0.2887 | 0.1959 | 0.7109 |
| 9 | Ike | 0.2222 | 0.2222 | 0.4286 | 0.2431 | 0.0481 | 0.7914 |
| 10 | Jane | 0.1111 | 0.0000 | 0.3103 | 0.2168 | 0.0112 | 0.6225 |
| ID | Name | Recovered Nodes | Iterations |
|---|---|---|---|
| 1 | Andre | 4.8015 | 3.3399 |
| 2 | Beverley | 4.7902 | 3.3392 |
| 3 | Carol | 4.3485 | 3.1529 |
| 4 | Diane | 5.3182 | 3.3684 |
| 5 | Ed | 4.3336 | 3.1455 |
| 6 | Fernando | 5.0856 | 3.3251 |
| 7 | Garth | 5.0338 | 3.2994 |
| 8 | Heather | 4.0765 | 2.9625 |
| 9 | Ike | 2.6060 | 2.1806 |
| 10 | Jane | 1.8086 | 1.7019 |
| Dataset Name | |E| | <k> | <d> | |C| | r | |H| | ||
|---|---|---|---|---|---|---|---|---|
| Club [47] | 34 | 78 | 4.5882 | 2.4082 | 0.5706 | −0.4756 | 1.6933 | 0.1477 |
| Dolphins [48] | 62 | 159 | 5.1290 | 3.3570 | 0.2590 | −0.0436 | 1.3268 | 0.1723 |
| 911 [49] | 69 | 159 | 4.6087 | 2.4672 | 0.4698 | −0.0380 | 1.7304 | 0.1434 |
| Lesmis [50] | 77 | 254 | 6.5974 | 2.6411 | 0.5731 | −0.1652 | 1.8273 | 0.0905 |
| Escherichia [51] | 97 | 212 | 4.3711 | 5.5369 | 0.3675 | 0.4116 | 1.2367 | 0.2270 |
| Eron [52] | 143 | 623 | 8.7133 | 2.9670 | 0.4339 | −0.0195 | 1.4829 | 0.0839 |
| Jazz [30] | 198 | 2742 | 27.6970 | 2.2350 | 0.6175 | 0.0202 | 1.3951 | 0.0266 |
| USAir [32] | 332 | 2126 | 12.8072 | 2.7381 | 0.6252 | −0.2079 | 3.4639 | 0.0231 |
| NS [31] | 379 | 914 | 4.8232 | 6.0419 | 0.7412 | −0.0817 | 1.6630 | 0.1424 |
| C.elegans [53] | 453 | 2032 | 9.0066 | 2.6638 | 0.6465 | −0.2197 | 4.4782 | 0.0254 |
| DMLC [54] | 659 | 1570 | 4.7648 | 2.6370 | 0.3279 | −0.1914 | 14.8897 | 0.0143 |
| Power [26] | 4941 | 6594 | 2.6691 | 18.9892 | 0.0801 | 0.0035 | 1.4504 | 0.3483 |
| Dataset Name | |E| | || | || | <k> | <d> | |C| | ||
|---|---|---|---|---|---|---|---|---|
| Padgett [55] | 2 | 26 | 46 | 35 | 11 | 3.5385 | 2.6923 | 0.1441 |
| Krackhardt [56] | 3 | 63 | 307 | 244 | 63 | 9.746 | 2.1731 | 0.3943 |
| Vickers [57] | 3 | 87 | 605 | 518 | 87 | 13.908 | 2.1802 | 0.4823 |
| Kapferer [58] | 4 | 150 | 769 | 552 | 217 | 10.2533 | 2.5889 | 0.3002 |
| Lazega [59,60] | 3 | 211 | 2051 | 1842 | 209 | 19.4408 | 2.3958 | 0.3938 |
| humanHIV1 [61] | 5 | 1195 | 1504 | 1269 | 235 | 2.5172 | 4.1385 | 0.0221 |
| CS-Aarhus [62] | 5 | 224 | 948 | 620 | 328 | 8.4643 | 3.1847 | 0.3603 |
| LondonTransport [63] | 3 | 399 | 472 | 441 | 31 | 2.3659 | 14.2989 | 0.0243 |
| EUAirTransportation [64] | 37 | 2034 | 15199 | 3588 | 11611 | 14.9449 | 3.5087 | 0.5969 |
| Dataset Name | DC | INF | LGR | BC | CC |
|---|---|---|---|---|---|
| Club | 0.2442 | 0.2513 | 0.1515 | 0.1016 | −0.0766 |
| Dolphins | 0.0238 | 0.0344 | −0.0196 | −0.0323 | −0.0354 |
| 911 | 0.0878 | 0.1918 | 0.0426 | 0.0409 | −0.0895 |
| Lesmis | 0.0909 | 0.1114 | 0.1032 | −0.1839 | −0.0519 |
| Escherichia | 0.0726 | 0.0692 | 0.0009 | −0.0808 | 0.0971 |
| Eron | 0.0454 | 0.0927 | −0.0056 | 0.0031 | 0.0107 |
| Jazz | 0.069 | 0.0838 | 0.0325 | 0.0244 | 0.0312 |
| USAir | −0.0358 | 0.0743 | −0.0402 | −0.0491 | −0.0311 |
| NS | −0.0378 | 0.0302 | −0.0089 | −0.0143 | −0.0462 |
| C.elegans | 0.0154 | 0.0697 | 0.0242 | 0.0185 | 0.0214 |
| Power | −0.005 | 0.0272 | 0.0024 | −0.0015 | 0.0209 |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Huang, X.; Chen, D.; Wang, D.; Ren, T. Identifying Influencers in Social Networks. Entropy 2020, 22, 450. https://doi.org/10.3390/e22040450
Huang X, Chen D, Wang D, Ren T. Identifying Influencers in Social Networks. Entropy. 2020; 22(4):450. https://doi.org/10.3390/e22040450
Chicago/Turabian StyleHuang, Xinyu, Dongming Chen, Dongqi Wang, and Tao Ren. 2020. "Identifying Influencers in Social Networks" Entropy 22, no. 4: 450. https://doi.org/10.3390/e22040450
APA StyleHuang, X., Chen, D., Wang, D., & Ren, T. (2020). Identifying Influencers in Social Networks. Entropy, 22(4), 450. https://doi.org/10.3390/e22040450






