Information Theoretic Modeling of High Precision Disparity Data for Lossy Compression and Object Segmentation
Abstract
:1. Introduction
1.1. Motivation
1.2. Related Work
1.2.1. Methods for Disparity Map Compression
1.2.2. Methods for Image Segmentation and Edge Detection
1.2.3. Methods Combining Image Compression and Image Segmentation
1.3. Contribution
2. Proposed Methods
2.1. Definitions and Statement of the Problem
2.1.1. Image Partition into Regions
2.1.2. Representing the Region’s Contours
2.1.3. Representing a Hierarchical Segmentation
2.1.4. Polynomial Surface for Approximating the Disparity Map over a Region
- the rate-distortion description , with and , should be competitive with the rate-distortion of lossy compression algorithms, at very low bitrates. The wish is to extract relevant information from , to encode it efficiently, and use it for obtaining a reconstruction with a small distortion, as in the lossy compression tasks, but with the next additional wish on the relevance of the segmentation for the objects in the image.
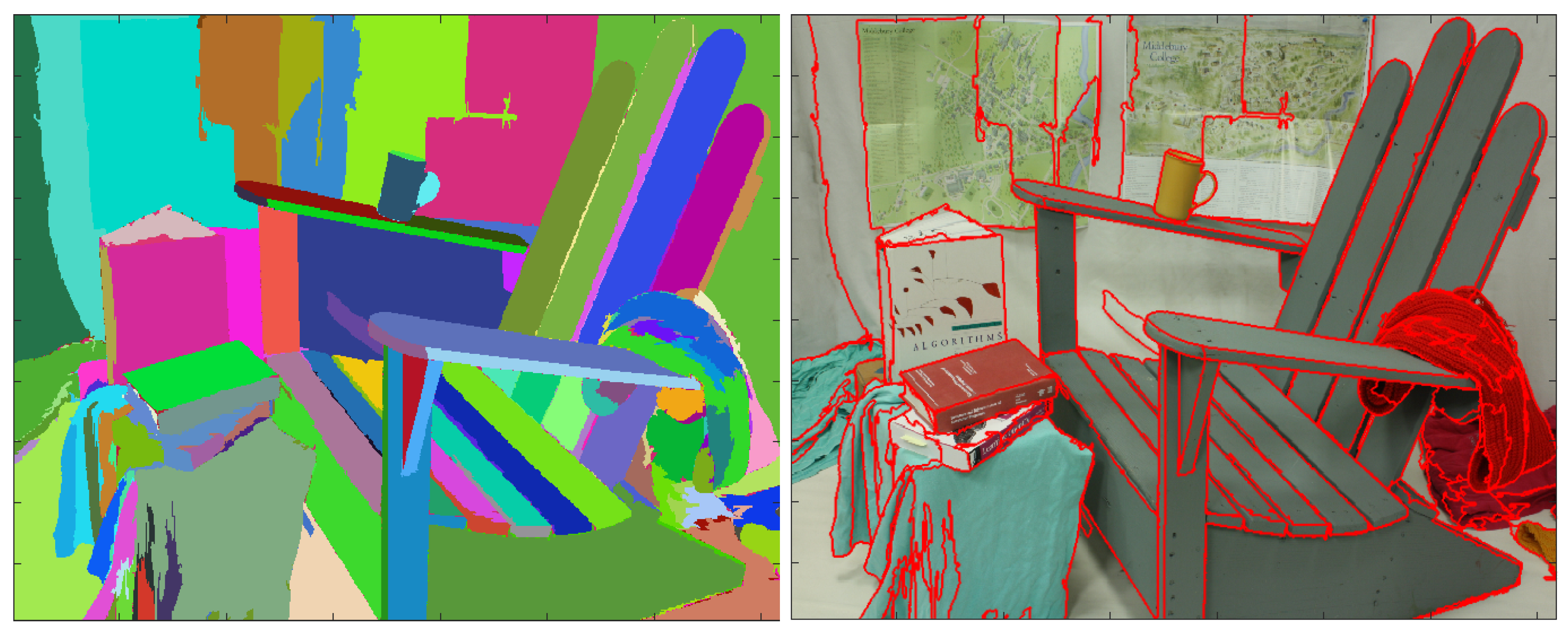
- The sequence of partitions should compare favorably with the hierarchical partitions obtained from the color information of the same scene, having the diagram (recall, precision) competitive with the existing state of the art boundary detection or segmentation algorithms for finding general structure in images.
2.1.5. Statement of the Problem
2.2. Algorithm for Hierarchical Segmentation based on Persistency of Contours of the Segmentations Generated by Iterative Piece-Wise Polynomial Modeling
| Algorithm 1 Hierarchical segmentations based on persistency of contours generated by iterative piece-wise polynomial modeling |
|
| Algorithm 2 Hierarchical partition based on (description length - distortion) optimization |
|
2.3. Algorithm for Hierarchical Segmentation based on (Description Length-Distortion) Optimization
3. Experimental Results
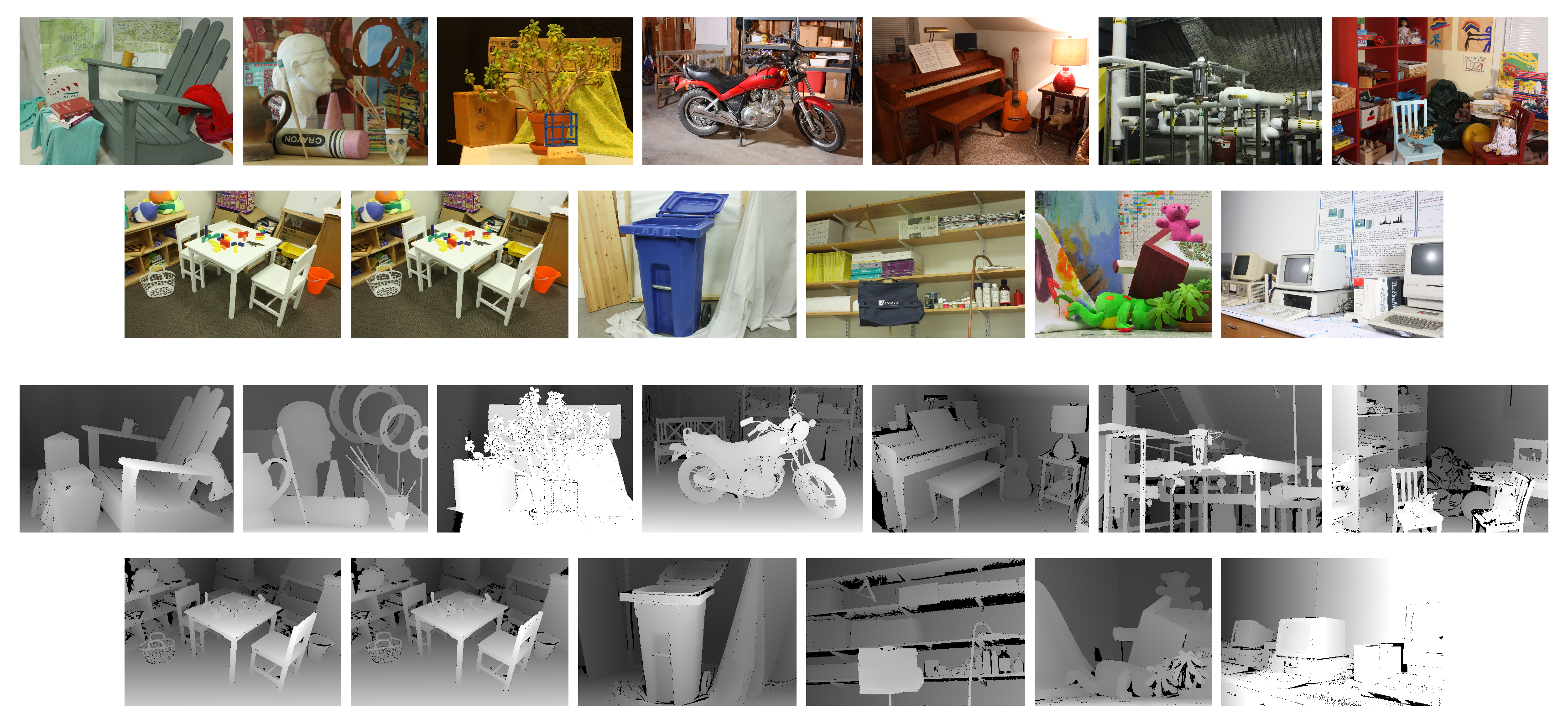
3.1. The Datasets
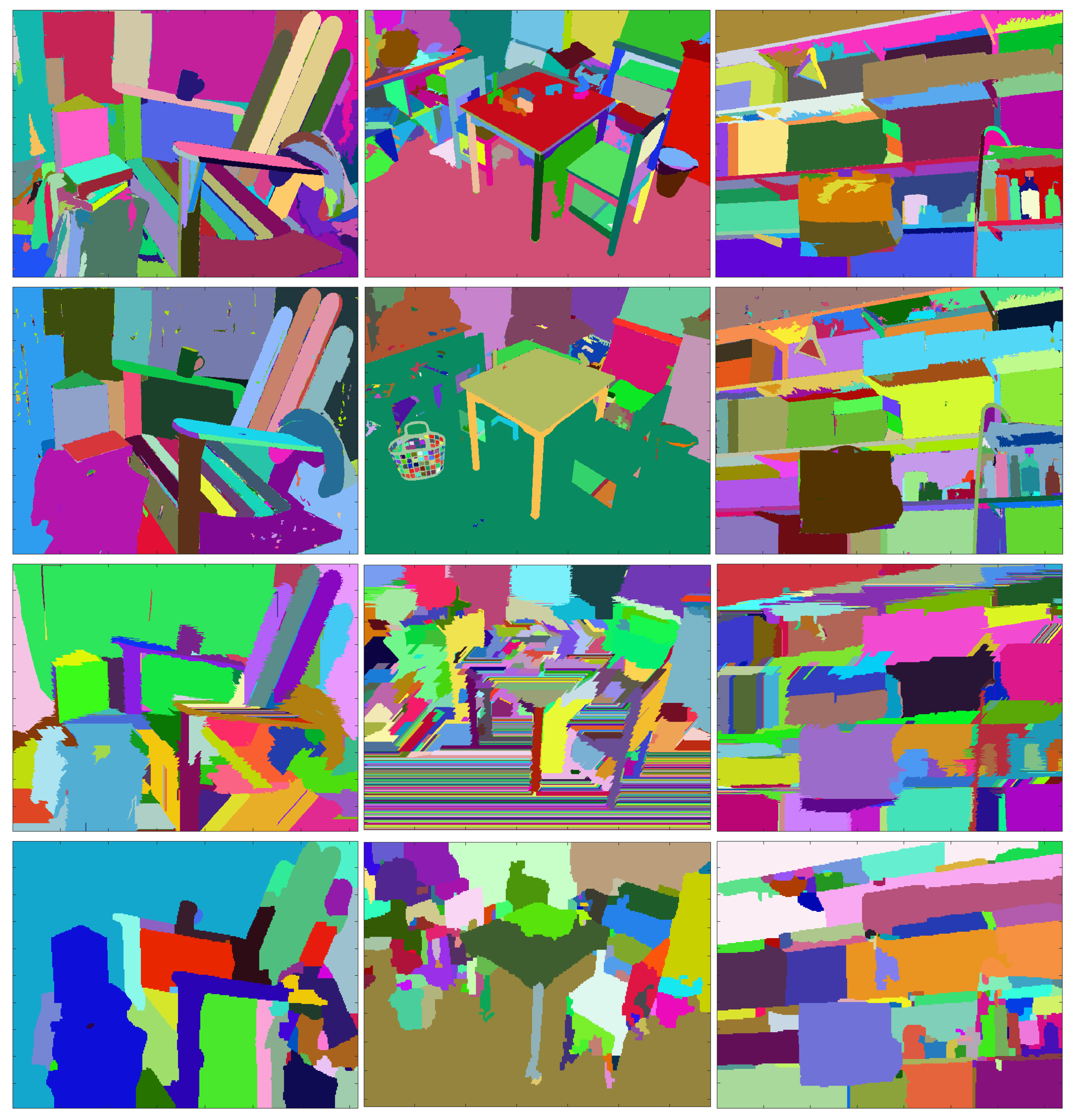
3.2. Obtaining the Sequences of Segmentations A and B
3.3. Benchmarking the Sequences of Segmentations against References Extracted from the Color Images
3.4. Rate-Distortion Performance of the Segmentation Algorithm
| Algorithm 3 Encoding based on the segmentation and polynomial models over each region |
|
4. Conclusions
Author Contributions
Funding
Conflicts of Interest
References
- Hartley, R.; Zisserman, A. Multiple View Geometry in Computer Vision; Cambridge University Press: New York, NY, USA, 2003. [Google Scholar]
- Ebrahimi, T.; Foessel, S.; Pereira, F.; Schelkens, P. JPEG Pleno: Toward an Efficient Representation of Visual Reality. IEEE MultiMedia 2016, 23, 14–20. [Google Scholar] [CrossRef]
- Schwarz, S.; Preda, M.; Baroncini, V.; Budagavi, M.; Cesar, P.; Chou, P.A.; Cohen, R.A.; Krivokuća, M.; Lasserre, S.; Li, Z.; et al. Emerging MPEG Standards for Point Cloud Compression. IEEE J. Emerg. Sel. Top. Circuits Syst. 2019, 9, 133–148. [Google Scholar] [CrossRef]
- Astola, P.; Tabus, I. Lossless Compression of High Resolution Disparity Map Images. In Proceedings of the 2017 International Symposium on Signals, Circuits and Systems (ISSCS), Iasi, Romania, 13–14 July 2017; pp. 1–4. [Google Scholar]
- Tabus, I.; Schiopu, I.; Astola, J. Context Coding of Depth Map Images Under the Piecewise-Constant Image Model Representation. IEEE Trans. Image Process. 2013, 22, 4195–4210. [Google Scholar] [CrossRef] [PubMed]
- Schiopu, I.; Tabus, I. Parametrizations of Planar models for Region-merging based Lossy Depth-map Compression. In Proceedings of the 2015 3DTV-Conference: The True Vision—Capture, Transmission and Display of 3D Video (3DTV-CON), Xi’an, China, 18–20 September 2015; pp. 1–4. [Google Scholar]
- Schiopu, I.; Saarinen, J.P.; Tabus, I. Lossy-to-lossless Progressive Coding of Depth-map Images Using Competing Constant and Planar Models. In Proceedings of the 2015 International Conference on 3D Imaging (IC3D), Liege, Belgium, 14–15 December 2015; pp. 1–7. [Google Scholar]
- Özkalaycı, B.O.; Alatan, A.A. 3D Planar Representation of Stereo Depth Images for 3DTV Applications. IEEE Trans. Image Process. 2014, 23, 5222–5232. [Google Scholar] [CrossRef]
- Kiani, V.; Harati, A.; Vahedian, A. Planelets—A Piecewise Linear Fractional Model for Preserving Scene Geometry in Intra-Coding of Indoor Depth Images. IEEE Trans. Image Process. 2017, 26, 590–602. [Google Scholar] [CrossRef] [PubMed]
- Mathew, R.; Taubman, D.; Zanuttigh, P. Scalable Coding of Depth Maps With R-D Optimized Embedding. IEEE Trans. Image Process. 2013, 22, 1982–1995. [Google Scholar] [CrossRef] [PubMed]
- Mathew, R.; Taubman, D. WaSP Encoder with Breakpoint Adaptive DWT Coding of Disparity Maps. In Proceedings of the 2019 IEEE International Conference on Image Processing (ICIP), Taipei, Taiwan, 22–25 September 2019; pp. 3172–3176. [Google Scholar]
- Davis, L.S. A Survey of Edge Detection Techniques. Comput. Graph. Image Process. 1975, 4, 248–270. [Google Scholar] [CrossRef]
- Canny, J. A Computational Approach to Edge Detection. IEEE Trans. Pattern Anal. Mach. Intell. 1986, 6, 679–698. [Google Scholar] [CrossRef]
- Dollár, P.; Zitnick, C.L. Fast Edge Detection Using Structured Forests. IEEE Trans. Pattern Anal. Mach. Intell. 2015, 37, 1558–1570. [Google Scholar] [CrossRef]
- Xie, S.; Tu, Z. Holistically-Nested Edge Detection. In Proceedings of the IEEE International Conference on Computer Vision, Santiago, Chile, 7–13 December 2015; pp. 1395–1403. [Google Scholar]
- Liu, Y.; Cheng, M.; Hu, X.; Bian, J.; Zhang, L.; Bai, X.; Tang, J. Richer Convolutional Features for Edge Detection. IEEE Trans. Pattern Anal. Mach. Intell. 2019, 41, 1939–1946. [Google Scholar] [CrossRef]
- Shen, W.; Wang, X.; Wang, Y.; Bai, X.; Zhang, Z. Deepcontour: A Deep Convolutional Feature Learned by Positive-sharing Loss for Contour Detection. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Boston, MA, USA, 7–12 June 2015; pp. 3982–3991. [Google Scholar]
- Van de Sande, K.E.A.; Uijlings, J.R.R.; Gevers, T.; Smeulders, A.W.M. Segmentation as Selective Search for Object Recognition. In Proceedings of the 2011 International Conference on Computer Vision, Tampa, FL, USA, 5–8 December 2011; pp. 1879–1886. [Google Scholar]
- Pont-Tuset, J.; Arbelaez, P.; Barron, J.T.; Marques, F.; Malik, J. Multiscale Combinatorial Grouping for Image Segmentation and Object Proposal Generation. IEEE Trans. Pattern Anal. Mach. Intell. 2017, 39, 128–140. [Google Scholar] [CrossRef] [PubMed]
- Farabet, C.; Couprie, C.; Najman, L.; LeCun, Y. Learning Hierarchical Features for Scene Labeling. IEEE Trans. Pattern Anal. Mach. Intell. 2012, 35, 1915–1929. [Google Scholar] [CrossRef] [PubMed]
- Fulkerson, B.; Vedaldi, A.; Soatto, S. Class Segmentation and Object Localization with Superpixel Neighborhoods. In Proceedings of the 2009 IEEE 12th International Conference on Computer Vision, Kyoto, Japan, 29 September–2 October 2009; pp. 670–677. [Google Scholar]
- Felzenszwalb, P.F.; Huttenlocher, D.P. Efficient Graph-based Image Segmentation. Int. J. Comput. Vis. 2004, 59, 167–181. [Google Scholar] [CrossRef]
- Arbelaez, P.; Maire, M.; Fowlkes, C.; Malik, J. Contour Detection and Hierarchical Image Segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 2011, 33, 898–916. [Google Scholar] [CrossRef] [PubMed]
- Leclerc, Y. Constructing Simple Stable Descriptions for Image Partitioning. Int. J. Comput. Vis. 1989, 3, 73–102. [Google Scholar] [CrossRef]
- Zhu, S.C.; Yuille, A. Region Competition: Unifying Snakes, Region Growing, and Bayes/MDL for Multiband Image Segmentation. IEEE Trans. Pattern Anal. Mach. Intell. 1996, 18, 884–900. [Google Scholar]
- Cardinale, J.; Paul, G.; Sbalzarini, I.F. Discrete Region Competition for Unknown Numbers of Connected Regions. IEEE Trans. Image Process. 2012, 21, 3531–3545. [Google Scholar] [CrossRef]
- Penalver Benavent, A.; Escolano Ruiz, F.; Saez, J.M. Learning Gaussian Mixture Models With Entropy-Based Criteria. IEEE Trans. Neural Networks 2009, 20, 1756–1771. [Google Scholar] [CrossRef]
- Gorelick, L.; Delong, A.; Veksler, O.; Boykov, Y. Recursive MDL via Graph Cuts: Application to Segmentation. In Proceedings of the 2011 International Conference on Computer Vision, Tampa, FL, USA, 5–8 December 2011; pp. 890–897. [Google Scholar]
- Helin, P.; Astola, P.; Rao, B.; Tabus, I. Minimum Description Length Sparse Modeling and Region Merging for Lossless Plenoptic Image Compression. IEEE J. Sel. Top. Signal Process. 2017, 11, 1146–1161. [Google Scholar] [CrossRef]
- Vereshchagin, N.K.; Vitanyi, P.M.B. Kolmogorov’s Structure Functions and Model Selection. IEEE Trans. Inf. Theory 2004, 50, 3265–3290. [Google Scholar] [CrossRef]
- Rissanen, J.; Tabus, I. Kolmogorov’s structure function in MDL theory and lossy data compression. In Advances in Minimum Description Length: Theory and Applications; The MIT Press: Cambridge, UK, 2003; pp. 245–264. [Google Scholar]
- Katkovnik, V.I.; Egiazarian, K.; Astola, J. Local Approximation Techniques in Signal and Image Processing; SPIE Bellingham: Bellingham, WA, USA, 2006. [Google Scholar]
- Yang, Q.; Boukerroui, D. Ultrasound Image Segmentation Using Local Statistics with an Adaptive Scale Selection. In Proceedings of the 2012 9th IEEE International Symposium on Biomedical Imaging (ISBI), Barcelona, Spain, 2–5 May 2012; pp. 1096–1099. [Google Scholar]
- Boukerroui, D. Optimal Spatial Scale for Local Region-based Active Contours. In Proceedings of the 2014 IEEE International Conference on Image Processing (ICIP), Paris, France, 27–30 Ocober 2014; pp. 4393–4397. [Google Scholar]
- Martin, D.R.; Fowlkes, C.C.; Malik, J. Learning to Detect Natural Image Boundaries Using Local Brightness, Color and Texture Cues. IEEE Trans. Pattern Anal. Mach. Intell. 2004, 26, 530–549. [Google Scholar] [CrossRef] [PubMed]
- Scharstein, D.; Hirschmüller, H.; Kitajima, Y.; Krathwohl, G.; Nesic, N.; Wang, X.; Westling, P. High-resolution Stereo Datasets with Subpixel-Accurate Ground Truth. In Proceedings of the German Conference on Pattern Recognition (GCPR 2014), Münster, Germany, 2–5 September 2014. [Google Scholar]

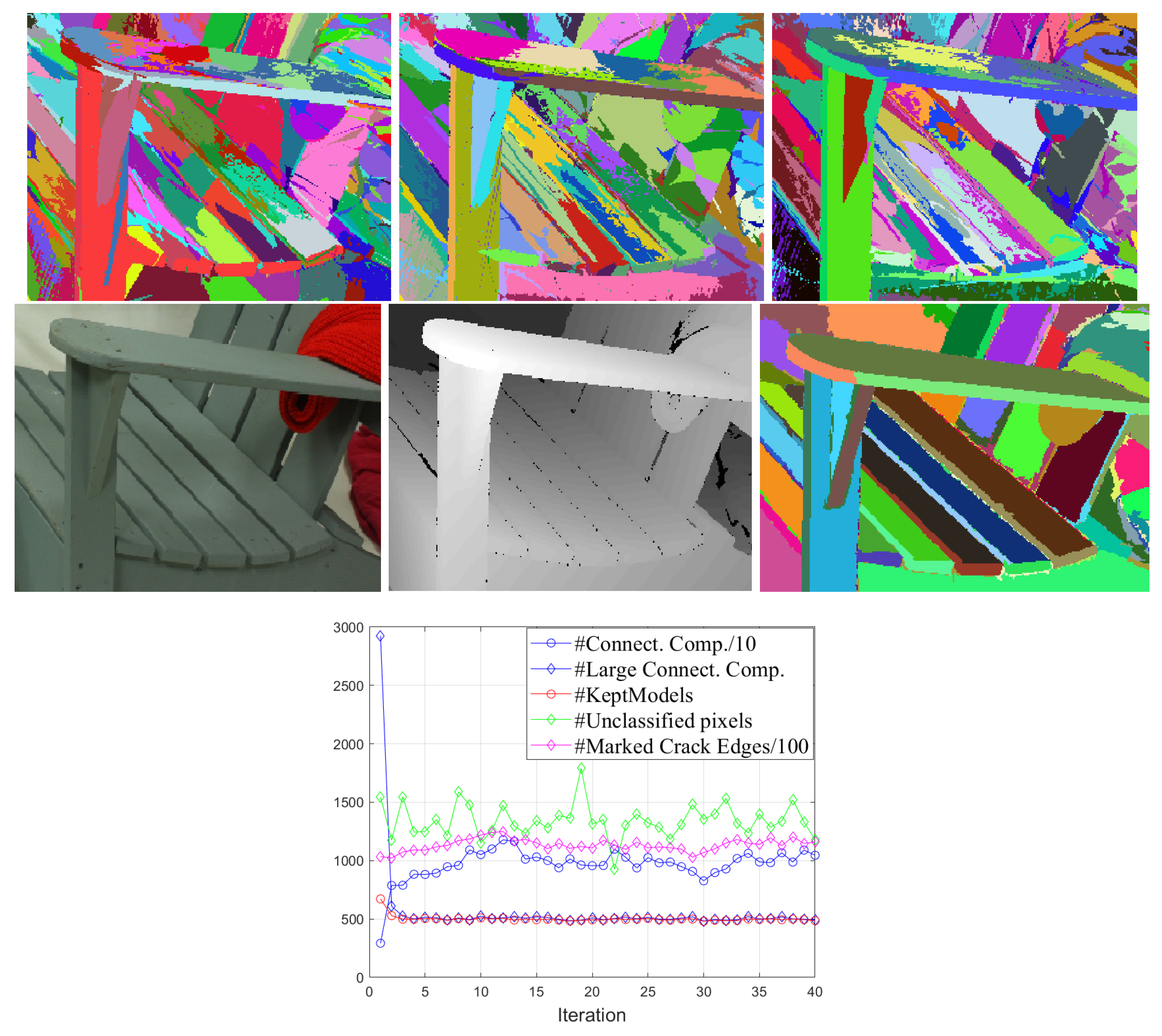


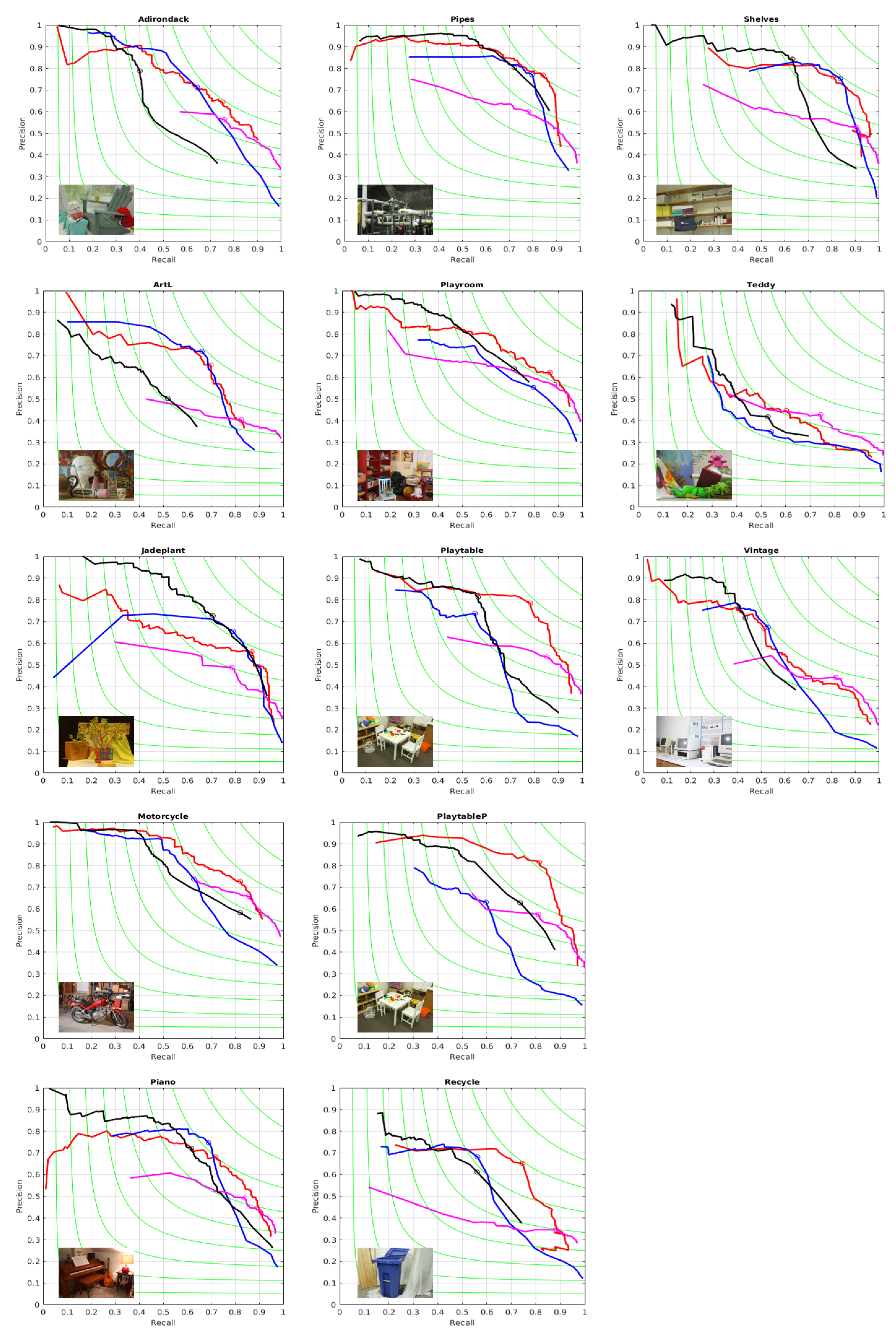


| Precision-Recall | F-Value | Bjøntegaard BD-PCNR (dB) | ||||||||||
|---|---|---|---|---|---|---|---|---|---|---|---|---|
| Scene | A1 | A2 | SSS | MCG | A1 | A2 | SSS | MCG | A1 | A2 | SSS | MCG |
| Adirondack | 0.71–0.64 | 0.65–0.75 | 0.56–0.76 | 0.79–0.40 | 0.68 | 0.69 | 0.65 | 0.53 | −5.11 | −2.73 | −6.62 | −3.21 |
| ArtL | 0.72–0.66 | 0.66–0.70 | 0.40–0.83 | 0.50–0.52 | 0.69 | 0.68 | 0.54 | 0.51 | −1.77 | −0.77 | −4.72 | −2.82 |
| Jadeplant | 0.66–0.79 | 0.56–0.87 | 0.49–0.79 | 0.73–0.71 | 0.72 | 0.68 | 0.60 | 0.72 | −1.83 | −1.94 | −5.82 | −2.77 |
| Motorcycle | 0.72–0.64 | 0.73–0.82 | 0.66–0.86 | 0.58-0.82 | 0.68 | 0.77 | 0.74 | 0.68 | −8.21 | −2.79 | −7.30 | −3.86 |
| Piano | 0.75–0.69 | 0.68–0.72 | 0.49–0.84 | 0.72–0.62 | 0.72 | 0.70 | 0.62 | 0.66 | −1.93 | −2.93 | −6.27 | −3.64 |
| Pipes | 0.77–0.80 | 0.77–0.84 | 0.60–0.78 | 0.80–0.72 | 0.78 | 0.80 | 0.68 | 0.76 | −1.83 | −1.29 | −6.64 | −3.49 |
| Playroom | 0.55–0.80 | 0.62–0.87 | 0.57-0.88 | 0.63–0.73 | 0.65 | 0.72 | 0.69 | 0.68 | −0.75 | −2.67 | −5.32 | −2.95 |
| Playtable | 0.74–0.55 | 0.78–0.78 | 0.54-0.85 | 0.81–0.57 | 0.63 | 0.78 | 0.66 | 0.67 | −6.92 | −3.25 | −7.49 | −4.00 |
| PlaytableP | 0.63–0.60 | 0.82–0.81 | 0.57–0.81 | 0.63–0.74 | 0.61 | 0.82 | 0.67 | 0.68 | −5.57 | −2.91 | −8.44 | −3.70 |
| Recycle | 0.68–0.56 | 0.65–0.74 | 0.35–0.88 | 0.65–0.53 | 0.62 | 0.70 | 0.50 | 0.58 | −7.87 | −2.51 | −6.39 | −2.59 |
| Shelves | 0.75–0.84 | 0.76–0.81 | 0.53–0.91 | 0.84–0.63 | 0.79 | 0.78 | 0.67 | 0.72 | 1.36 | −1.24 | −6.88 | −3.32 |
| Teddy | 0.35–0.54 | 0.45–0.60 | 0.43–0.74 | 0.42–0.53 | 0.42 | 0.51 | 0.54 | 0.47 | −0.62 | −2.42 | −5.01 | −3.09 |
| Vintage | 0.67–0.53 | 0.66–0.52 | 0.44–0.82 | 0.72–0.43 | 0.59 | 0.58 | 0.57 | 0.54 | 0.32 | −1.56 | −11.59 | −2.11 |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Tăbuş, I.; Kaya, E.C. Information Theoretic Modeling of High Precision Disparity Data for Lossy Compression and Object Segmentation. Entropy 2019, 21, 1113. https://doi.org/10.3390/e21111113
Tăbuş I, Kaya EC. Information Theoretic Modeling of High Precision Disparity Data for Lossy Compression and Object Segmentation. Entropy. 2019; 21(11):1113. https://doi.org/10.3390/e21111113
Chicago/Turabian StyleTăbuş, Ioan, and Emre Can Kaya. 2019. "Information Theoretic Modeling of High Precision Disparity Data for Lossy Compression and Object Segmentation" Entropy 21, no. 11: 1113. https://doi.org/10.3390/e21111113
APA StyleTăbuş, I., & Kaya, E. C. (2019). Information Theoretic Modeling of High Precision Disparity Data for Lossy Compression and Object Segmentation. Entropy, 21(11), 1113. https://doi.org/10.3390/e21111113





