Big Data Blind Separation
Abstract
1. Introduction
- Every column of the source matrix is non-negative.
- Source matrix has a full row rank.
- Mixing matrix has a full column rank, and .
- The rows of the source matrix, and columns of the mixing matrix have unit norm.
- Source matrix is sparse.
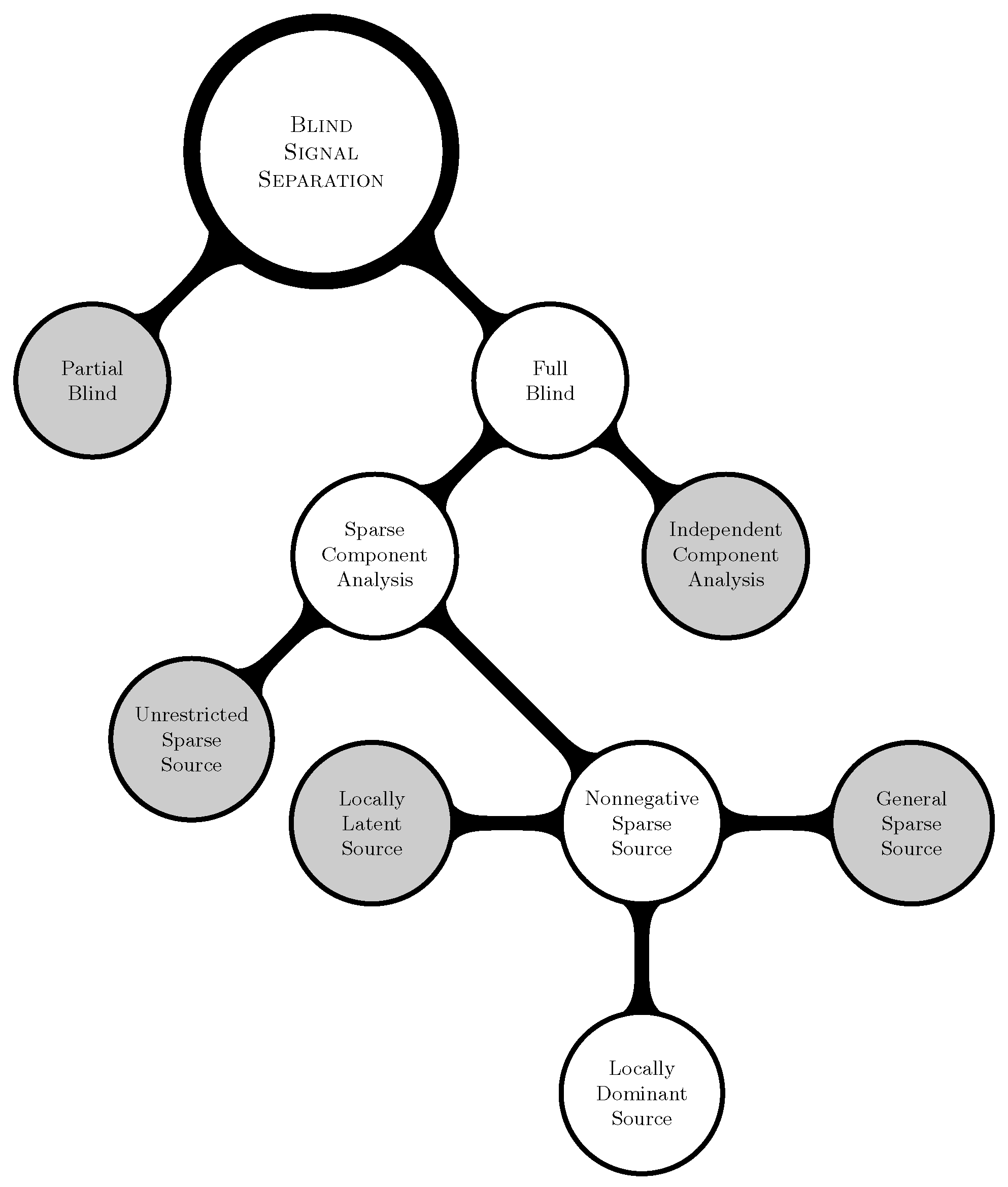
- Locally Dominant Case: In addition to the basic assumptions, for a given row r of , there exists at least one unique column c such that:
- Locally Latent Case: In addition to the basic assumptions, for a given row r of , there exists at least linearly independent and unique columns such that:
- General Sparse Case: This is the default case.
2. Locally Dominant Case
2.1. Conventional Formulations
2.2. Envelope Formulation
3. Point Correntropy
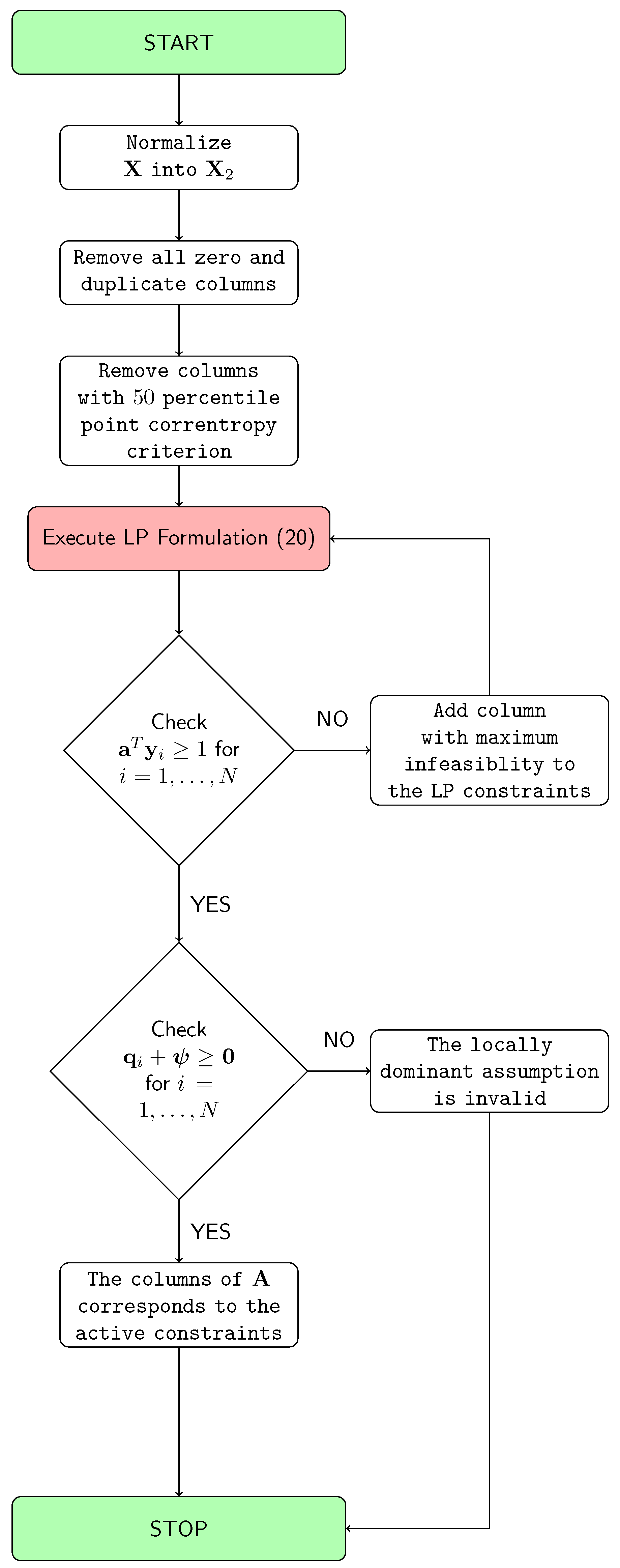
4. Solution Methodology
| Algorithm 1: The Proposed Algorithm. |
| Data: Given Result: Find and such that = normalize(); Remove all zero columns and duplicate columns from , and say ; Estimate from ; Obtain by removing all columns with the 50 percentile point correntropy criterion from ; Let ; Let be the ith column of ; = Solution of LP Formulation (20) with respect to data ; while do 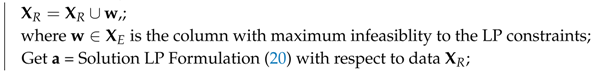 end Let be the matrix containing the columns of corresponding to the active constraints at the optimal solution of Formulation (20); Calculate for ; Set equal to 0 for non-noisy non-image data mixing, equal to for non-noisy image data mixing, or equal to the user-specified value for noisy mixing.; if then 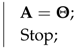 else  end |
5. Numerical Experiments
5.1. Simulated Data Separation
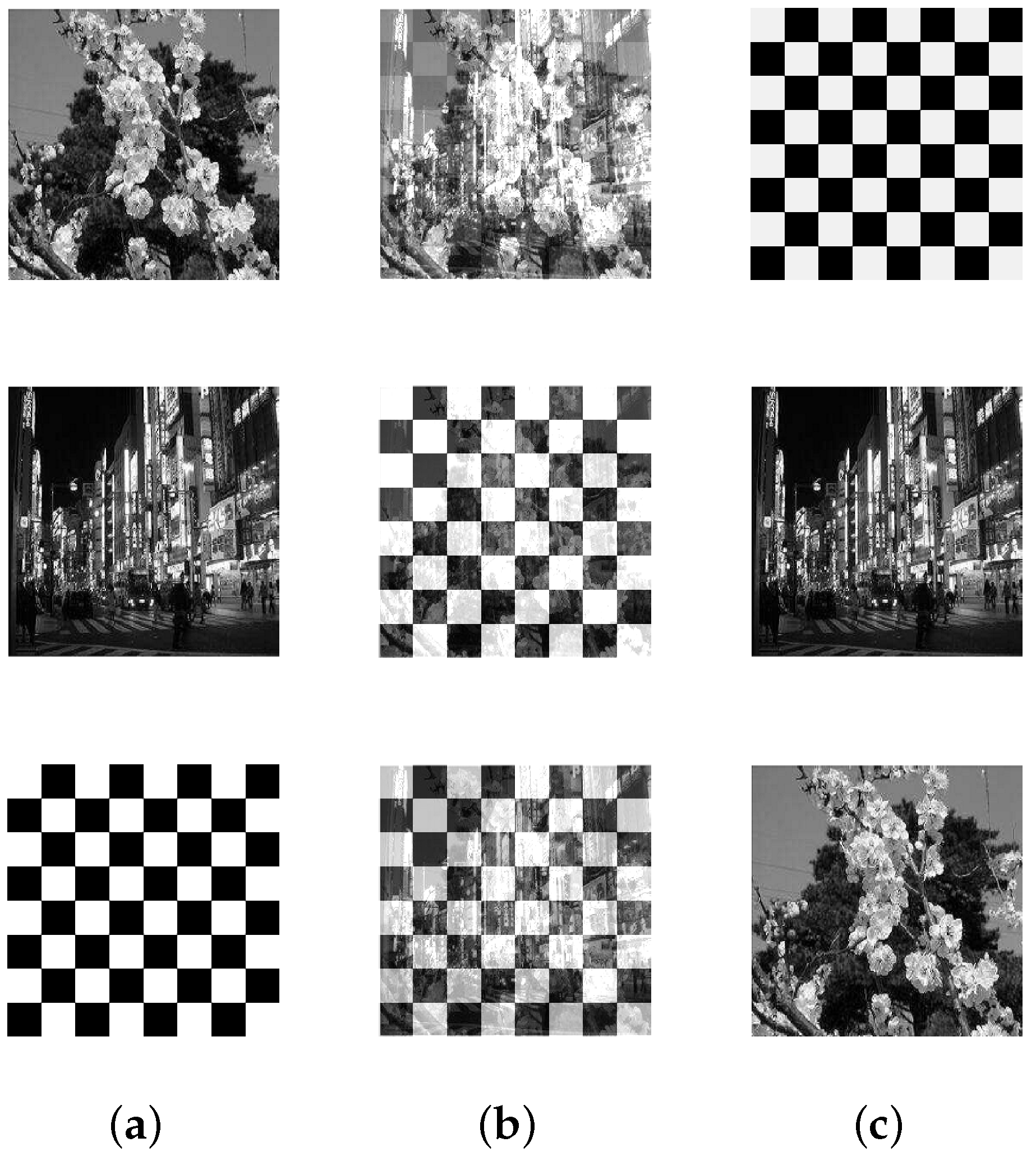
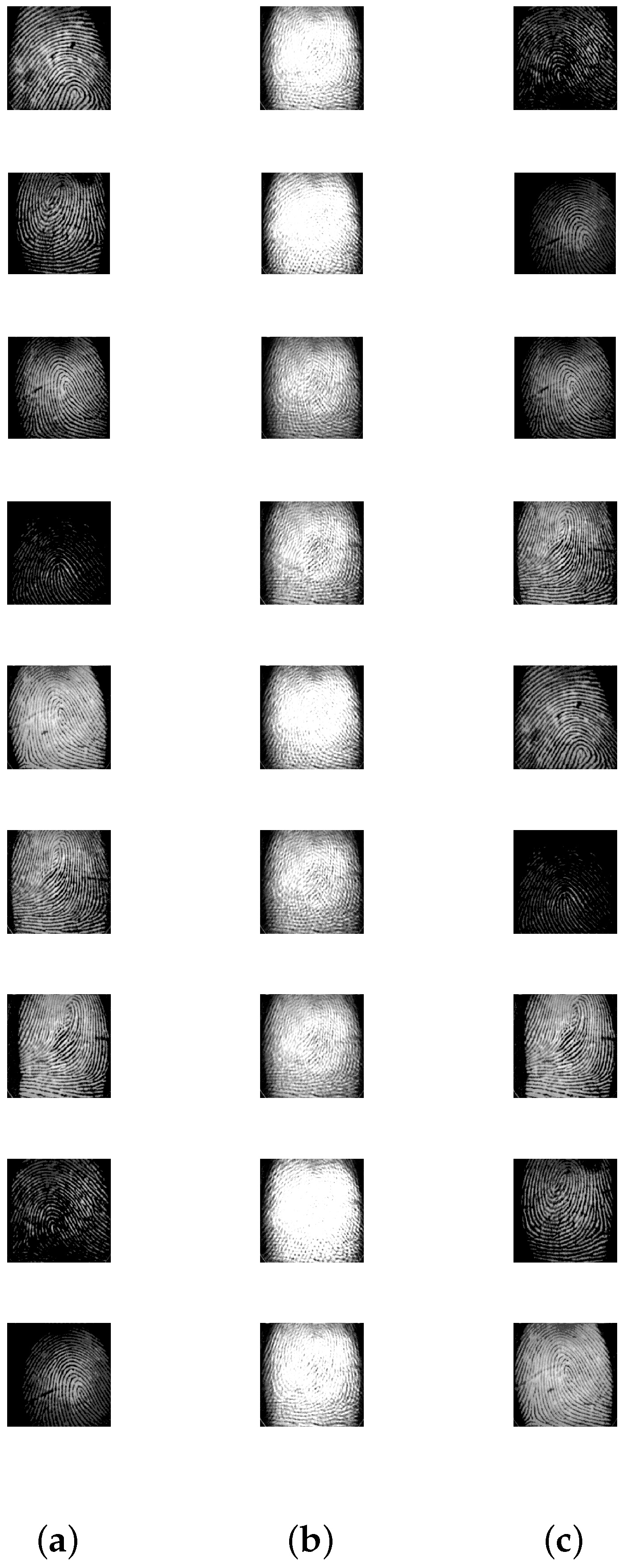
5.2. Image Mixture Separation
5.3. Comparative Experiment-I
5.4. Comparative Experiment-II
6. Discussion and Conclusions
Acknowledgments
Conflicts of Interest
References
- Joho, M.; Mathis, H.; Lambert, R.H. Overdetermined blind source separation: Using more sensors than source signals in a noisy mixture. In Proceedings of the Independent Component Analysis and Blind Signal Separation, Helsinki, Finlan, 19–22 June 2000; pp. 81–86. [Google Scholar]
- Winter, S.; Sawada, H.; Makino, S. Geometrical interpretation of the PCA subspace approach for overdetermined blind source separation. EURASIP J. Adv. Signal Process. 2006, 2006, 071632. [Google Scholar]
- Bofill, P.; Zibulevsky, M. Underdetermined blind source separation using sparse representations. Signal Process. 2001, 81, 2353–2362. [Google Scholar]
- Zhen, L.; Peng, D.; Yi, Z.; Xiang, Y.; Chen, P. Underdetermined blind source separation using sparse coding. IEEE Trans. Neural Netw. Learn. Syst. 2017, 28, 3102–3108. [Google Scholar] [CrossRef] [PubMed]
- Herault, J.; Jutten, C.; Ans, B. Detection de Grandeurs Primitives dans un Message Composite par une Architecture de Calcul Neuromimetique en Apprentissage Non Supervise. In 1985—GRETSI—Actes de Colloques; Groupe d’Etudes du Traitement du Signal et des Images: Juan-les-Pins, France, 1985; pp. 1017–1022. [Google Scholar]
- Syed, M.; Georgiev, P.; Pardalos, P. A hierarchical approach for sparse source blind signal separation problem. Comput. Oper. Res. 2012, 41, 386–398. [Google Scholar] [CrossRef]
- Hyvärinen, A.; Karhunen, J.; Oja, E. Independent Component Analysis; John Wiley & Sons: Hoboken, NJ, USA, 2004; Volume 46. [Google Scholar]
- Amari, S.I.; Cichocki, A.; Yang, H.H. A new learning algorithm for blind signal separation. In Proceedings of the 8th International Conference on Neural Information Processing Systems, Denver, CO, USA, 27 November–2 December 1996; pp. 757–763. [Google Scholar]
- Comon, P. Independent component analysis, a new concept? Signal Process. 1994, 36, 287–314. [Google Scholar] [CrossRef]
- Hyvärinen, A. New approximations of differential entropy for independent component analysis and projection pursuit. In Proceedings of the 1997 Conference on Advances in Neural Information Processing Systems, Denver, CO, USA, 1–6 December 1998; pp. 273–279. [Google Scholar]
- Bell, A.J.; Sejnowski, T.J. An information-maximization approach to blind separation and blind deconvolution. Neural Comput. 1995, 7, 1129–1159. [Google Scholar] [CrossRef] [PubMed]
- Chai, R.; Naik, G.R.; Nguyen, T.N.; Ling, S.H.; Tran, Y.; Craig, A.; Nguyen, H.T. Driver fatigue classification with independent component by entropy rate bound minimization analysis in an EEG-based system. IEEE J. Biomed. Health Inform. 2017, 21, 715–724. [Google Scholar] [CrossRef] [PubMed]
- Naik, G.R.; Baker, K.G.; Nguyen, H.T. Dependence independence measure for posterior and anterior EMG sensors used in simple and complex finger flexion movements: Evaluation using SDICA. IEEE J. Biomed. Health Inform. 2015, 19, 1689–1696. [Google Scholar] [CrossRef] [PubMed]
- Naik, G.R.; Al-Timemy, A.H.; Nguyen, H.T. Transradial amputee gesture classification using an optimal number of sEMG sensors: An approach using ICA clustering. IEEE Trans. Neural Syst. Rehabil. Eng. 2016, 24, 837–846. [Google Scholar]
- Deslauriers, J.; Ansado, J.; Marrelec, G.; Provost, J.S.; Joanette, Y. Increase of posterior connectivity in aging within the Ventral Attention Network: A functional connectivity analysis using independent component analysis. Brain Res. 2017, 1657, 288–296. [Google Scholar] [CrossRef] [PubMed]
- O’Muircheartaigh, J.; Jbabdi, S. Concurrent white matter bundles and grey matter networks using independent component analysis. NeuroImage 2017. [Google Scholar] [CrossRef] [PubMed]
- Hand, B.N.; Dennis, S.; Lane, A.E. Latent constructs underlying sensory subtypes in children with autism: A preliminary study. Autism Res. 2017, 10, 1364–1371. [Google Scholar] [CrossRef] [PubMed]
- Arya, Y. AGC performance enrichment of multi-source hydrothermal gas power systems using new optimized FOFPID controller and redox flow batteries. Energy 2017, 127, 704–715. [Google Scholar]
- Al-Ali, A.K.H.; Senadji, B.; Naik, G.R. Enhanced forensic speaker verification using multi-run ICA in the presence of environmental noise and reverberation conditions. In Proceedings of the 2017 IEEE International Conference on Signal and Image Processing Applications (ICSIPA), Kuching, Malaysia, 12–14 September 2017; pp. 174–179. [Google Scholar]
- Comon, P.; Jutten, C. Handbook of Blind Source Separation: Independent Component Analysis and Applications; Academic Press: Cambridge, MA, USA, 2010. [Google Scholar]
- Nascimento, J.M.; Dias, J.M. Does independent component analysis play a role in unmixing hyperspectral data? IEEE Trans. Geosci. Remote Sens. 2005, 43, 175–187. [Google Scholar] [CrossRef]
- Syed, M.; Georgiev, P.; Pardalos, P. Robust Physiological Mappings: From Non-Invasive to Invasive. Cybern. Syst. Anal. 2015, 1, 96–104. [Google Scholar]
- Georgiev, P.; Theis, F.; Cichocki, A.; Bakardjian, H. Sparse component analysis: A new tool for data mining. In Data Mining Biomedicine; Springer: Boston, MA, USA, 2007; pp. 91–116. [Google Scholar]
- Naanaa, W.; Nuzillard, J. Blind source separation of positive and partially correlated data. Signal Process. 2005, 85, 1711–1722. [Google Scholar] [CrossRef]
- Chan, T.H.; Ma, W.K.; Chi, C.Y.; Wang, Y. A convex analysis framework for blind separation of non-negative sources. IEEE Trans. Signal Process. 2008, 56, 5120–5134. [Google Scholar] [CrossRef]
- Chan, T.H.; Chi, C.Y.; Huang, Y.M.; Ma, W.K. A convex analysis-based minimum-volume enclosing simplex algorithm for hyperspectral unmixing. IEEE Trans. Signal Process. 2009, 57, 4418–4432. [Google Scholar] [CrossRef]
- Syed, M.N.; Georgiev, P.G.; Pardalos, P.M. Blind Signal Separation Methods in Computational Neuroscience. In Modern Electroencephalographic Assessment Techniques: Theory and Applications; Humana Press: New York, NY, USA, 2015; pp. 291–322. [Google Scholar]
- Ma, W.K.; Bioucas-Dias, J.M.; Chan, T.H.; Gillis, N.; Gader, P.; Plaza, A.J.; Ambikapathi, A.; Chi, C.Y. A signal processing perspective on hyperspectral unmixing: Insights from remote sensing. IEEE Signal Process. Mag. 2014, 31, 67–81. [Google Scholar] [CrossRef]
- Bioucas-Dias, J.M.; Plaza, A.; Dobigeon, N.; Parente, M.; Du, Q.; Gader, P.; Chanussot, J. Hyperspectral unmixing overview: Geometrical, statistical, and sparse regression-based approaches. IEEE J. Sel. Top. Appl. Earth Obs. Remote Sens. 2012, 5, 354–379. [Google Scholar] [CrossRef]
- Yin, P.; Sun, Y.; Xin, J. A geometric blind source separation method based on facet component analysis. Signal Image Video Process. 2016, 10, 19–28. [Google Scholar] [CrossRef]
- Lin, C.H.; Chi, C.Y.; Wang, Y.H.; Chan, T.H. A Fast Hyperplane-Based Minimum-Volume Enclosing Simplex Algorithm for Blind Hyperspectral Unmixing. IEEE Trans. Signal Process. 2016, 64, 1946–1961. [Google Scholar] [CrossRef]
- Zhang, S.; Agathos, A.; Li, J. Robust Minimum Volume Simplex Analysis for Hyperspectral Unmixing. IEEE Trans. Geosci. Remote Sens. 2017, 55, 6431–6439. [Google Scholar] [CrossRef]
- Naanaa, W.; Nuzillard, J.M. Extreme direction analysis for blind separation of nonnegative signals. Signal Process. 2017, 130, 254–267. [Google Scholar]
- Sun, Y.; Ridge, C.; Del Rio, F.; Shaka, A.; Xin, J. Postprocessing and sparse blind source separation of positive and partially overlapped data. Signal Process. 2011, 91, 1838–1851. [Google Scholar] [CrossRef]
- Aharon, M.; Elad, M.; Bruckstein, A. On the uniqueness of overcomplete dictionaries, and a practical way to retrieve them. Linear Algebra Appl. 2006, 416, 48–67. [Google Scholar] [CrossRef]
- Georgiev, P.; Theis, F.; Ralescu, A. Identifiability conditions and subspace clustering in sparse BSS. In Independent Component Analysis and Signal Separation; Springer: Berlin/Heidelberg, Germany, 2007; pp. 357–364. [Google Scholar]
- Drumetz, L.; Veganzones, M.A.; Henrot, S.; Phlypo, R.; Chanussot, J.; Jutten, C. Blind hyperspectral unmixing using an Extended Linear Mixing Model to address spectral variability. IEEE Trans. Image Process. 2016, 25, 3890–3905. [Google Scholar] [CrossRef] [PubMed]
- Amini, F.; Hedayati, Y. Underdetermined blind modal identification of structures by earthquake and ambient vibration measurements via sparse component analysis. J. Sound Vib. 2016, 366, 117–132. [Google Scholar] [CrossRef]
- Gribonval, R.; Schnass, K. Dictionary Identification—Sparse Matrix-Factorization via l1-Minimization. IEEE Trans. Inf. Theory 2010, 56, 3523–3539. [Google Scholar]
- Kreutz-Delgado, K.; Murray, J.; Rao, B.; Engan, K.; Lee, T.; Sejnowski, T. Dictionary learning algorithms for sparse representation. Neural Comput. 2003, 15, 349–396. [Google Scholar] [CrossRef] [PubMed]
- Nascimento, J.M.; Bioucas-Dias, J.M. Blind hyperspectral unmixing. In Proceedings of the SPIE Conference on Image and Signal Processing for Remote Sensing XIII, Florence, Italy, 18–20 September 2007. [Google Scholar]
- Duarte, L.T.; Moussaoui, S.; Jutten, C. Source separation in chemical analysis: Recent achievements and perspectives. IEEE Signal Process. Mag. 2014, 31, 135–146. [Google Scholar] [CrossRef]
- Sun, Y.; Xin, J. Nonnegative Sparse Blind Source Separation for NMR Spectroscopy by Data Clustering, Model Reduction, and ℓ1 Minimization. SIAM J. Imag. Sci. 2012, 5, 886–911. [Google Scholar] [CrossRef]
- Winter, M.E. N-FINDR: An algorithm for fast autonomous spectral end-member determination in hyperspectral data. In Proceedings of the Imaging Spectrometry V, Denver, CO, USA, 19–21 July 1999; pp. 266–275. [Google Scholar]
- Nascimento, J.M.; Dias, J.M. Vertex component analysis: A fast algorithm to unmix hyperspectral data. IEEE Trans. Geosci. Remote Sens. 2005, 43, 898–910. [Google Scholar] [CrossRef]
- Santamaria, I.; Pokharel, P.P.; Principe, J.C. Generalized correlation function: Definition, properties, and application to blind equalization. IEEE Trans. Signal Process. 2006, 54, 2187–2197. [Google Scholar] [CrossRef]
- Liu, W.; Pokharel, P.P.; Príncipe, J.C. Correntropy: Properties and applications in non-Gaussian signal processing. IEEE Trans. Signal Process. 2007, 55, 5286–5298. [Google Scholar] [CrossRef]
- Singh, A.; Principe, J.C. Using correntropy as a cost function in linear adaptive filters. In Proceedings of the 2009 International Joint Conference on Neural Networks, Atlanta, GA, USA, 14–19 June 2009; pp. 2950–2955. [Google Scholar]
- Zhao, S.; Chen, B.; Principe, J.C. Kernel adaptive filtering with maximum correntropy criterion. In Proceedings of the 2011 International Joint Conference on Neural Networks, San Jose, CA, USA, 31 July–5 August 2011; pp. 2012–2017. [Google Scholar]
- Chen, B.; Xing, L.; Liang, J.; Zheng, N.; Principe, J.C. Steady-state mean-square error analysis for adaptive filtering under the maximum correntropy criterion. IEEE Signal Process. Lett. 2014, 21, 880–884. [Google Scholar]
- Chen, B.; Xing, L.; Zhao, H.; Zheng, N.; Principe, J.C. Generalized correntropy for robust adaptive filtering. IEEE Trans. Signal Process. 2016, 64, 3376–3387. [Google Scholar] [CrossRef]
- Chen, B.; Liu, X.; Zhao, H.; Principe, J.C. Maximum correntropy Kalman filter. Automatica 2017, 76, 70–77. [Google Scholar] [CrossRef]
- Syed, M.N.; Pardalos, P.M.; Principe, J.C. On the optimization properties of the correntropic loss function in data analysis. Optim. Lett. 2014, 8, 823–839. [Google Scholar] [CrossRef]
- Kuhn, H.W. The hungarian metRhod for the assignment problem. In 50 Years of Integer Programming 1958–2008; Springer: Berlin, Germany, 2010; pp. 29–47. [Google Scholar]
- Cplex, I. User-Manual CPLEX; IBM Software Group: New York, NY, USA, 2011; Volume 12. [Google Scholar]
- Li, J.; Bioucas-Dias, J.M. Minimum volume simplex analysis: A fast algorithm to unmix hyperspectral data. In Proceedings of the 2008 IEEE International Geoscience and Remote Sensing Symposium, Boston, MA, USA, 7–11 July 2008; Volume 3. [Google Scholar] [CrossRef]
- Terlaky, T. Interior Point Methods of Mathematical Programming; Springer Science & Business Media: Berlin, Germany, 2013; Volume 5. [Google Scholar]
- Dantzig, G.B.; Thapa, M.N. Linear Programming 2: Theory and Extensions; Springer Science & Business Media: Berlin, Germany, 2006. [Google Scholar]
- Andersen, E.D.; Andersen, K.D. Presolving in linear programming. Math. Program. 1995, 71, 221–245. [Google Scholar] [CrossRef]
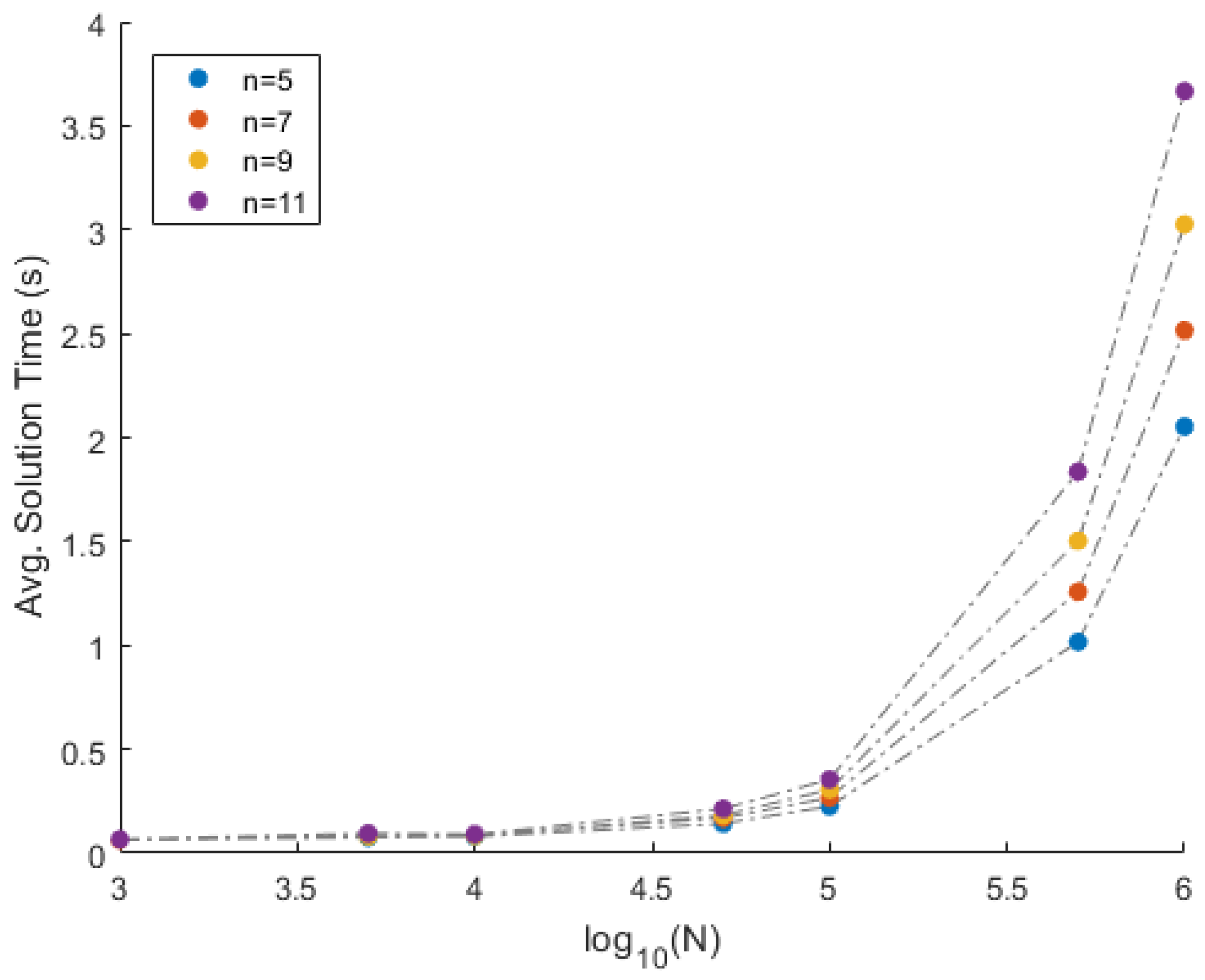



| n × N | mErrA | vErrA | mErrS | vErrS | mTime | vTime | mRed | vRed | nMiss |
|---|---|---|---|---|---|---|---|---|---|
| 5 × 1000 | 1.04 × 10 | 9.78 × 10 | 1.17 × 10 | 4.76 × 10 | 0.0647 | 0.07095 | 50 | 0 | 0 |
| 5 × 5000 | 1.08 × 10 | 1.16 × 10 | 2.03 × 10 | 5.24 × 10 | 0.0756 | 0.04114 | 50 | 0 | 0 |
| 5 × 10,000 | 1.07 × 10 | 1.26 × 10 | 9.50 × 10 | 1.88 × 10 | 0.08 | 0.16436 | 50 | 0 | 0 |
| 5 × 50,000 | 1.10 × 10 | 1.13 × 10 | 5.10 × 10 | 5.39 × 10 | 0.1427 | 0.09452 | 50 | 0 | 0 |
| 5 × 100,000 | 1.02 × 10 | 8.22 × 10 | 4.85 × 10 | 3.21 × 10 | 0.2254 | 0.12459 | 50 | 0 | 0 |
| 5 × 500,000 | 1.01 × 10 | 9.28 × 10 | 1.81 × 10 | 6.50 × 10 | 1.0166 | 2.2 | 50 | 0 | 0 |
| 5 × 1,000,000 | 1.03 × 10 | 1.11 × 10 | 1.19 × 10 | 8.55 × 10 | 2.0526 | 1.9 | 50 | 0 | 0 |
| 7 × 1000 | 7.28 × 10 | 3.47 × 10 | 1.30 × 10 | 3.41 × 10 | 0.0641 | 0.05835 | 50 | 0 | 0 |
| 7 × 5000 | 6.85 × 10 | 2.72 × 10 | 1.71 × 10 | 1.08 × 10 | 0.0816 | 0.06362 | 50 | 0 | 0 |
| 7 × 10,000 | 6.86 × 10 | 2.28 × 10 | 1.01 × 10 | 7.70 × 10 | 0.0891 | 0.11015 | 50 | 0 | 0 |
| 7 × 50,000 | 7.08 × 10 | 1.89 × 10 | 7.05 × 10 | 4.27 × 10 | 0.1689 | 0.14786 | 50 | 0 | 0 |
| 7 × 100,000 | 6.78 × 10 | 2.46 × 10 | 1.14 × 10 | 5.67 × 10 | 0.2675 | 0.14104 | 50 | 0 | 0 |
| 7 × 500,000 | 7.26 × 10 | 2.61 × 10 | 6.11 × 10 | 1.80 × 10 | 1.2584 | 1.3 | 50 | 0 | 0 |
| 7 × 1,000,000 | 6.85 × 10 | 2.62 × 10 | 9.93 × 10 | 1.77 × 10 | 2.5154 | 3.3 | 50 | 0 | 0 |
| 9 × 1000 | 5.50 × 10 | 1.05 × 10 | 1.59 × 10 | 2.94 × 10 | 0.067 | 0.10896 | 50 | 0 | 0 |
| 9 × 5000 | 5.82 × 10 | 1.42 × 10 | 2.44 × 10 | 1.96 × 10 | 0.0812 | 0.05954 | 50 | 0 | 0 |
| 9 × 10,000 | 5.62 × 10 | 1.30 × 10 | 8.13 × 10 | 4.63 × 10 | 0.0835 | 0.08635 | 50 | 0 | 0 |
| 9 × 50,000 | 5.52 × 10 | 1.38 × 10 | 2.62 × 10 | 5.69 × 10 | 0.1821 | 0.12421 | 50 | 0 | 0 |
| 9 × 100,000 | 5.57 × 10 | 1.31 × 10 | 3.79 × 10 | 1.68 × 10 | 0.3066 | 0.1855 | 50 | 0 | 0 |
| 9 × 500,000 | 5.40 × 10 | 1.36 × 10 | 9.78 × 10 | 1.84 × 10 | 1.5029 | 1 | 50 | 0 | 0 |
| 9 × 1,000,000 | 5.32 × 10 | 1.21 × 10 | 1.49 × 10 | 2.10 × 10 | 3.0258 | 3.1 | 50 | 0 | 0 |
| 11 × 1000 | 4.05 × 10 | 4.77 × 10 | 1.75 × 10 | 3.13 × 10 | 0.0672 | 0.02826 | 50 | 0 | 0 |
| 11 × 5000 | 4.16 × 10 | 5.04 × 10 | 2.03 × 10 | 9.47 × 10 | 0.096 | 0.10778 | 50 | 0 | 0 |
| 11 × 10,000 | 4.21 × 10 | 4.87 × 10 | 1.31 × 10 | 7.33 × 10 | 0.0916 | 0.00928 | 50 | 0 | 0 |
| 11 × 50,000 | 4.09 × 10 | 5.02 × 10 | 6.61 × 10 | 2.97 × 10 | 0.2137 | 0.05729 | 50 | 0 | 0 |
| 11 × 100,000 | 4.12 × 10 | 3.79 × 10 | 1.90 × 10 | 6.92 × 10 | 0.355 | 0.25758 | 50 | 0 | 0 |
| 11 × 500,000 | 4.14 × 10 | 3.96 × 10 | 1.18 × 10 | 1.66 × 10 | 1.8358 | 0.69888 | 50 | 0 | 0 |
| 11 × 1,000,000 | 4.21 × 10 | 4.37 × 10 | 8.91 × 10 | 8.11 × 10 | 3.667 | 5.4 | 50 | 0 | 0 |
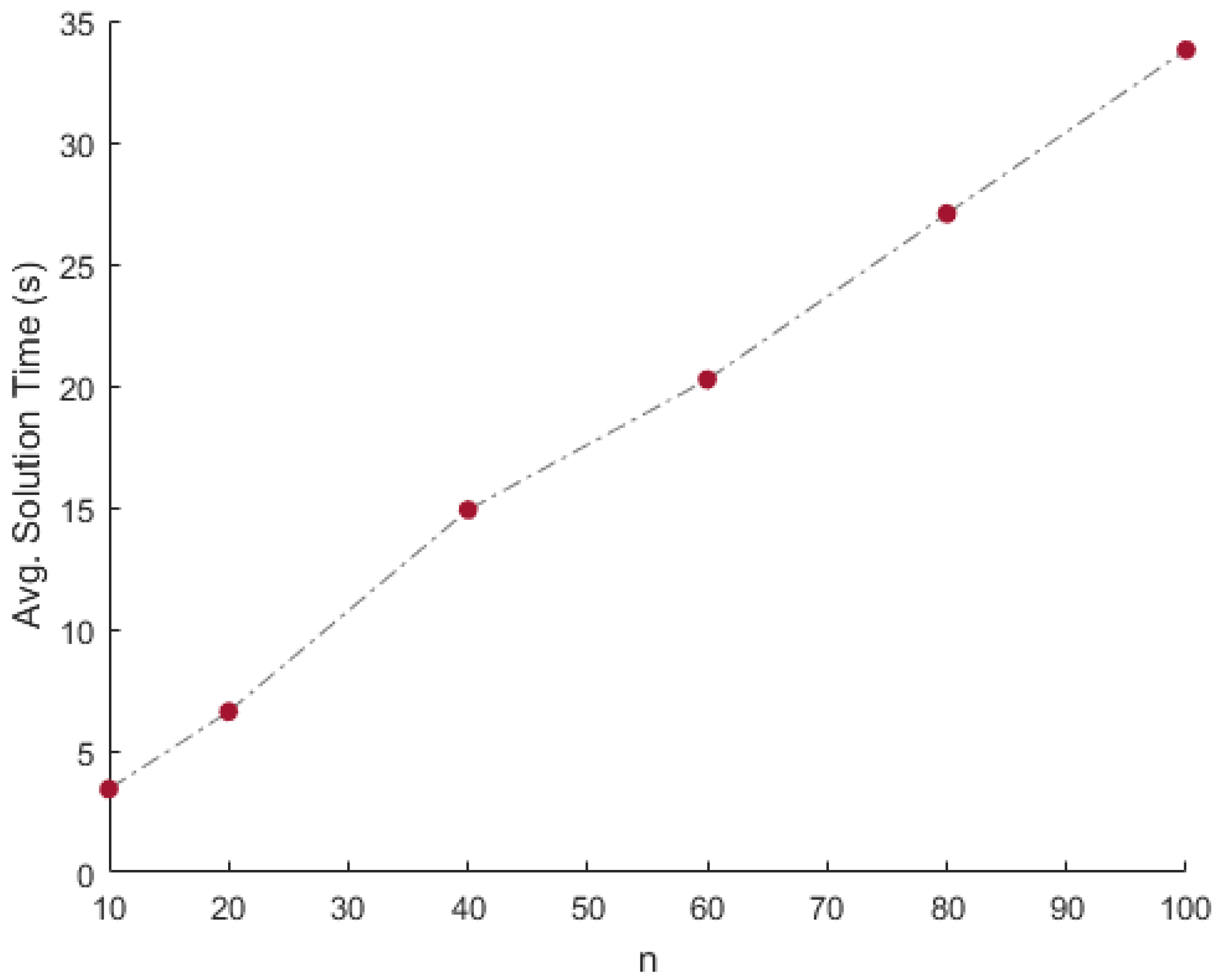
| 10 × 1,000,000 | 4.71 × 10 | 7.54 × 10 | 8.46 × 10 | 8.89 × 10 | 3.4245 | 2.5 | 50 | 0 | 0 |
| 20 × 1,000,000 | 1.83 × 10 | 5.10 × 10 | 1.12 × 10 | 6.32 × 10 | 6.6053 | 9.3 | 50 | 0 | 0 |
| 40 × 1,000,000 | 7.84 × 10 | 5.33 × 10 | 4.22 × 10 | 9.61 × 10 | 14.8988 | 64.8 | 50 | 0 | 0 |
| 60 × 1,000,000 | 7.32 × 10 | 4.62 × 10 | 1.05 × 10 | 2.18 × 10 | 20.2509 | 628.9 | 50 | 0 | 0 |
| 80 × 1,000,000 | 6.51 × 10 | 6.39 × 10 | 7.51 × 10 | 1.40 × 10 | 27.0654 | 101.7 | 50 | 0 | 0 |
| 100 × 1,000,000 | 5.13 × 10 | 2.83 × 10 | 4.65 × 10 | 5.57 × 10 | 33.7978 | 136.6 | 50 | 0 | 0 |
| Image Set | n | N |
|---|---|---|
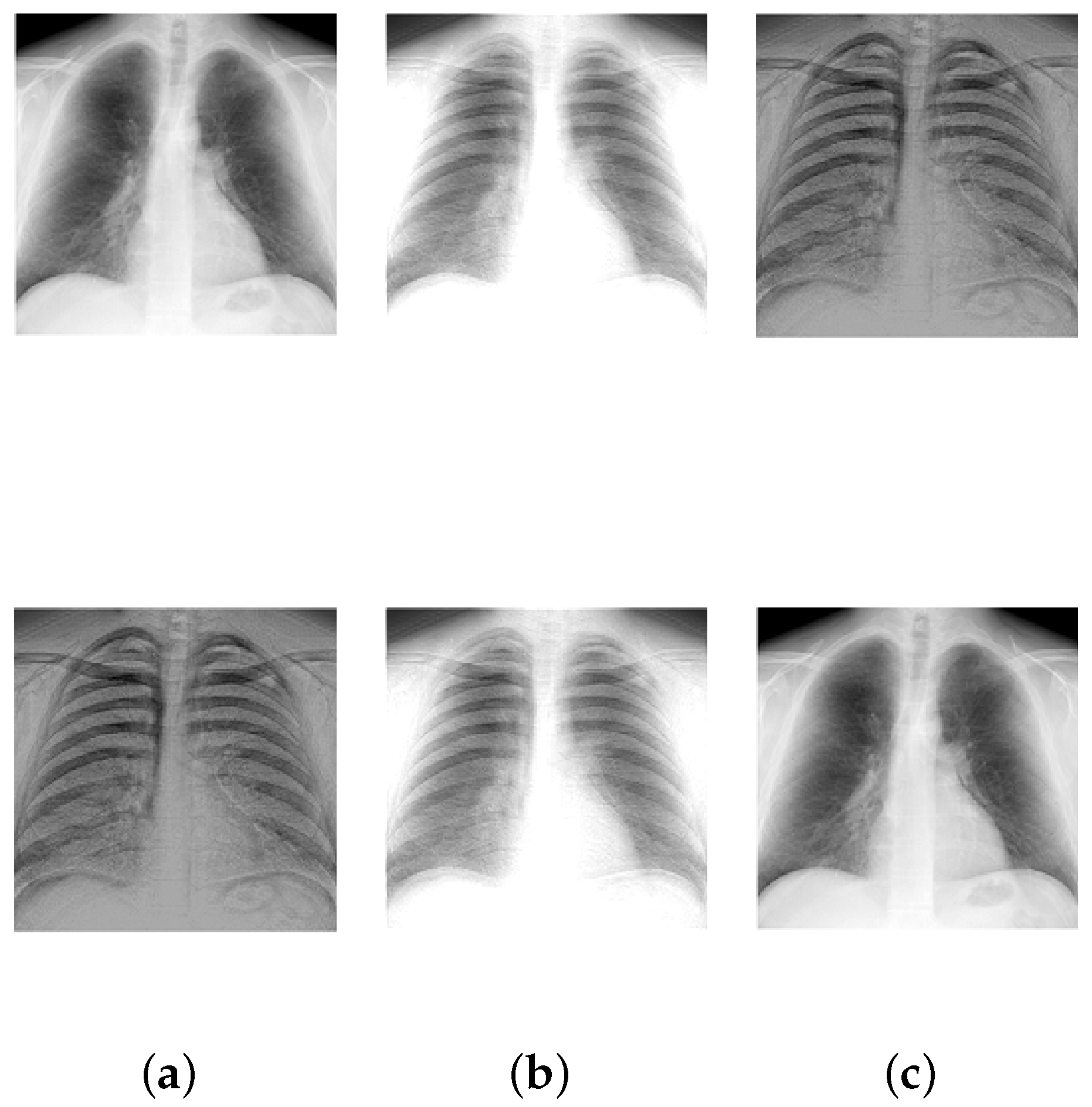
| Chest X-rays | 2 | 26,896 |
| Scenery | 3 | 65,536 |
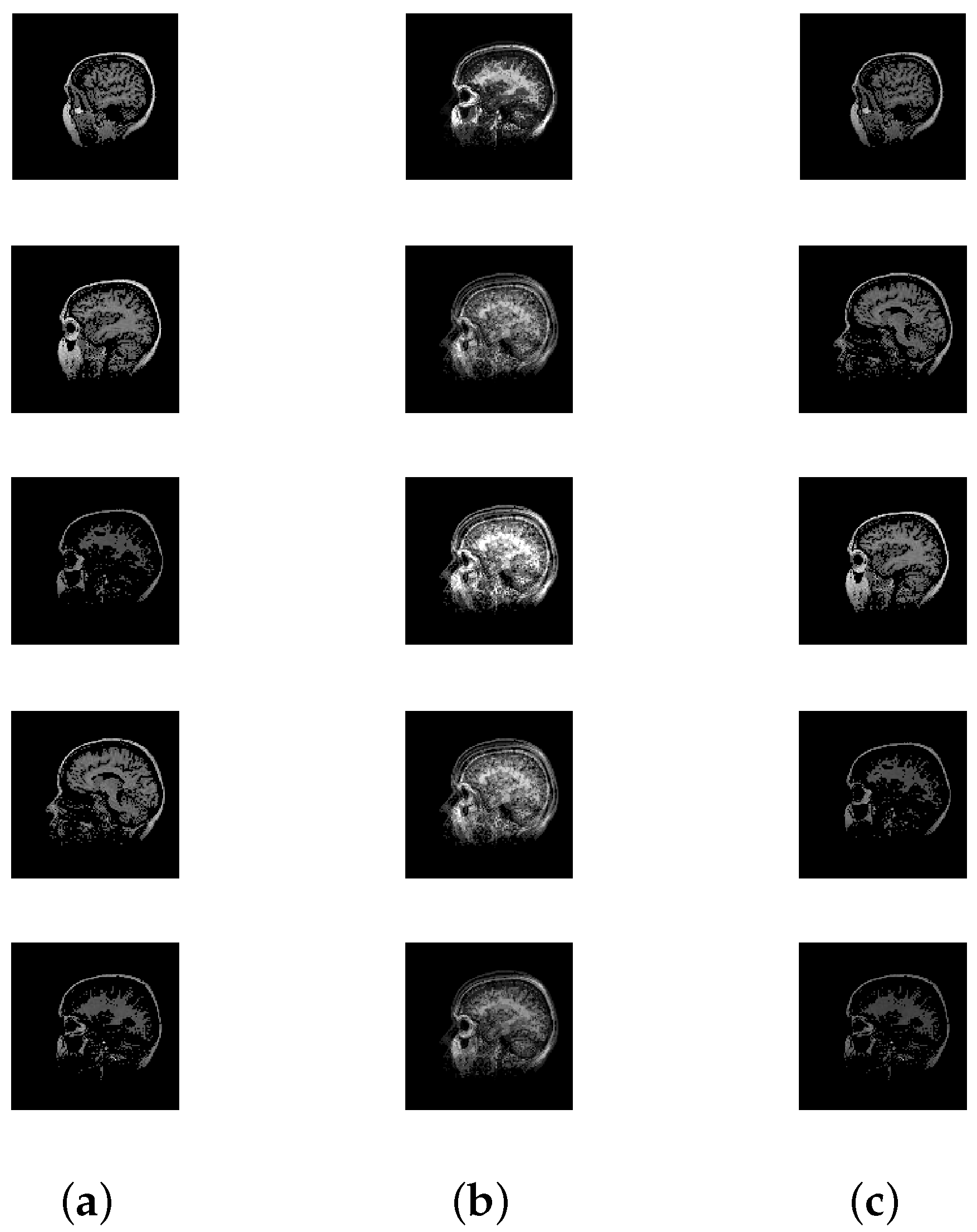
| CT Scans | 5 | 16,384 |
| Zip Codes | 7 | 12,672 |
| Finger Print | 9 | 90,000 |
| Image Set | mErrA | vErrA | mErrS | vErrS | mTime | vTime | mRed | vRed | nMiss |
|---|---|---|---|---|---|---|---|---|---|
| Chest X-rays | 2.45 × 10 | 2.81 × 10 | 1.33 × 10 | 2.02 × 10 | 0.0755 | 4.89 × 10 | 81.632 | 0.0101 | 0 |
| Scenery | 2.06 × 10 | 5.30 × 10 | 4.10 × 10 | 4.12 × 10 | 0.109 | 9.81 × 10 | 73.2417 | 0.0033 | 7 |
| CT Scan | 1.19 × 10 | 9.09 × 10 | 4.80 × 10 | 1.20 × 10 | 0.0679 | 1.34 × 10 | 89.7026 | 0.0012 | 4 |
| Zip Codes | 7.36 × 10 | 2.61 × 10 | 4.96 × 10 | 8.17 × 10 | 0.0787 | 3.74 × 10 | 74.0513 | 0.0047 | 6 |
| Finger Print | 5.72 × 10 | 1.19 × 10 | 1.67 × 10 | 2.23 × 10 | 0.2716 | 1.57 × 10 | 55.952 | 6.49 × 10 | 0 |
| n | N | VCA | MVSA | N-FINDR | Proposed | ||||
|---|---|---|---|---|---|---|---|---|---|
| mErrA | vErrA | mErrA | vErrA | mErrA | vErrA | ErrA | TnMiss | ||
| 5 | 10,000 | 0.0755 | 7.08 × 10 | 0.0813 | 6.65 × 10 | 0.0905 | 4.14 × 10 | — | 100 |
| 7 | 10,000 | 0.056 | 9.63 × 10 | 0.0567 | 2.6 × 10 | 0.0604 | 7.23 × 10 | — | 100 |
| 9 | 10,000 | 0.0422 | 2.96 × 10 | 0.0402 | 1.13 × 10 | 0.0441 | 1.34 × 10 | — | 100 |
| 11 | 10,000 | 0.0333 | 8.56 × 10 | 0.0314 | 4.16 × 10 | 0.0342 | 7.16 × 10 | — | 100 |
| 13 | 10,000 | 0.0269 | 3.28 × 10 | 0.0252 | 1.56 × 10 | 0.0274 | 3.6 × 10 | — | 100 |
| 15 | 10,000 | 0.0223 | 1.52 × 10 | 0.0212 | 7.53 × 10 | 0.0226 | 1.8 × 10 | — | 100 |
| n | N | VCA | MVSA | N-FINDR | Proposed |
|---|---|---|---|---|---|
| 5 | 10,000 | 0.0798 | 0.0872 | 0.0954 | 6.43 × |
| 7 | 10,000 | 0.0573 | 0.045 | 0.0658 | 4.73 × |
| 9 | 10,000 | 0.0407 | 0.0309 | 0.0467 | 3.77 × |
| 11 | 10,000 | 0.0321 | 0.0265 | 0.0357 | 0.0012 |
| 13 | 10,000 | 0.0255 | 0.0231 | 0.0284 | 0.0007 |
| 15 | 10,000 | 0.0212 | 0.0197 | 0.0235 | 0.0021 |
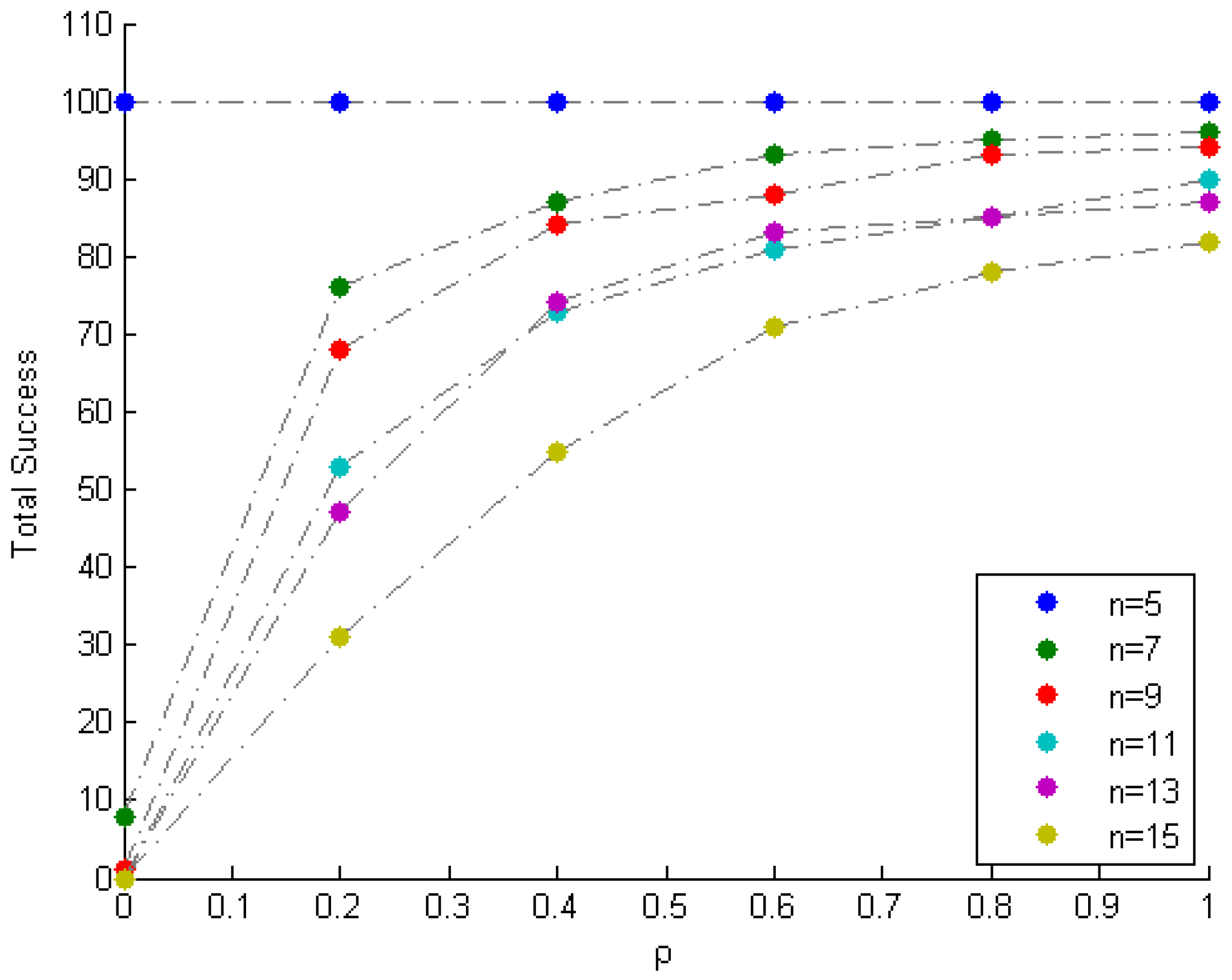
| n | N | = 0 | = 0.2 | = 0.4 | = 0.6 | = 0.8 | = 1 |
|---|---|---|---|---|---|---|---|
| 5 | 10,000 | 0 | 0 | 0 | 0 | 0 | 0 |
| 7 | 10,000 | 92 | 24 | 13 | 7 | 5 | 4 |
| 9 | 10,000 | 99 | 32 | 16 | 12 | 7 | 6 |
| 11 | 10,000 | 100 | 47 | 27 | 19 | 15 | 10 |
| 13 | 10,000 | 100 | 53 | 26 | 17 | 15 | 13 |
| 15 | 10,000 | 100 | 69 | 45 | 29 | 22 | 18 |
© 2018 by the author. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Syed, M.N. Big Data Blind Separation. Entropy 2018, 20, 150. https://doi.org/10.3390/e20030150
Syed MN. Big Data Blind Separation. Entropy. 2018; 20(3):150. https://doi.org/10.3390/e20030150
Chicago/Turabian StyleSyed, Mujahid N. 2018. "Big Data Blind Separation" Entropy 20, no. 3: 150. https://doi.org/10.3390/e20030150
APA StyleSyed, M. N. (2018). Big Data Blind Separation. Entropy, 20(3), 150. https://doi.org/10.3390/e20030150





