DPP9 as a Potential Novel Mediator in Gastrointestinal Virus Infection
Abstract
:1. Introduction
2. Materials and Methods
2.1. Colon Cohort
2.2. Jejunum Cohort
2.3. Metabolomics Validation Cohort
2.4. Clinical Biochemistry of Colon Cohort
2.5. Clinical Biochemistry of Jejunum Cohort
2.6. Colon Transcriptomics
2.7. Jejunum Transcriptomics
2.8. Circulating DPP9 Expression Analysis for Metabolomics Validation Cohort
2.9. Plasma Non-Targeted Metabolomics for Colon Cohort
2.10. Plasma Non-Targeted Metabolomics for Metabolomics Validation Cohort
2.11. Statistical Analysis
2.11.1. Gene Co-Expression Patterns
2.11.2. Enrichment Networks
2.11.3. Gene-Gene Interaction Networks
2.11.4. Machine Learning Variable Selection Analysis
2.11.5. Gene-Drug Enrichment Analyses
3. Results
3.1. The DPP9 Expression Levels in Jejunum, Colon, and Blood Are Strongly Associated with Biomarkers and Genes Involved in Oxidative Stress and Antioxidant Defense
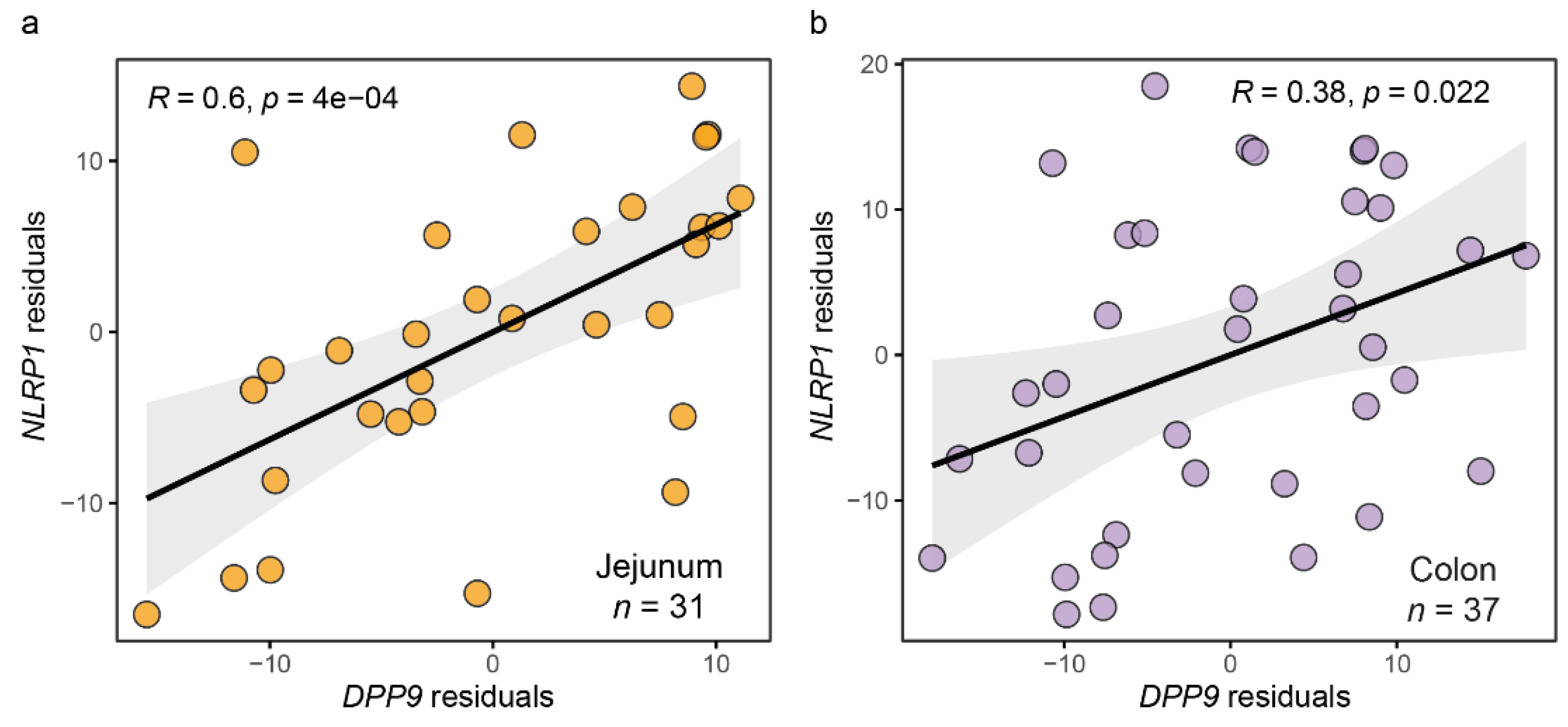
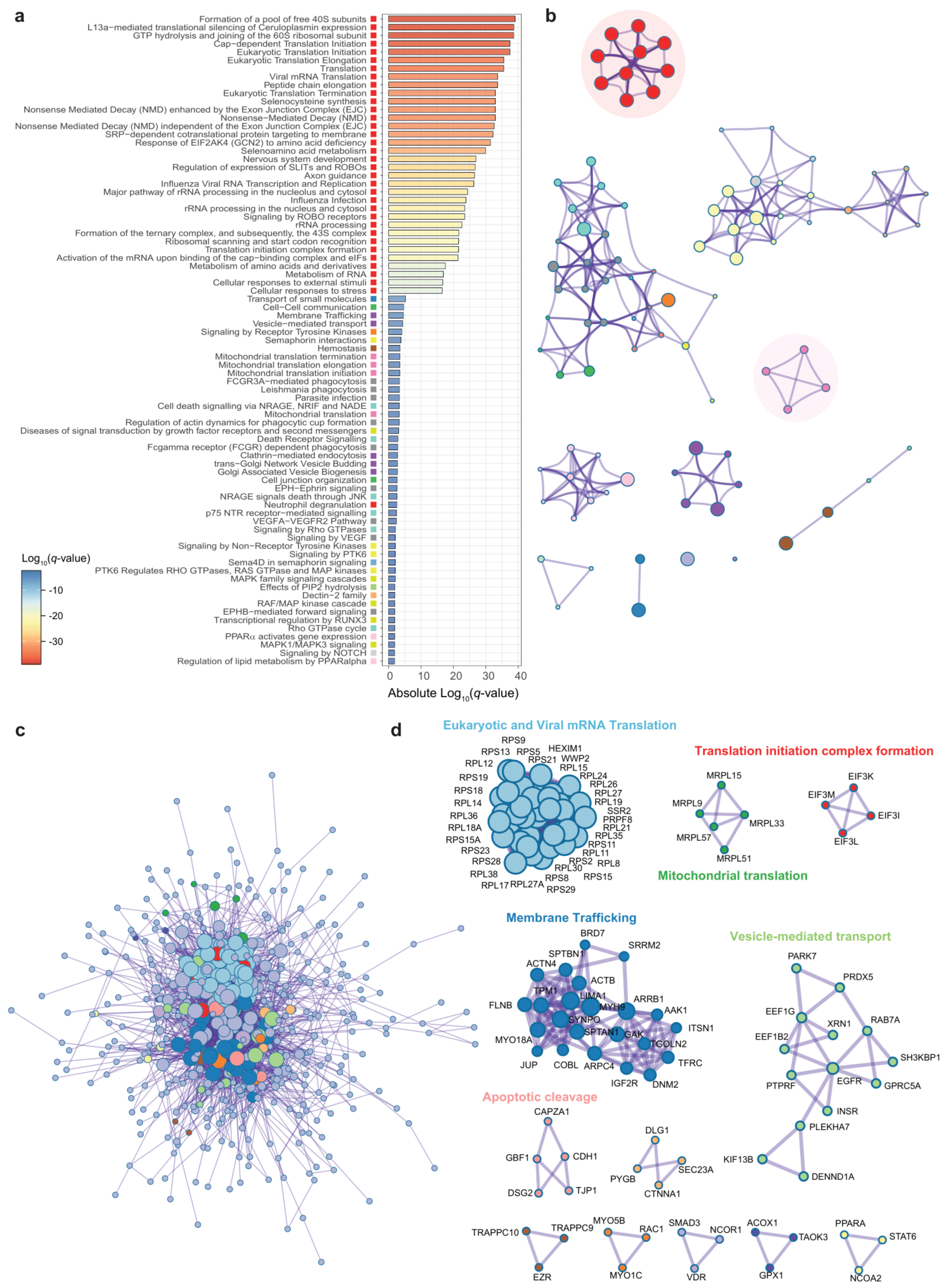
3.2. Genes Associated to DPP9 Expression in the Colon Are Involved in Viral Replication, Antiviral Mechanisms and Metabolism of Nucleotides
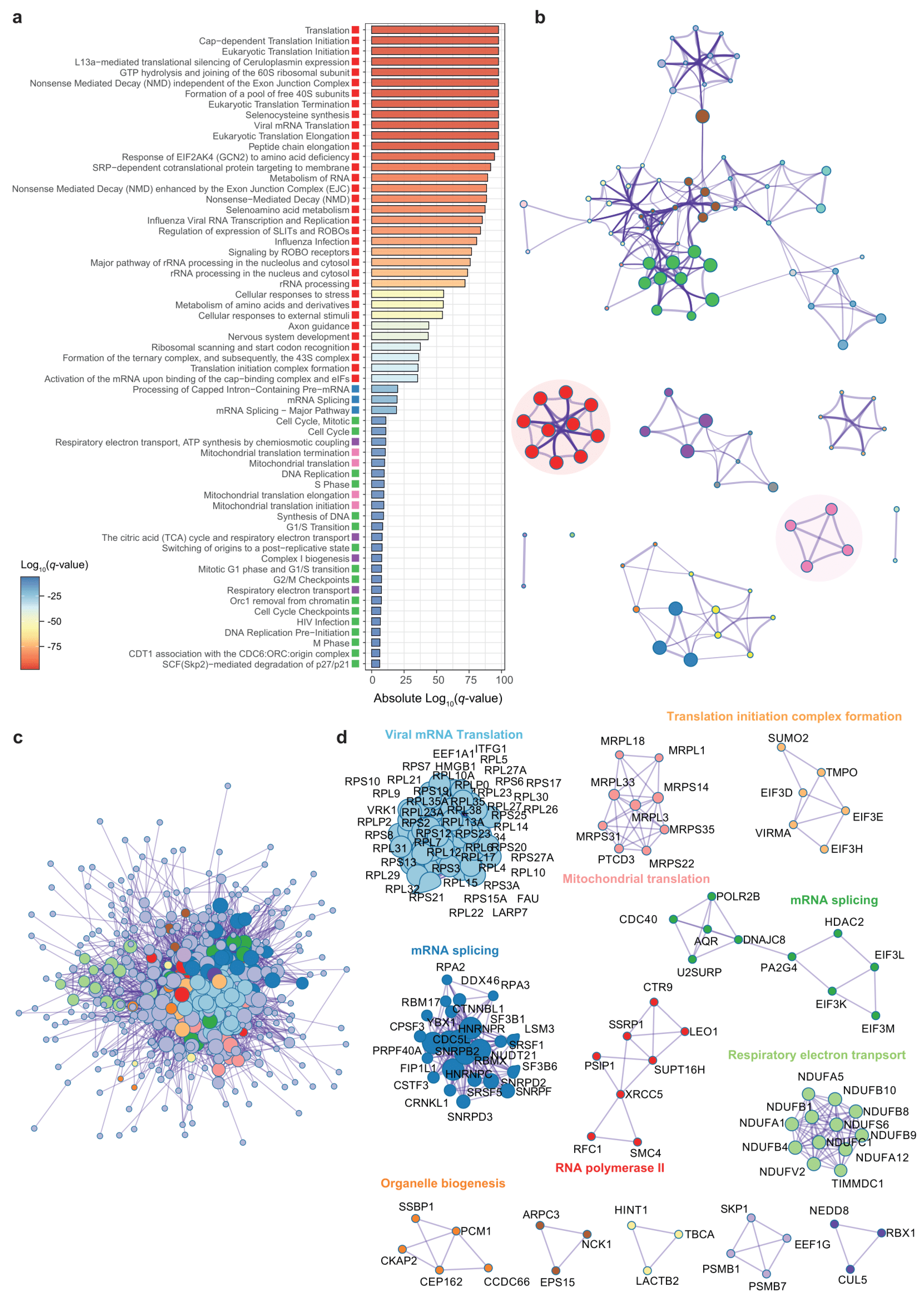
3.3. Genes Associated to DPP9 Expression in the Jejunum Are Also Involved in Viral Replication and Antiviral Mechanisms
3.4. Metabolomics Signatures Associated with DPP9 in Plasma
3.5. Promising Drugs Targeting DPP9-Associated Genes
4. Discussion and Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Jones, J.D.G.; Vance, R.E.; Dangl, J.L. Intracellular Innate Immune Surveillance Devices in Plants and Animals. Science 2016, 354, aaf6395. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Martinon, F.; Burns, K.; Tschopp, J. The Inflammasome: A Molecular Platform Triggering Activation of Inflammatory Caspases and Processing of ProIL-Beta. Mol. Cell 2002, 10, 417–426. [Google Scholar] [CrossRef]
- Robinson, K.S.; Teo, D.E.T.; Tan, K.S.; Toh, G.A.; Ong, H.H.; Lim, C.K.; Lay, K.; Au, B.V.; Lew, T.S.; Chu, J.J.H.; et al. Enteroviral 3C Protease Activates the Human NLRP1 Inflammasome in Airway Epithelia. Science 2020, 370, eaay2002. [Google Scholar] [CrossRef] [PubMed]
- Bauernfried, S.; Scherr, M.J.; Pichlmair, A.; Duderstadt, K.E.; Hornung, V. Human NLRP1 Is a Sensor for Double-Stranded RNA. Science 2021, 371, eabd0811. [Google Scholar] [CrossRef]
- Waumans, Y.; Baerts, L.; Kehoe, K.; Lambeir, A.M.; de Meester, I. The Dipeptidyl Peptidase Family, Prolyl Oligopeptidase, and Prolyl Carboxypeptidase in the Immune System and Inflammatory Disease, Including Atherosclerosis. Front. Immunol. 2015, 6, 387. [Google Scholar] [CrossRef] [Green Version]
- Zhang, H.; Chen, Y.; Keane, F.M.; Gorrell, M.D. Advances in Understanding the Expression and Function of Dipeptidyl Peptidase 8 and 9. Mol. Cancer Res. 2013, 11, 1487–1496. [Google Scholar] [CrossRef] [Green Version]
- Raj, V.S.; Mou, H.; Smits, S.L.; Dekkers, D.H.W.; Müller, M.A.; Dijkman, R.; Muth, D.; Demmers, J.A.A.; Zaki, A.; Fouchier, R.A.M.; et al. Dipeptidyl Peptidase 4 Is a Functional Receptor for the Emerging Human Coronavirus-EMC. Nature 2013, 495, 251–254. [Google Scholar] [CrossRef] [Green Version]
- Andrieu, T.; Thibault, V.; Malet, I.; Laporte, J.; Bauvois, B.; Agut, H.; Cahour, A. Similar Increased Serum Dipeptidyl Peptidase IV Activity in Chronic Hepatitis C and Other Viral Infections. J. Clin. Virol. 2003, 27, 59–68. [Google Scholar] [CrossRef]
- Inn, K.S.; Kim, Y.; Aigerim, A.; Park, U.; Hwang, E.S.; Choi, M.S.; Kim, Y.S.; Cho, N.H. Reduction of Soluble Dipeptidyl Peptidase 4 Levels in Plasma of Patients Infected with Middle East Respiratory Syndrome Coronavirus. Virology 2018, 518, 324–327. [Google Scholar] [CrossRef]
- Huang, M.; Zhang, X.; Toh, G.A.; Gong, Q.; Wang, J.; Han, Z.; Wu, B.; Zhong, F.; Chai, J. Structural and Biochemical Mechanisms of NLRP1 Inhibition by DPP9. Nature 2021, 592, 773–777. [Google Scholar] [CrossRef]
- Hollingsworth, L.R.; Sharif, H.; Griswold, A.R.; Fontana, P.; Mintseris, J.; Dagbay, K.B.; Paulo, J.A.; Gygi, S.P.; Bachovchin, D.A.; Wu, H. DPP9 Sequesters the C Terminus of NLRP1 to Repress Inflammasome Activation. Nature 2021, 592, 778–783. [Google Scholar] [CrossRef]
- Park, J.; Knott, H.M.; Nadvi, N.A.; Collyer, C.A.; Wang, X.M.; Church, B.W.; Gorrell, M.D. Reversible Inactivation of Human Dipeptidyl Peptidases 8 and 9 by Oxidation. Open Enzym. Inhib. J. 2008, 1, 52–60. [Google Scholar] [CrossRef]
- Pairo-Castineira, E.; Clohisey, S.; Klaric, L.; Bretherick, A.D.; Rawlik, K.; Pasko, D.; Walker, S.; Parkinson, N.; Fourman, M.H.; Russell, C.D.; et al. Genetic Mechanisms of Critical Illness in COVID-19. Nature 2020, 591, 92–98. [Google Scholar] [CrossRef]
- Genomewide Association Study of Severe Covid-19 with Respiratory Failure. N. Engl. J. Med. 2020, 383, 1522–1534. [CrossRef]
- Wang, L.; Balmat, T.J.; Antonia, A.L.; Constantine, F.J.; Henao, R.; Burke, T.W.; Ingham, A.; McClain, M.T.; Tsalik, E.L.; Ko, E.R.; et al. An Atlas Connecting Shared Genetic Architecture of Human Diseases and Molecular Phenotypes Provides Insight into COVID-19 Susceptibility. Genome Med. 2020, 13, 83. [Google Scholar] [CrossRef]
- Kappel, B.A.; de Angelis, L.; Heiser, M.; Ballanti, M.; Stoehr, R.; Goettsch, C.; Mavilio, M.; Artati, A.; Paoluzi, O.A.; Adamski, J.; et al. Cross-Omics Analysis Revealed Gut Microbiome-Related Metabolic Pathways Underlying Atherosclerosis Development after Antibiotics Treatment. Mol. Metab. 2020, 36, 100976. [Google Scholar] [CrossRef]
- Puig, J.; Biarnes, C.; Pedraza, S.; Vilanova, J.C.; Pamplona, R.; Fernández-Real, J.M.; Brugada, R.; Ramos, R.; Coll-de-Tuero, G.; Calvo-Perxas, L.; et al. The Aging Imageomics Study: Rationale, Design and Baseline Characteristics of the Study Population. Mech. Ageing Dev. 2020, 189, 111257. [Google Scholar] [CrossRef]
- Mayneris-Perxachs, J.; Moreno-Navarrete, J.M.; Ballanti, M.; Monteleone, G.; Alessandro Paoluzi, O.; Mingrone, G.; Lefebvre, P.; Staels, B.; Federici, M.; Puig, J.; et al. Lipidomics and Metabolomics Signatures of SARS-CoV-2 Mediators/Receptors in Peripheral Leukocytes, Jejunum and Colon. Comput. Struct. Biotechnol. J. 2021, 19, 6080–6089. [Google Scholar] [CrossRef]
- Dobin, A.; Davis, C.A.; Schlesinger, F.; Drenkow, J.; Zaleski, C.; Jha, S.; Batut, P.; Chaisson, M.; Gingeras, T.R. STAR: Ultrafast Universal RNA-Seq Aligner. Bioinformatics 2013, 29, 15–21. [Google Scholar] [CrossRef]
- Li, B.; Dewey, C.N. RSEM: Accurate Transcript Quantification from RNA-Seq Data with or without a Reference Genome. BMC Bioinform. 2011, 12, 323. [Google Scholar] [CrossRef]
- Wikoff, W.R.; Pendyala, G.; Siuzdak, G.; Fox, H.S. Metabolomic Analysis of the Cerebrospinal Fluid Reveals Changes in Phospholipase Expression in the CNS of SIV-Infected Macaques. J. Clin. Investig. 2008, 118, 2661–2669. [Google Scholar] [CrossRef] [Green Version]
- Mayneris-Perxachs, J.; Castells-Nobau, A.; Arnoriaga-Rodríguez, M.; Martin, M.; de la Vega-Correa, L.; Zapata, C.; Burokas, A.; Blasco, G.; Coll, C.; Escrichs, A.; et al. Microbiota Alterations in Proline Metabolism Impact Depression. Cell Metab. 2022, 34, 681–701.e10. [Google Scholar] [CrossRef]
- Love, M.I.; Huber, W.; Anders, S. Moderated Estimation of Fold Change and Dispersion for RNA-Seq Data with DESeq2. Genome Biol. 2014, 15, 550. [Google Scholar] [CrossRef] [Green Version]
- Zhou, Y.; Zhou, B.; Pache, L.; Chang, M.; Khodabakhshi, A.H.; Tanaseichuk, O.; Benner, C.; Chanda, S.K. Metascape Provides a Biologist-Oriented Resource for the Analysis of Systems-Level Datasets. Nat. Commun. 2019, 10, 1523. [Google Scholar] [CrossRef]
- Shannon, P.; Markiel, A.; Ozier, O.; Baliga, N.S.; Wang, J.T.; Ramage, D.; Amin, N.; Schwikowski, B.; Ideker, T. Cytoscape: A Software Environment for Integrated Models of Biomolecular Interaction Networks. Genome Res. 2003, 13, 2498–2504. [Google Scholar] [CrossRef]
- Bader, G.D.; Hogue, C.W.V. An Automated Method for Finding Molecular Complexes in Large Protein Interaction Networks. BMC Bioinform. 2003, 4, 2–27. [Google Scholar] [CrossRef] [Green Version]
- Kursa, M.B.; Rudnicki, W.R. Feature Selection with the Boruta Package. J. Stat. Softw. 2010, 36, 1–13. [Google Scholar] [CrossRef] [Green Version]
- Lundberg, S.M.; Erion, G.; Chen, H.; DeGrave, A.; Prutkin, J.M.; Nair, B.; Katz, R.; Himmelfarb, J.; Bansal, N.; Lee, S.-I. From Local Explanations to Global Understanding with Explainable AI for Trees. Nat. Mach. Intell. 2020, 2, 56–67. [Google Scholar] [CrossRef]
- Liao, Y.; Wang, J.; Jaehnig, E.J.; Shi, Z.; Zhang, B. WebGestalt 2019: Gene Set Analysis Toolkit with Revamped UIs and APIs. Nucleic. Acids Res. 2019, 47, W199–W205. [Google Scholar] [CrossRef] [Green Version]
- Araki, H.; Knapp, C.; Tsai, P.; Print, C. GeneSetDB: A Comprehensive Meta-Database, Statistical and Visualisation Framework for Gene Set Analysis. FEBS Open Bio 2012, 2, 76–82. [Google Scholar] [CrossRef]
- Kand’ár, R.; Žáková, P.; Mužáková, V. Monitoring of Antioxidant Properties of Uric Acid in Humans for a Consideration Measuring of Levels of Allantoin in Plasma by Liquid Chromatography. Clin. Chim. Acta 2006, 365, 249–256. [Google Scholar] [CrossRef]
- Ames, B.N.; Cathcart, R.; Schwiers, E.; Hochstein, P. Uric Acid Provides an Antioxidant Defense in Humans against Oxidant- and Radical-Caused Aging and Cancer: A Hypothesis. Proc. Natl. Acad. Sci. USA 1981, 78, 6858–6862. [Google Scholar] [CrossRef] [Green Version]
- Nieto, F.J.; Iribarren, C.; Gross, M.D.; Comstock, G.W.; Cutler, R.G. Uric Acid and Serum Antioxidant Capacity: A Reaction to Atherosclerosis? Atherosclerosis 2000, 148, 131–139. [Google Scholar] [CrossRef]
- Jansen, T.; Daiber, A. Direct Antioxidant Properties of Bilirubin and Biliverdin. Is There a Role for Biliverdin Reductase? Front. Pharmacol. 2012, 3, 30. [Google Scholar] [CrossRef] [Green Version]
- Miller, S.; Krijnse-Locker, J. Modification of Intracellular Membrane Structures for Virus Replication. Nat. Rev. Microbiol. 2008, 6, 363–374. [Google Scholar] [CrossRef]
- Schmidt, N.; Lareau, C.A.; Keshishian, H.; Ganskih, S.; Schneider, C.; Hennig, T.; Melanson, R.; Werner, S.; Wei, Y.; Zimmer, M.; et al. The SARS-CoV-2 RNA–Protein Interactome in Infected Human Cells. Nat. Microbiol. 2020, 6, 339–353. [Google Scholar] [CrossRef]
- Balistreri, G.; Horvath, P.; Schweingruber, C.; Zünd, D.; McInerney, G.; Merits, A.; Mühlemann, O.; Azzalin, C.; Helenius, A. The Host Nonsense-Mediated MRNA Decay Pathway Restricts Mammalian RNA Virus Replication. Cell Host Microbe 2014, 16, 403–411. [Google Scholar] [CrossRef] [Green Version]
- Wada, M.; Lokugamage, K.G.; Nakagawa, K.; Narayanan, K.; Makino, S. Interplay between Coronavirus, a Cytoplasmic RNA Virus, and Nonsense-Mediated MRNA Decay Pathway. Proc. Natl. Acad. Sci. USA 2018, 115, E10157–E10166. [Google Scholar] [CrossRef] [Green Version]
- Mazumder, B.; Poddar, D.; Basu, A.; Kour, R.; Verbovetskaya, V.; Barik, S. Extraribosomal L13a Is a Specific Innate Immune Factor for Antiviral Defense. J. Virol. 2014, 88, 9100–9110. [Google Scholar] [CrossRef] [Green Version]
- Thoms, M.; Buschauer, R.; Ameismeier, M.; Koepke, L.; Denk, T.; Hirschenberger, M.; Kratzat, H.; Hayn, M.; MacKens-Kiani, T.; Cheng, J.; et al. Structural Basis for Translational Shutdown and Immune Evasion by the Nsp1 Protein of SARS-CoV-2. Science 2020, 369, 1249–1256. [Google Scholar] [CrossRef]
- Ariav, Y.; Chng, J.H.; Christofk, H.R.; Ron-Harel, N.; Erez, A. Targeting Nucleotide Metabolism as the Nexus of Viral Infections, Cancer, and the Immune Response. Sci. Adv. 2021, 7, eabg6165. [Google Scholar] [CrossRef]
- Kojima, S.; Sakamoto, T.; Ishihara, M.; Kimura, K.; Miyazaki, S.; Yamagishi, M.; Tei, C.; Hiraoka, H.; Sonoda, M.; Tsuchihashi, K.; et al. Prognostic Usefulness of Serum Uric Acid after Acute Myocardial Infarction (the Japanese Acute Coronary Syndrome Study). Am. J. Cardiol. 2005, 96, 489–495. [Google Scholar] [CrossRef]
- Kim, S.Y.; Park, S.C. Physiological Antioxidative Network of the Bilirubin System in Aging and Age-Related Diseases. Front. Pharmacol. 2012, 3, 45. [Google Scholar] [CrossRef] [Green Version]
- Marrocco, I.; Altieri, F.; Peluso, I. Measurement and Clinical Significance of Biomarkers of Oxidative Stress in Humans. Oxid. Med. Cell. Longev. 2017, 2017, 6501046. [Google Scholar] [CrossRef]
- Lee, J.M.; Johnson, J.A. An Important Role of Nrf2-ARE Pathway in the Cellular Defense Mechanism. J. Biochem. Mol. Biol. 2004, 37, 139–143. [Google Scholar] [CrossRef] [Green Version]
- Higgins, P.; Dawson, J.; Lees, K.R.; Mcarthur, K.; Quinn, T.J.; Walters, M.R. Xanthine Oxidase Inhibition for the Treatment of Cardiovascular Disease: A Systematic Review and Meta-Analysis. Cardiovasc. Ther. 2012, 30, 217–226. [Google Scholar] [CrossRef]
- Drummond, G.R.; Selemidis, S.; Griendling, K.K.; Sobey, C.G. Combating Oxidative Stress in Vascular Disease: NADPH Oxidases as Therapeutic Targets. Nat. Rev. Drug Discov. 2011, 10, 453–471. [Google Scholar] [CrossRef] [Green Version]
- Cappola, T.P.; Kass, D.A.; Nelson, G.S.; Berger, R.D.; Rosas, G.O.; Kobeissi, Z.A.; Marbán, E.; Hare, J.M. Allopurinol Improves Myocardial Efficiency in Patients with Idiopathic Dilated Cardiomyopathy. Circulation 2001, 104, 2407–2411. [Google Scholar] [CrossRef] [Green Version]
- Goodwin, C.M.; Xu, S.; Munger, J. Stealing the Keys to the Kitchen: Viral Manipulation of the Host Cell Metabolic Network. Trends Microbiol. 2015, 23, 789–798. [Google Scholar] [CrossRef]
- Lee, H.N.; Surh, Y.J. Resolvin D1-Mediated NOX2 Inactivation Rescues Macrophages Undertaking Efferocytosis from Oxidative Stress-Induced Apoptosis. Biochem. Pharmacol. 2013, 86, 759–769. [Google Scholar] [CrossRef]
- Saito, P.; Melo, C.P.B.; Martinez, R.M.; Fattori, V.; Cezar, T.L.C.; Pinto, I.C.; Bussmann, A.J.C.; Vignoli, J.A.; Georgetti, S.R.; Baracat, M.M.; et al. The Lipid Mediator Resolvin D1 Reduces the Skin Inflammation and Oxidative Stress Induced by UV Irradiation in Hairless Mice. Front. Pharmacol. 2018, 9, 1242. [Google Scholar] [CrossRef] [Green Version]
- Kasuga, K.; Yang, R.; Porter, T.F.; Agrawal, N.; Petasis, N.A.; Irimia, D.; Toner, M.; Serhan, C.N. Rapid Appearance of Resolvin Precursors in Inflammatory Exudates: Novel Mechanisms in Resolution. J. Immunol. 2008, 181, 8677–8687. [Google Scholar] [CrossRef] [Green Version]
- Lee, D.H.; Blomhoff, R.; Jacobs, D.R. Is Serum Gamma Glutamyltransferase a Marker of Oxidative Stress? Free Radic. Res. 2004, 38, 535–539. [Google Scholar] [CrossRef]
- Gordon, D.E.; Jang, G.M.; Bouhaddou, M.; Xu, J.; Obernier, K.; White, K.M.; O’Meara, M.J.; Rezelj, V.V.; Guo, J.Z.; Swaney, D.L.; et al. A SARS-CoV-2 Protein Interaction Map Reveals Targets for Drug Repurposing. Nature 2020, 583, 459–468. [Google Scholar] [CrossRef]
- Sehailia, M.; Chemat, S. Antimalarial-Agent Artemisinin and Derivatives Portray More Potent Binding to Lys353 and Lys31-Binding Hotspots of SARS-CoV-2 Spike Protein than Hydroxychloroquine: Potential Repurposing of Artenimol for COVID-19. J. Biomol. Struct. Dyn. 2020, 39, 6184–6194. [Google Scholar] [CrossRef]



Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
del Castillo-Izquierdo, Á.; Moreno-Navarrete, J.M.; Latorre, J.; Arnoriaga-Rodríguez, M.; Ballanti, M.; Monteleone, G.; Alessandro Paoluzi, O.; Mingrone, G.; Puig, J.; Ramos, R.; et al. DPP9 as a Potential Novel Mediator in Gastrointestinal Virus Infection. Antioxidants 2022, 11, 2177. https://doi.org/10.3390/antiox11112177
del Castillo-Izquierdo Á, Moreno-Navarrete JM, Latorre J, Arnoriaga-Rodríguez M, Ballanti M, Monteleone G, Alessandro Paoluzi O, Mingrone G, Puig J, Ramos R, et al. DPP9 as a Potential Novel Mediator in Gastrointestinal Virus Infection. Antioxidants. 2022; 11(11):2177. https://doi.org/10.3390/antiox11112177
Chicago/Turabian Styledel Castillo-Izquierdo, Ángela, José María Moreno-Navarrete, Jessica Latorre, María Arnoriaga-Rodríguez, Marta Ballanti, Giovanni Monteleone, Omero Alessandro Paoluzi, Geltrude Mingrone, Josep Puig, Rafael Ramos, and et al. 2022. "DPP9 as a Potential Novel Mediator in Gastrointestinal Virus Infection" Antioxidants 11, no. 11: 2177. https://doi.org/10.3390/antiox11112177




