1. Introduction
The discovery of Asgard archaea—first isolated from deep-sea sediments near hydrothermal vents at Loki’s Castle by Spang and colleagues in 2015 [
1]—provided compelling support for a long-standing hypothesis in evolutionary biology: that eukaryotes emerged from within the archaeal domain [
2]. Prior to this discovery, multiple lines of evidence pointed to deep homologies between Archaea and Eukaryota, including shared features such as histone-like proteins, complex RNA polymerases, and cytoskeletal protein homologues [
3,
4,
5]. The identification of the Asgard superphylum, particularly members such as Lokiarchaea, did not originate this hypothesis but rather strengthened it by revealing extant organisms with a richer complement of eukaryotic signature proteins (ESPs) and cellular complexity than previously seen in Archaea [
4,
6]. Although extant Asgardians are not direct ancestors of eukaryotes, they are likely descendants of a lineage closely related to the Last Universal Common Ancestor (LUCA), which existed over 4 billion years ago [
7]. Thus, Asgardians may serve as plausible modern analogs that illuminate the evolutionary steps bridging prokaryotic simplicity and eukaryotic complexity. These findings have contributed to the increasing acceptance of a two-domain model of life, in which Eukarya are nested within Archaea [
3,
4].
The implications of this discovery extend far beyond the redrawing of evolutionary trees. They raise the possibility that descendants of even more ancient transitional forms—organisms that bridged the earliest stages of life—may persist today in the Earth’s extreme environments. Considering this, we propose the POLET hypothesis (Pre-prokaryotic Organismal Lifeforms Existing Today), which suggests that direct descendants of pre-prokaryotic, membrane-bound entities resembling protocells could have survived through geological time, hidden in deep anoxic sediments and mud. We refer to these hypothesized organisms as “POLETicians”. Unlike the “Shadow Life” hypothesis proposed by Paul Davies and colleagues, which suggests that life may have arisen multiple times independently on Earth [
8,
9], the POLET hypothesis asserts a single origin of life, with POLETicians representing a living remnant of lineages that preceded the Last Universal Common Ancestor (LUCA), effectively serving as a missing link between abiogenesis and LUCA.
Building on other theories for the origin of life discussed in this review, we envision POLETicians as ancient, slowly metabolizing membranous vesicles filled with primitive biochemical machinery. Instead of fast-acting enzymes, they may employ RNA–amino acid aptamers—relics of a primordial molecular system that predated the ribosome and even enzymes [
10,
11]. Lacking sophisticated metabolic networks, these entities can still grow and reproduce over extended periods using blebbing-like mechanisms for division [
12]. Their distinctive biochemical composition—potentially incorporating both L- and D-amino acids as well as L- and D-RNAs due to their pre-enzymatic nature—along with extraordinarily long lifespans that could last years, decades, or even millennia in their dormant states, sets them apart from all known forms of life. Energy may be directly derived from non-traditional sources, such as radioactive decay or mineral-catalyzed redox gradients [
13].
A widely accepted general definition of life, for example, NASA’s working definition, written by Steven Benner in 2010, contends that “
Life is a self-sustaining chemical system capable of Darwinian evolution” [
14]. This definition is broad enough to encompass both extant life and hypothetical entities like the POLETicians. Our goal is not to challenge this foundational definition but rather to highlight features that may expand our working understanding of how life-like systems can emerge and persist before reaching the threshold of prokaryotes or archaea. Specifically, POLETicians may not possess stable genomes or high-fidelity replication machinery, but they may exhibit the following features:
Energetic self-maintenance via simple enzymatic functions,
Compartmentalization and molecular turnover (e.g., blebbing), and
Adaptation through stochastic compositional variation and selection.
In ecological terms, POLETicians may occupy a unique niche. Unlike all known life, which is largely restricted to metabolizing homochiral biomolecules, these hypothetical racemic organisms could utilize geologically generated racemic mixtures of amino acids and sugars that accumulate in deep sediments [
15]. This would provide them with exclusive access to a reservoir of otherwise inaccessible prebiotic carbon and nitrogen sources. Such a capability would represent an alternative evolutionary solution to the chemical constraints of early life, one that bypasses the need for homochirality.
In modern biology, homochirality is a defining feature that is essential for the consistent function of biopolymers, such as RNA and proteins. However, the origin of this asymmetry remains unclear. Recent work has demonstrated that magnetic surfaces like magnetite (Fe
3O
4) can act as chiral agents via the chiral-induced spin selectivity (CISS) effect, enabling spin-selective crystallization of racemic precursors, such as ribo-aminooxazoline (RAO)—an RNA building block. In shallow lake environments rich in sedimentary magnetite, this process can produce a high enantiomeric excess and ultimately homochirality, offering a plausible route for chiral selection on the early Earth [
15].
However, some modern organisms suggest that racemic metabolism is chemically feasible. Several bacteria—including
Bacillus subtilis,
Vibrio cholerae, and the extremophile
Thermus thermophilus—can synthesize and utilize D-amino acids using amino acid racemases [
16]. These enzymes interconvert D- and L-enantiomers, aiding cell wall remodeling, biofilm regulation, and stress adaptation. While Archaea generally lack such metabolic flexibility, the presence of these traits in extremophilic bacteria supports the plausibility of racemic metabolism in extreme environments. This biochemical precedent adds weight to the possibility that POLETicians may directly exploit racemic substrates in habitats such as deep-sea sediments or hydrothermal vent systems [
16].
Moreover, POLETicians may engage in syntrophic relationships with Asgard archaea or other extremophiles, possibly providing reduced carbon compounds, cofactors, or redox buffering in exchange for structural protection or additional resources. We do not propose that POLETicians will have all the aspects that we describe in this paper; rather, we present possibilities based on established Origin of Life (OOL) hypotheses.
The discovery of Asgardians also has ramifications for astrobiology. Panspermia (the origins of life on Earth were hitchhikers on asteroids) has been largely debunked because most present-day “primitive” life cannot survive the intense heat and pressure experienced when entering the Earth’s atmosphere [
17,
18]. However, Asgardians and POLETicians, which may possess spore-like durability and extreme metabolic flexibility, have not been rigorously tested under such conditions.
We propose that POLETicians can be identified using advanced imaging and analytical techniques, such as chiral-sensitive mass spectrometry, fluorescence microscopy using membrane-selective dyes, and proteomics, which can detect non-standard amino acids and aptamer-based translation. Both mirror L- and standard D-RNAs can be detected using existing RNA polymerases engineered to recognize non-standard bases [
19] or nanopores that can directly sequence both standard and modified RNAs [
20]. A mirror version of T7 RNA polymerase, a workhorse for molecular biology, has been constructed, which allows the direct synthesis of mirror RNAs and polymerase chain reaction of mirror RNAs [
21,
22]. In addition, the use of mirror D-amino acid versions of thermostable polymerases has enabled PCR of mirror L-RNAs [
23]. Generation of a mirror D-amino acid of a reverse transcriptase would allow the synthesis of mirror L-cDNAs from environmental RNA (eRNA) from extreme environments, which could be used in massively parallel next-generation cDNA sequencing experiments to identify minuscule amounts of environmental mirror L-RNAs, should they exist. This is analogous to the discovery of Asgaridians by sequencing large amounts of environmental DNA (eDNA) from the mud near Loki’s Castle [
2].
If discovered, POLETicians with mirror RNAs and mirror proteins, or unknown types of linear polymers (ULPs), would redefine our assumptions about the boundaries between prebiotic chemistry and life, positioning POLETicians as living fossils from the pre-prokaryotic world. In this review, we examine the evidence for this hypothesis, discuss potential detection strategies, and explore the implications of the POLET hypothesis for our understanding of the origins and continuity of life.
2. Pre-Prokaryotic Organismal Lifeforms Existing Today (POLET) Hypothesis
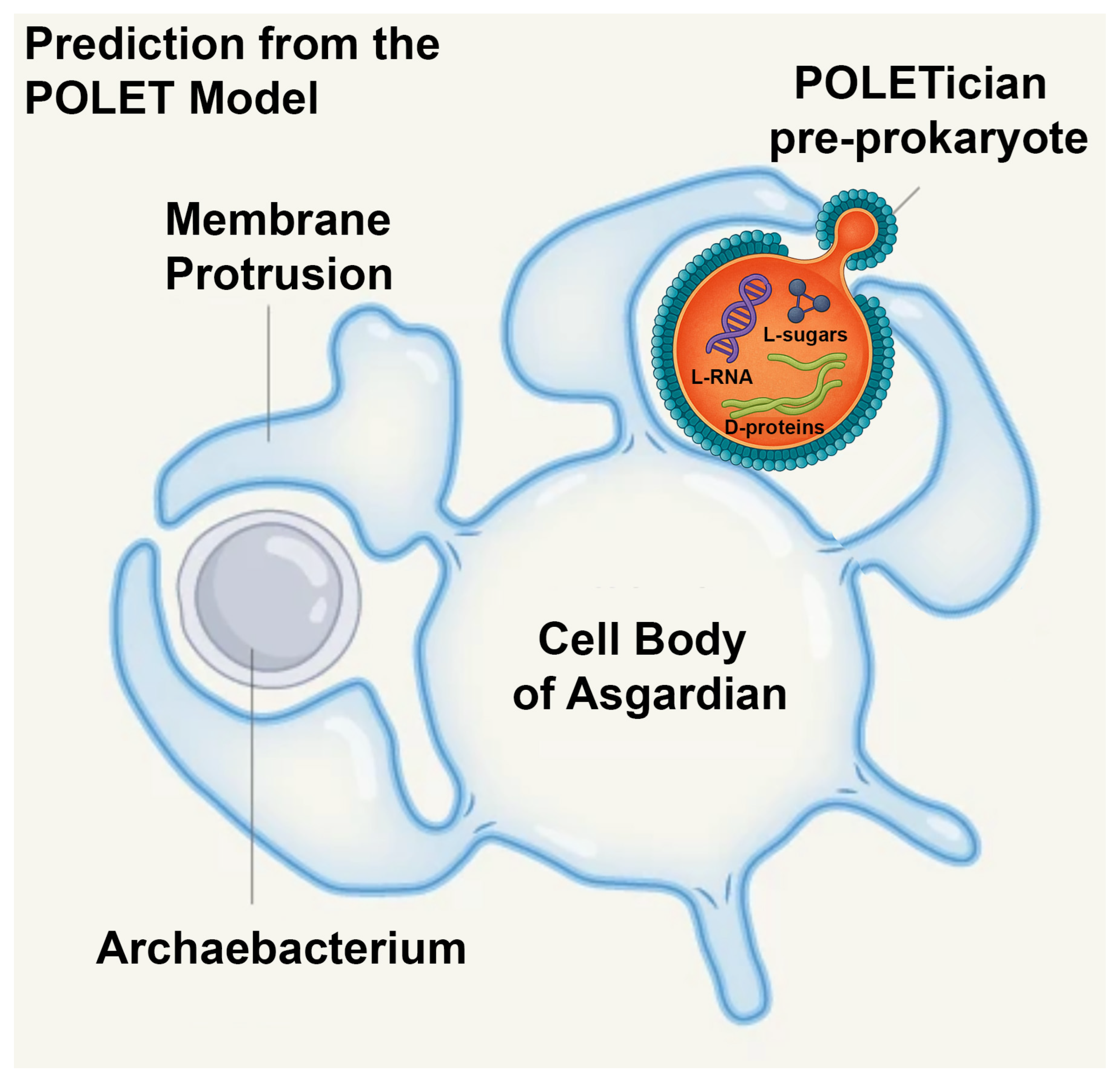
This section explores the conceptual and hypothetical basis of the POLET hypothesis, which proposes the continued existence of ancient, pre-prokaryotic organisms known as POLETicians (
Figure 1). These theoretical life forms may have survived in deep, anoxic environments and exhibited a range of unique traits that distinguish them from known cellular life. We categorize these hypothetical features in the following subsections.
2.1. Longevity and Metabolic Quiescence
Lifespan Characteristics
POLETicians, if they exist today, may possess remarkably long cell cycles, potentially lasting weeks or even months, and lifespans that could last years, decades, or even millennia in dormant states. Their long cell cycles and lifespans are hypothesized to be a consequence of minimal metabolic activity and extremely slow rates of molecular turnover, as seen in some extremophiles [
24]. Unlike modern organisms, which rely on rapid biochemical cycles, POLETicians may persist in a near-static or geochemically persistent state, allowing for chemical longevity under extreme conditions without requiring evolved repair mechanisms.
We propose that their metabolism is likely extremely slow because they lack the highly evolved and efficient enzymes characteristic of modern life. Instead of relying on enzyme-driven catalytic acceleration, these hypothetical organisms may depend on spontaneous chemical reactions and slow, geochemical processes. Membrane formation and other structural assemblies can occur passively, with precursors generated over long timescales through radioactive decay, redox gradients, or mineral surface catalysis [
12,
25,
26]. Although such processes operate at glacial rates, they could still support a viable metabolic cycle when extended over centuries or millennia, particularly in stable and extreme environments.
Analogous metabolic resilience can be observed in modern extremophiles—organisms that persist in high-radiation deserts, deep-sea hydrothermal vents, or subzero hypersaline lakes—where biological activity is often paused or drastically slowed, awaiting transient favorable conditions [
24]. For example,
Halicephalobus mephisto, the ‘devil worm,’ a nematode found deep underground, exhibits extremely low metabolic rates and long survival times under high pressure and limited nutrient availability [
24]. Remarkably, radiocarbon dating of the groundwater where they live suggests that these worms can persist for 3000–12,000 years [
24]. Similarly,
Deinococcus radiodurans and Antarctic cryophiles maintain viability despite intense radiation and prolonged dormancy [
27]. These extremophiles demonstrate that slow, durable metabolic strategies are not only plausible but already exist in life today, lending credibility to the idea that a primitive, enzyme-poor POLETician lineage could persist in extreme niches where competition from modern organisms is minimal.
The long dormancy of hypothetical POLETicains could be similar to that of water fleas in the genus Daphnia. When environmental conditions signal unfavorable (unstable) environmental conditions in the future, Daphnia metabolism is reduced to near zero, and they enter a prolonged diapause (up to 700 years) [
28]. In several studies, researchers have sampled populations from soil cores that were formed by digging from the bottom of lakes, determined their age with isotopic analyses (some can be hundreds of years old), and “resurrect” and expose “ancient” genetic variants to present-day environmental conditions to see how old genotypes fare in present-day environments [
29]. Analogously, laboratory revival of ancient POLET specimens—if obtained—could allow direct experimental validation of prebiotic chemical systems operating under modern geochemical conditions.
The hypothesis that POLETicians may possess extraordinarily long cell cycles and lifespans is partly inspired by the limited but intriguing data on the growth dynamics of Asgard archaea. These organisms, named after Norse mythology due to their initial discovery near Loki’s Castle hydrothermal vent [
1], exhibit exceptionally slow growth rates. In a landmark 2020 study, Imachi et al. successfully cultured the Lokiarchaeon strain MK-D1 from methane-seep sediments using a methane-fed continuous-flow bioreactor operated for over 2000 days [
4]. Following inoculation with casamino acids and antibiotics at 20 °C, faint cell turbidity was observed only after approximately one year. Strain MK-D1 displayed an extended lag phase of 30–60 days and required more than three months to reach cell densities of ~10
5 16S rRNA gene copies/mL, with a doubling time of 14–25 days [
4]. After six transfers, MK-D1 reached 13% abundance within a tri-culture that also included
Halodesulfovibrio (85%) and
Methanogenium (2%). Fluorescence in situ hybridization and scanning electron microscopy have revealed close physical associations among these three organisms [
4]. Subsequent enrichment steps eventually eliminated
Halodesulfovibrio, resulting in a stable co-culture of MK-D1 and
Methanogenium, which was achieved after 12 years of continuous cultivation [
4]. These findings suggest that Asgard archaea may grow and divide at extremely slow rates under natural conditions. By analogy, we hypothesize that POLETicians—putative pre-prokaryotic life forms—may exhibit even longer lifespans and slower metabolic activity, consistent with life in stable, low-energy environments, such as the deep biosphere or those found in acidic environments in hot springs and geysers [
30].
2.2. Primitive Biochemistry and Genetic Mechanisms
2.2.1. Pre-Ribosomal RNA–Amino Acid Interactions
In the absence of modern enzymes, POLETicians may have relied on spontaneous chemical catalysis and structural RNA molecules to initiate basic biochemical functions. Ribozymes, which are capable of limited catalytic activity, and RNA aptamers that bind amino acids could have supported primitive translational systems without ribosomes [
11]. These mechanisms imply a plausible pre-ribosomal stage in which RNA–amino acid interactions form the biochemical foundation of early life.
One prominent hypothesis addressing the emergence of the genetic code is the stereochemical model. Yarus and colleagues [
11], among others, have experimentally demonstrated that certain RNA sequences exhibit selective binding to their cognate amino acids. These findings suggest that physicochemical affinities may have shaped early codon assignments. However, while these associations appear robust, they do not yet offer a complete mechanistic explanation for the origin of the genetic code and are more likely to be relevant to its elaboration or stabilization (
Table 1) [
10,
31].
The discovery or engineering of POLET-like systems could provide experimental platforms to test these theories, offering empirical access to plausible prebiotic RNA–peptide interactions that operated before the emergence of ribosomal translation.
2.2.2. Peptide Self-Assembly and Structural Functions
In addition to RNA-based interactions, POLETicians may utilize non-enzymatically synthesized peptides for rudimentary structural roles. However, in the absence of ribosomal control or enzymatic specificity, such peptides would likely contain a mixture of L- and D-amino acids, which significantly reduces the likelihood of stable and ordered folding [
32]. Most random peptide mixtures, especially those that are racemic or compositionally heterogeneous, tend to form amorphous hydrated aggregates rather than defined supramolecular structures [
32].
Nevertheless, experimental studies have shown that under certain conditions—such as localized enrichment of homochiral amino acids or repetitive sequences—peptides can self-assemble into ordered structures like beta-sheets, fibrils, or nanotubes [
33,
34]. While these scenarios are unlikely to have been widespread in early Earth conditions, they may have occurred in specific microenvironments, such as mineral surfaces, drying pools, or lipid interfaces, where prebiotic selection and concentration effects could have favored partial sequence regularity or chirality.
In
Table 1, we summarize the known amino acid homopolymers whose codons or anticodons correspond to their RNA aptamer-binding sequences. Several of these homopolymers, when synthesized under laboratory conditions with controlled chirality, form regular nanostructures resembling cytoskeletal filaments. Although these findings do not imply that such structures spontaneously emerge from fully random mixtures, they raise the possibility that selective pressures in early POLET-like systems could have stabilized rare peptide assemblies with structural or compartmentalizing functions. These self-organizing properties, even if infrequent, could have provided scaffolding or phase-separated domains in protocell evolution—particularly before the emergence of lipid membranes or protein-based cytoskeletons.
Table 1.
Probabilities that cognate coding triplets are unconcentrated in amino acid binding sites and fiber-forming potentials of poly-amino acid homopolymers.
Table 1.
Probabilities that cognate coding triplets are unconcentrated in amino acid binding sites and fiber-forming potentials of poly-amino acid homopolymers.
| Amino Acid | Codon |
*PCodon | Anticodon | PAnticodon | Fiber Forming Potential | Notes | Refs. |
|---|
| L-Arg | CGG | 4.0 × 10−3 | CCG | 1 | Weakly with PEG-Q11 | Electrostatic repulsion may prevent stable fiber formation alone | [35] |
| L-Arg | CGA | 1 | UCG | 3.4 × 10−5 | weak | See above. | [35] |
| L-Gln | CAA | 0.16 | UUG | 1 | Amyloid fibers | PolyQ diseases | [36] |
| L-His | CAC | 0.40 | GUG | 6.4 × 10−8 | Has cell penetrating properties | Aggregate under certain pH conditions | [37] |
| L-Ile | AUU | 4.8 × 10−109 | AAU | 1 | β-sheets and amyloids | Contributes to amyloid-like structures | [38] |
| L-Leu | CUA | 1 | UAG | 0.07 | β-sheets | hydrophobic | |
| Phe | UUU | 1 | AAA | 0.047 | Nanotubes/fibers | Aromatic stacking | [39] |
| Phe | UUC | 1 | GAA | 2.2 × 10−4 | Nanotubes/fibers | π–π stacking drives aggregation | [39] |
| Trp | UGG | 1 | CCA | 5.5 × 10−13 | Forms nanostructured micelles | π–π stacking drives aggregation | [40] |
| Tyr | UAU | 0.10 | AUA | 2.4 × 10−5 | Fibrils | Forms fibrils via aromatic and hydrogen bonding interactions. | [39] |
| Tyr | UAC | 0.016 | GUA | 8.0 × 10−3 | Fibrils | See above | [39] |
2.2.3. Hypothetical Ribosome-Free Translation-like Processes
Although ribosomes are universal among modern life forms, POLETicians may employ an alternative system for amino acid polymerization that is slower and structurally distinct. Ribozymes or small RNA–protein complexes may have facilitated non-ribosomal peptide formation before the evolution of modern ribosomes. This idea is supported by several lines of experimental and theoretical research suggesting that aminoacylated tRNA-like structures can form without enzymes and that peptide bond formation can occur under prebiotic conditions [
11]. Further research is required to determine whether such systems exist in modern organisms or can be reconstructed in vitro.
In this context, POLETicians may serve as living models of intermediate translation systems—between random polymerization and full ribosomal translation, shedding light on the evolutionary pathway from chemistry to biology.
2.3. Reproductive Strategy
2.3.1. Blebbing over Binary Fission: A Primitive Mode of Reproduction
Rather than dividing through the highly coordinated process of binary fission observed in modern prokaryotes, POLETicians may have reproduced via a far simpler and more stochastic process—membrane blebbing. In this mode of replication, vesicle-like structures gradually accumulate primitive biochemical components such as short nucleic acid fragments, amino acids, and rudimentary catalytic oligomers. Once a critical threshold is reached, spontaneous budding produces irregular daughter vesicles with heterogeneous molecular content [
41].
Blebbing does not require cytoskeletal machinery or cell wall synthesis, making it a plausible precursor of regulated cell division. This aligns with contemporary models of the origin of life, where fatty acid vesicles have been observed to divide under simulated prebiotic conditions, especially in the presence of catalytic mineral surfaces like clays or metal sulfides [
42,
43]. Charras (2008) and others have suggested that blebbing-like behaviors in modern cells might echo ancient division mechanisms [
41].
Crucially, although blebbing does not ensure faithful genetic replication, it may still facilitate compositional inheritance. If certain oligomeric complexes, such as RNA aptamers, ribozymes, or proto-peptides, localize within specific regions of a vesicle, these molecular clusters may co-segregate into daughter blebs. This could result in a rudimentary form of phenotypic memory and selection across generations, even in the absence of template replication.
Such models have precedents in origin-of-life theories, such as Gánti’s chemoton model [
44] and Kauffman’s autocatalytic sets [
45], where systems evolve through the persistence of functional reaction networks rather than precise sequences. Blebs that pass on advantageous molecular constellations can grow more efficiently or resist environmental stress, introducing selectable variations into an otherwise non-genetic system.
Over time, iterative selection acting on these emergent properties may have driven the evolution of increased fidelity and functionality—paving the way for digital genetics. Thus, blebbing, while inefficient, may represent the first evolutionary scaffold on which true biological reproduction is eventually built.
Recently, Cho and colleagues demonstrated that protocells could form through a simple, prebiotically plausible reaction in which the amino acid cysteine spontaneously reacts with short-chain (C8) thioesters to produce diacyl lipids. These lipids self-assemble into stable membrane vesicles even under aqueous conditions with low reactant concentrations. Remarkably, this three-component reaction is compatible with functional ribozymes, supporting its relevance to early-stage biochemistry. Importantly, silica surfaces, which are chemically related to clays previously implicated in prebiotic models, can catalyze vesicle formation through basic electrostatic interactions. These findings suggest that early cell-like compartments may have arisen from minimal and readily available components, with environmental surfaces such as silica playing a key organizational role in facilitating the emergence of membrane-bound protocells [
41].
2.3.2. Environmental Catalysis and Structural Innovation
The plausibility of blebbing-based reproduction is enhanced by this environmental catalysis. Mineral-rich niches—particularly those involving montmorillonite clays and metal sulfide surfaces—likely acted as dual-function platforms that promoted both membrane formation and biochemical polymerization. Montmorillonite clays, which contain microscopic crystals of silicate minerals, have been shown to catalyze RNA polymerization while simultaneously templating fatty acid vesicle formation [
43].
Furthermore, incorporating non-homochiral or synthetic lipids could offer both functional flexibility and chemical resilience. Unlike modern cell membranes composed of chiral phospholipids, POLETician membranes may incorporate racemic fatty acids, monoglycerides, and alcohols, which are products of abiotic processes such as Fischer–Tropsch-type synthesis [
46]. This could enable adaptability to extreme niches, including hypersaline lakes, sub-ice brine pockets, and hydrothermal vent systems with fluctuating redox gradients.
After blebbing, daughter vesicles may remain loosely tethered, forming filamentous chains or vesicle aggregates. These transient groupings could enhance structural stability or serve as primitive colonies, allowing for shared catalytic advantages and collective resilience—early hints of multicellular behavior.
Together, these features reinforce the idea that POLETicians inhabited geochemically active environments where reproduction was driven not by internal genome replication but by compositional persistence, environmental templating, and selective retention of functional assemblies.
2.4. Membrane Composition and Origin
2.4.1. Mineral-Assisted Membrane Assembly
The formation of POLETician membranes may rely heavily on abiotic interactions with mineral surfaces. Catalytic clays (e.g., montmorillonite) and transition metal oxides, commonly found near hydrothermal vents, would have provided both chemical energy and physical scaffolding. These surfaces can align amphiphilic molecules, thereby facilitating membrane formation and catalyzing key polymerization reactions, such as the formation of RNA oligomers.
Rather than using modern phospholipids, POLETician membranes may consist of simple prebiotically plausible amphiphiles, such as fatty acids and monoglycerides. These molecules can spontaneously self-assemble into micelles or vesicles over a broad range of pH, ionic strength, and temperature conditions. Critically, they are robust in acidic, metal-rich, and redox-active environments, which are believed to be common on early Earth—and possibly other planetary bodies [
47].
Moreover, mineral-induced elongation or chemical modification of amphiphiles can contribute to increased membrane stability, selectivity, and functional persistence over time [
45]. Selection may favor vesicles whose membranes retain key catalytic elements or exclude destabilizing components. This feedback between membrane composition and metabolic potential has been suggested to represent an early form of genotype-phenotype coupling—albeit without digital genetic encoding [
45].
2.4.2. Racemic Chirality and Chemical Adaptability
In contrast to modern life, which exhibits strict homochirality—D-sugars in nucleic acids and L-amino acids in proteins—POLETicians may contain racemic mixtures of chiral molecules. This reflects the abiotic, geochemically driven synthesis of their abiotic ancestors, where no selective enzymatic machinery existed to enforce stereospecificity. Racemic mixtures of fatty acids, amino acids, and sugar derivatives would have been the default products of prebiotic chemistry, as supported by numerous simulations and Miller–Urey-type experiments [
48,
49,
50].
While racemization is generally viewed as destabilizing biological polymers and membranes, certain physicochemical contexts suggest that chirally mixed systems may confer unique adaptive advantages. For example, racemic fatty acid vesicles have been shown to form stable membranes under simulated prebiotic conditions and may even exhibit enhanced permeability or environmental responsiveness compared to homochiral analogs [
12]. Furthermore, vesicle formation from simple amphiphiles—such as short-chain monocarboxylic acids and alcohols—can be promoted in racemic mixtures, particularly when aided by mineral surfaces, pH fluctuations, or cycles of hydration and dehydration.
Importantly, these chiral-flexible membranes may have been better suited to the extreme conditions that likely characterized many early Earth niches, including hydrothermal vents, subglacial brine channels, and acidic geothermal ponds. Racemic flexibility could buffer POLETician membranes against pH shifts, salinity extremes, and fluctuating temperatures—traits now found in modern extremophiles in several genera, such as
Thermoplasma,
Halorubrum, and
Methanopyrus, whose membrane chemistries have evolved to tolerate such hostile environments [
51].
Moreover, while the inclusion of sugars in primitive membranes remains speculative, it is plausible that early sugar-like amphiphiles—formed via abiotic pathways, such as formose reactions or borate-stabilized intermediates—could have contributed to membrane functionality. Although modern glycosylation is a sophisticated biosynthetic process, prebiotic analogs of sugar-based amphiphiles may have added chemical versatility, especially in racemic forms [
47].
Together, these features suggest that POLETician membranes may not only be chemically robust but also evolutionarily permissive. Their racemic, modular architecture could potentially support diverse chemical interactions and environmental adaptability, allowing them to persist and evolve in geochemically active habitats where early life first gained a foothold.
2.5. Detectability of POLETicians
2.5.1. Analytical Chemistry and Mass Spectrometry Approaches
Detection of POLETicians may require specialized analytical chemistry tools tailored to recognize non-standard biochemistries. Mass spectrometry techniques, such as time-of-flight (TOF) systems, which are capable of resolving chiral molecules, can distinguish unusual amino acid compositions, including racemic mixtures. A Matrix-Assisted Laser Desorption Ionization—Time of Flight (MALDI-TOF/TOF) approach has been used to demonstrate that D-amino acids can be detected in peptides and proteins [
52]. Chiral derivatization protocols coupled with ultra-high-performance liquid chromatography (UHPLC) may further enhance resolution.
Non-canonical lipid assemblies, such as those composed of branched, racemic, or prebiotically plausible amphiphiles, can be identified using advanced lipidomics workflows employing high-resolution techniques like electrospray ionization mass spectrometry (ESI-MS) and Fourier transform ion cyclotron resonance mass spectrometry (FT-ICR-MS) [
53]. These methods enable precise characterization of lipid molecular masses, isotopic patterns, and fragmentation spectra, making it possible to detect unusual lipid species that deviate from the standard ester- or ether-linked phospholipids found in extant life. ESI-MS allows for sensitive, soft ionization of intact lipid molecules in complex mixtures, while FT-ICR-MS provides ultra-high mass accuracy and resolving power, facilitating the identification of lipids with non-standard chain lengths, head groups, chirality, or elemental compositions [
53]. When combined with chromatographic separation (e.g., LC-ESI-MS or direct infusion with ion mobility), these workflows can distinguish racemic or non-biogenic lipid structures from background environmental organics [
53]. Such analytical strategies are essential for probing potential POLETician membranes, which may incorporate primitive or abiotic lipid-like molecules synthesized under geochemical conditions, offering insights into early compartmentalization mechanisms predating canonical cell biology.
2.5.2. Microscopy and Spectroscopy Techniques
Fluorescent membrane dyes selective for non-standard lipid assemblies could facilitate in situ microscopic identification of POLETician cells. Techniques like Raman spectroscopy, nanoscale Secondary Ion Mass Spectrometry (NanoSIMS) [
54,
55], or high-resolution cryo-electron microscopy may also reveal subcellular structures, unusual morphologies, or metabolic gradients [
56], helping to differentiate POLETicians from abiotic organic material or conventional microbial life. Machine learning–assisted image analysis can also support the high-throughput detection of morphologically distinct entities.
2.5.3. Detection of Mirror L-RNAs
Perhaps the most promising strategy for detecting POLETicians—should they exist—would involve the identification of mirror L-RNAs using engineered RNA-directed RNA polymerases or reverse transcriptases. All known extant terrestrial life relies exclusively on D-ribose nucleotides in RNA, a universal feature resulting from homochirality imposed by early enzymatic selection pressure [
15,
57]. However, in hypothetical pre-enzymatic systems, such as POLETicians, this chiral constraint may not apply. As discussed by Yarus and colleagues, primitive ribozymes or RNA–amino acid aptamers could mediate early translation-like processes without a strict preference for D- or L-nucleotides [
11]. In such a system, mirror L-RNAs could function as informational molecules just as effectively as their D-counterparts. Therefore, detecting these alternative chiral forms would provide a powerful marker for the presence of pre-prokaryotic biochemistry.
2.5.4. Lessons from Asgard Archaea and the Role of Environmental Metagenomics
Asgard archaea offer a precedent for discovering unusual lineages through non-traditional means. They were first identified not through microscopy or cultivation but via metagenomic sequencing of environmental DNA extracted from deep-sea hydrothermal sediments near Loki’s Castle, near the Arctic Mid-Ocean Ridge [
1,
2]. It took over a decade to obtain the first cultured images of a member of this group, strain MK-D1, which revealed unprecedented cellular structures and membrane features [
4]. Similarly, the discovery of POLETicians may depend not on direct imaging but on the development of next-generation sequencing tools tailored to unconventional biomolecules. One promising route involves adapting Oxford Nanopore Technologies’ (ONT’s) direct RNA sequencing platform to analyze mirror L-RNAs [
58].
2.5.5. Engineering Nanopore Platforms for Mirror-RNA Detection
The least expensive of the ONT sequencers, the MINION, is a handheld device that plugs into a laptop USB port and can sequence a drop of DNA or RNA in 24 to 48 h. The major issue with MINION sequencing is coverage, as there would likely be large excesses of non-mirror D-RNAs contaminating the sample, potentially interfering with signals from mirror L-RNAs. Bioinformatics pipelines must also be developed to distinguish and analyze mirror L-RNA signals.
This would require the development of novel mirror D-amino acid RNA helicases or helicase adapters capable of recognizing mirror L-RNAs and feeding them through the nanopore—an engineering challenge, but not an insurmountable one given the precedent for redesigning other polymerases [
59,
60]. Specifically, the RNA helicase used in the ONT nanopore sequencer, a mutated version of Hel308 helicase [
61], could be re-engineered in the mirror form using D-amino acids, as recently done with T7 RNA polymerase [
22]. The resulting nanopore signals, or “squigglegrams,” would differ from those produced by natural D-RNA due to reversed backbone polarity and chirality. AI-based signal interpretation models can be retrained to decode mirrored signal patterns [
62]. In a later section, we describe the concept of a generalized nanopore sequencing instrument that can potentially sequence any unknown linear polymer (ULP).
2.5.6. Mirror-RNA Stability and Fossil Potential
Mirror L-RNAs, also known as “Spiegelmers,” from the German word Spiegel (mirror), are of particular interest because they resist degradation by RNases and other cellular enzymes, making them attractive candidates for drug development [
63,
64]. Their biochemical stability and mirror-image architecture also make them ideal molecular fossils if such entities persist in modern extreme environments. If POLETicians store or express information via Spiegelmer-like molecules, detecting them could offer not only evidence of a novel domain of life but also a window into prebiotic information systems. The successful identification of mirror L-RNAs in deep sediment samples would revolutionize our understanding of the origin of life and expand the biochemical space considered habitable—on Earth and beyond.
2.6. Ecological Niches and Interactions
2.6.1. Symbiotic Roles in Sediment Microbiomes
The discovery of Asgard archaea as obligate symbionts, particularly the
Lokiarchaeota strain MK-D1 co-cultured with
Methanogenium, provides a compelling precedent for seeking POLETicians in similar symbiotic contexts [
4]. These ancient associations, revealed after over a decade of meticulous enrichment, suggest that such partnerships are not transient but are deeply rooted in sediment microbiome ecology. POLETicians may occupy comparable ecological roles, acting as ultraslow-growing symbionts within microbial consortia in anoxic sediments. Similar to the MK-D1/
Methanogenium system, POLETicians may depend on close metabolic integration with cohabiting archaea, bacteria, or even abiotic mineral surfaces. They may interact metabolically with Asgard archaea or methanogens, exchanging small molecules or serving as low turnover scaffolds for redox buffering. Their hypothesized roles include stabilizing microenvironments by gradually absorbing and releasing redox-active species, thereby moderating geochemical fluctuations over timescales ranging from months to years.
2.6.2. Racemic Substrate Utilization and Ecosystem Niche Expansion
Unlike modern life, which exhibits strict homochirality, POLETicians may be uniquely equipped to exploit racemic pools of amino acids and sugars generated abiotically through geochemical and extraterrestrial processes. These racemic substrates are often inaccessible to extant organisms due to their chiral specificity. By metabolizing both D- and L-forms of key organic molecules, POLETicians occupy a distinct trophic niche, functioning as ecological recyclers of otherwise inert carbon and nitrogen sources. In turn, they may provide rare or energetically expensive metabolites to surrounding organisms, potentially including methanogens, sulfate reducers, or Asgard archaea. This mutualistic relationship could extend beyond mere metabolic exchange to include structural roles, with POLET biofilms acting as diffusion buffers or anchoring platforms within porous sediment matrices. Thus, the existence of racemate-adapted POLETicians would not only redefine our understanding of biochemistry but also expand the functional biodiversity within ancient microbial ecosystems.
2.7. Alternative Energy Acquisition
2.7.1. Redox and Radiolytic Metabolism
Unlike modern phototrophs and chemotrophs, POLETicians, if they exist, likely bypass both photosynthesis and enzymatically catalyzed respiration. Instead, they are hypothesized to exploit extremely slow and low-yield abiotic energy sources, such as electrons from radioactive decay, mineral-sourced redox gradients, or reactive radicals formed through radiolysis or geochemical reactions. These processes may be sufficient only to maintain molecular integrity and enable sparse biosynthetic events over long periods—potentially months to centuries—rather than fueling growth or cellular division.
While some modern deep subsurface bacteria can survive on radiolysis-derived hydrogen and sulfate for millions of years [
65,
66], these organisms rely on elaborate enzyme systems and metabolic networks. In contrast, POLETicians are postulated to lack enzymes entirely or possess only rudimentary catalytic systems (e.g., mineral surface–mediated chemistry). They would rely on passive, non-enzymatic electron flow through environmental redox couples such as Fe
2+/Fe
3+ and Mn
2+/Mn
4+ without the complex transport and regulation machinery seen in modern cells.
Thus, although both POLETicians and some extremophilic bacteria may inhabit low-energy environments, POLETicians differ in both degree and kind; they represent a prebiotic or proto-biological mode of energy utilization, operating at a lower complexity threshold and metabolic tempo. This distinction positions POLETicians not merely as slow bacteria but as a fundamentally different category of life-like systems rooted in geochemistry rather than biochemistry.
2.7.2. Distinctions from Known Extremophile Bacteria
Although some extremophile bacteria can survive under energy-limited conditions using alternative sources—such as hydrogen, sulfur compounds, or metal ions—these organisms still rely on complex enzymes and homochiral biochemistry. In contrast, POLETicians may operate without such enzymatic sophistication, instead relying on slow geochemical fluxes or radiolytic particles as direct energy sources. They may function more like quasi-biotic electrochemical reactors, harvesting ambient gradients rather than actively catalyzing reactions. Their metabolism resembles abiotic mineral oxidation more than cellular respiration, placing them at the conceptual boundary between life and pre-life systems. This fundamental distinction underscores the need to explore new models of energy acquisition that do not assume the presence of enzyme-catalyzed metabolic cycles.
2.7.3. Energy Required for Repair Mechanisms in POLETs
One of the fundamental challenges for any living or proto-living system is to maintain its structure and function in the face of continuous molecular-level damage. To better frame this challenge for POLETs, we now explicitly discuss the concept of an “energy per damage repair unit” threshold.
We argue that POLETicians are likely to have the following characteristics:
Lower complexity (e.g., fewer distinct macromolecular components),
Slower turnover rates, and
Reduced information density, which collectively lowers the daily “hit rate” from environmental stressors (e.g., UV, hydrolysis, oxidation).
In this framework, even if POLETicians are subjected to 5–10 damaging events per day (rather than hundreds as in more complex cells), they would still require a minimum sustainable energy flux to drive rudimentary repair or replacement pathways. Radioactive decay, redox gradients in hydrothermal vents, and thioester chemistry could serve as plausible energy sources, even in the absence of ATP or proton gradients.
While the quantitative energy yield of such sources remains speculative, we argue that the energy input from these sources could have been sufficient to sustain life-like persistence in POLETicians, precisely because their repair costs were much lower than those of modern prokaryotic and eukaryotic cells.
2.8. Implications for the Search for Life
2.8.1. A Broader Framework for Life’s Emergence
The possibility that POLETicians exist challenges the traditional view that life emerged linearly through a single, rare origin-of-life (OOL) event in the Earth’s early history. Rather than invoking modern de novo abiogenesis, we propose that POLETicians are descendants of ancient, pre-LUCA systems that persisted through deep time. The ancestors of POLETicains may include precellular or non-cellular entities—metabolically active, quasi-living systems that originated under Archean or pre-Archean geochemical conditions and subsequently occupied ecological refugia.
Such POLETician entities could represent a “shadow biosphere” an alternative biochemical system that never progressed to full cellularity or homochirality but nonetheless exhibits information storage, energy coupling, and environmental responsiveness [
8,
9]. Crucially, their persistence does not require modern OOL conditions but only ecological continuity in energy-limited, geologically stable environments. These POLETicians may function at extremely low metabolic rates, similar to dormant archaea or viruses in permafrost and abyssal clays. Rather than competing with canonical life, they may operate in distinct thermodynamic and ecological environments.
If so, POLETicians may provide a distributed substrate for evolutionary experimentation—evolutionary “dead ends” or alternate paths that could inform the early trajectory of life’s emergence. The study of POLETicians may thus yield insights into pre-Darwinian complexity and the phase space of early biological systems, including those that failed to achieve vertical heredity or robust replication.
2.8.2. Terrestrial and Extraterrestrial Search Strategies
Asgard archaea offer a key precedent: originally discovered in deep-sea hydrothermal vents, they were later found in diverse anaerobic environments through environmental DNA (eDNA) surveys, including anoxic sediments, methane seeps, and hypersaline lakes. Similarly, if POLETicians exist, they may be widespread but undetected and adapted to extreme or cryptic ecological niches. These could include deep biosphere habitats, radiolytically active mineral surfaces, hydrothermal chimney interiors, or symbiotic relationships within LUCA-based life, where they escape detection by canonical probes.
Detection strategies should extend beyond conventional metagenomics. Novel tools are needed because POLETicians may exhibit non-canonical biochemistry, such as racemic chirality, alternate base-pairing systems, or L-sugar backbones. These include mirror-PCR, enantiomer-selective fluorescence probes, and single-molecule sequencing, which are optimized for non-standard sugars and bases. Enantiomeric separation techniques and nanopore-based chirality detection can further expand the detection sensitivity.
The discovery of POLETicians on Earth would also redefine strategies for life detection beyond Earth. Mars, Europa, Enceladus, and Titan—worlds with cryogenic, anoxic, or radiolytic conditions [
67]—may host POLETician-like systems that are similar. Data from past missions should be re-analyzed with this broader biosignature framework in mind. Oxidized crusts or subsurface niches, once deemed too marginal, may harbor slow-metabolizing or dormant POLETicians.
2.9. Redefining the Hallmarks of Life
2.9.1. Toward a POLET-Compatible Life Definition
Contemporary definitions of life are often based on criteria such as rapid replication, enzymatic metabolism, cellular compartmentalization, and responsiveness to stimuli. However, POLETicians would challenge these norms by embodying life-like systems that operate on geological timescales, with minimal structural complexity and no requirement for high-speed enzymatic catalysis. Their existence would demand a more inclusive definition of life, one that allows for slow, equilibrium-nearing metabolic states, racemic biochemistries, and non-genomic reproduction through blebbing or autocatalytic cycles.
This expanded view of life prioritizes persistence, information transfer, and environmental coupling over replication speed or genetic complexity. It would also recognize that early prebiotic systems may have passed through a prolonged POLET-like phase before transitioning to canonical cellular life.
2.9.2. A New Domain Between Chemistry and Biology
The study of POLETicians could uncover a missing evolutionary stratum—a previously unrecognized domain of quasi-life bridging prebiotic chemistry and biological evolution. Rather than viewing abiogenesis as a binary transition, POLETicians offer a model of a gradual continuum populated by stable, semi-autonomous metabolic systems that failed to—or never needed to—cross into full-fledged life [
44]. Their inclusion in our conceptual framework would necessitate a revised taxonomy of living systems, encompassing non-cellular, racemic, and ultraslow entities alongside the traditional domains of life.
Collectively, these expanded characteristics and implications reinforce the plausibility and scientific value of the POLET hypothesis. If POLETicians, loosely defined as any pre-prokaryotic direct descendant that exists today anywhere on the planet, are confirmed to exist, they would revolutionize our understanding of the origins of life and provide a powerful new lens through which to explore both terrestrial biology and the search for extraterrestrial life [
68].
2.10. Evidence Supporting the POLET Hypothesis
While POLETicians are proposed as hypothetical pre-prokaryotic systems predating LUCA, multiple empirical lines of evidence lend plausibility to their existence and suggest that some POLETician-like features may persist in marginal or cryptic modern contexts. These lines of support span microbiology, geochemistry, astrobiology and prebiotic chemistry.
First, laboratory experiments simulating prebiotic Earth environments have robustly demonstrated that amino acids, nucleobases, simple sugars, and amphiphilic molecules can form spontaneously under plausible early Earth conditions—particularly through spark discharge, UV irradiation, or hydrothermal chemistry [
48,
49,
50]. These abiotic syntheses typically yield racemic mixtures and heterogeneous polymers, aligning with the predicted properties of POLETician systems. Notably, many of these molecules spontaneously assemble into higher-order structures, such as micelles, vesicles, fibrils, or coacervates, even without enzymatic control.
Second, modern extremophiles offer indirect support for the persistence or re-emergence of POLETician-like characteristics. Organisms thriving in hypersaline lakes, hydrothermal vents, and subglacial brines tolerate physicochemical conditions resembling those predicted on early Earth. Some extremophilic archaea and bacteria exhibit metabolic and structural minimalism, suggesting an evolutionary descent from more rudimentary forms [
69,
70]. Additionally, organisms such as
Nanoarchaeum equitans, with only 536 genes, and Candidate Phyla Radiation (CPR) bacteria exhibit highly reduced genomes, mosaic gene content, and syntrophic lifestyles [
71], which echo the hypothesized modularity and compartment-sharing among POLETs.
Third, biochemical and structural studies have revealed that peptides composed of racemic amino acids can still form ordered beta-sheet-rich fibrils and nanotubes, offering structural integrity and compartmentalization independent of membrane lipids [
72]. Moreover, racemic fatty acid vesicles formed in prebiotic simulations have demonstrated robustness under fluctuating environmental conditions [
46]. These findings support the idea that POLET membranes and scaffolds may be chemically tolerant, non-homochiral, and dynamically assembled.
Fourth, rare but persistent biological fossils or metabolic signatures, such as kerogen-like structures, isotopic fractionation patterns, and abiotic methane production, suggest that pre-LUCA chemistry may still leave detectable imprints in ancient rocks and extraterrestrial settings [
73]. Kerogen is a solid, waxy organic material found in sedimentary rocks, particularly oil shales, and is insoluble in common organic solvents. It is essentially a precursor to petroleum and natural gas, formed from the remains of dead microorganisms, plants, and animals [
74]. Ongoing and future missions to Mars, Europa, and Enceladus seek precisely such prebiotic or transitional chemistries [
67], which could be interpreted through the POLET framework if discovered.
Finally, synthetic biology is beginning to recreate life-like behavior in non-enzymatic systems. Experiments using self-replicating RNAs, ribozymes with primitive peptide bond activity, and artificial protocells with dynamic membranes provide proof of principle that life-like properties can emerge without the full suite of modern cellular machinery [
75]. These synthetic efforts echo key attributes of POLETicians and may offer an experimental testbed for refining the model.
In summary, while POLETicians are theoretical constructs, converging evidence from disparate fields supports the plausibility of such intermediate systems bridging geochemistry and cellular life and warrants further investigation. Recognizing POLETician-like behavior in either extant extremophiles or synthetic systems may not only illuminate the origins of life, but also guide astrobiological investigations and the design of minimal life forms.
While POLETicians are hypothesized to be primordial life-like entities, it is conceivable that some of their features persist in rare or cryptic modern contexts. Several empirical observations from microbiology, geochemistry, and prebiotic chemistry support the existence or feasibility of such systems.
We summarize the existing evidence for POLETicians in
Table 2, including both positive experimental observations (e.g., long-lived mirror RNAs and unexplained D-amino acid metabolism) and process-level plausibility (e.g., spontaneous membrane dynamics, geothermal or radiogenic energy inputs, and extremophile persistence in nutrient-poor environments).
Some highlights of
Table 2 are as follows:
These lines of evidence—although we admit they are individually circumstantial—collectively support the plausibility of ancient and perhaps even surviving POLETician-like systems.
Table 2.
Summary of empirical and theoretical evidence supporting the POLET hypothesis.
Table 2.
Summary of empirical and theoretical evidence supporting the POLET hypothesis.
| Evidence Type | Observation or Phenomenon | Relevance to POLET Hypothesis | References/Notes |
|---|
| Extant Precellular Forms | Asgard archaea as intermediates between archaea and eukaryotes | Suggests pre-prokaryotic entities (POLETs) may once have existed and left descendants | [2] |
| Mirror RNA stability | Long mirror-RNA (L-RNA) persists in pond water for weeks | Suggests a lack of natural degrading enzymes; intrinsic stability or presence of rare mirror-life RNases | [22]; proposed assay using deep-sea sediments |
| Spontaneous membrane blebbing | Fatty acid vesicles form and divide spontaneously in thermal and osmotic gradients | Supports leaky, dynamic compartmentalization in early cells | [22] |
| D-Amino acid or L-sugar metabolism by microbes | Bacteria reported to grow on D-amino acids or L-sugars | Usually interpreted as conversion enzymes, but could indicate primitive or mirror metabolic pathways | Various, e.g., Pasteur noted selective sugar metabolism [77] |
| Low-energy environments | Radioactive decay, radiolysis of water, and redox gradients can provide usable energy in the absence of sunlight | Compatible with POLETs’ low complexity and metabolic demands | [76,78] |
| Persistent abiotic chemistry | Ribose, peptides, and phosphates persist under geothermal and high-pressure conditions | Supports plausibility of prebiotic synthesis sustaining POLET-like systems | [79] |
| Unusual extremophile organisms | Life persists in ultradeep, oligotrophic, and high-radiation environments | Suggests ancient adaptive mechanisms or relic metabolic flexibility | Reviewed in [80] |
2.11. Experiment Approaches Needed to Validate the POLET Hypothesis
To assess the scientific validity of the POLET hypothesis, we propose a framework for experimental investigations capable of detecting remnants or modern analogs of pre-prokaryotic life forms. These approaches are summarized in
Table 3, which outlines the necessary experiments for unknown linear polymer (ULP) detection with a generalized nanopore device, mirror biomolecule detection, polymer sequencing, and functional assays of chirality-based metabolism.
For example, the use of mirror PCR with mirror L-DNA primers and mirror L-dNTPs or mirror nanopore sequencing could enable the detection of long L-RNA or L-DNA sequences from environmental samples, such as deep-sea sediments, where POLETicians may persist. Development of generalized nanopores capable of sequencing polymers regardless of base chemistry or chirality is an essential goal, both to explore noncanonical life and to address the broader search for xenobiology.
Mass spectrometry (MS) approaches targeting mirror peptides and mirror sugars can complement this effort by identifying unnatural D-proteins or L-sugars. Finally, environmental screens for organisms capable of metabolizing exclusively mirror amino acids or mirror sugars (e.g., D-glucose) should be revisited using modern genomic tools to exclude artifact explanations, such as isomerase activity.
These proposed experiments offer practical avenues for confirming or refuting the core predictions of the POLET hypothesis.
2.12. Clarifying the Nature and Diversity of POLETicians
The concept of POLETicians was introduced to provide a theoretical framework for molecular systems that existed before or parallel to the last universal common ancestor (LUCA) and may still persist in rare or shielded environments. POLETicians are defined broadly as molecular lineages that exhibit partial or complete hallmarks of life (e.g., boundedness, metabolism, heredity, variation, and evolution) but do not rely on the canonical ribosome-DNA central dogma. This section expands on their possible modes of existence, the ecological niches they occupy, and their evolutionary relevance.
We clarify that POLETicians are not restricted to a single chemical or biological model. Instead, they encompass a spectrum of pre-ribosomal and pre-LUCA systems, some of which may be extinct, others extant but cryptic, and some potentially reconstructable through synthetic biology. We subdivided POLETicians into distinct categories, as presented in
Table 4, and discussed their potential relationship with known biological systems.
2.12.1. Abiotic-Chemical POLETicians: Surface-Templated Systems
These POLETicians represent non-cellular chemical replicators bound to mineral surfaces, such as montmorillonite clays, pyrite, or hydroxyapatite, where the surface charge and geometry may catalyze polymerization and selective retention of molecular species [
84]. They could consist of short peptides, racemic polymers, or proto-RNAs that evolve within geochemical gradients in hydrothermal vents or mineral matrices. These systems fall short of life by modern definitions but may have been precursors to biological POLETicians or contributors to the origin of life.
2.12.2. Protocellular POLETicians: Non-DNA-Based Self-Replicators
This class includes membrane-bound protocells that encapsulate non-DNA informational molecules, such as
Ribozymes or mirror-RNA (based on L-ribose instead of D-ribose),
Peptide-nucleic acid hybrids, and
Xeno-nucleic acids (XNAs) are synthesized through non-enzymatic or early enzymatic means [
85]. XNA stands for xeno nucleic acid. They are a class of synthetic nucleic acid polymers that are structurally similar to DNA and RNA but with modifications to their sugar-phosphate backbone. These modifications can include changes to the sugar molecule itself or to the way the sugar is attached to the phosphate group [
86].
These systems may have achieved heredity and selection without DNA or ribosomes and could plausibly exist today in deep subsurface environments, sea ice brine, or biofilm interstices.
2.12.3. Host-Dependent POLETicians: Living Within Archaea or Bacteria
Some POLETicians may persist today in symbiotic, parasitic, or endosymbiotic relationships with archaeal or bacterial hosts, avoiding competition with faster-replicating organisms by occupying metabolically minimal niches. For example, nanoarchaea and other ultramicrobacteria possess reduced genomes that may reflect ancestral metabolic simplicity [
87]. Likewise, defective or semi-autonomous replicators may be hidden in the pangenomes of archaea or remain unclassified in metagenomic dark matter.
2.12.4. Viral POLETicians: The Case for Giant and Extinct RNA Viruses
An intriguing possibility is that the extinct lineages of large or giant RNA viruses once represented autonomous or semi-autonomous POLETician-like systems [
88]. These ancient viral forms may have exhibited transient independence, encoding partial translation systems, metabolic enzymes, and capsid-free life stages. Modern descendants may include
Narnaviridae,
Mitoviridae, or other capsid-less RNA replicators, which still replicate within host mitochondria or fungi and share features with minimal pre-cellular life forms [
89].
Additionally, Mimiviruses and their relatives, although DNA-based, demonstrate how complexity and large genome size can evolve in non-cellular replicators. Their deep evolutionary roots suggest possible links to ancient RNA-based viral forms that now exist only as molecular fossils [
90].
These RNA-based viral POLETicians could be the modern equivalents of ‘root dwellers’ in the Tree of Life, representing lineages that diverged before LUCA and survived by evolving symbiotic or parasitic strategies.
2.12.5. Synthetic and Reconstructable POLETs
The final category includes systems that have not yet been observed in nature but can be reconstructed or simulated in the laboratory using xenobiology.
Examples include:
Mirror-life based on L-amino acids and L-ribose RNA,
Fully racemic protocells, and
Autocatalytic chemical networks that reproduce functional motifs in synthetic vesicles [
91].
These various types of POLETicians provide testable hypotheses regarding minimal life, prebiotic chemistry, and alternative biologies (
Table 4).
2.13. A Rooted Model: Connecting POLETicians to the Tree of Life and the Origin of Life
In
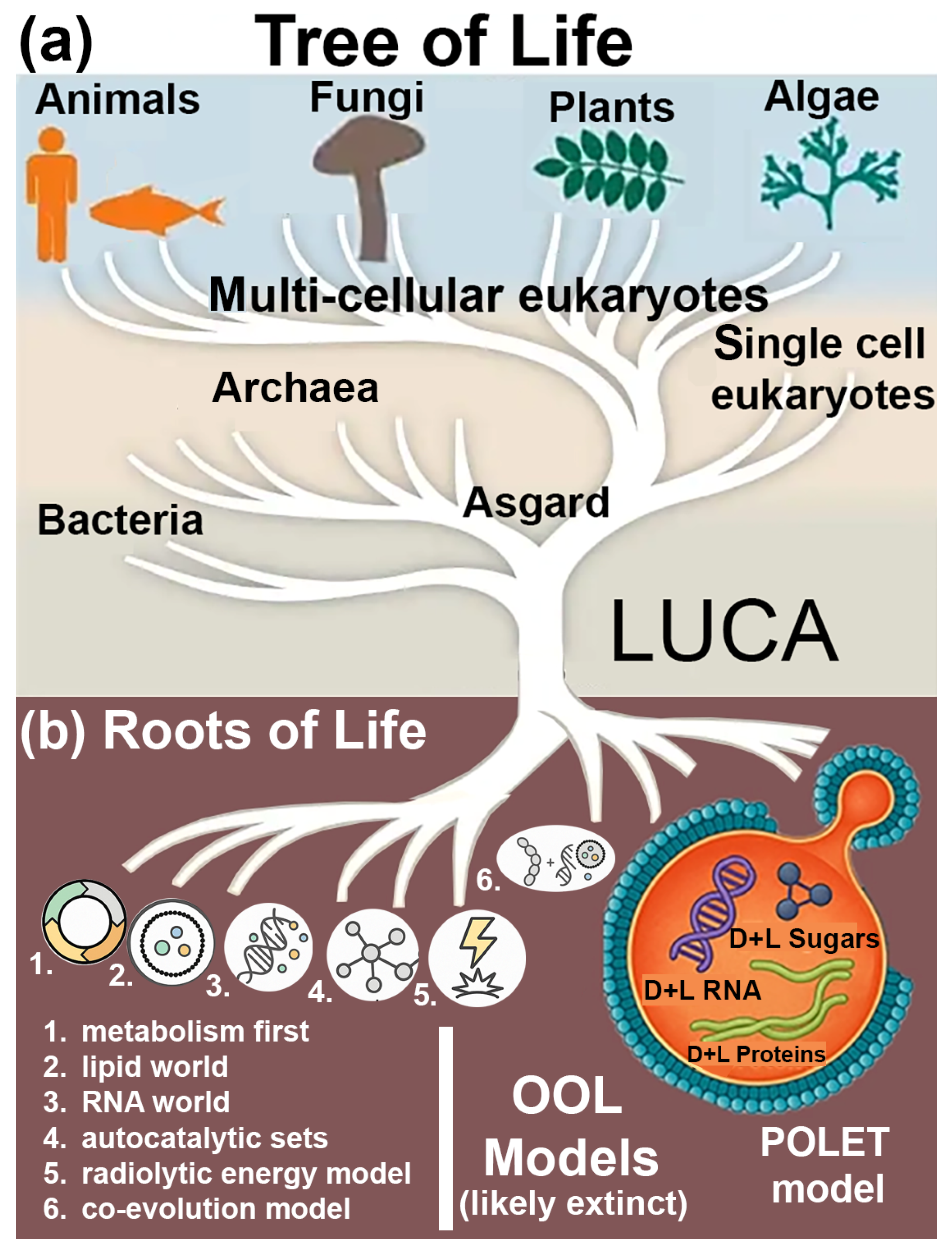
Figure 2, we present a rooted Tree of Life with LUCA (the Last Universal Common Ancestor) at the trunk and modern domains (Archaea, Bacteria, and Eukarya) branching upward. Critically, the roots of the tree extend downward into deeper evolutionary times and biochemical simplicity. These roots represent hypothetical pre-LUCA life forms—POLETs. These entities may include extinct lineages or cryptic extant life forms within a “shadow biosphere,” that have not yet been detected by conventional life-detection methods.
POLETicians are proposed to have existed prior to the advent of enzymatic systems, relying on spontaneous or mineral-catalyzed biochemistry using racemic RNA, peptides, and alternative membrane structures. This framework, shown as an extension below LUCA in
Figure 2, suggests that the roots of life are best explored through the intersection of ancient geochemistry, synthetic prebiotic chemistry, and models of non-enzymatic life.
To integrate this view, we summarize the major models of the Origin of Life (OOL) in
Table 5. Each model proposes a distinct pathway from abiotic chemistry to protocellular systems. The first six models listed in
Table 5 are illustrated along the base of
Figure 2, aligned with the POLET framework as possible ancestral precursors or parallel paths that may have contributed modularly to LUCA.
2.13.1. Origin of Life (OOL) Models
We highlight six leading theories for the Origin of Life (OOL), each contributing a different viewpoint from which to study POLETicians (
Figure 2 and
Table 5):
Metabolism-First: This model posits that self-sustaining networks of metabolic reactions arose first, often driven by mineral catalysts on surfaces, such as iron–sulfur minerals. These networks can generate energy-rich intermediates and eventually support more complex biochemistry [
92].
Lipid World: Amphiphilic molecules form spontaneously in aqueous environments, producing micelles or vesicles capable of encapsulating and concentrating organic molecules. These pre-cellular compartments provide a scaffold for primitive biochemical reactions, enhancing molecular stability and selectivity [
93].
RNA World: A popular model suggesting that RNA molecules, capable of both storing genetic information and catalyzing chemical reactions, were the first biomolecules to achieve Darwinian evolution. Ribozymes can self-replicate and undergo selection, eventually giving rise to DNA and proteins (reviewed in [
94]).
Autocatalytic Sets: This model centers on the emergence of chemical networks in which the products of reactions catalyze the production of more components within the set. These systems can self-sustain, adapt, and exhibit rudimentary evolution without requiring specific polymers [
95].
Radiolytic Energy Model: Radiolysis of water and organic molecules by natural radioactivity (e.g., from U, Th, or K isotopes) can generate key reactants and energy gradients. These gradients may have powered early redox reactions, polymer formation, and membrane synthesis in deep subsurface or hydrothermal environments [
96].
Co-evolution Model: Rather than a single origin, this model proposes that multiple molecular systems—membranes, nucleic acids, and peptides—co-evolved in tandem. Cross-stabilizing interactions among these systems increase the overall fitness, enabling gradual complexity increases without requiring dominance by any one biopolymer [
97].
We emphasize that these models are not mutually exclusive. Instead, they may represent complementary or sequential stages in chemical evolution. For example, lipid compartments from the Lipid World may have housed autocatalytic networks, while radiolytic energy could have powered early metabolism. POLETicians can be envisioned as mosaics of these models, representing early life-like systems before the rise of enzymatic control and template-based replication.
2.13.2. The POLET Hypothesis in the Context of OOL Models
The POLET hypothesis integrates and extends traditional OOL models by proposing that pre-LUCA entities were neither fully cellular nor governed by modern enzymatic fidelity. Instead, POLETicians may exist as modular, low-fidelity systems capable of partial self-replication, compartmentalization, and energy utilization.
Unlike models that emphasize a singular dominant pathway (e.g., the RNA World), the POLET framework envisions a prebiotic ecosystem in which multiple OOL pathways co-occurred. Each POLET lineage may have emerged from a different combination of features drawn from the six core models—autocatalysis, lipid encapsulation, redox energy, or partial nucleic acid replication.
This conceptual shift frames LUCA not as the first life form, but as the first successful system that unified and stabilized diverse OOL modules under a robust enzymatic architecture. Thus, LUCA marks the transition from distributed chemical evolution to centralized biological evolution. In contrast, POLETicians represent the direct descendants of transitional states between geochemistry and modern biology.
By grounding POLETicians in well-studied OOL frameworks, we can design experiments to simulate plausible chemistries and identify relic biosignatures. For instance, autocatalytic reaction networks can be explored in the presence of radiolytic byproducts or housed in di-lipid vesicles. These systems can then be interrogated for robustness, heritability, variation, and capacity for selection, which are hallmarks of life in the absence of enzymes [
103].
2.14. A Generalized Nanopore System: RNA Tethering for Universal Polymer Sequencing
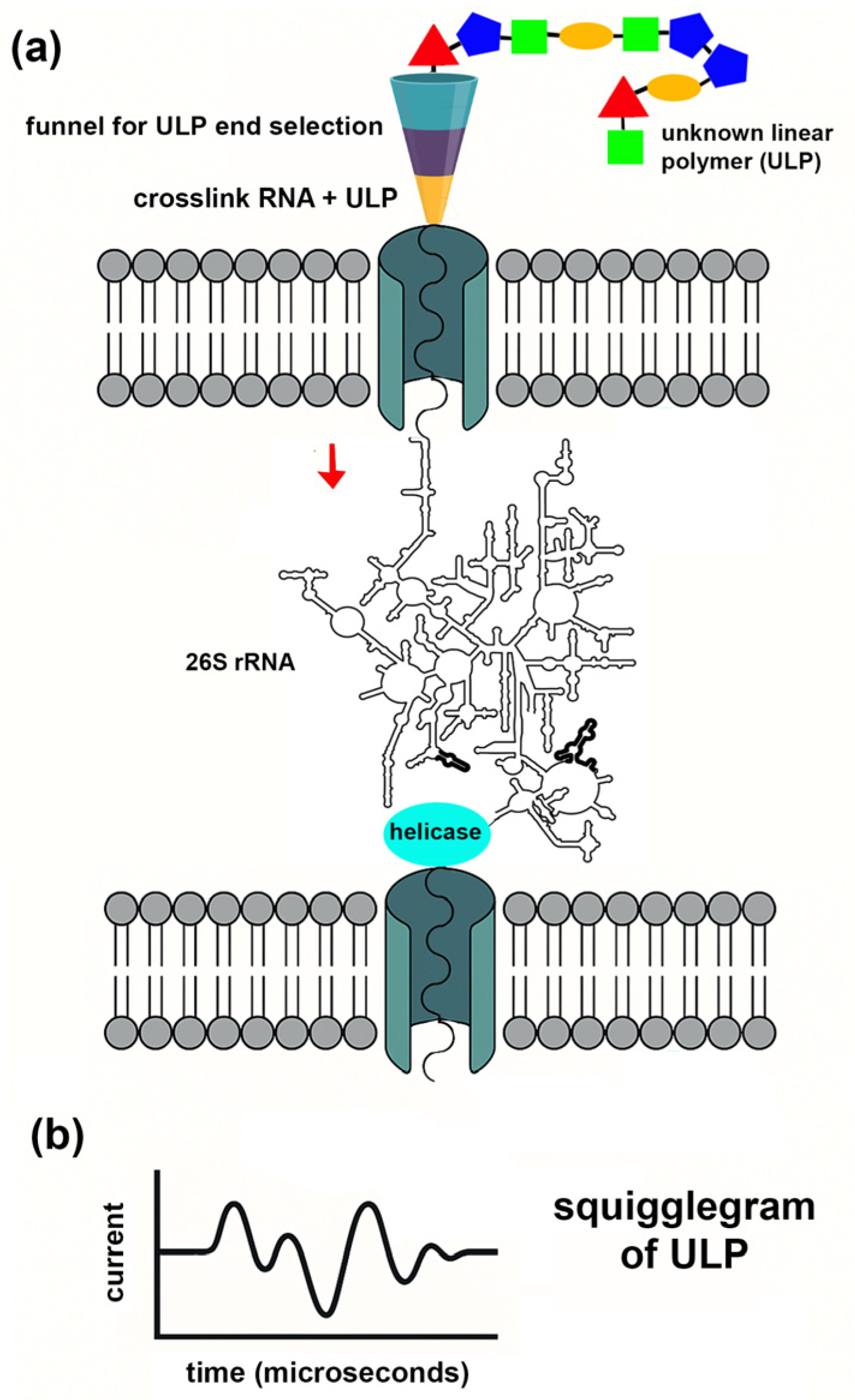
In
Figure 3, we present a conceptual design for a generalized nanopore platform that leverages the compact structure of 26S ribosomal RNA (rRNA) and the translational force of RNA helicases to sequence unknown linear polymers (ULPs)—including natural, synthetic, or extraterrestrial polymers—that can be drawn through a nanopore interface.
2.14.1. Overview of the Generalized Nanopore System
At the core of this system is a vertically oriented three-chamber device with two nanopores embedded in a synthetic or biomimetic membrane. The middle chamber contains a fully folded 26S rRNA complex in a compact conformation. The 5′ end of the 26S rRNA is engineered to reside in the top chamber, positioned at the base of a tapered molecular funnel above the top nanopore. The 3′ end threads through the bottom nanopore, where electrophoretic or motor protein forces pull the rRNA downward, applying directional tension.
This directional force is transmitted from the 3′ end of the rRNA to the 5′ end, which is crosslinked to an unknown linear polymer (ULP) in the molecular funnel in the top chamber. As the 26S rRNA moves into the bottom chamber, it drags the attached ULP through the top nanopore in a linear, unidirectional manner in a controlled manner. Additional sub-chambers under the bottom chamber can be added to sequence the ULP sequentially through several nanopores as a means of error correction.
2.14.2. Caged Crosslinking at the 5′ End
To ensure the specific end-to-end attachment of ULP to rRNA, a caged crosslinker was positioned at the 5′ end of the 26S rRNA. The ULP is loaded into the top chamber, where it encounters the 5′ terminus of rRNA in a sterically constrained funnel environment. Upon localized UV exposure, the caging group—such as a nitrobenzyl-protected azide or diazirine—undergoes photolysis to reveal a reactive site [
104], enabling bioorthogonal covalent bonding to the terminal chemical group of the ULP.
This configuration ensures that only linear, terminally available polymers are ligated, avoiding nonspecific interactions or internal crosslinking. After ligation, the ULP is anchored to the 5′ end of the 26S rRNA, positioning it for force-driven translocation through the top nanopore.
2.14.3. Helicase-Driven Translocation for Sequencing the ULP
With the ULP covalently linked, a biological motor similar or identical to the helicase in the MINION direct RNA sequencing system [
58] initiates the translocation of compact 26S rRNA through the bottom nanopore. The resulting tension propagates upward, pulling the ULP through the top nanopore in a controlled, linear manner.
As the ULP moves through the top nanopore, the ULP is sequenced by ionic current disruption, allowing monomer-by-monomer characterization of ULPs that would otherwise evade conventional sequencing techniques.
This 26S rRNA-tethered system combines the force-transducing ability of large ribonucleoprotein complexes with precise chemical tethering and nanopore confinement. Key advantages include:
Compatibility with both biological and synthetic polymers
Selective covalent attachment via UV-activated caged chemistries
Dual (or more) nanopore configuration for bidirectional sensing or redundancy, and
Modular adaptation to study mirror RNAs, proteins, protein nucleic acids (PNAs), peptoids, also known as poly N-substituted glycines [
105], xeno nucleic acids (XNAs) [
85], deep-sea extracted, or extraterrestrial ULPs is also possible.
Future versions may integrate optical traps, multi-rRNA arrays for parallel processing, or automated caging/uncaging control, forming a foundation for de novo polymer sequencing in astrobiology, xenobiochemistry and synthetic biology.
2.14.4. Advantages and Applications
Once developed, a generalized nanopore sequencing platform can be systematically optimized using well-characterized linear polymers—including DNA, RNA, proteins, and synthetic polymers with defined sequences and structures. By benchmarking the resolution, accuracy, and signal-to-noise performance of the system with these known molecules, researchers can calibrate the signal patterns and refine base-calling or monomer identification algorithms.
Following this validation phase, the nanopore system can be deployed to analyze complex, uncharacterized environmental samples that may contain novel biopolymers or uncharacterized entities, such as POLETicians. These targets could originate from bioorganic matter or sediment samples collected from extreme environments, such as deep-sea mud near hydrothermal vents, where chemical gradients, high pressure, and geothermal energy provide favorable conditions for the emergence or preservation of ancient or exotic life-like polymers.
This approach balances technical rigor with exploratory potential, paving the way for the high-throughput detection of unknown linear polymers that may hold clues to the origin of life or even point to non-canonical biological systems.
3. Discussion
“The originally inactive liquid possesses a sensible rotative power to the left, which increases little by little and reaches a maximum. At this point, fermentation is suspended. There is no longer a trace of right-acid in the liquid.”
Louis Pasteur (1860) [
77].
The above quote from Louis Pasteur reflects one of the earliest empirical observations of molecular chirality in biology [
77]. He discovered that yeast could ferment L-ammonium tartrate but not its mirror image (D-ammonium tartrate). This asymmetric metabolic preference led Pasteur to propose the biological importance of chirality, earning him the title of “Father of Molecular Chirality” [
106].
The POLET hypothesis suggests that Pasteur’s conclusions, while foundational, may not be universally applicable to all forms of life. Specifically, Pasteur never studied organisms inhabiting extreme or anoxic environments—niches where mirror-image (D-chiral) proteins and metabolites could have evolved independently. POLETicians may utilize D-amino acid proteins, L-sugars, and mirror metabolites, escaping detection by conventional microbiological approaches designed for normal chiral organisms.
This hypothesis also builds upon the discovery of Asgard archaea, which have narrowed the evolutionary gap between archaea and eukaryotes [
4]. If such evolutionary intermediates exist today, it is plausible that even more primitive pre-prokaryotic forms—POLETicians— persist in geochemically stable, anoxic environments. These hypothetical organisms offer a paradigm shift: rather than viewing the origin of life solely through synthetic reconstruction or ancient fossils, we might identify relic life forms that still operate under near-prebiotic conditions.
However, it must be emphasized that extant POLETicians, if they exist, are unlikely to be identical to their primordial ancestors. Evolutionary drift, horizontal gene transfer, and localized environmental pressures may have altered their characteristics, potentially incorporating partial chirality or even merging with prokaryotic forms. The POLET hypothesis allows for these possibilities while maintaining its core assertion that direct descendants of pre-prokaryotic life may still exist.
The metabolic strategies proposed for POLETicians—such as radiolytic hydrogen production, redox gradient utilization, and non-enzymatic catalysis—are supported by observations of extremophiles in deep subsurface environments [
65,
66]. Unlike typical bacteria that use alternative energy sources, such as sulfur, hydrogen, or methane oxidation, in active metabolisms, POLETicians are predicted to operate on ultra-slow timescales, prioritizing stability over replication. This distinction from known extremophiles is critical, as it underscores the POLET model’s focus on “life at the edge of metabolism.”
This concept also forces a re-examination of what qualifies as “life.” POLETicians may challenge traditional definitions by existing in a near metabolic stasis, reproducing over millennia, and lacking enzymes altogether. Their existence would demand a revised biological framework—one that accommodates geologically slow processes and non-chiral biochemistry.
Emerging technologies, such as chiral mass spectrometry, super-resolution fluorescence microscopy, and deep metagenomic sequencing—have made it increasingly feasible to detect such organisms. Deep marine sediments, serpentinite-hosted vents, and impact-fractured cratons offer accessible natural laboratories.
The implications of this extend far beyond Earth. If POLET-like organisms are detected in Earth’s deep biosphere, similar entities may exist in the subsurface oceans of Europa or Enceladus or in Martian salt deposits. Thus, POLETicians provide a potential bridge between origin-of-life research and astrobiological exploration.
Importantly, the 2024 proposal of a “mirror life moratorium,” published in
Science by Adamala et al., raised ethical and biosafety concerns regarding artificially creating organisms with mirror-image biochemistry [
107]. These concerns stem from fears that mirror organisms could evade natural immune responses and destabilize systems at multiple levels of organization. The POLET hypothesis suggests an alternative, safer path forward: instead of constructing mirror life, we should search for it in nature, where it may already exist, naturally constrained by isolation and geological timescales.
This approach avoids the bioethical dilemmas of synthetic mirror life while still addressing its core scientific questions. If POLETicians are discovered, their biology will inform both the safe development of mirror systems and the natural boundary conditions under which such life can exist.
Given their hypothesized metabolic stasis, laboratory culture may be impractical for these bacteria. Instead, observational approaches, such as detecting radiolytic energy gradients, vesicle aging, or chiral macromolecules, will be essential. Age gradients or molecular divergence within vesicle populations may allow for indirect lineage tracing and evolutionary reconstruction.
4. Tree of Life Revisioned: Implications for Last Universal Common Ancestor (LUCA)
A conceptual model of life’s deep ancestry is illustrated in
Figure 2, where the conventional tree of life arises from a system of deeper, prebiotic ‘roots’—the POLETicians and the hypothetical, extinct pre-prokaryotic life forms (PPKs). These root organisms form interconnected structures beneath the trunk, representing systems that contribute key components to the emergence of modern cellular life. Some PPKs may have contributed to the evolution of proto-mitochondria, centrioles, or primitive endomembrane systems, while POLETicians may reflect surviving analogs of primitive compartmentalized chemistry.
This framework clarifies a key conceptual distinction noted by a reviewer: if discovered, the Asgard archaea and extant POLETs are not direct ancestors of modern life but rather evolutionary holdovers—lineages that have themselves continued evolving over billions of years. Just as extant Asgard are not the ancestors of eukaryotes but their closest known relatives, modern POLETicians would not be the original source of cellular life but representatives of a primordial branch that never gave rise to the dominant clades.
Positioned just above these roots is the Last Universal Common Ancestor (LUCA), representing the trunk of the tree from which all extant Bacteria, Archaea, and Eukarya diverged. LUCA is not believed to be the first life form on Earth but rather the last shared ancestor of all modern cellular organisms. LUCA likely possessed a genetic code, ribosomes, and ATP-based metabolism, as indicated by conserved features across life. The LUCA model, as elaborated in the 2024 study by Moody et al., suggests an age between 4.09 and 4.33 billion years ago, placing it firmly within the early Earth environment shaped by tectonic activity, hydrothermal gradients, and a declining heavy bombardment phase [
7].
The addition of POLETicians and PPKs to this extended model does not contradict LUCA but complements it by providing a pre-LUCA context. These hypothetical life forms—or proto-life forms—may not meet classical definitions of life based on rapid metabolism and replication but nonetheless offer plausible models for compartmentalized, racemic, blebbing-based systems capable of limited heredity and selective catalysis [
44].
Figure 2 further illustrates how these POLET roots may have diversified into multiple lineages, most of which are extinct, but some may persist in niche environments. Just as the discovery of Asgards began at deep-sea vents but was later expanded to numerous other extreme environments through eDNA sequencing, POLETs, if they exist, may also be widely distributed. Future detection strategies, including mirror-PCR or single-molecule mirror-RNA sequencing, could reveal their presence across Earth’s unexplored niches, helping us map the ‘roots of life.’
This paradigm not only reshapes our understanding of how life emerged, but also expands the range of systems we consider ‘alive’ and worth searching for in astrobiological investigations. By acknowledging that the tree of life may have grown from many roots, we open ourselves to detecting diverse lineages that challenge conventional dogma—and perhaps help us glimpse the prebiotic past still lingering beneath our feet.
5. Conclusions
The POLET hypothesis posits that direct descendants of pre-prokaryotic, ultra-slow, and potentially mirror-chiral entities—termed POLETicians—may persist in isolated anoxic niches on Earth. These hypothesized life forms challenge foundational biological assumptions by potentially combining non-enzymatic catalysis, radiolytic energy utilization, and extreme metabolic latency.
In contrast to known extremophiles that deploy active metabolic pathways, such as sulfur reduction or methanogenesis, POLETicians might rely on passive and geochemically slow energy acquisition mechanisms, with dormant lifespans potentially spanning decades or longer. Their persistence does not depend on frequent replication but on molecular and membrane stability in protected microenvironments.
If discovered, POLETicians could redefine our understanding of the tree of life by providing a living link to the prebiotic world and the conditions preceding the Last Universal Common Ancestor (LUCA). Such a discovery would profoundly impact evolutionary theory, geobiology and astrobiology.
Moreover, by focusing on natural rather than synthetic mirror lifeforms, the POLET framework offers a bioethically grounded path for investigating mirror chirality without venturing into ecologically risky synthetic biology. Detecting POLETicians could address key questions in origin-of-life research while remaining within the scope of natural biology.
Future research should prioritize the development of analytical tools capable of detecting non-standard chiral molecules, long-lived vesicular populations, and radiolytic or geochemical energy signatures in subsurface and extreme environments. These advances may one day uncover the possibility that the emergence of life is not a singular past event but a continuing, cryptic presence within the Earth’s deep biosphere.
6. Caveats and Future Directions
While the POLET hypothesis offers a provocative extension to the origin-of-life framework, several key caveats must be considered. First, the biochemical feasibility of sustaining molecular replication and heredity through uncatalyzed or mineral-assisted reactions remains uncertain. Although mineral surfaces can facilitate certain prebiotic reactions, maintaining such systems over geological timescales without enzymatic machinery remains uncertain. The racemic nature of abiotically synthesized compounds, including amino acids and nucleotides, also poses challenges to the emergence of heritable functions and structures in POLET-like systems.
Second, the concept of mirror-chiral biomolecules, while chemically plausible, remains empirically unconfirmed in natural systems. The detection of such molecules requires highly specialized instrumentation, such as nanopore or spectroscopic platforms capable of resolving stereochemistry at the single-molecule resolution, none of which are yet in routine environmental use. Although pharmaceutical and astrobiological incentives may eventually drive the development of such tools, their application to POLETician detection remains largely aspirational. In
Figure 3, we present a concept for a “generalized nanopore” that can potentially sequence any linear polymer that can fit through a pore. Such a device could be used on space probes to characterize possible extraterrestrial genetic material prior to returning the specimens to Earth.
Third, ecological feasibility must be considered. POLETicians are likely to be extremely limited in both habitat and population size, constrained to microenvironments such as subsurface mineral fissures, anoxic brine pockets, and geochemically active rock-water interfaces. In these settings, their ability to persist without being outcompeted by faster-growing microbial life may depend on ecological isolation and metabolic non-interference.
Finally, there is the issue of falsifiability. The traits that define POLETs—slow turnover, rare occurrence, and unconventional biochemistry—make them inherently difficult to detect or refute. This places the hypothesis in a precarious position with respect to empirical validation.
Nonetheless, several future directions may help bring the POLET hypothesis into the realm of testable sciences. These include:
Development of non-chiral-biased sequencing and mass spectrometry tools.
Sampling of extreme and underexplored niches, such as serpentinized rock systems, deep brines, and subglacial lakes.
Experimental recreation of POLETician-like vesicular chemistries under simulated radiolytic and prebiotic conditions.
Search for long-lived, low-diversity vesicle populations that may act as metabolic fossils.
Integration of POLET-like models into astrobiological mission planning to guide instrumentation that is sensitive to alternate biochemistries.
In conclusion, although the POLET model is unconventional and highly speculative, it opens new lines of inquiry into the nature of life, its origins, and its persistence in forms and places previously overlooked. A concerted interdisciplinary effort—bridging geochemistry, synthetic biology, astrobiology, and biophysics is essential to assess the plausibility and detectability of these enigmatic shadow life forms.