Trends in Synthetic Biology in the Bioeconomy of Non-Food-Competing Biofuels
Abstract
:1. Introduction
2. Results
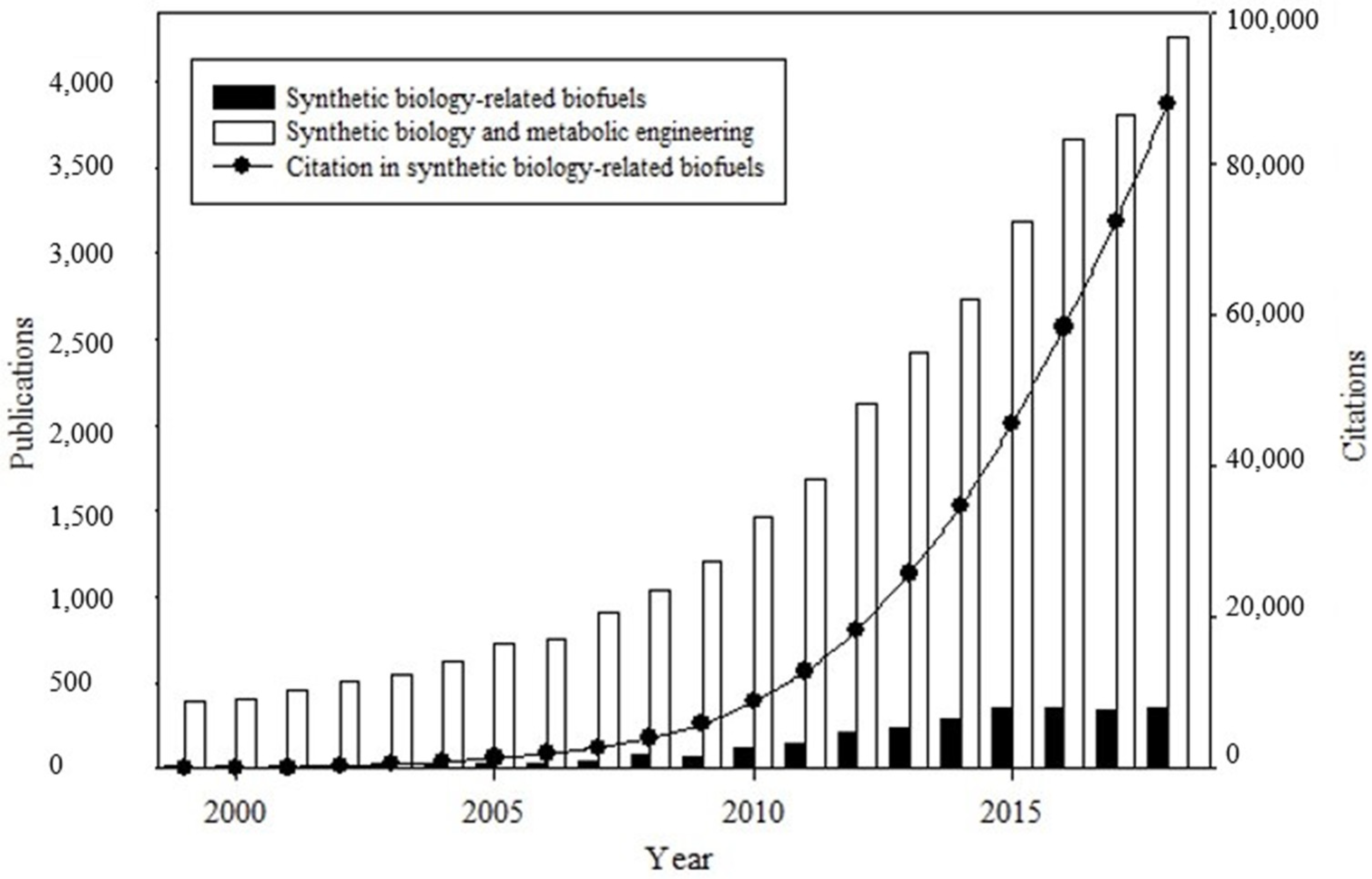
2.1. Articles, Reviews, and Citations on Synthetic-Biology-Related Biofuels
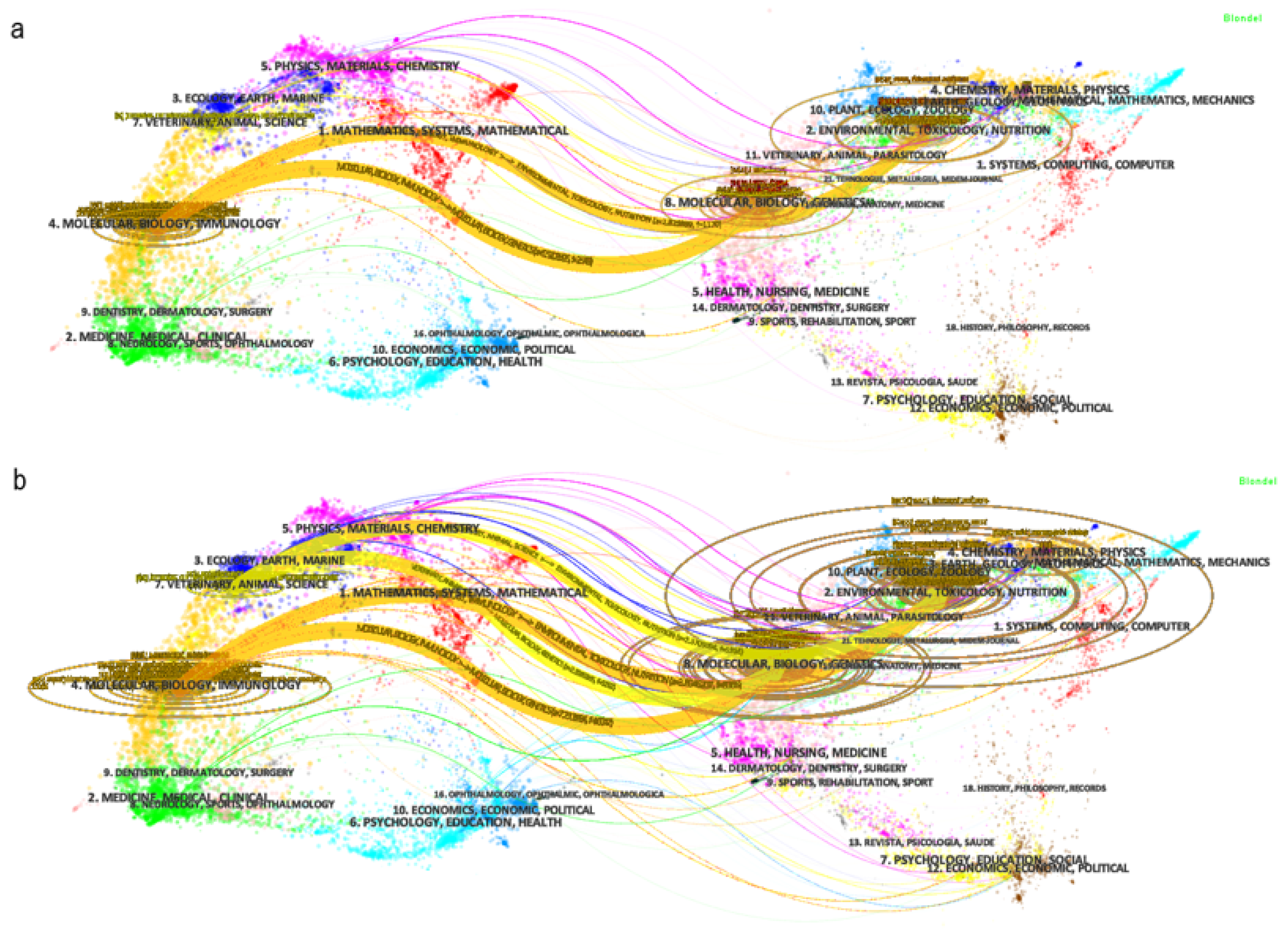
2.2. Interdisciplinary Knowledge on Synthetic-Biology-Related Biofuels
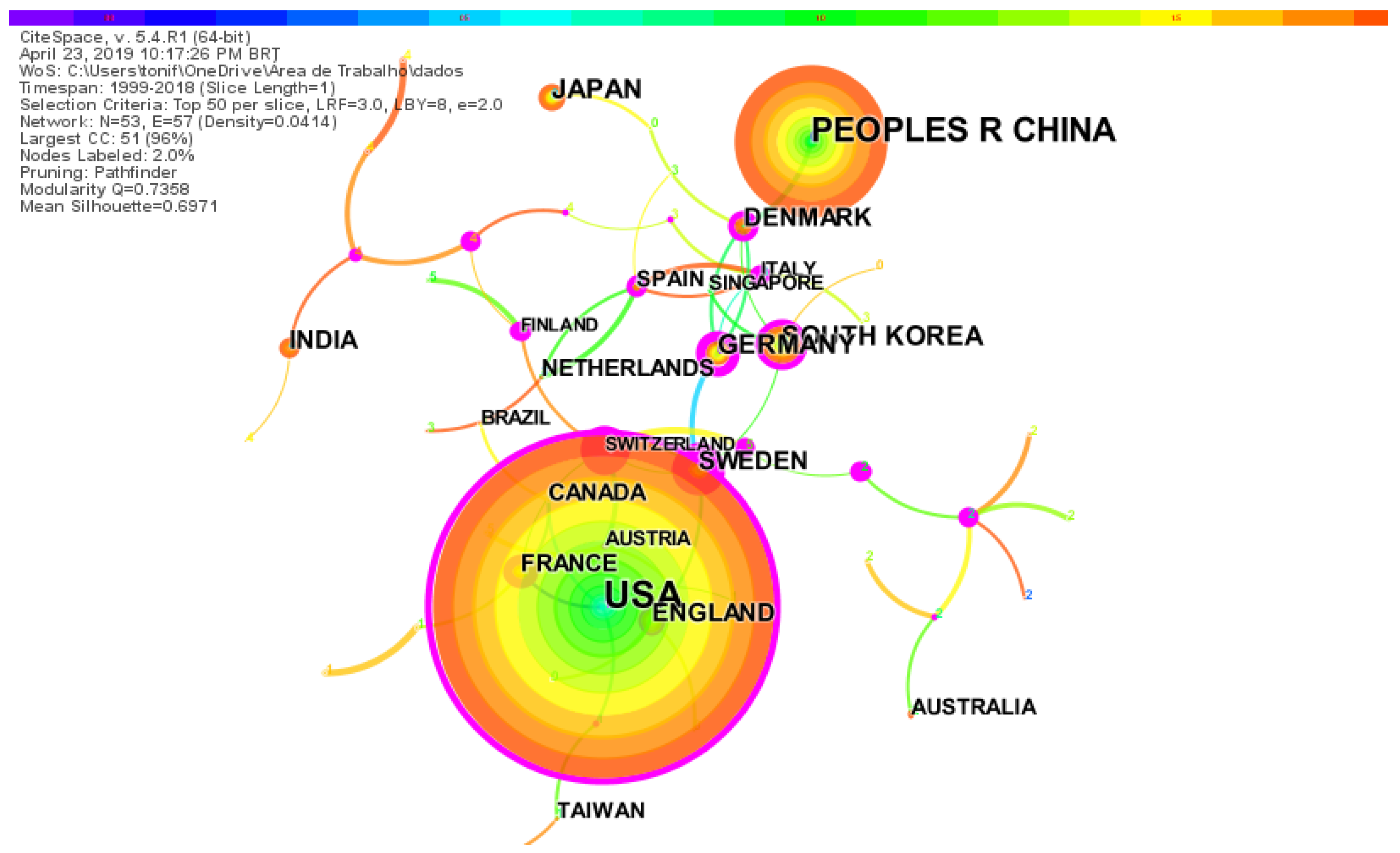
2.3. Worldwide Distribution of Research on Synthetic-Biology-Related Biofuels
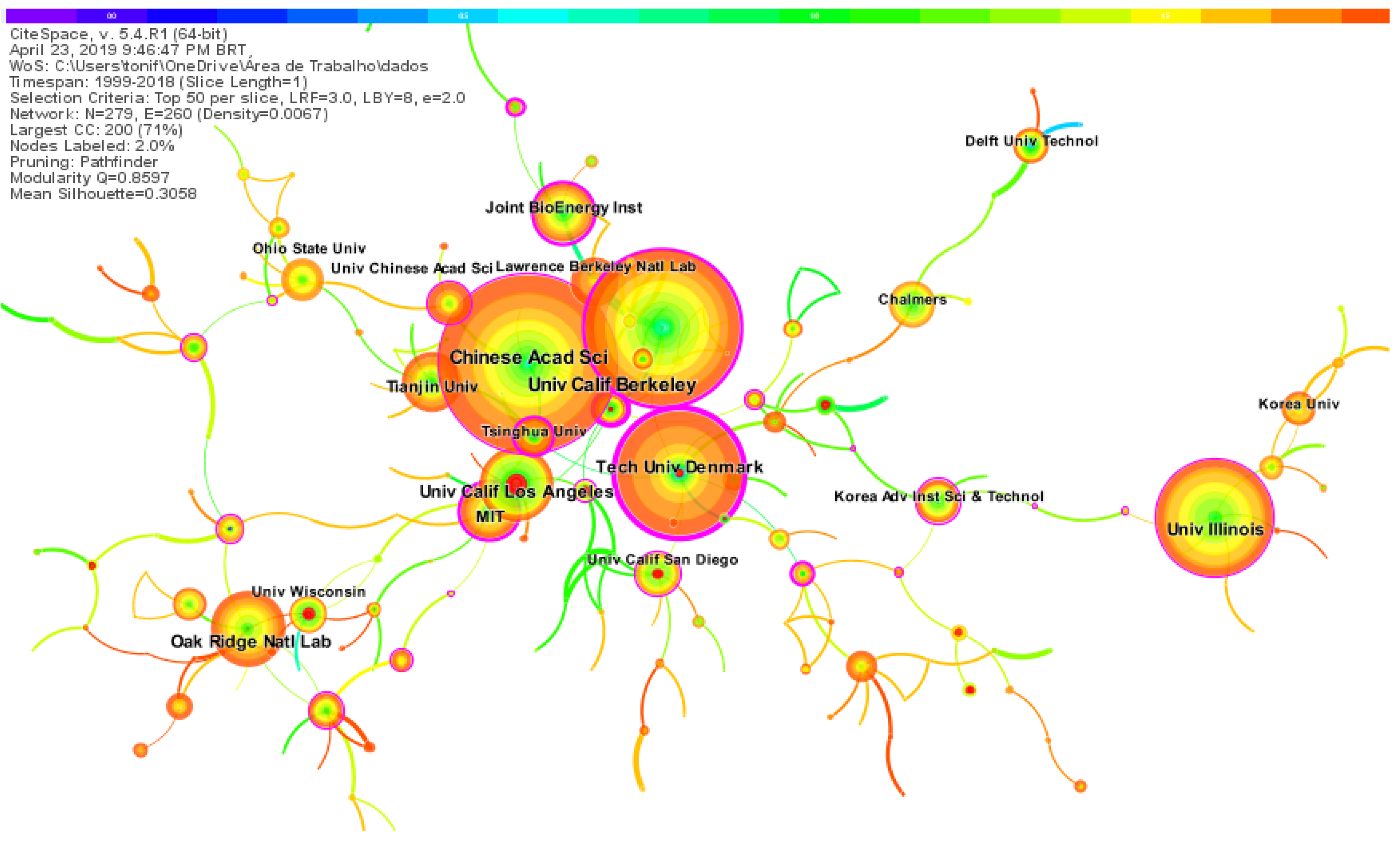
2.4. Institutions Co-Citation Network
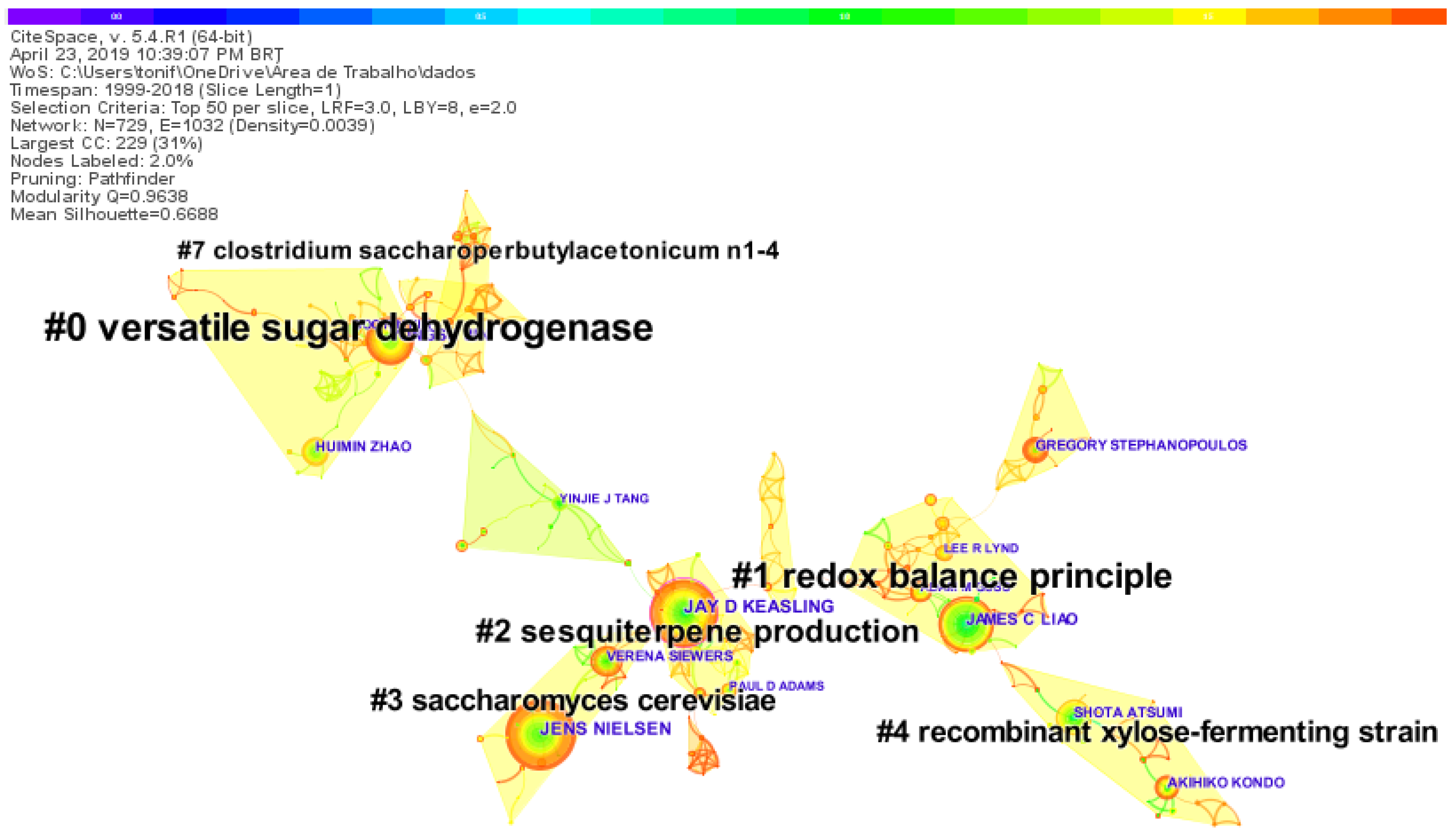
2.5. Clusters Co-Citation Map of Authors and Research Topics on Synthetic-Biology-Related Biofuels
2.6. Papers with the Strongest Citation Bursts on Synthetic-Biology-Related Biofuels
2.7. Emerging Trends and New Developments in the Research on Synthetic-Biology-Related Biofuels
3. Discussion and Future Perspectives
4. Materials and Methods
4.1. Research Data
4.2. Data Analysis
4.3. Networks Properties
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Balat, M. Potential alternatives to edible oils for biodiesel production—A review of current work. Energy Convers. Manag. 2011, 52, 1479–1492. [Google Scholar] [CrossRef]
- Coleman, M.A.; Goold, H.D. Harnessing synthetic biology for kelp forest conservation 1. J. Phycol. 2019, 55, 745–751. [Google Scholar] [CrossRef] [PubMed]
- French, K.E. Harnessing synthetic biology for sustainable development. Nat. Sustain. 2019, 2, 250–252. [Google Scholar] [CrossRef]
- Rudorff, B.F.T.; Aguiar, D.A.; Silva, W.F.; Sugawara, L.M.; Adami, M.; Moreira, M.A. Studies on the Rapid Expansion of Sugarcane for Ethanol Production in São Paulo State (Brazil) Using Landsat Data. Remote Sens. 2010, 2, 1057–1076. [Google Scholar] [CrossRef]
- Pimentel, D.; Patzek, T.W. Ethanol Production Using Corn, Switchgrass, and Wood; Biodiesel Production Using Soybean and Sunflower. Nonrenew. Resour. 2005, 14, 65–76. [Google Scholar] [CrossRef]
- Bergmann, J.C.; Tupinambá, D.; Costa, O.; Almeida, J.; Barreto, C.; Quirino, B. Biodiesel production in Brazil and alternative biomass feedstocks. Renew. Sustain. Energy Rev. 2013, 21, 411–420. [Google Scholar] [CrossRef]
- Souza, S.P.; Seabra, J.E.A.; Nogueira, L.H. Feedstocks for biodiesel production: Brazilian and global perspectives. Biofuels 2018, 9, 455–478. [Google Scholar] [CrossRef]
- Ayres, A. Germany’s water footprint of transport fuels. Appl. Energy 2014, 113, 1746–1751. [Google Scholar] [CrossRef]
- Nanda, S.; Mohammad, J.; Reddy, S.N.; Kozinski, J.A.; Dalai, A.K. Pathways of lignocellulosic biomass conversion to renewable fuels. Biomass Convers. Bioref. 2014, 4, 157–191. [Google Scholar] [CrossRef]
- Ribeiro, B.E.; Quintanilla, M.A. Transitions in biofuel technologies: An appraisal of the social impacts of cellulosic ethanol using the Delphi method. Technol. Forecast. Soc. Chang. 2015, 92, 53–68. [Google Scholar] [CrossRef]
- Wang, M.-Y.; Fang, S.-C.; Chang, Y.-H. Exploring technological opportunities by mining the gaps between science and technology: Microalgal biofuels. Technol. Forecast. Soc. Chang. 2015, 92, 182–195. [Google Scholar] [CrossRef]
- Anto, S.; Mukherjee, S.S.; Muthappa, R.; Mathimani, T.; Deviram, G.; Kumar, S.; Verma, T.N.; Pugazhendhi, A. Algae as green energy reserve: Technological outlook on biofuel production. Chemosphere 2019, 242, 125079. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.-H.P. A sweet out-of-the-box solution to the hydrogen economy: Is the sugar-powered car science fiction? Energy Environ. Sci. 2009, 2, 272–282. [Google Scholar] [CrossRef]
- Zhu, P.; Abdelaziz, O.; Hulteberg, C.P.; Riisager, A. New synthetic approaches to biofuels from lignocellulosic biomass. Curr. Opin. Green Sustain. Chem. 2020, 21, 16–21. [Google Scholar] [CrossRef]
- Trumbo, J.L.; Tonn, B.E. Biofuels: A sustainable choice for the United States’ energy future? Technol. Forecast. Soc. Chang. 2016, 104, 147–161. [Google Scholar] [CrossRef]
- Srivastava, N.; Srivastava, M.; Ramteke, P.W.; Mishra, P.K. Synthetic biology strategy for microbial cellulases. In New and Future Developments in Microbial Biotechnology and Bioengineering; Elsevier: Amsterdam, The Netherlands, 2019; pp. 229–238. [Google Scholar]
- Kalluri, U.C.; Yin, H.; Yang, X.; Davison, B.H. Systems and synthetic biology approaches to alter plant cell walls and reduce biomass recalcitrance. Plant Biotechnol. J. 2014, 12, 1207–1216. [Google Scholar] [CrossRef] [PubMed]
- Prasad, R.K.; Chatterjee, S.; Mazumder, P.B.; Gupta, S.K.; Sharma, S.; Vairale, M.G.; Datta, S.; Dwivedi, S.K.; Gupta, D.K. Bioethanol production from waste lignocelluloses: A review on microbial degradation potential. Chemosphere 2019, 231, 588–606. [Google Scholar] [CrossRef]
- Mukherji, S.; van Oudenaarden, A. Synthetic biology: Understanding biological design from synthetic circuits. Nat. Rev. Genet. 2009, 10, 859–871. [Google Scholar] [CrossRef]
- Shapira, P.; Kwon, S.; Youtie, J.L. Tracking the emergence of synthetic biology. Scientometrics 2020, 112, 1439–1469. [Google Scholar] [CrossRef]
- Porter, A.L.; Chiavetta, D.; Newman, N.C. Measuring tech emergence: A contest. Technol. Forecast. Soc. Chang. 2020, 159, 120176. [Google Scholar] [CrossRef]
- Benner, S.A.; Sismour, A.M. Synthetic biology. Nat. Rev. Genet. 2005, 6, 533–543. [Google Scholar] [CrossRef] [PubMed]
- Philp, J. Balancing the bioeconomy: Supporting biofuels and bio-based materials in public policy. Energy Environ. Sci. 2015, 8, 3063–3068. [Google Scholar] [CrossRef]
- Aro, E.-M. From first generation biofuels to advanced solar biofuels. Ambio 2016, 45, 24–31. [Google Scholar] [CrossRef]
- Ribeiro, B.; Shapira, P. Anticipating governance challenges in synthetic biology: Insights from biosynthetic menthol. Technol. Forecast. Soc. Chang. 2019, 139, 311–320. [Google Scholar] [CrossRef] [PubMed]
- Wehrs, M.; Tanjore, D.; Eng, T.; Lievense, J.; Pray, T.R.; Mukhopadhyay, A. Engineering robust production microbes for large-scale cultivation. Trends Microbiol. 2019, 27, 524–537. [Google Scholar] [CrossRef] [PubMed]
- Xu, N.; Wei, L.; Liu, J. Recent advances in the applications of promoter engineering for the optimization of metabolite biosynthesis. World J. Microbiol. Biotechnol. 2019, 35, 33. [Google Scholar] [CrossRef] [PubMed]
- Buschke, N.; Schäfer, R.; Becker, J.; Wittmann, C. Metabolic engineering of industrial platform microorganisms for biorefinery applications—Optimization of substrate spectrum and process robustness by rational and evolutive strategies. Bioresour. Technol. 2013, 135, 544–554. [Google Scholar] [CrossRef]
- Moysés, D.N.; Reis, V.C.B.; de Almeida, J.R.M.; de Moraes, L.M.P.; Torres, F.A.G. Xylose Fermentation by Saccharomyces cerevisiae: Challenges and Prospects. Int. J. Mol. Sci. 2016, 17, 207. [Google Scholar] [CrossRef]
- Lee, W.-H.; Jin, Y.-S. Evaluation of Ethanol Production Activity by Engineered Saccharomyces cerevisiae Fermenting Cellobiose through the Phosphorolytic Pathway in Simultaneous Saccharification and Fermentation of Cellulose. J. Microbiol. Biotechnol. 2017, 27, 1649–1656. [Google Scholar] [CrossRef]
- Bracher, J.M.; Verhoeven, M.; Wisselink, H.W.; Crimi, B.; Nijland, J.G.; Driessen, A.J.M.; Klaassen, P.; van Maris, A.J.A.; Daran, J.-M.; Pronk, J.T. The Penicillium chrysogenum transporter PcAraT enables high-affinity, glucose-insensitive l-arabinose transport in Saccharomyces cerevisiae. Biotechnol. Biofuels 2018, 11, 63. [Google Scholar] [CrossRef]
- Koppolu, V.; Vasigala, V.K. Role of Escherichia coli in Biofuel Production. Microbiol. Insights 2016, 9, MBI.S10878. [Google Scholar] [CrossRef] [PubMed]
- Sherkhanov, S.; Korman, T.P.; Clarke, S.G.; Bowie, J.U. Production of FAME biodiesel in E. coli by direct methylation with an insect enzyme. Sci. Rep. 2016, 6, 24239. [Google Scholar] [CrossRef] [PubMed]
- Wenning, L.; Ejsing, C.S.; David, F.; Sprenger, R.R.; Nielsen, J.; Siewers, V. Increasing jojoba-like wax ester production in Saccharomyces cerevisiae by enhancing very long-chain, monounsaturated fatty acid synthesis. Microb. Cell Factories 2019, 18, 49. [Google Scholar] [CrossRef] [PubMed]
- Bergman, A.; Vitay, D.; Hellgren, J.; Chen, Y.; Nielsen, J.; Siewers, V. Effects of overexpression of STB5 in Saccharomyces cerevisiae on fatty acid biosynthesis, physiology and transcriptome. FEMS Yeast Res. 2019, 19, foz027. [Google Scholar] [CrossRef]
- Jagadevan, S.; Banerjee, A.; Banerjee, C.; Guria, C.; Tiwari, R.; Baweja, M.; Shukla, P. Recent developments in synthetic biology and metabolic engineering in microalgae towards biofuel production. Biotechnol. Biofuels 2018, 11, 185. [Google Scholar] [CrossRef] [PubMed]
- Arora, N.; Tripathi, S.; Poluri, K.M.; Pruthi, V. Advanced gene technology and synthetic biology approaches to custom design microalgae for biodiesel production. In Microalgae Biotechnology for Development of Biofuel and Wastewater Treatment; Springer: Singapore, 2019; pp. 147–175. [Google Scholar] [CrossRef]
- Bellefleur, M.P.A.; Wanda, S.-Y.; Curtiss, R. Characterizing active transportation mechanisms for free fatty acids and antibiotics in Synechocystis sp. PCC 6803. BMC Biotechnol. 2019, 19, 5. [Google Scholar] [CrossRef]
- Markham, K.A.; Alper, H.S. Synthetic Biology Expands the Industrial Potential of Yarrowia lipolytica. Trends Biotechnol. 2018, 36, 1085–1095. [Google Scholar] [CrossRef]
- Chisti, Y. Biodiesel from microalgae beats bioethanol. Trends Biotechnol. 2008, 26, 126–131. [Google Scholar] [CrossRef]
- Davis, R.W.; Siccardi, A.J.; Huysman, N.D.; Wyatt, N.B.; Hewson, J.; Lane, T. Growth of mono- and mixed cultures of Nannochloropsis salina and Phaeodactylum tricornutum on struvite as a nutrient source. Bioresour. Technol. 2015, 198, 577–585. [Google Scholar] [CrossRef]
- Popko, J.; Herrfurth, C.; Feussner, K.; Ischebeck, T.; Iven, T.; Haslam, R.; Hamilton, M.; Sayanova, O.; Napier, J.; Khozin-Goldberg, I.; et al. Metabolome Analysis Reveals Betaine Lipids as Major Source for Triglyceride Formation, and the Accumulation of Sedoheptulose during Nitrogen-Starvation of Phaeodactylum tricornutum. PLoS ONE 2016, 11, e0164673. [Google Scholar] [CrossRef]
- Atsumi, S.; Higashide, W.; Liao, J.C. Direct photosynthetic recycling of carbon dioxide to isobutyraldehyde. Nat. Biotechnol. 2009, 27, 1177–1180. [Google Scholar] [CrossRef] [PubMed]
- Strong, P.J.; Xie, S.; Clarke, W. Methane as a Resource: Can the Methanotrophs Add Value? Environ. Sci. Technol. 2015, 49, 4001–4018. [Google Scholar] [CrossRef] [PubMed]
- Atsumi, S.; Cann, A.F.; Connor, M.R.; Shen, C.R.; Smith, K.M.; Brynildsen, M.P.; Chou, K.J.; Hanai, T.; Liao, J.C. Metabolic engineering of Escherichia coli for 1-butanol production. Metab. Eng. 2008, 10, 305–311. [Google Scholar] [CrossRef] [PubMed]
- Atsumi, S.; Hanai, T.; Liao, J.C. Non-fermentative pathways for synthesis of branched-chain higher alcohols as biofuels. Nature 2008, 451, 86–89. [Google Scholar] [CrossRef] [PubMed]
- Rude, M.A.; Schirmer, A. New microbial fuels: A biotech perspective. Curr. Opin. Microbiol. 2009, 12, 274–281. [Google Scholar] [CrossRef] [PubMed]
- Singh, B.K.; Trivedi, P. Microbiome and the future for food and nutrient security. Microb. Biotechnol. 2017, 10, 50–53. [Google Scholar] [CrossRef]
- Narnoliya, L.K.; Jadaun, J.S.; Singh, S.P. Management of agro-industrial wastes with the aid of synthetic biology. In Biosynthetic Technology and Environmental Challenges; Springer: Singapore, 2018; pp. 11–28. [Google Scholar] [CrossRef]
- Paul, S.; Dutta, A. Challenges and opportunities of lignocellulosic biomass for anaerobic digestion. Resour. Conserv. Recycl. 2018, 130, 164–174. [Google Scholar] [CrossRef]
- Dahmen, N.; Lewandowski, I.; Zibek, S.; Weidtmann, A. Integrated lignocellulosic value chains in a growing bioeconomy: Status quo and perspectives. GCB Bioenergy 2019, 11, 107–117. [Google Scholar] [CrossRef]
- Talamini, E.; Caldarelli, C.; Wubben, E.F.; Dewes, H. The composition and impact of stakeholders’ agendas on US ethanol production. Energy Policy 2012, 50, 647–658. [Google Scholar] [CrossRef]
- Talamini, E.; Wubben, E.F.; Padula, A.D.; Dewes, H. Scanning the macro-environment for liquid biofuels: A comparative analysis from public policies in Brazil, United States and Germany. J. Strat. Manag. 2013, 6, 40–60. [Google Scholar] [CrossRef]
- Gomes, J.; Dewes, H. Disciplinary dimensions and social relevance in the scientific communications on biofuels. Scientometrics 2017, 110, 1173–1189. [Google Scholar] [CrossRef]
- Befort, N. Going beyond definitions to understand tensions within the bioeconomy: The contribution of sociotechnical regimes to contested fields. Technol. Forecast. Soc. Chang. 2020, 153, 119923. [Google Scholar] [CrossRef]
- Clomburg, J.M.; Gonzalez, R. Biofuel production in Escherichia coli: The role of metabolic engineering and synthetic biology. Appl. Microbiol. Biotechnol. 2010, 86, 419–434. [Google Scholar] [CrossRef] [PubMed]
- Berla, B.M.; Esaha, R.; Immethun, C.M.; Maranas, C.D.; Emoon, T.S.; Pakrasi, H.B. Synthetic biology of cyanobacteria: Unique challenges and opportunities. Front. Microbiol. 2013, 4, 246. [Google Scholar] [CrossRef] [PubMed]
- Zilberman, D.; Kim, E.; Kirschner, S.; Kaplan, S.; Reeves, J. Technology and the future bioeconomy. Agric. Econ. 2013, 44, 95–102. [Google Scholar] [CrossRef]
- Jonsson, R.; Rinaldi, F.; Pilli, R.; Fiorese, G.; Hurmekoski, E.; Cazzaniga, N.; Robert, N.; Camia, A. Boosting the EU forest-based bioeconomy: Market, climate, and employment impacts. Technol. Forecast. Soc. Chang. 2021, 163, 120478. [Google Scholar] [CrossRef]
- Chen, C.; Leydesdorff, L. Patterns of connections and movements in dual-map overlays: A new method of publication portfolio analysis. J. Assoc. Inf. Sci. Technol. 2014, 65, 334–351. [Google Scholar] [CrossRef]
- Freeman, L.C. Centrality in social networks conceptual clarification. Soc. Netw. 1978, 1, 215–239. [Google Scholar] [CrossRef]
- Chen, C.M. CiteSpace II: Detecting and visualizing emerging trends and transient patterns in scientific literature. J. Am. Soc. Inf. Sci. Technol. 2006, 57, 359–377. [Google Scholar] [CrossRef]
- Chen, C.; Ibekwe-SanJuan, F.; Hou, J. The structure and dynamics of cocitation clusters: A multiple-perspective cocitation analysis. J. Am. Soc. Inf. Sci. Technol. 2010, 61, 1386–1409. [Google Scholar] [CrossRef]
- Liu, H.; Valdehuesa, K.N.G.; Ramos, K.R.M.; Nisola, G.M.; Lee, W.-K.; Chung, W.-J. L-arabonate and d-galactonate production by expressing a versatile sugar dehydrogenase in metabolically engineered Escherichia coli. Bioresour. Technol. 2014, 159, 455–459. [Google Scholar] [CrossRef]
- Valdehuesa, K.N.G.; Liu, H.; Ramos, K.R.M.; Park, S.J.; Nisola, G.M.; Lee, W.-K.; Chung, W.-J. Direct bioconversion of d-xylose to 1,2,4-butanetriol in an engineered Escherichia coli. Process Biochem. 2014, 49, 25–32. [Google Scholar] [CrossRef]
- Zhang, S.; Skerker, J.M.; Rutter, C.D.; Maurer, M.J.; Arkin, A.P.; Rao, C.V. Engineering Rhodosporidium toruloides for increased lipid production. Biotechnol. Bioeng. 2016, 113, 1056–1066. [Google Scholar] [CrossRef] [PubMed]
- Oh, E.J.; Skerker, J.M.; Kim, S.R.; Wei, N.; Turner, T.L.; Maurer, M.J.; Arkin, A.P.; Jin, Y.-S. Gene Amplification on Demand Accelerates Cellobiose Utilization in Engineered Saccharomyces cerevisiae. Appl. Environ. Microbiol. 2016, 82, 3631–3639. [Google Scholar] [CrossRef] [PubMed]
- Zhang, L.; King, E.; Luo, R.; Li, H. Development of a High-Throughput, In Vivo Selection Platform for NADPH-Dependent Reactions Based on Redox Balance Principles. ACS Synth. Biol. 2018, 7, 1715–1721. [Google Scholar] [CrossRef] [PubMed]
- Black, W.B.; King, E.; Wang, Y.; Jenic, A.; Rowley, A.T.; Seki, K.; Luo, R.; Li, H. Engineering a Coenzyme A Detour to Expand the Product Scope and Enhance the Selectivity of the Ehrlich Pathway. ACS Synth. Biol. 2018, 7, 2758–2764. [Google Scholar] [CrossRef]
- Black, W.B.; Zhang, L.; Kamoku, C.; Liao, J.C.; Li, H. Rearrangement of Coenzyme A-Acylated Carbon Chain Enables Synthesis of Isobutanol via a Novel Pathway in Ralstonia eutropha. ACS Synth. Biol. 2018, 7, 794–800. [Google Scholar] [CrossRef]
- Rydzak, T.; Garcia, D.; Stevenson, D.M.; Sladek, M.; Klingeman, D.M.; Holwerda, E.; Amador-Noguez, D.; Brown, S.D.; Guss, A.M. Deletion of Type I glutamine synthetase deregulates nitrogen metabolism and increases ethanol production in Clostridium thermocellum. Metab. Eng. 2017, 41, 182–191. [Google Scholar] [CrossRef]
- Biswas, R.; Zheng, T.; Olson, D.G.; Lynd, L.R.; Guss, A.M. Elimination of hydrogenase active site assembly blocks H2 production and increases ethanol yield in Clostridium thermocellum. Biotechnol. Biofuels 2015, 8, 8. [Google Scholar] [CrossRef]
- Alonso-Gutierrez, J.; Koma, D.; Hu, Q.; Yang, Y.; Chan, L.J.G.; Petzold, C.J.; Adams, P.D.; Vickers, C.E.; Nielsen, L.K.; Keasling, J.D.; et al. Toward industrial production of isoprenoids in Escherichia coli: Lessons learned from CRISPR-Cas9 based optimization of a chromosomally integrated mevalonate pathway. Biotechnol. Bioeng. 2018, 115, 1000–1013. [Google Scholar] [CrossRef]
- Brunk, E.; George, K.W.; Alonso-Gutierrez, J.; Thompson, M.; Baidoo, E.; Wang, G.; Petzold, C.; McCloskey, D.; Monk, J.M.; Yang, L.; et al. Characterizing Strain Variation in Engineered E. coli Using a Multi-Omics-Based Workflow. Cell Syst. 2016, 2, 335–346. [Google Scholar] [CrossRef] [PubMed]
- Scalcinati, G.; Partow, S.; Siewers, V.; Schalk, M.; Daviet, L.; Nielsen, J. Combined metabolic engineering of precursor and co-factor supply to increase α-santalene production by Saccharomyces cerevisiae. Microb. Cell Factories 2012, 11, 117. [Google Scholar] [CrossRef] [PubMed]
- Hammar, P.; Angermayr, S.A.; Sjostrom, S.L.; van der Meer, J.; Hellingwerf, K.J.; Hudson, E.P.; Joensson, H.N. Single-cell screening of photosynthetic growth and lactate production by cyanobacteria. Biotechnol. Biofuels 2015, 8, 8. [Google Scholar] [CrossRef] [PubMed]
- Partow, S.; Siewers, V.; Daviet, L.; Schalk, M.; Nielsen, J. Reconstruction and Evaluation of the Synthetic Bacterial MEP Pathway in Saccharomyces cerevisiae. PLoS ONE 2012, 7, e52498. [Google Scholar] [CrossRef]
- Yamada, R.; Wakita, K.; Mitsui, R.; Nishikawa, R.; Ogino, H. Efficient production of 2,3-butanediol by recombinant Saccharomyces cerevisiae through modulation of gene expression by cocktail δ-integration. Bioresour. Technol. 2017, 245, 1558–1566. [Google Scholar] [CrossRef] [PubMed]
- Yamada, R.; Wakita, K.; Mitsui, R.; Ogino, H. Enhanced d -lactic acid production by recombinant Saccharomyces cerevisiae following optimization of the global metabolic pathway. Biotechnol. Bioeng. 2017, 114, 2075–2084. [Google Scholar] [CrossRef] [PubMed]
- Hasunuma, T.; Kondo, A. Development of yeast cell factories for consolidated bioprocessing of lignocellulose to bioethanol through cell surface engineering. Biotechnol. Adv. 2012, 30, 1207–1218. [Google Scholar] [CrossRef]
- Sanda, T.; Hasunuma, T.; Matsuda, F.; Kondo, A. Repeated-batch fermentation of lignocellulosic hydrolysate to ethanol using a hybrid Saccharomyces cerevisiae strain metabolically engineered for tolerance to acetic and formic acids. Bioresour. Technol. 2011, 102, 7917–7924. [Google Scholar] [CrossRef]
- Balamurugan, S.; Wang, X.; Wang, H.-L.; An, C.-J.; Li, H.; Li, D.-W.; Yang, W.-D.; Liu, J.-S.; Li, H.-Y. Occurrence of plastidial triacylglycerol synthesis and the potential regulatory role of AGPAT in the model diatom Phaeodactylum tricornutum. Biotechnol. Biofuels 2017, 10, 97. [Google Scholar] [CrossRef]
- Yao, Y.; Lu, Y.; Peng, K.-T.; Huang, T.; Niu, Y.-F.; Xie, W.-H.; Yang, W.-D.; Liu, J.-S.; Li, H.-Y. Glycerol and neutral lipid production in the oleaginous marine diatom Phaeodactylum tricornutum promoted by overexpression of glycerol-3-phosphate dehydrogenase. Biotechnol. Biofuels 2014, 7, 110. [Google Scholar] [CrossRef]
- Ma, Y.-H.; Wang, X.; Niu, Y.-F.; Yang, Z.-K.; Zhang, M.-H.; Wang, Z.-M.; Yang, W.-D.; Liu, J.-S.; Li, H.-Y. Antisense knockdown of pyruvate dehydrogenase kinase promotes the neutral lipid accumulation in the diatom Phaeodactylum tricornutum. Microb. Cell Factories 2014, 13, 100. [Google Scholar] [CrossRef] [PubMed]
- Wang, M.; Liu, L.; Fan, L.; Tan, T. CRISPRi based system for enhancing 1-butanol production in engineered Klebsiella pneumoniae. Process Biochem. 2017, 56, 139–146. [Google Scholar] [CrossRef]
- Alonso-Gutierrez, J.; Kim, E.-M.; Batth, T.S.; Cho, N.; Hu, Q.; Chan, L.J.G.; Petzold, C.; Hillson, N.J.; Adams, P.; Keasling, J.; et al. Principal component analysis of proteomics (PCAP) as a tool to direct metabolic engineering. Metab. Eng. 2015, 28, 123–133. [Google Scholar] [CrossRef] [PubMed]
- Kleinberg, J. Bursty and Hierarchical Structure in Streams. Data Min. Knowl. Discov. 2003, 7, 373–397. [Google Scholar] [CrossRef]
- Peralta-Yahya, P.P.; Zhang, F.; del Cardayre, S.B.; Keasling, J.D. Microbial engineering for the production of advanced biofuels. Nature 2012, 488, 320–328. [Google Scholar] [CrossRef] [PubMed]
- Gibson, D.G.; Young, L.; Chuang, R.-Y.; Venter, J.C.; Hutchison, C.A., III; Smith, H.O. Enzymatic assembly of DNA molecules up to several hundred kilobases. Nat. Methods 2009, 6, 343–345. [Google Scholar] [CrossRef]
- Shen, C.R.; Lan, E.I.; Dekishima, Y.; Baez, A.; Cho, K.M.; Liao, J.C. Driving Forces Enable High-Titer Anaerobic 1-Butanol Synthesis in Escherichia coli. Appl. Environ. Microbiol. 2011, 77, 2905–2915. [Google Scholar] [CrossRef]
- Connor, M.R.; Atsumi, S. Synthetic Biology Guides Biofuel Production. J. Biomed. Biotechnol. 2010, 2010, 541698. [Google Scholar] [CrossRef]
- Tai, M.; Stephanopoulos, G. Engineering the push and pull of lipid biosynthesis in oleaginous yeast Yarrowia lipolytica for biofuel production. Metab. Eng. 2013, 15, 1–9. [Google Scholar] [CrossRef]
- Ruffing, A.M.; Chen, R.R. Metabolic engineering of Agrobacterium sp. strain ATCC 31749 for production of an α-Gal epitope. Microb. Cell Factories 2010, 9, 94. [Google Scholar] [CrossRef]
- Zhang, F.; Carothers, J.M.; Keasling, J.D. Design of a dynamic sensor-regulator system for production of chemicals and fuels derived from fatty acids. Nat. Biotechnol. 2012, 30, 354–359. [Google Scholar] [CrossRef] [PubMed]
- Runguphan, W.; Keasling, J.D. Metabolic engineering of Saccharomyces cerevisiae for production of fatty acid-derived biofuels and chemicals. Metab. Eng. 2014, 21, 103–113. [Google Scholar] [CrossRef] [PubMed]
- Pandey, S. Prospects of Metagenomic Cellulases for Converting Lignocellulosic Biomass into Bioethanol. J. Pure Appl. Microbiol. 2017, 11, 1079–1090. [Google Scholar] [CrossRef]
- Yu, A.; Zhao, Y.; Pang, Y.; Hu, Z.; Zhang, C.; Xiao, N.; Chang, M.W.; Leong, S.S.J. An oleaginous yeast platform for renewable 1-butanol synthesis based on a heterologous CoA-dependent pathway and an endogenous pathway. Microb. Cell Factories 2018, 17, 166. [Google Scholar] [CrossRef]
- Löbs, A.-K.; Schwartz, C.; Wheeldon, I. Genome and metabolic engineering in non-conventional yeasts: Current advances and applications. Synth. Syst. Biotechnol. 2017, 2, 198–207. [Google Scholar] [CrossRef]
- Hagen, L.H.; Frank, J.A.; Zamanzadeh, M.; Eijsink, V.G.H.; Pope, P.B.; Horn, S.J.; Arntzen, M. Quantitative Metaproteomics Highlight the Metabolic Contributions of Uncultured Phylotypes in a Thermophilic Anaerobic Digester. Appl. Environ. Microbiol. 2017, 83, e01955-16. [Google Scholar] [CrossRef]
- Jayakody, L.N.; Johnson, C.W.; Whitham, J.M.; Giannone, R.J.; Black, B.A.; Cleveland, N.S.; Klingeman, D.M.; Michener, W.E.; Olstad, J.L.; Vardon, D.R.; et al. Thermochemical wastewater valorization via enhanced microbial toxicity tolerance. Energy Environ. Sci. 2018, 11, 1625–1638. [Google Scholar] [CrossRef]
- Chen, Y.; Wang, Y.; Chen, T.-H.; Yao, M.-D.; Xiao, W.-H.; Li, B.-Z.; Yuan, Y.-J. Identification and manipulation of a novel locus to improve cell tolerance to short-chain alcohols in Escherichia coli. J. Ind. Microbiol. Biotechnol. 2018, 45, 589–598. [Google Scholar] [CrossRef]
- Bervoets, I.; van Brempt, M.; van Nerom, K.; van Hove, B.; Maertens, J.; de Mey, M.; Charlier, D. A sigma factor toolbox for orthogonal gene expression in Escherichia coli. Nucleic Acids Res. 2018, 46, 2133–2144. [Google Scholar] [CrossRef]
- Speda, J.; Jonsson, B.-H.; Carlsson, U.; Karlsson, M. Metaproteomics-guided selection of targeted enzymes for bioprospecting of mixed microbial communities. Biotechnol. Biofuels 2017, 10, 128. [Google Scholar] [CrossRef]
- Ehollinshead, W.; Ehe, L.; Tang, Y.J. Biofuel production: An odyssey from metabolic engineering to fermentation scale-up. Front. Microbiol. 2014, 5, 344. [Google Scholar] [CrossRef]
- Toor, M.; Kumar, S.; Malyan, S.K.; Bishnoi, N.R.; Mathimani, T.; Rajendran, K.; Pugazhendhi, A. An overview on bioethanol production from lignocellulosic feedstocks. Chemosphere 2020, 242, 125080. [Google Scholar] [CrossRef] [PubMed]
- Akinosho, H.; Eyee, K.; Close, D.; Eragauskas, A. The emergence of Clostridium thermocellum as a high utility candidate for consolidated bioprocessing applications. Front. Chem. 2014, 2, 66. [Google Scholar] [CrossRef] [PubMed]
- Ko, J.K.; Um, Y.; Woo, H.M.; Kim, K.H.; Lee, S.-M. Ethanol production from lignocellulosic hydrolysates using engineered Saccharomyces cerevisiae harboring xylose isomerase-based pathway. Bioresour. Technol. 2016, 209, 290–296. [Google Scholar] [CrossRef] [PubMed]
- Radecka, D.; Mukherjee, V.; Mateo, R.Q.; Stojiljkovic, M.; Foulquié-Moreno, M.R.; Thevelein, J.M. Looking beyond Saccharomyces: The potential of non-conventional yeast species for desirable traits in bioethanol fermentation. FEMS Yeast Res. 2015, 15, fov053. [Google Scholar] [CrossRef]
- Zhou, S.; Du, G.; Kang, Z.; Li, J.; Chen, J.; Li, H.; Zhou, J. The application of powerful promoters to enhance gene expression in industrial microorganisms. World J. Microbiol. Biotechnol. 2017, 33, 23. [Google Scholar] [CrossRef] [PubMed]
- Dasgupta, C.N.; Suseela, M.; Mandotra, S.; Kumar, P.; Pandey, M.K.; Toppo, K.; Lone, J. Dual uses of microalgal biomass: An integrative approach for biohydrogen and biodiesel production. Appl. Energy 2015, 146, 202–208. [Google Scholar] [CrossRef]
- Park, J.M.; Rathnasingh, C.; Song, H. Metabolic engineering of Klebsiella pneumoniae based on in silico analysis and its pilot-scale application for 1,3-propanediol and 2,3-butanediol co-production. J. Ind. Microbiol. Biotechnol. 2017, 44, 431–441. [Google Scholar] [CrossRef]
- Liu, C.; Zhang, K.; Cao, W.; Zhang, G.; Chen, G.; Yang, H.; Wang, Q.; Liu, H.; Xian, M.; Zhang, H. Genome mining of 2-phenylethanol biosynthetic genes from Enterobacter sp. CGMCC 5087 and heterologous overproduction in Escherichia coli. Biotechnol. Biofuels 2018, 11, 305. [Google Scholar] [CrossRef]
- Hwang, H.J.; Lee, S.Y.; Lee, P.C. Engineering and application of synthetic nar promoter for fine-tuning the expression of metabolic pathway genes in Escherichia coli. Biotechnol. Biofuels 2018, 11, 103. [Google Scholar] [CrossRef]
- Buhaescu, I.; Izzedine, H. Mevalonate pathway: A review of clinical and therapeutical implications. Clin. Biochem. 2007, 40, 575–584. [Google Scholar] [CrossRef] [PubMed]
- Guo, X.-W.; Zhang, Y.; Li, L.-L.; Guan, X.-Y.; Guo, J.; Wu, D.-G.; Chen, Y.-F.; Xiao, D.-G. Improved xylose tolerance and 2,3-butanediol production of Klebsiella pneumoniae by directed evolution of rpoD and the mechanisms revealed by transcriptomics. Biotechnol. Biofuels 2018, 11, 307. [Google Scholar] [CrossRef] [PubMed]
- Rhie, M.N.; Kim, H.T.; Jo, S.Y.; Chu, L.L.; Baritugo, K.A.; Baylon, M.G.; Lee, J.; Na, J.-G.; Kim, L.H.; Kim, T.W.; et al. Recent Advances in the Metabolic Engineering of Klebsiella pneumoniae: A Potential Platform Microorganism for Biorefineries. Biotechnol. Bioprocess Eng. 2019, 24, 48–64. [Google Scholar] [CrossRef]
- Allen, R.; Tilbrook, K.; Warden, A.; Campbell, P.C.; Rolland, V.; Singh, S.P.; Wood, C.C. Expression of 16 Nitrogenase Proteins within the Plant Mitochondrial Matrix. Front. Plant Sci. 2017, 8, 287. [Google Scholar] [CrossRef]
- Veerabadhran, M.; Natesan, S.; MubarakAli, D.; Xu, S.; Yang, F. Using different cultivation strategies and methods for the production of microalgal biomass as a raw material for the generation of bioproducts. Chemosphere 2021, 285, 131436. [Google Scholar] [CrossRef]
- Maeda, Y.; Yoshino, T.; Matsunaga, T.; Matsumoto, M.; Tanaka, T. Marine microalgae for production of biofuels and chemicals. Curr. Opin. Biotechnol. 2018, 50, 111–120. [Google Scholar] [CrossRef]
- Si, T.; Zhao, H. A brief overview of synthetic biology research programs and roadmap studies in the United States. Synth. Syst. Biotechnol. 2016, 1, 258–264. [Google Scholar] [CrossRef]
- Katz, L.; Chen, Y.Y.; Gonzalez, R.; Peterson, T.C.; Zhao, H.; Baltz, R.H. Synthetic biology advances and applications in the biotechnology industry: A perspective. J. Ind. Microbiol. Biotechnol. 2018, 45, 449–461. [Google Scholar] [CrossRef]
- Adler-Agnon, Z.; Leu, S.; Zarka, A.; Boussiba, S.; Khozin-Goldberg, I. Novel promoters for constitutive and inducible expression of transgenes in the diatom Phaeodactylum tricornutum under varied nitrate availability. J. Appl. Phycol. 2018, 30, 2763–2772. [Google Scholar] [CrossRef]
- Keasling, J.D. Synthetic biology and the development of tools for metabolic engineering. Metab. Eng. 2012, 14, 189–195. [Google Scholar] [CrossRef]
- Ekelwick, R.; MacDonald, J.; Webb, A.; Efreemont, P. Developments in the Tools and Methodologies of Synthetic Biology. Front. Bioeng. Biotechnol. 2014, 2, 60. [Google Scholar] [CrossRef]
- Paddon, C.J.; Keasling, J. Semi-synthetic artemisinin: A model for the use of synthetic biology in pharmaceutical development. Nat. Rev. Genet. 2014, 12, 355–367. [Google Scholar] [CrossRef] [PubMed]
- Zou, X.; Wang, L.; Li, Z.; Luo, J.; Wang, Y.; Deng, Z.; Du, S.; Chen, S. Genome Engineering and Modification Toward Synthetic Biology for the Production of Antibiotics. Med. Res. Rev. 2018, 38, 229–260. [Google Scholar] [CrossRef] [PubMed]
- Sengupta, A.; Pakrasi, H.B.; Wangikar, P.P. Recent advances in synthetic biology of cyanobacteria. Appl. Microbiol. Biotechnol. 2018, 102, 5457–5471. [Google Scholar] [CrossRef]
- Smanski, M.J.; Zhou, H.; Claesen, J.; Shen, B.; Fischbach, M.A.; Voigt, C.A. Synthetic biology to access and expand nature’s chemical diversity. Nat. Rev. Microbiol. 2016, 14, 135–149. [Google Scholar] [CrossRef]
- Bueso, Y.F.; Tangney, M. Synthetic Biology in the Driving Seat of the Bioeconomy. Trends Biotechnol. 2017, 35, 373–378. [Google Scholar] [CrossRef]
- Gronvall, G.K. US Competitiveness in Synthetic Biology. Health Secur. 2015, 13, 378–389. [Google Scholar] [CrossRef]
- Darvishi, F.; Ariana, M.; Marella, E.R.; Borodina, I. Advances in synthetic biology of oleaginous yeast Yarrowia lipolytica for producing non-native chemicals. Appl. Microbiol. Biotechnol. 2018, 102, 5925–5938. [Google Scholar] [CrossRef]
- Xu, G.; Wu, A.; Xiao, L.; Han, R.; Ni, Y. Enhancing butanol tolerance of Escherichia coli reveals hydrophobic interaction of multi-tasking chaperone SecB. Biotechnol. Biofuels 2019, 12, 164. [Google Scholar] [CrossRef]
- Cheng, F.; Tang, X.-L.; Kardashliev, T. Transcription Factor-Based Biosensors in High-Throughput Screening: Advances and Applications. Biotechnol. J. 2018, 13, e1700648. [Google Scholar] [CrossRef]
- Chen, Y.; Banerjee, D.; Mukhopadhyay, A.; Petzold, C.J. Systems and synthetic biology tools for advanced bioproduction hosts. Curr. Opin. Biotechnol. 2020, 64, 101–109. [Google Scholar] [CrossRef] [PubMed]
- Schwartz, C.M.; Hussain, M.S.; Blenner, M.; Wheeldon, I. Synthetic RNA polymerase III promoters facilitate high-efficiency CRISPR–Cas9-mediated genome editing in Yarrowia lipolytica. ACS Synth. Biol. 2016, 5, 356–359. [Google Scholar] [CrossRef] [PubMed]
- Gao, D.; Smith, S.; Spagnuolo, M.; Rodriguez, G.; Blenner, M. Dual CRISPR-Cas9 Cleavage Mediated Gene Excision and Targeted Integration in Yarrowia lipolytica. Biotechnol. J. 2018, 13, e1700590. [Google Scholar] [CrossRef] [PubMed]
- Santos-Merino, M.; Garcillán-Barcia, M.P.; de la Cruz, F. Engineering the fatty acid synthesis pathway in Synechococcus elongatus PCC 7942 improves omega-3 fatty acid production. Biotechnol. Biofuels 2018, 11, 239. [Google Scholar] [CrossRef]
- Maheshwari, N.V. Agro-industrial lignocellulosic waste: An alternative to unravel the future bioenergy. In Biofuels: Greenhouse Gas Mitigation and Global Warming; Springer: New Delhi, India, 2018; pp. 291–305. [Google Scholar] [CrossRef]
- González-García, S.; Gullón, P.; Gullón, B. Bio-compounds production from agri-food wastes under a biorefinery approach: Exploring environmental and social sustainability. In Quantification of Sustainability Indicators in the Food Sector; Springer: Singapore, 2019; pp. 25–53. [Google Scholar] [CrossRef]
- Siddiqui, M.R.; Miranda, A.; Mouradov, A. Microalgae as bio-converters of wastewater into biofuel and food. In Water Scarcity and Ways to Reduce the Impact; Springer International Publishing: Cham, Switzerland, 2019; pp. 75–94. [Google Scholar]
- Ko, J.K.; Lee, S.-M. Advances in cellulosic conversion to fuels: Engineering yeasts for cellulosic bioethanol and biodiesel production. Curr. Opin. Biotechnol. 2018, 50, 72–80. [Google Scholar] [CrossRef]
- Kim, N.M.; Sinnott, R.W.; Sandoval, N.R. Transcription factor-based biosensors and inducible systems in non-model bacteria: Current progress and future directions. Curr. Opin. Biotechnol. 2020, 64, 39–46. [Google Scholar] [CrossRef]
- Shi, T.-Q.; Huang, H.; Kerkhoven, E.J.; Ji, X.-J. Advancing metabolic engineering of Yarrowia lipolytica using the CRISPR/Cas system. Appl. Microbiol. Biotechnol. 2018, 102, 9541–9548. [Google Scholar] [CrossRef]
- Gujjala, L.K.S.; Kumar, S.P.J.; Talukdar, B.; Dash, A.; Sherpa, K.; Banerjee, R. Biodiesel from oleaginous microbes: Opportunities and challenges. Biofuels 2019, 10, 45–59. [Google Scholar] [CrossRef]
- Albers, S.C.; Berklund, A.M.; Graff, G. The rise and fall of innovation in biofuels. Nat. Biotechnol. 2016, 34, 814–821. [Google Scholar] [CrossRef]
- Goold, H.D.; Wright, P.; Hailstones, D. Emerging Opportunities for Synthetic Biology in Agriculture. Genes 2018, 9, 341. [Google Scholar] [CrossRef]
- Pixley, K.V.; Falck-Zepeda, J.B.; Giller, K.E.; Glenna, L.L.; Gould, F.; Mallory-Smith, C.A.; Stelly, D.M.; Stewart, C.N. Genome Editing, Gene Drives, and Synthetic Biology: Will They Contribute to Disease-Resistant Crops, and Who Will Benefit? Annu. Rev. Phytopathol. 2019, 57, 165–188. [Google Scholar] [CrossRef] [PubMed]
- Liu, G.; Gilding, E.K.; Kerr, E.D.; Schulz, B.L.; Tabet, B.; Hamaker, B.R.; Godwin, I.D. Increasing protein content and digestibility in sorghum grain with a synthetic biology approach. J. Cereal Sci. 2019, 85, 27–34. [Google Scholar] [CrossRef]
- Wurtzel, E.T.; Vickers, C.E.; Hanson, A.D.; Millar, A.H.; Cooper, M.; Voss-Fels, K.P.; Nikel, P.I.; Erb, T.J. Revolutionizing agriculture with synthetic biology. Nat. Plants 2019, 5, 1207–1210. [Google Scholar] [CrossRef] [PubMed]
- Tyagi, A.; Kumar, A.; Aparna, S.V.; Mallappa, R.H.; Grover, S.; Batish, V.K. Synthetic Biology: Applications in the Food Sector. Crit. Rev. Food Sci. Nutr. 2016, 56, 1777–1789. [Google Scholar] [CrossRef]
- Mortimer, J.C. Plant synthetic biology could drive a revolution in biofuels and medicine. Exp. Biol. Med. 2019, 244, 323–331. [Google Scholar] [CrossRef]
- Sekurova, O.N.; Schneider, O.; Zotchev, S.B. Novel bioactive natural products from bacteria via bioprospecting, genome mining and metabolic engineering. Microb. Biotechnol. 2019, 12, 828–844. [Google Scholar] [CrossRef]
- Aguilar, A.; Twardowski, T.; Wohlgemuth, R. Bioeconomy for Sustainable Development. Biotechnol. J. 2019, 14, e1800638. [Google Scholar] [CrossRef]
- Bengyella, L.; Iftikhar, S.; Nawaz, K.; Fonmboh, D.J.; Yekwa, E.L.; Jones, R.C.; Njanu, Y.M.T.; Roy, P. Biotechnological application of endophytic filamentous bipolaris and curvularia: A review on bioeconomy impact. World J. Microbiol. Biotechnol. 2019, 35, 69. [Google Scholar] [CrossRef]
- Zabed, H.M.; Akter, S.; Yun, J.; Zhang, G.; Awad, F.; Qi, X.; Sahu, J. Recent advances in biological pretreatment of microalgae and lignocellulosic biomass for biofuel production. Renew. Sustain. Energy Rev. 2019, 105, 105–128. [Google Scholar] [CrossRef]
- Havlík, P.; Schneider, U.; Schmid, E.; Böttcher, H.; Fritz, S.; Skalský, R.; Aoki, K.; de Cara, S.; Kindermann, G.E.; Kraxner, F.; et al. Global land-use implications of first and second generation biofuel targets. Energy Policy 2011, 39, 5690–5702. [Google Scholar] [CrossRef]
- Wallington, T.J.; Anderson, J.E.; de Kleine, R.D.; Kim, H.C.; Maas, H.; Brandt, A.R.; Keoleian, G.A. When Comparing Alternative Fuel-Vehicle Systems, Life Cycle Assessment Studies Should Consider Trends in Oil Production. J. Ind. Ecol. 2017, 21, 244–248. [Google Scholar] [CrossRef]
- Synnestvedt, M.B.; Chen, C.; Holmes, J.H. CiteSpace II: Visualization and knowledge discovery in bibliographic databases. AMIA Annu. Symp. Proc. AMIA Symp. 2005, 2005, 724–728. [Google Scholar]
- Yu, X.; Zhang, B. Obtaining advantages from technology revolution: A patent roadmap for competition analysis and strategy planning. Technol. Forecast. Soc. Chang. 2019, 145, 273–283. [Google Scholar] [CrossRef]
- Daim, T.; Lai, K.K.; Yalcin, H.; Alsoubie, F.; Kumar, V. Forecasting technological positioning through technology knowledge redundancy: Patent citation analysis of IoT, cybersecurity, and Blockchain. Technol. Forecast. Soc. Chang. 2020, 161, 120329. [Google Scholar] [CrossRef]
- Huang, D.; Jin, X.; Coghlan, A. Advances in consumer innovation resistance research: A review and research agenda. Technol. Forecast. Soc. Chang. 2021, 166, 120594. [Google Scholar] [CrossRef]
- Mogoutov, A.; Kahane, B. Data search strategy for science and technology emergence: A scalable and evolutionary query for nanotechnology tracking. Res. Policy 2007, 36, 893–903. [Google Scholar] [CrossRef]
- Jarboe, L.R.; Zhang, X.; Wang, X.; Moore, J.C.; Shanmugam, K.T.; Ingram, L.O. Metabolic Engineering for Production of Biorenewable Fuels and Chemicals: Contributions of Synthetic Biology. J. Biomed. Biotechnol. 2010, 2010, 761042. [Google Scholar] [CrossRef]
- Nielsen, J.; Keasling, J.D. Synergies between synthetic biology and metabolic engineering. Nat. Biotechnol. 2011, 29, 693–695. [Google Scholar] [CrossRef]
- Jullesson, D.; David, F.; Pfleger, B.; Nielsen, J. Impact of synthetic biology and metabolic engineering on industrial production of fine chemicals. Biotechnol. Adv. 2015, 33, 1395–1402. [Google Scholar] [CrossRef]
- Raimbault, B.; Cointet, J.-P.; Joly, P.-B. Mapping the Emergence of Synthetic Biology. PLoS ONE 2016, 11, e0161522. [Google Scholar] [CrossRef]
- Yeung, A.W.K.; Tzvetkov, N.T.; Gupta, V.K.; Gupta, S.C.; Orive, G.; Bonn, G.K.; Fiebich, B.; Bishayee, A.; Efferth, T.; Xiao, J.; et al. Current research in biotechnology: Exploring the biotech forefront. Curr. Res. Biotechnol. 2019, 1, 34–40. [Google Scholar] [CrossRef]
- Hou, J.; Yang, X.; Chen, C. Emerging trends and new developments in information science: A document co-citation analysis (2009–2016). Scientometrics 2018, 115, 869–892. [Google Scholar] [CrossRef]
- Guangfen, Y.; Dongke, Z. An Analysis Based on Citespace III Knowledge Maps of Chinese Vocational Education Research. Chin. Educ. Soc. 2017, 50, 499–519. [Google Scholar] [CrossRef]
- Ma, S.; Yu, X.; Chen, Q. Hotspots Analysis and Its Applications in Vocational Education with CiteSpace. In Proceedings of the 2019 10th International Conference on Information Technology in Medicine and Education (ITME), Qingdao, China, 23–25 August 2019; IEEE: Piscataway, NJ, USA; pp. 394–398. [Google Scholar]
- Rousseeuw, P.J. Silhouettes: A graphical aid to the interpretation and validation of cluster analysis. J. Comput. Appl. Math. 1987, 20, 53–65. [Google Scholar] [CrossRef]

| Cluster | Size | Silhouette | Label (LLR) | Year | Citation Coverage |
|---|---|---|---|---|---|
| #0 | 41 | 1 | Versatile Sugar Dehydrogenase (219.06, 1.0 × 10−4); D-Galactonate Production (219.06, 1.0 × 10−4); Engineered Saccharomyces cerevisiae (204.24, 1.0 × 10−4) | 2014 | [64,65,66,67] |
| #1 | 34 | 0.911 | Redox Balance Principle (197.02, 1.0 × 10−4); Nadph-Dependent Reaction (197.02, 1.0 × 10−4); Vivo Selection Platform (197.02, 1.0 × 10−4) | 2014 | [68,69,70,71,72] |
| #2 | 29 | 0.946 | Sesquiterpene Production (313.39, 1.0 × 10−4); Principal Component Analysis (209.89, 1.0 × 10−4); Characterizing Strain Variation (189.08, 1.0 × 10−4); | 2014 | [73,74] |
| #3 | 25 | 0.977 | Saccharomyces cerevisiae (360.26, 1.0 × 10−4); Co-Factor Supply (207.54, 1.0 × 10−4); Alpha-Santalene Production (207.54, 1.0 × 10−4) | 2014 | [75,76,77] |
| #4 | 24 | 0.984 | Recombinant xylose-fermenting strain (191.79, 1.0 × 10−4); Metabolic Pathway Engineering (191.79, 1.0 × 10−4); Formic Acid Tolerance (191.79, 1.0 × 10−4) | 2014 | [78,79,80,81] |
| #7 | 16 | 0.984 | Clostridium saccharoperbutylacetonicum n1-4 (275.82, 1.0 × 10−4); Model Diatom Phaeodactylum tricornutum (197.67, 1.0 × 10−4); Potential Regulatory Role (197.67, 1.0 × 10−4) | 2015 | [82,83,84,85] |
| Articles | Year | Strength | Begin | End | 1999–2018 |
|---|---|---|---|---|---|
| Peralta-Yahya et al. [88] | 2012 | 371.738 | 2013 | 2018 |  |
| Atsumi et al. [46] | 2008 | 367.477 | 2010 | 2016 |  |
| Gibson et al. [89] | 2009 | 270.418 | 2013 | 2018 |  |
| Atsumi et al. [43] | 2008 | 269.481 | 2010 | 2014 |  |
| Shen et al. [90] | 2011 | 264.006 | 2012 | 2018 |  |
| Connor and Atsumi [91] | 2009 | 256.165 | 2011 | 2016 |  |
| Tai and Stephanopoulos [92] | 2013 | 216.846 | 2014 | 2018 |  |
| Dellomonaco et al. [93] | 2011 | 214.931 | 2012 | 2018 |  |
| Zhang et al. [94] | 2012 | 212.798 | 2013 | 2018 |  |
| Runguphan and Keasling [95] | 2014 | 210.905 | 2015 | 2018 |  |
| Classes | Keywords | Strength | Year | Begin | End | 1999–2018 |
|---|---|---|---|---|---|---|
| Organisms | Yarrowia lipolytica | 160.346 | 1999 | 2016 | 2018 |  |
| Oleaginous yeast | 82.659 | 1999 | 2016 | 2018 |  | |
| E. coli | 54.287 | 1999 | 2014 | 2018 |  | |
| Klebsiella pneumoniae | 54.065 | 1999 | 2016 | 2018 |  | |
| Phaeodactylum tricornutum | 51.463 | 1999 | 2016 | 2018 |  | |
| Microalgae | 45.048 | 1999 | 2016 | 2018 |  | |
| Processes and products | Anaerobic digestion | 83.171 | 1999 | 2016 | 2018 |  |
| Advanced biofuel | 82.808 | 1999 | 2014 | 2018 |  | |
| Bioethanol | 82.265 | 1999 | 2015 | 2018 |  | |
| Lignocellulosic biomass | 62.484 | 1999 | 2015 | 2018 |  | |
| Synthetic promoters | 58.377 | 1999 | 2015 | 2018 |  | |
| Biorefinery | 43.827 | 1999 | 2016 | 2018 |  | |
| Genetic sequencing | 43.701 | 1999 | 2016 | 2018 |  | |
| Nitrogen starvation | 39.342 | 1999 | 2015 | 2018 |  | |
| Wastewater | 38.613 | 1999 | 2016 | 2018 |  | |
| Mevalonate pathways | 38.257 | 1999 | 2016 | 2018 |  | |
| Lipids accumulation | 33.134 | 1999 | 2016 | 2018 |  | |
| Biotechnology | 31.641 | 1999 | 2016 | 2018 |  | |
| N-butanol | 5.198 | 1999 | 2016 | 2018 |  | |
| Transcription factor | 4.045 | 1999 | 2015 | 2018 |  |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Fantinel, A.L.; Margis, R.; Talamini, E.; Dewes, H. Trends in Synthetic Biology in the Bioeconomy of Non-Food-Competing Biofuels. SynBio 2023, 1, 33-53. https://doi.org/10.3390/synbio1010003
Fantinel AL, Margis R, Talamini E, Dewes H. Trends in Synthetic Biology in the Bioeconomy of Non-Food-Competing Biofuels. SynBio. 2023; 1(1):33-53. https://doi.org/10.3390/synbio1010003
Chicago/Turabian StyleFantinel, Antônio Luiz, Rogério Margis, Edson Talamini, and Homero Dewes. 2023. "Trends in Synthetic Biology in the Bioeconomy of Non-Food-Competing Biofuels" SynBio 1, no. 1: 33-53. https://doi.org/10.3390/synbio1010003
APA StyleFantinel, A. L., Margis, R., Talamini, E., & Dewes, H. (2023). Trends in Synthetic Biology in the Bioeconomy of Non-Food-Competing Biofuels. SynBio, 1(1), 33-53. https://doi.org/10.3390/synbio1010003








