Efficient Parameter Estimation for Oscillatory Biochemical Reaction Networks via a Genetic Algorithm with Adaptive Simulation Termination
Abstract
1. Introduction
2. Proposed Framework
2.1. Problem Formulation
2.2. Log-Transformed Parameterization
2.3. Global Exploration Using a GA
- Offspring generation (AREX): In each generation, offspring are generated in -space using AREX with parents selected from . Each offspring is assigned a fitness value, evaluated as using either full-horizon simulation (baseline) or adaptive termination (proposed; Section 2.5).
- Generational replacement (JGG with partial replacement): In JGG, only a subset of parents is replaced in each generation. The parents selected for AREX are treated as replacement targets. After evaluating all offspring, we collect the subset of valid offspring for which the simulation is completed over the full horizon, and defineWe then perform partial replacement by: (i) sorting the target parents by fitness (worst first); (ii) sorting by fitness (best first); and (iii) replacing the worst r target parents with the best r valid offspring to form .
2.4. Local Refinement Using the Modified Powell Method
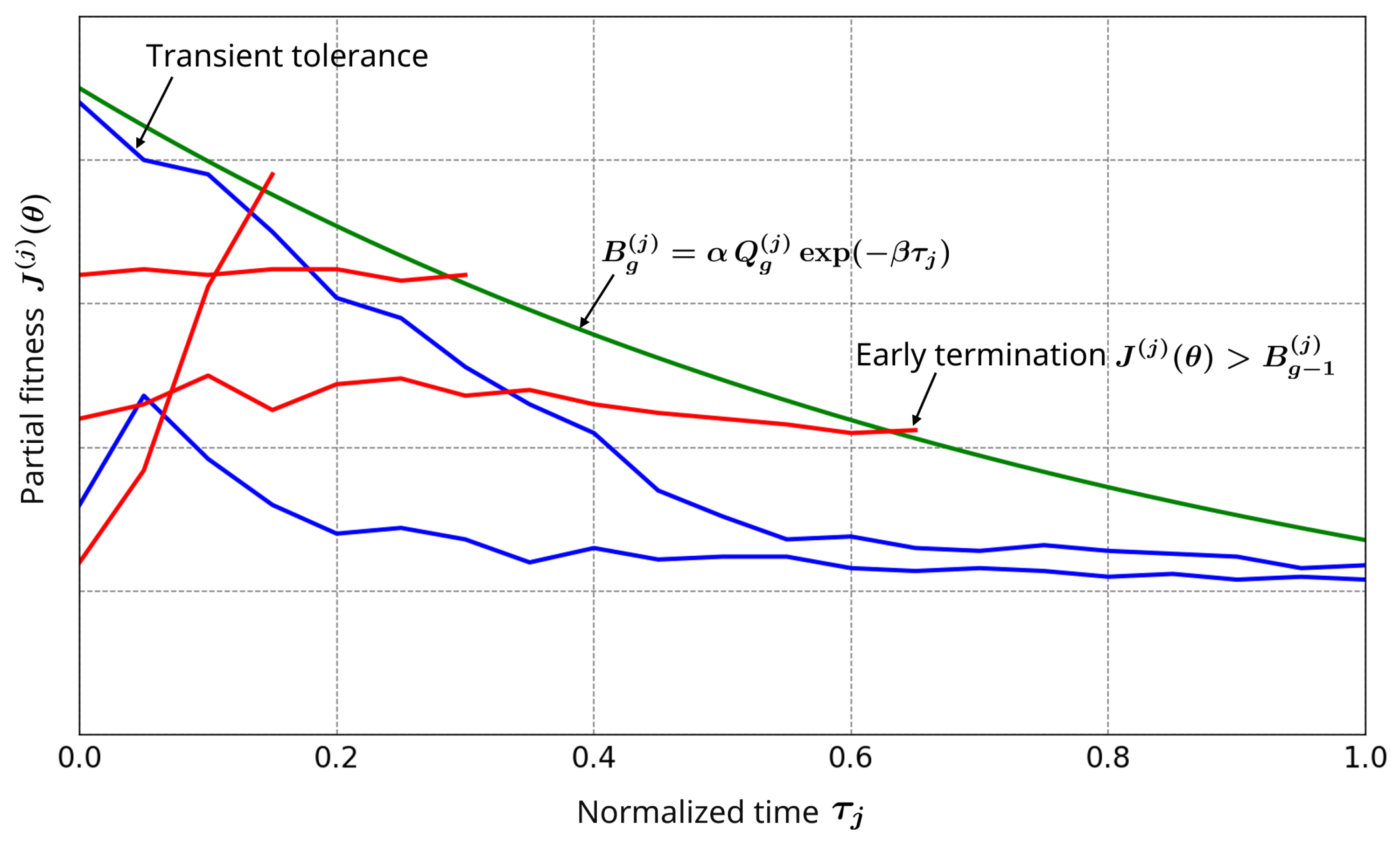
2.5. Partial Objective for Adaptive Simulation Termination
2.6. Quantile-Based Reference and Updating Strategy
2.6.1. Trade-Offs and Strategies for Oscillatory Systems
- Early Phases (Transient): The population distribution is broad, i.e., partial fitness values have a wide range, yielding permissive thresholds that tolerate transient deviations.
- Late Phases (Stabilized): The population distribution becomes more concentrated, leading to stricter thresholds.
2.6.2. Population-Based Update with Generation Lag
3. Numerical Experiments
3.1. Test Models and Synthetic Data
- (a)
- Lotka–Volterra (fully observed)
- (b)
- Goodwin oscillator (partially observed)
3.2. Compared Frameworks and Settings
- Baseline (GA + Powell; full-horizon simulation): A real-coded GA explores the log-transformed parameter space, followed by local refinement using the modified Powell method. Each candidate is evaluated with the full-horizon objective (Equation (3)).
- GA settings: The population size is ; offspring are generated per generation using AREX with parents. The GA runs for generations with JGG partial replacement (Equation (7)). In both models, each kinetic parameter is constrained to lie between and 10 times its ground truth value . All GA operations are performed in the log-transformed parameter space. The penalized fitness for rejected candidates is set to .
- Powell method settings: After the GA terminates, the modified Powell method is applied, starting from the top GA individuals ranked by the full objective . Each Powell run stops when improvements fall below a specified tolerance or when a maximum number of objective evaluations is reached; these stopping criteria are the same for both frameworks. During Powell refinement, the adaptive termination boundary is held fixed at the final GA boundary .
- Adaptive termination settings: Unless otherwise stated, we use quantile level , tolerance factor , and decay rate in Equations (9)–(12) (Section 3.6.5 and Section 4.2). The initial population is evaluated without adaptive termination to obtain and (Equations (9) and (10)). Offspring in generation g are tested against with a one-generation lag (Equation (11)); if no valid offspring are obtained, the references are retained (Equation (12)).
3.3. Evaluation Metrics
3.4. Implementation
3.4.1. Time-Sequential Evaluation Without Restarting from
3.4.2. Computational Cost Accounting
3.5. Summary of Experimental Settings
3.6. Results
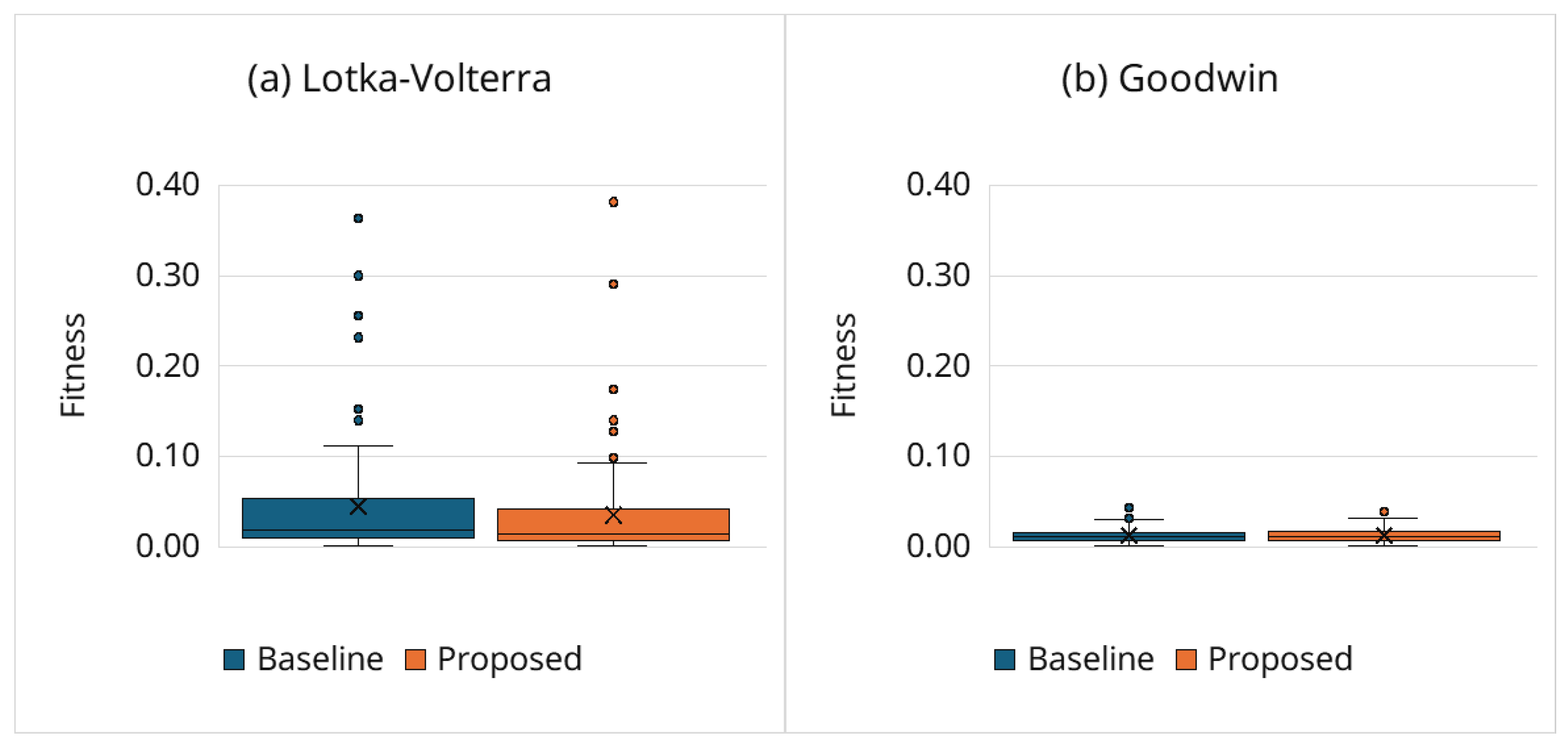
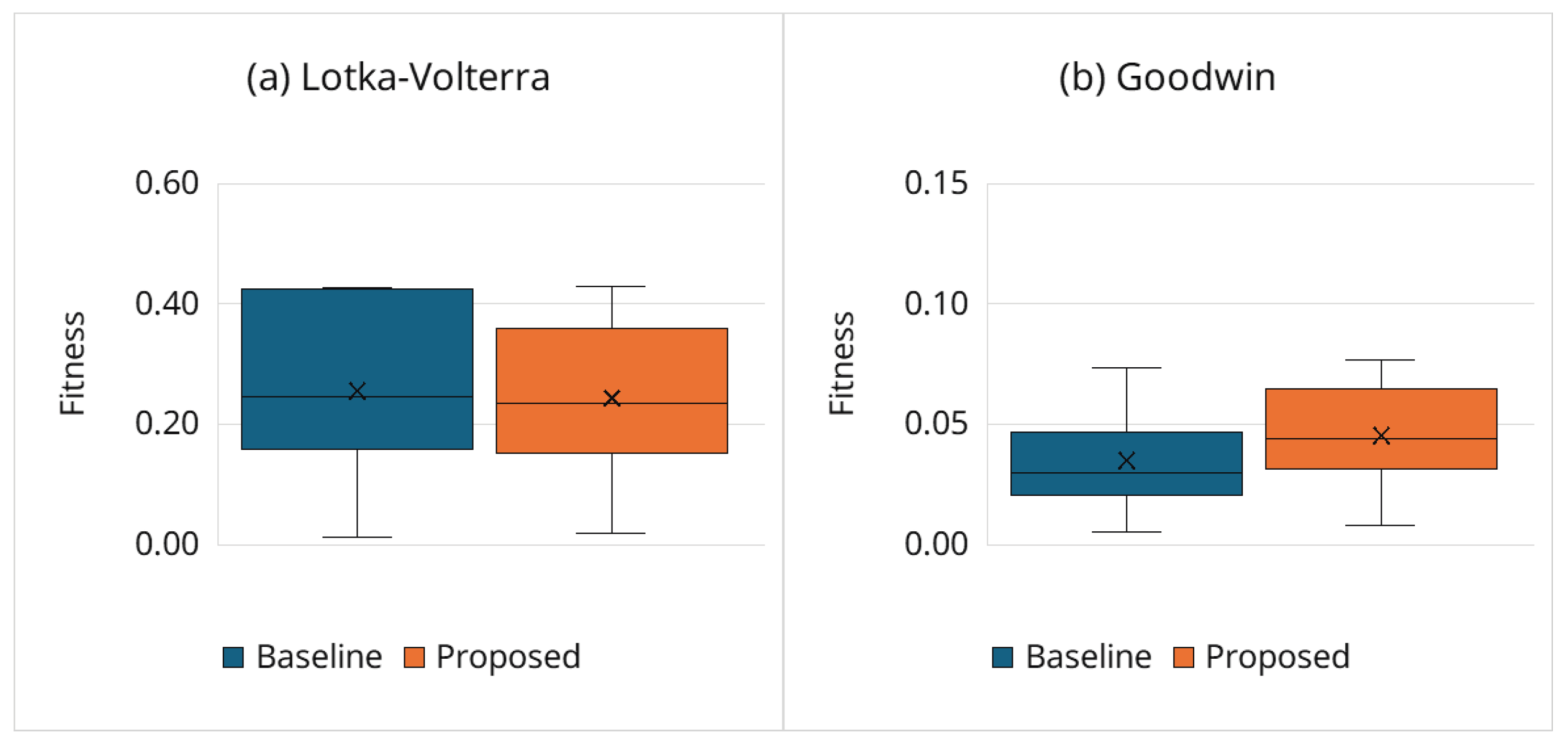
3.6.1. Estimation Accuracy
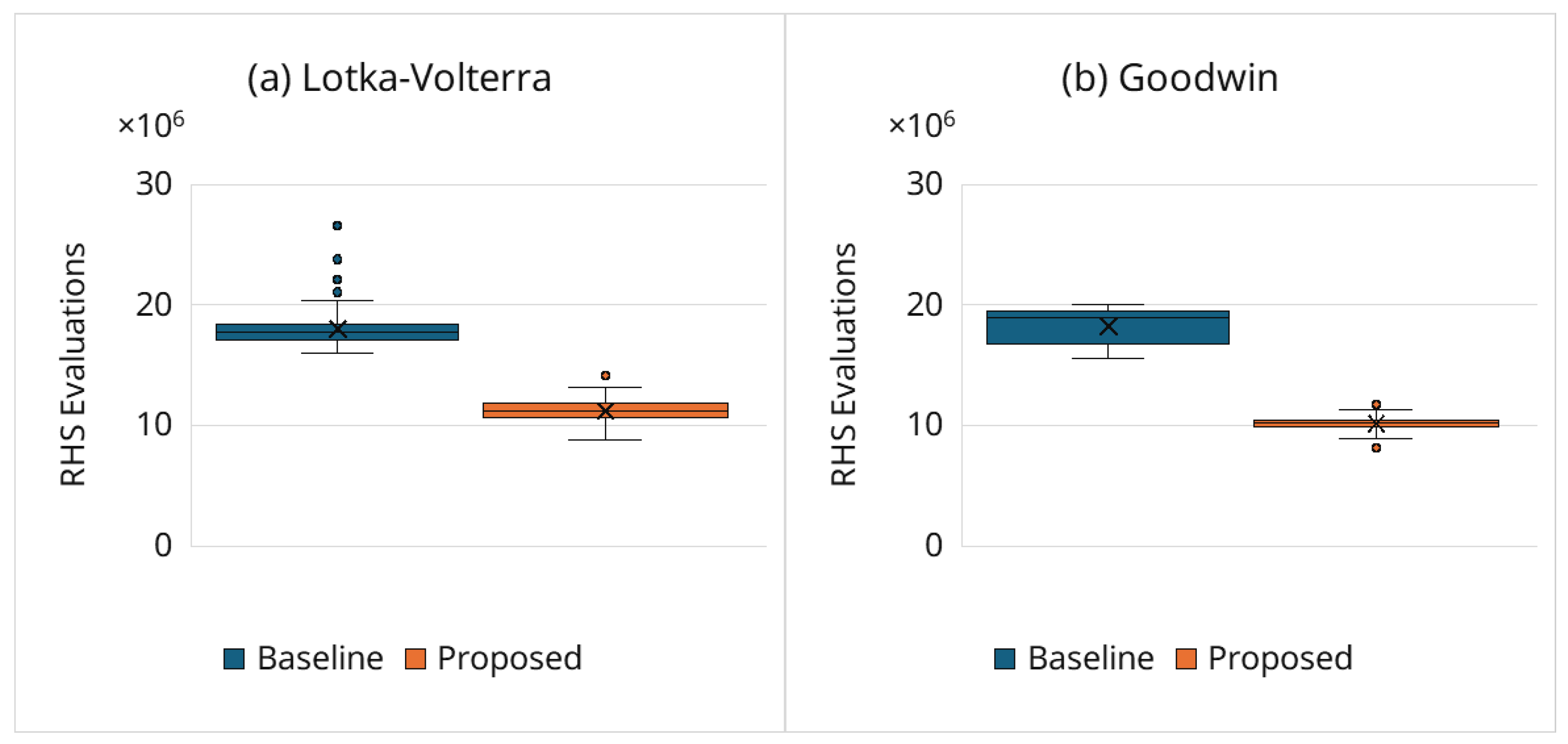
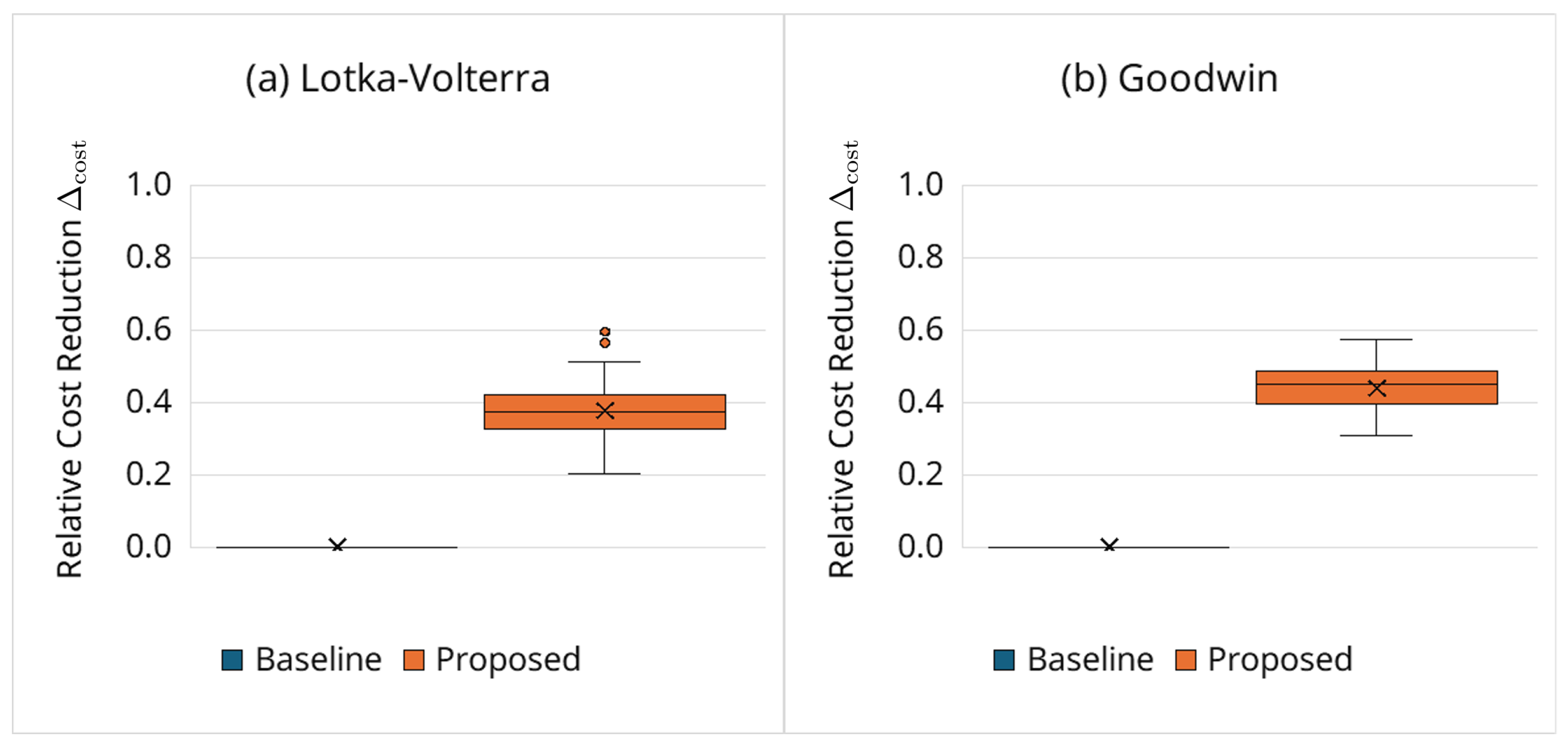
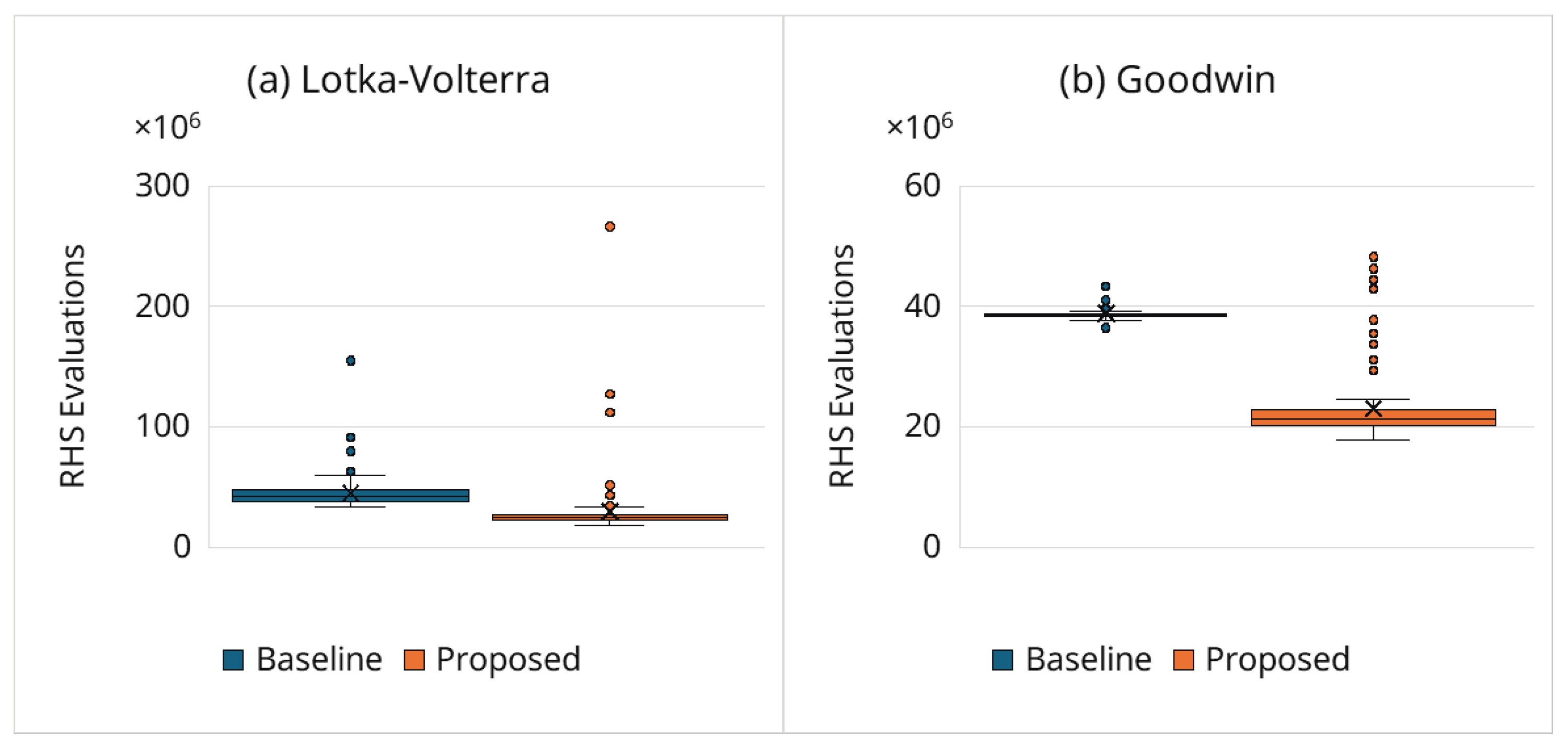
3.6.2. Computational Cost Reduction
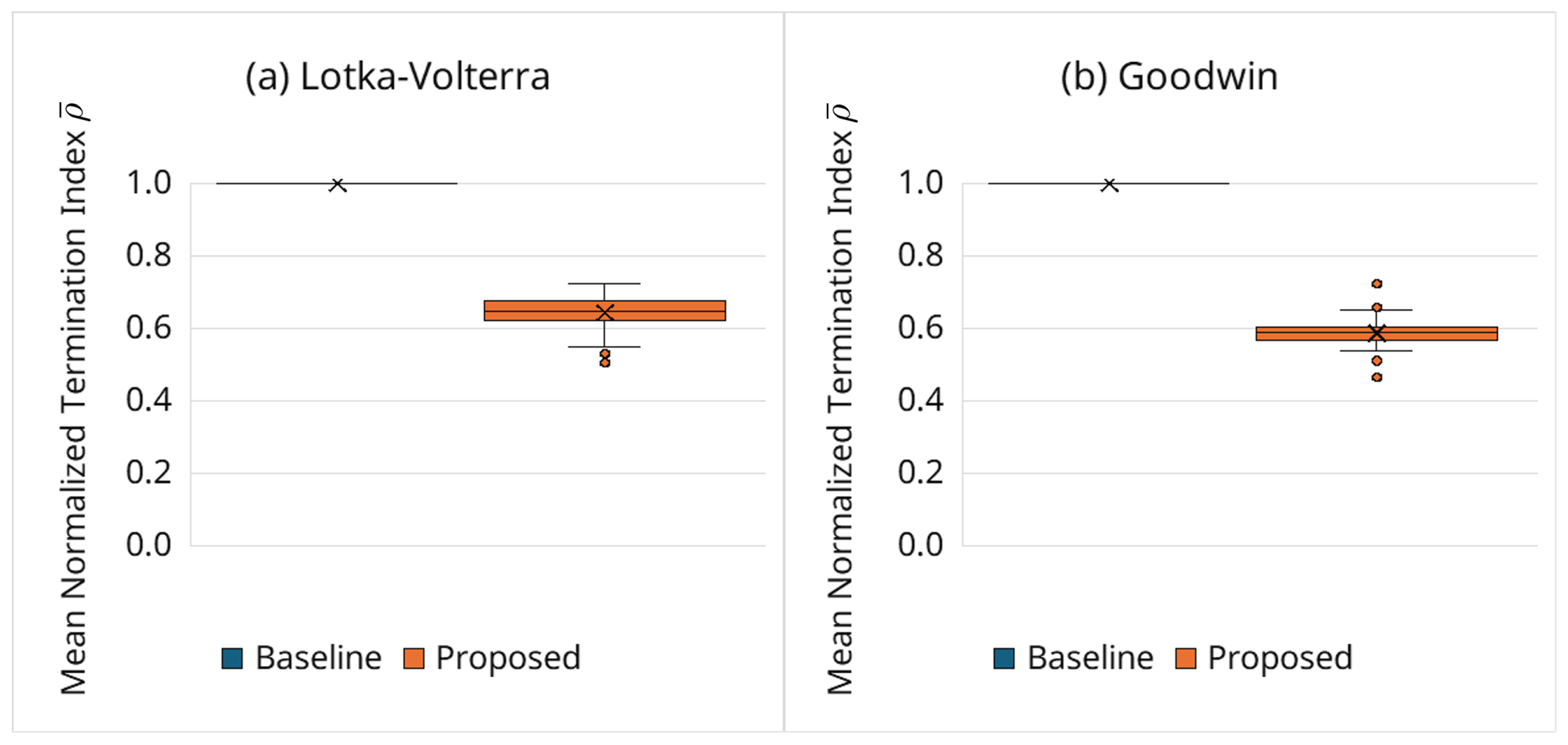
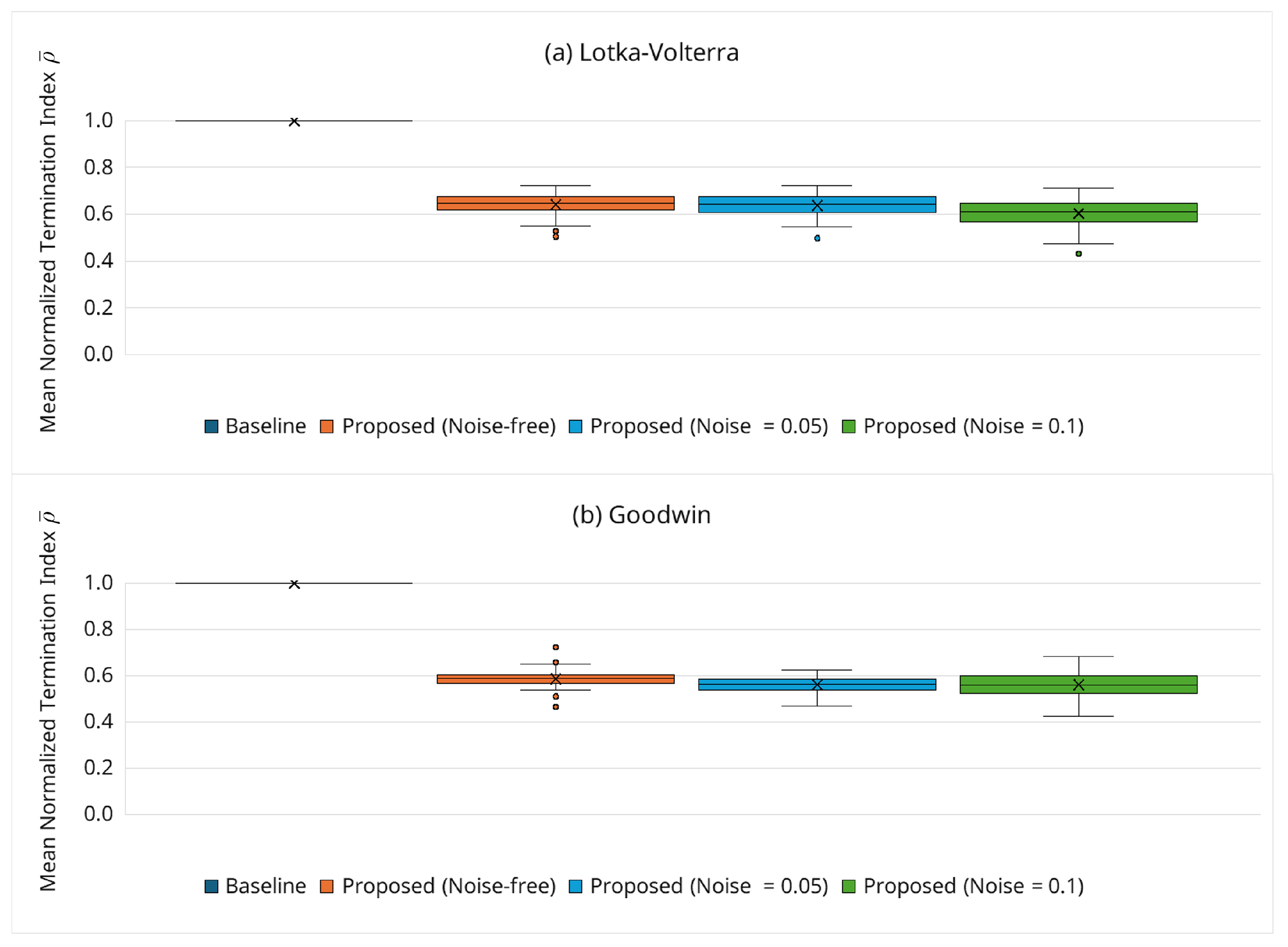
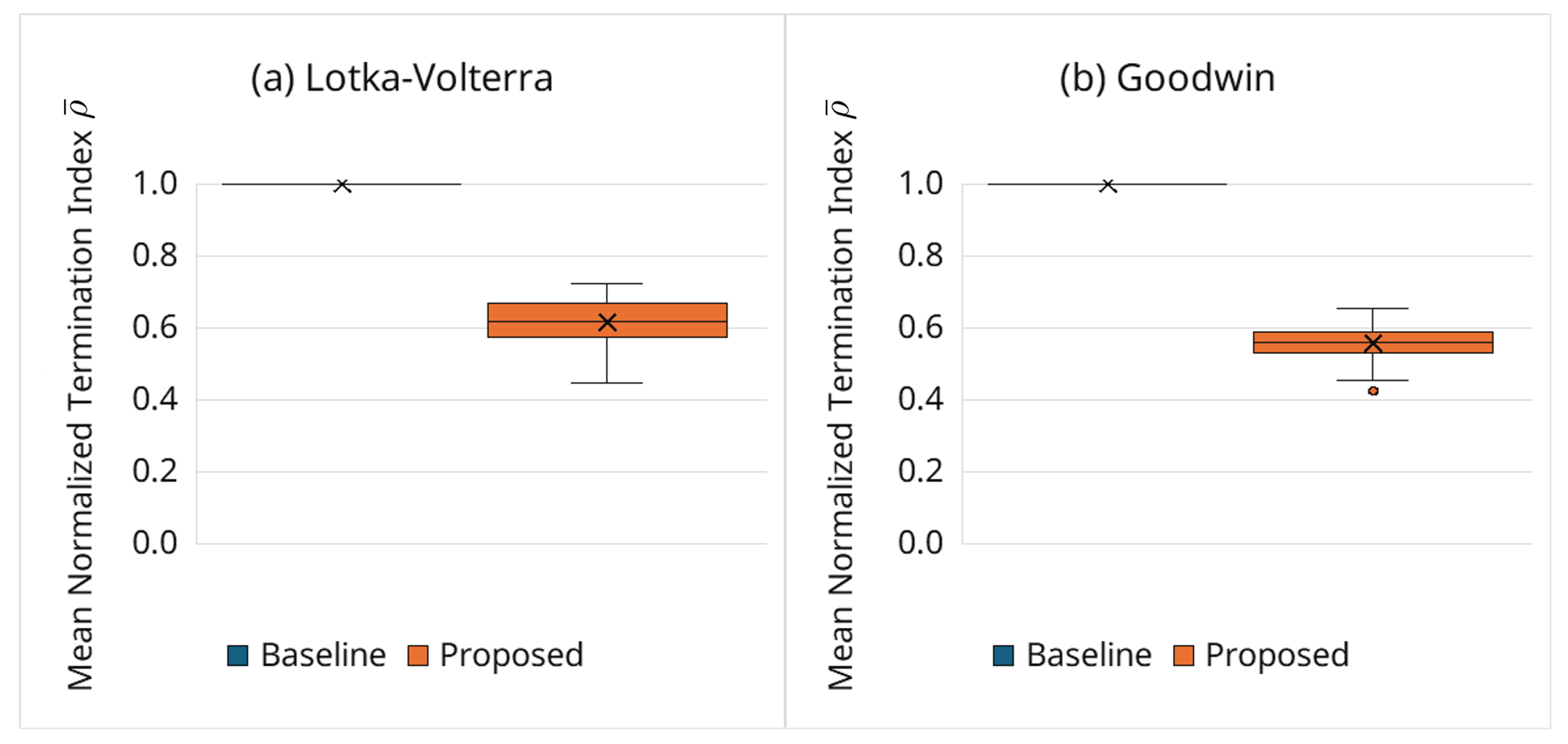
3.6.3. Behavior of Adaptive Termination via the Normalized Index
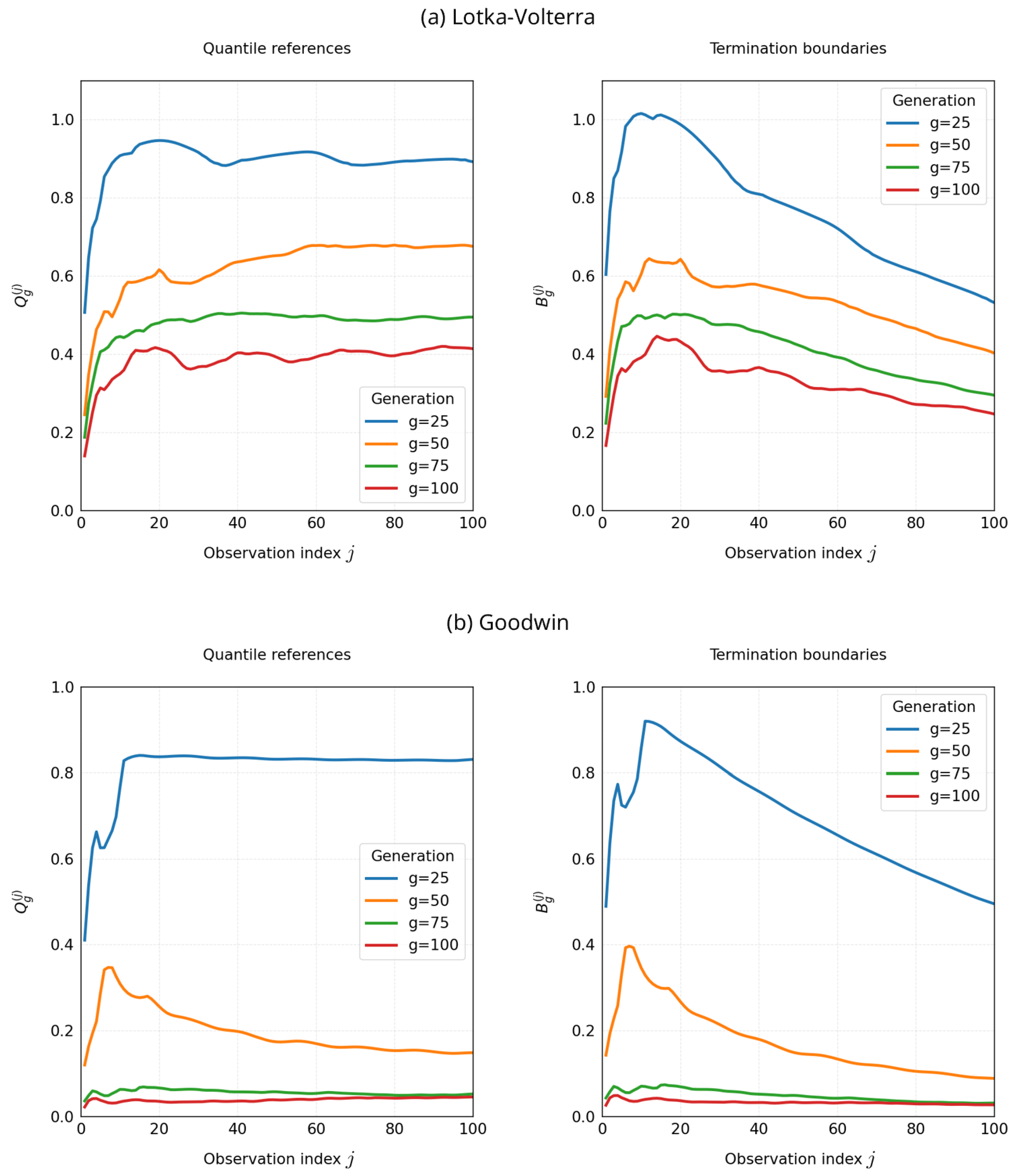
3.6.4. Evolution of Quantile References and Termination Boundaries
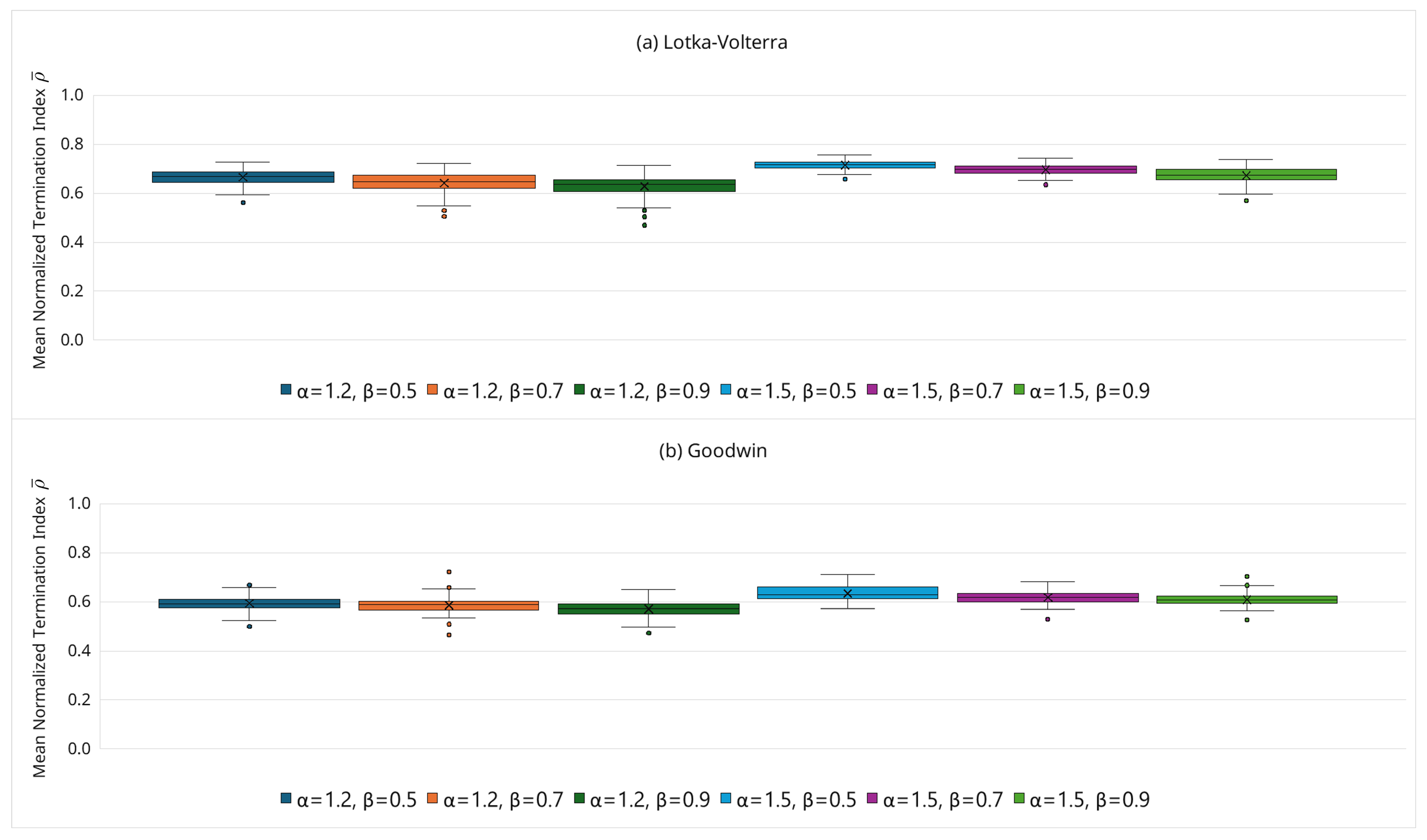
3.6.5. Sensitivity Analysis of Hyperparameters
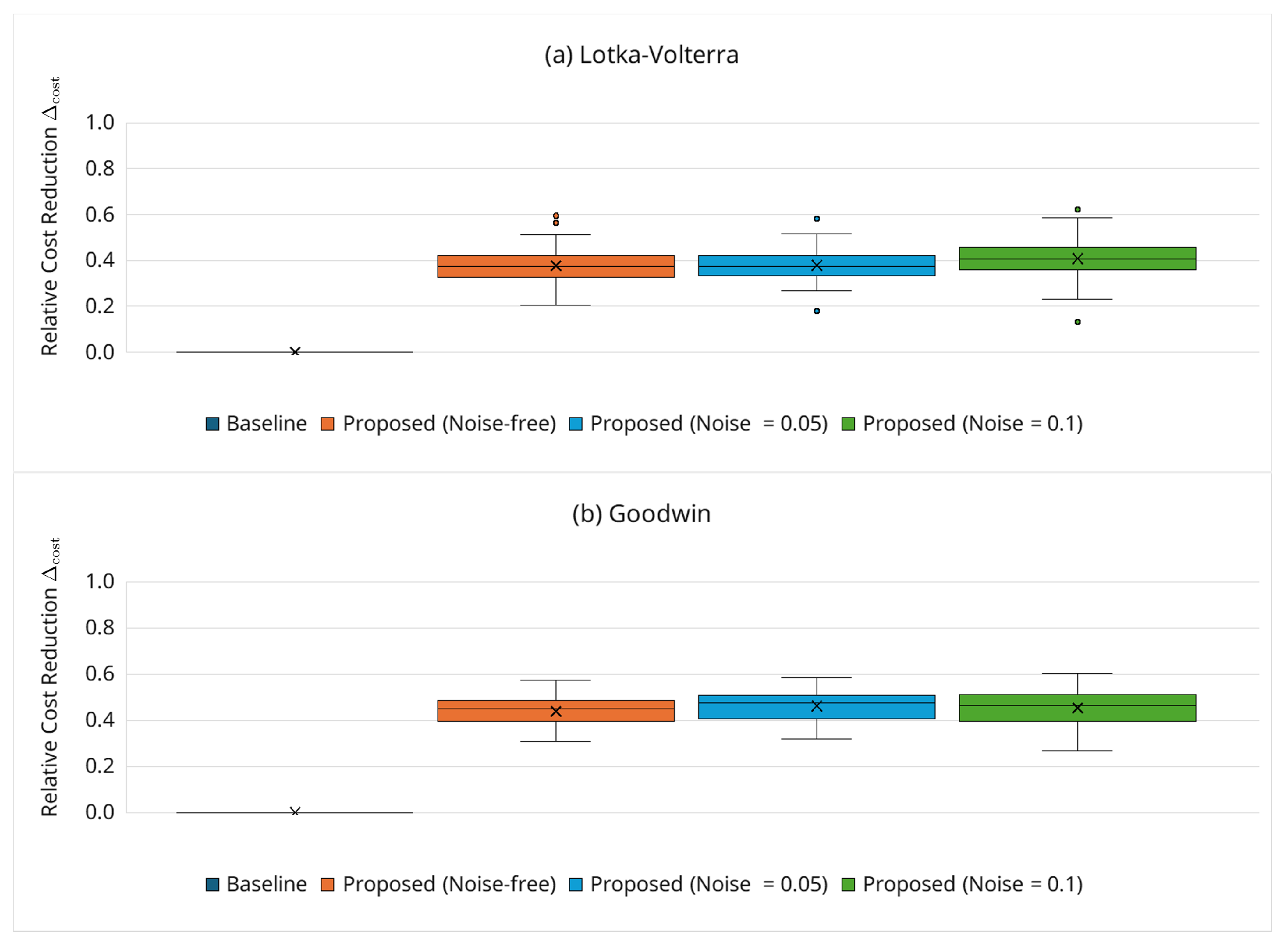
3.6.6. Robustness Under Noisy Observations
3.6.7. Effect of Broader Search Ranges
3.6.8. Effect of a Grace Period
3.6.9. Ability to Distinguish Damped Oscillations
- Early termination rate by n: Figure 22a shows the early termination rate, which is 56.1% total. This demonstrates the proposed framework’s ability to distinguish damped oscillations. Not all such candidates are eliminated immediately; instead, they are gradually replaced across GA generations through the JGG partial replacement scheme, which reinjects fitter offspring. A non-zero termination rate is also observed for , indicating that even when n satisfies the oscillatory condition, mismatched kinetic parameters can still produce decay or large discrepancies that are filtered out by the adaptive boundary.
- When termination occurs: Figure 22b reports the mean among the terminated candidates for each n. For , termination occurs earlier in the horizon (small ); for and 8, termination tends to occur later (larger ), consistent with gradually decaying trajectories that initially appear plausible but only reveal decay in the second half of the observation window. This behavior aligns with the time-adaptive boundary, which is permissive early on and becomes stricter later (Equation (10)), exactly where it is designed to act.
- Best solution n at the end of estimation: Table 3 lists the Hill coefficient of the best final solution over the 100 trials for both the baseline and proposed frameworks. In both cases, most of the best solutions satisfy (91% for the baseline; 96% for the proposed), indicating that both frameworks reliably identify oscillatory regimes. However, the proposed framework achieves substantially lower computational cost (e.g., the median is reduced from 25.5 million to 16.6 million RHS evaluations; ; Section 3.6.2). The few best solutions with suggest that some kinetic parameter combinations can transiently mimic sustained oscillations within ; conclusively resolving such borderline cases would require longer horizons. Even in these cases, the adaptive simulation termination strategy can reject many damped candidates before a full-length simulation, reducing the computational cost while maintaining accuracy.
4. Discussion
4.1. Effectiveness of the Adaptive Simulation Termination Strategy
4.2. Limitations and Practical Guidance
- Observation design and horizon length: For partially observed systems (e.g., the Goodwin model with only, as in Section 3.1 (b)), the objective function may be less constrained, and borderline cases can appear plausible within limited simulation horizons. When borderline regimes (e.g., Hill coefficients and 8 in the Goodwin model, as shown in Section 3.6.9) are of particular interest, using a longer simulation horizon or multiple complementary initial conditions can improve discrimination. When transients dominate most of the observation horizon (e.g., short horizons or strongly underdamped regimes), the partial objective may remain weakly informative until late times; in such cases, using a longer horizon and/or multiple initial conditions improves robustness of the termination decision.
- Hyperparameter selection (, ): The sensitivity study (Figure 8) demonstrates stable behavior for and . As a default, we recommend (moderately permissive early) and (moderate tightening). If false negatives are suspected (over pruning), can be increased slightly or tightening slowed (decrease ); if cost reduction is the priority and accuracy is stable, can be decreased or tightening accelerated (increase ). In addition, our noisy observation experiments (Section 3.6.6) indicate that, for the tested moderate noise levels ( and ), substantial cost reduction was retained without substantially widening the boundary (i.e., using the same default and settings). For stronger noise levels or different noise models, however, retuning the boundary parameters may be necessary, and the cost–accuracy trade-off may change.
- Stiff or large-scale ODE systems: For very stiff systems or large networks, step size control and event handling may interact with the time-sequential evaluation. In such cases, it is advisable to set the maximum step size conservatively to ensure the stability of the numerical integration (Section 3.4). This interaction becomes more apparent when broader search domains are used; as shown in Section 3.6.7 (experiments for broader search range), some trials enter numerically difficult regions where the solver workload increases substantially, highlighting the importance of conservative step size control and, if needed, additional solver/termination tuning.
4.3. Future Work
5. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Kitano, H. Systems biology: A brief overview. Science 2002, 295, 1662–1664. [Google Scholar] [CrossRef] [PubMed]
- Kikuchi, S.; Tominaga, D.; Arita, M.; Takahashi, K.; Tomita, M. Dynamic modeling of genetic networks using genetic algorithm and S-system. Bioinformatics 2003, 19, 643–650. [Google Scholar] [CrossRef]
- Crampin, E.J.; Schnell, S.; McSharry, P.E. Mathematical and computational techniques to deduce complex biochemical reaction mechanisms. Prog. Biophys. Mol. Biol. 2004, 86, 77–112. [Google Scholar] [CrossRef]
- Fernandez, M.R.; Mendes, P.; Banga, J.R. A hybrid approach for efficient and robust parameter estimation in biochemical pathways. BioSystems 2006, 83, 248–265. [Google Scholar] [CrossRef]
- Tangherloni, A.; Spolaor, S.; Cazzaniga, P.; Besozzi, D.; Rundo, L.; Mauri, G.; Nobile, M.S. Biochemical parameter estimation vs. benchmark functions: A comparative study of optimization performance and representation design. Appl. Soft Comput. 2019, 81, 105494. [Google Scholar] [CrossRef]
- El Yazidi, Y.; Ellabib, A. Reconstruction of the Depletion Layer in MOSFET by Genetic Algorithms. Math. Model. Comput. 2020, 7, 96–103. [Google Scholar] [CrossRef]
- Zaki, G.; Akbarfam, A.J.; Irandoust-Pakchin, S. On the Inverse Problem of Time Fractional Heat Equation Using Crank-Nicholson Type Method and Genetic Algorithm. Filomat 2024, 38, 6829–6849. [Google Scholar] [CrossRef]
- Okamoto, M.; Nonaka, T.; Ochiai, S.; Tominaga, D. Nonlinear numerical optimization with use of a hybrid Genetic Algorithm incorporating the Modified Powell method. Appl. Math. Comput. 1998, 91, 63–72. [Google Scholar] [CrossRef]
- Akimoto, Y.; Nagata, Y.; Sakuma, J.; Ono, I.; Kobayashi, S. Analysis of The Behavior of MGG and JGG As A Selection Model for Real-coded Genetic Algorithms. Trans. Jpn. Soc. Artif. Intell. 2010, 25, 281–289. [Google Scholar] [CrossRef]
- Akimoto, Y.; Nagata, Y.; Sakuma, J.; Ono, I.; Kobayashi, S. Proposal and Evaluation of Adaptive Real-coded Crossover AREX. Trans. Jpn. Soc. Artif. Intell. 2009, 24, 446–458. [Google Scholar] [CrossRef]
- Komori, A.; Maki, Y.; Nakatsui, M.; Ono, I.; Okamoto, M. Efficient Numerical Optimization Algorithm Based on New Real-Coded Genetic Algorithm, AREX + JGG, and Application to the Inverse Problem in Systems Biology. Appl. Math. 2012, 3, 1463–1470. [Google Scholar] [CrossRef]
- Goodwin, B.C. Oscillatory behavior in enzymatic control processes. Adv. Enzyme Regul. 1965, 3, 425–438. [Google Scholar] [CrossRef] [PubMed]
- Higgins, J. The Theory of Oscillating Reactions. Ind. Eng. Chem. 1967, 59, 18–62. [Google Scholar] [CrossRef]
- Griffith, J.S. Mathematics of cellular control processes I. Negative feedback to one gene. J. Theor. Biol. 1968, 20, 202–208. [Google Scholar] [CrossRef] [PubMed]
- Gonze, D.; Abou-Jaoudé, W. The Goodwin Model: Behind the Hill Function. PLoS ONE 2013, 8, e69573. [Google Scholar] [CrossRef]
- Gonze, D.; Ruoff, P. The Goodwin Oscillator and its Legacy. Acta Biotheor. 2021, 69, 857–874. [Google Scholar] [CrossRef]
- Powell, M.J.D. An efficient method for finding the minimum of a function of several variables without calculating derivatives. Comput. J. 1964, 7, 155–162. [Google Scholar] [CrossRef]
- Lotka, A.J. Analytical Note on Certain Rhythmic Relations in Organic Systems. Proc. Natl. Acad. Sci. USA 1920, 6, 410–415. [Google Scholar] [CrossRef]
- Volterra, V. Fluctuations in the Abundance of a Species considered Mathematically. Nature 1926, 118, 558–560. [Google Scholar] [CrossRef]
- Koza, J.R.; Mydlowec, W.; Lanza, G.; Yu, J.; Keane, A.M. Reverse engineering of metabolic pathways from observed data using genetic programming. Pac. Symp. Biocomput. 2001, 434–445. [Google Scholar] [CrossRef]
- Sugimoto, N.; Sakamoto, E.; Iba, H. Inference of Differential Equations by Using Genetic Programming. J. Jpn. Soc. Artif. Intell. 2004, 19, 450–459. [Google Scholar] [CrossRef]
- Sugimoto, M.; Kikuchi, S.; Tomita, M. Reverse engineering of biochemical equations from time-course data by means of genetic programming. BioSystems 2005, 80, 155–164. [Google Scholar] [CrossRef] [PubMed]
- Iba, H. Inference of differential equation models by genetic programming. Inf. Sci. 2008, 178, 4453–4468. [Google Scholar] [CrossRef]
- Sekiguchi, T.; Hamada, H.; Okamoto, M. Inference of General Mass Action-Based State Equations for Oscillatory Biochemical Reaction Systems Using k-Step Genetic Programming. Appl. Math. 2019, 10, 627–645. [Google Scholar] [CrossRef]










| Aspect | Early Update | Delayed Update |
|---|---|---|
| Computational efficiency | High | Moderate |
| Risk of false negatives | High | Low |
| Preservation of solution diversity | Low | High |
| Category | Name | Symbol/Value | Notes |
|---|---|---|---|
| Genetic algorithm | Population size | Real-coded GA in log space (Equation (6)) | |
| Offspring | Generated via AREX | ||
| Parents | JGG partial replacement (Equation (7)) | ||
| Generations | Maximum generation | ||
| Parameter bounds | Kinetic parameter bounded relative to ground truth | ||
| Powell method | Function | SciPy minimize | Maximum number of iterations is set to 200 |
| Starts | Initialized from top-ranked GA solutions | ||
| Termination boundary | Fixed (no updates during Powell) | ||
| Adaptive termination | Quantile level | For (Equation (9)) | |
| Tolerance factor | Controls the tolerance boundary (Equation (10)) | ||
| Decay rate | Controls time-dependent strictness (Equation (10)) | ||
| Warm-up | — | evaluated without adaptive termination | |
| Boundary lag | 1 generation | Evaluate at g using (Equation (11)) | |
| Reference retention rule | — | If no valid offspring, retain , (Equation (12)) | |
| Rejection penalty | For ranking; excluded from quantile computation | ||
| Objective | Stabilization constant | Treatment of zero/near-zero measurements (Equation (4)) | |
| Solver | Integrator | SciPy RK45 | Explicit Runge–Kutta; shared across frameworks |
| Tolerances | rtol , atol | Fixed across runs | |
| Maximum step | Equal to the sampling interval (Equation (17)) | ||
| Data | Sampling grid | 100 points over | Per condition; excludes |
| Observables (Lotka–Volterra) | Fully observed | ||
| Observables (Goodwin) | Partially observed (single variable) |
| Hill Coefficient n | 1–6 | 7 | 8 | 9 | 10 | 11 | 12 |
|---|---|---|---|---|---|---|---|
| Baseline | 0 | 1 | 8 | 42 | 44 | 5 | 0 |
| Proposed | 0 | 0 | 4 | 42 | 45 | 9 | 0 |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2026 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license.
Share and Cite
Sekiguchi, T.; Hamada, H.; Okamoto, M. Efficient Parameter Estimation for Oscillatory Biochemical Reaction Networks via a Genetic Algorithm with Adaptive Simulation Termination. AppliedMath 2026, 6, 47. https://doi.org/10.3390/appliedmath6030047
Sekiguchi T, Hamada H, Okamoto M. Efficient Parameter Estimation for Oscillatory Biochemical Reaction Networks via a Genetic Algorithm with Adaptive Simulation Termination. AppliedMath. 2026; 6(3):47. https://doi.org/10.3390/appliedmath6030047
Chicago/Turabian StyleSekiguchi, Tatsuya, Hiroyuki Hamada, and Masahiro Okamoto. 2026. "Efficient Parameter Estimation for Oscillatory Biochemical Reaction Networks via a Genetic Algorithm with Adaptive Simulation Termination" AppliedMath 6, no. 3: 47. https://doi.org/10.3390/appliedmath6030047
APA StyleSekiguchi, T., Hamada, H., & Okamoto, M. (2026). Efficient Parameter Estimation for Oscillatory Biochemical Reaction Networks via a Genetic Algorithm with Adaptive Simulation Termination. AppliedMath, 6(3), 47. https://doi.org/10.3390/appliedmath6030047





