Appendix B. Experimental Results
This appendix presents the detailed experimental results for the 18 benchmark functions used in this study. For each benchmark problem, the performance of all compared algorithms is summarized in a dedicated table. Each table reports the average, maximum, and minimum of the best objective values found in 30 trials, as well as the standard deviation and the average computation time. In all tables, the performance values of all algorithms are compared, and the best value in each row is highlighted in bold.
Tables in this appendix are labeled as
Table A2,
Table A3,
Table A4,
Table A5,
Table A6,
Table A7,
Table A8,
Table A9,
Table A10,
Table A11,
Table A12,
Table A13,
Table A14,
Table A15,
Table A16,
Table A17,
Table A18 and
Table A19 corresponding to each benchmark function in the order listed in
Appendix A.
Table A2.
Experimental results on F1, the sphere function, for , 30, 50, and 100 (30 trials).
Table A2.
Experimental results on F1, the sphere function, for , 30, 50, and 100 (30 trials).
| Metric | PSO | PSO-C | ELPSO | ELPSO-C | GA | ACOR |
|---|
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Variance | | | | | | |
| Time [s] | 49.28 | 148.16 | 23.61 | 70.57 | 14.23 | 638.44 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Variance | | | | | | |
| Time [s] | 67.17 | 315.60 | 36.16 | 155.27 | 21.18 | 1766.12 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Variance | | | | | | |
| Time [s] | 90.78 | 484.99 | 53.79 | 233.88 | 27.85 | 2779.22 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Variance | | | | | | |
| Time [s] | 125.35 | 891.54 | 102.50 | 427.83 | 39.80 | 5894.17 |
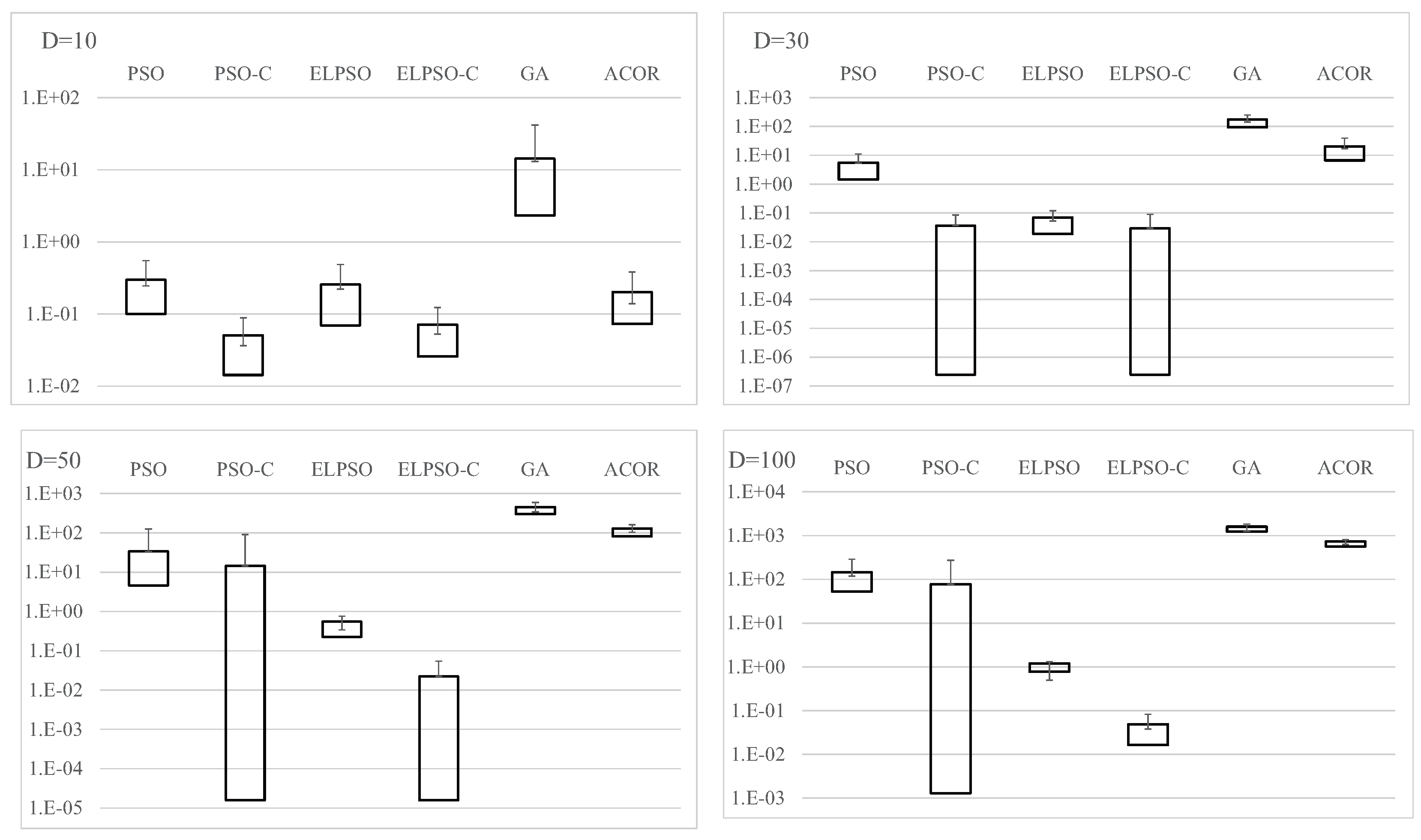
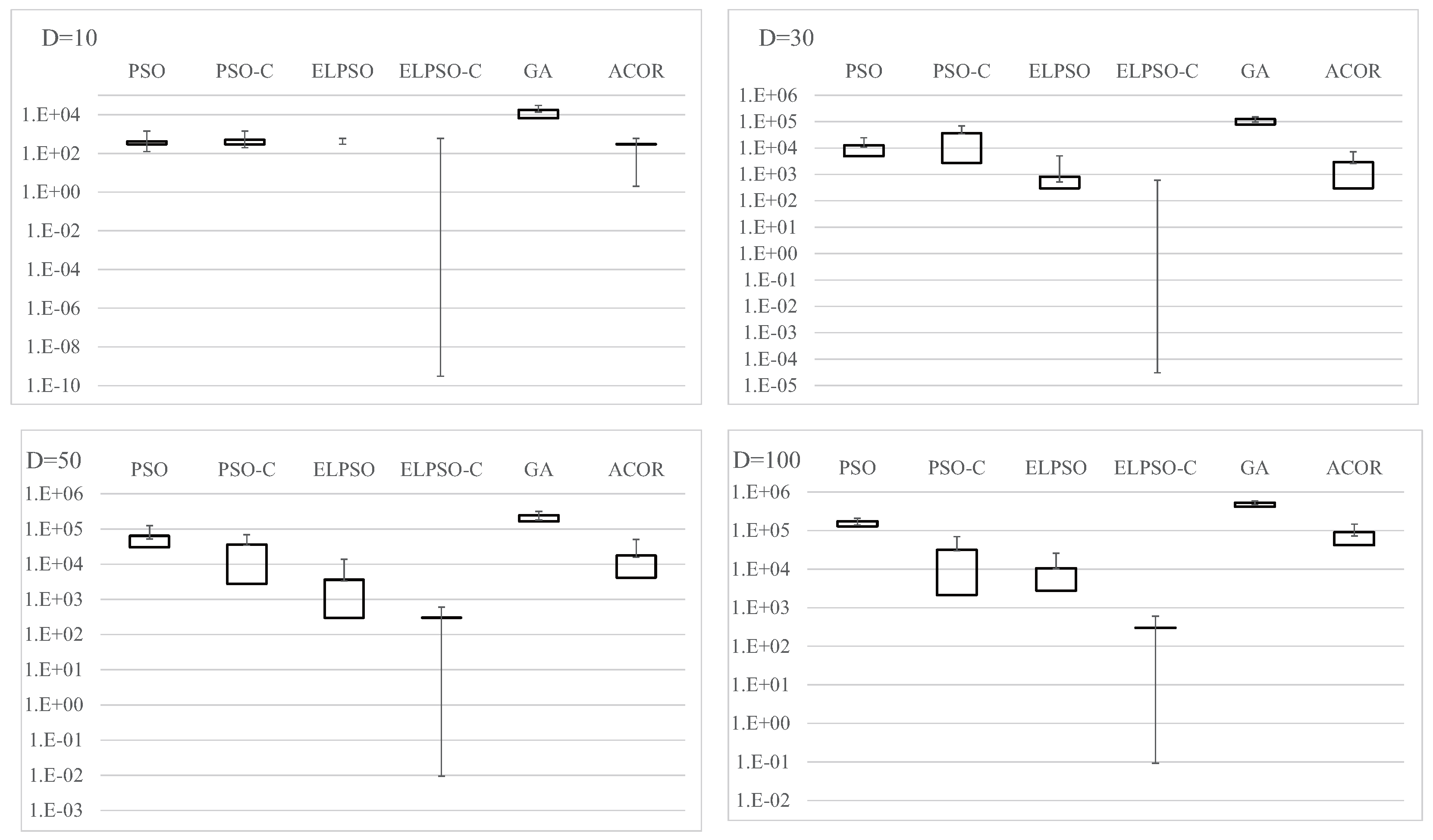
Figure A1.
Experimental results on F1, the sphere function, for , 30, 50, and 100 (30 trials).
Figure A1.
Experimental results on F1, the sphere function, for , 30, 50, and 100 (30 trials).
Table A3.
Experimental results on F2, the weighted sphere function, for , 30, 50, and 100 (30 trials).
Table A3.
Experimental results on F2, the weighted sphere function, for , 30, 50, and 100 (30 trials).
| Metric | PSO | PSO-C | ELPSO | ELPSO-C | GA | ACOR |
|---|
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Variance | | | | | | |
| Time [s] | 54.88 | 149.40 | 27.41 | 74.66 | 15.60 | 596.94 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Variance | | | | | | |
| Time [s] | 73.97 | 315.08 | 42.25 | 155.37 | 24.37 | 1718.49 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Variance | | | | | | |
| Time [s] | 72.15 | 468.87 | 68.32 | 237.27 | 31.69 | 3038.35 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Variance | | | | | | |
| Time [s] | 137.34 | 887.10 | 122.08 | 418.39 | 53.65 | 5498.25 |
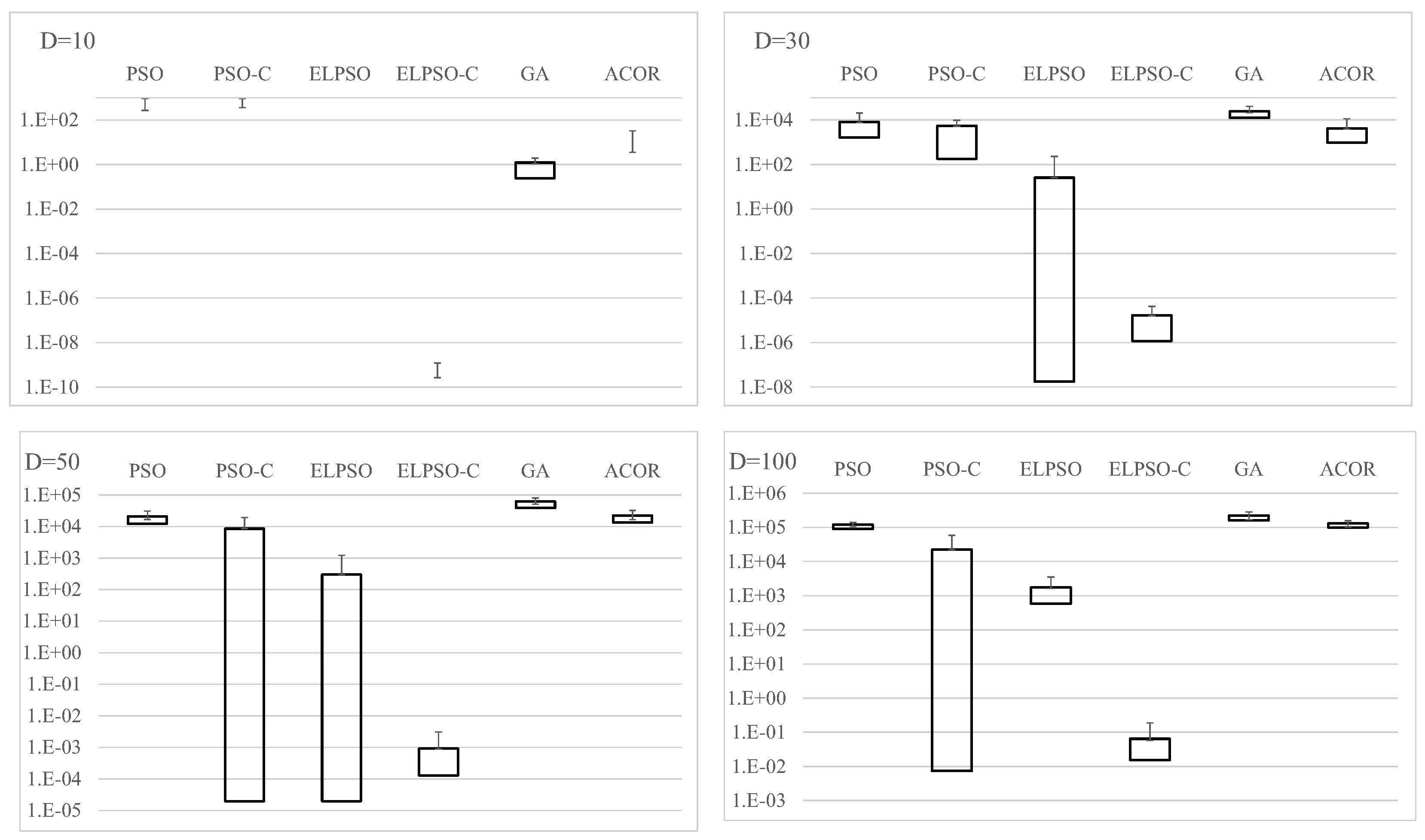
Figure A2.
Experimental results on F2, the weighted sphere function, for , 30, 50, and 100 (30 trials).
Figure A2.
Experimental results on F2, the weighted sphere function, for , 30, 50, and 100 (30 trials).
Table A4.
Experimental results on F3, the Rosenbrock function, for , 30, 50, and 100 (30 trials).
Table A4.
Experimental results on F3, the Rosenbrock function, for , 30, 50, and 100 (30 trials).
| Metric | PSO | PSO-C | ELPSO | ELPSO-C | GA | ACOR |
|---|
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Variance | | | | | | |
| Time [s] | 62.29 | 146.49 | 30.70 | 72.18 | 20.98 | 601.12 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Variance | | | | | | |
| Time [s] | 97.76 | 298.33 | 60.19 | 169.77 | 45.57 | 1732.89 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Variance | | | | | | |
| Time [s] | 130.64 | 465.47 | 101.32 | 271.14 | 69.62 | 2767.56 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Variance | | | | | | |
| Time [s] | 215.43 | 837.74 | 236.69 | 565.32 | 132.11 | 5555.77 |
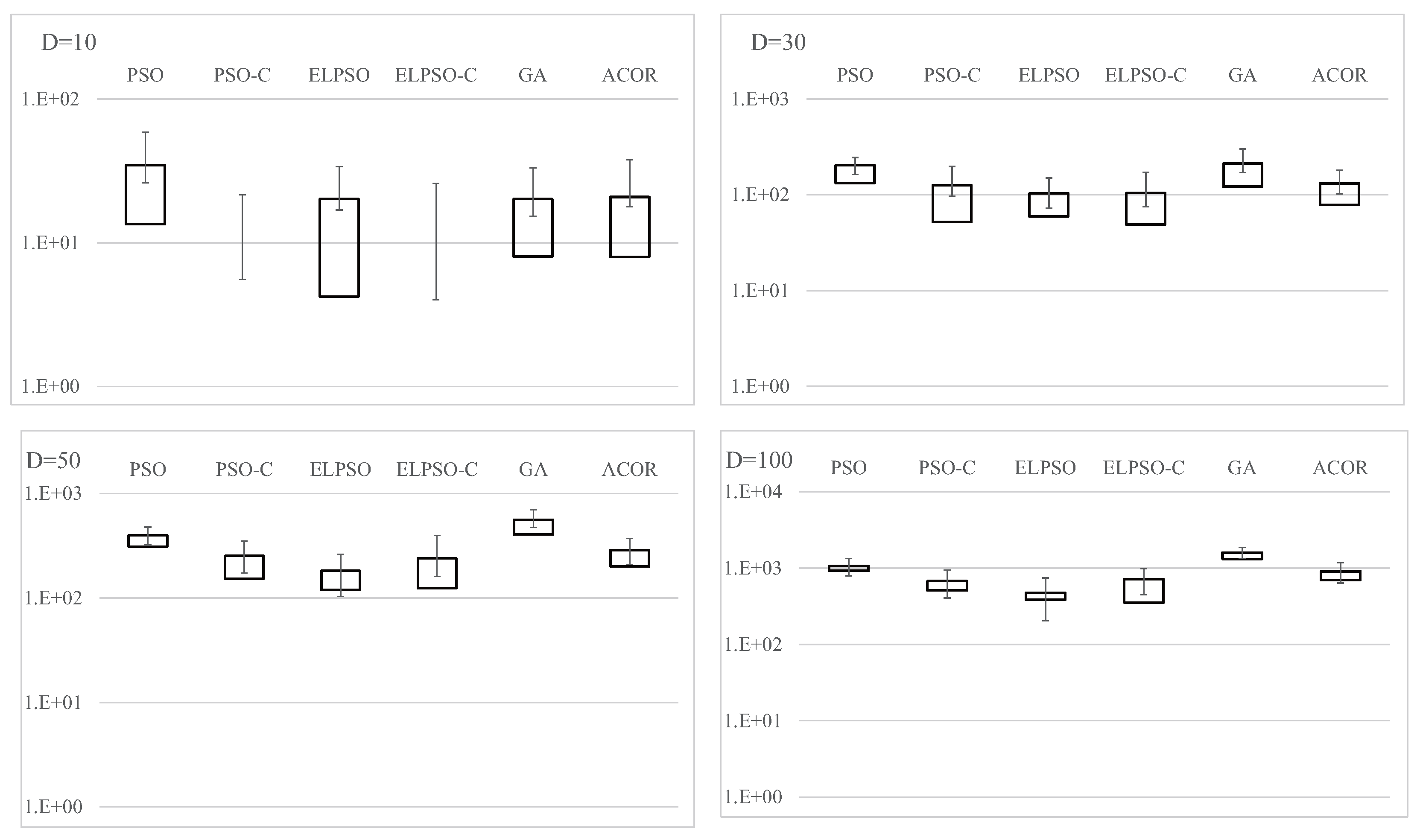
Figure A3.
Experimental results on F3, the Rosenbrock function, for , 30, 50, and 100 (30 trials).
Figure A3.
Experimental results on F3, the Rosenbrock function, for , 30, 50, and 100 (30 trials).
Table A5.
Results for F4, Schwefel’s problem 2.22, for , 30, 50, and 100 (30 trials).
Table A5.
Results for F4, Schwefel’s problem 2.22, for , 30, 50, and 100 (30 trials).
| Metric | PSO | PSO-C | ELPSO | ELPSO-C | GA | ACOR |
|---|
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 54.27 | 153.45 | 28.00 | 71.97 | 15.54 | 637.79 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 78.66 | 464.12 | 45.79 | 159.96 | 27.99 | 1773.50 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 96.81 | 464.12 | 77.04 | 247.27 | 42.48 | 2791.94 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 159.22 | 845.76 | 152.73 | 515.90 | 78.26 | 5932.17 |
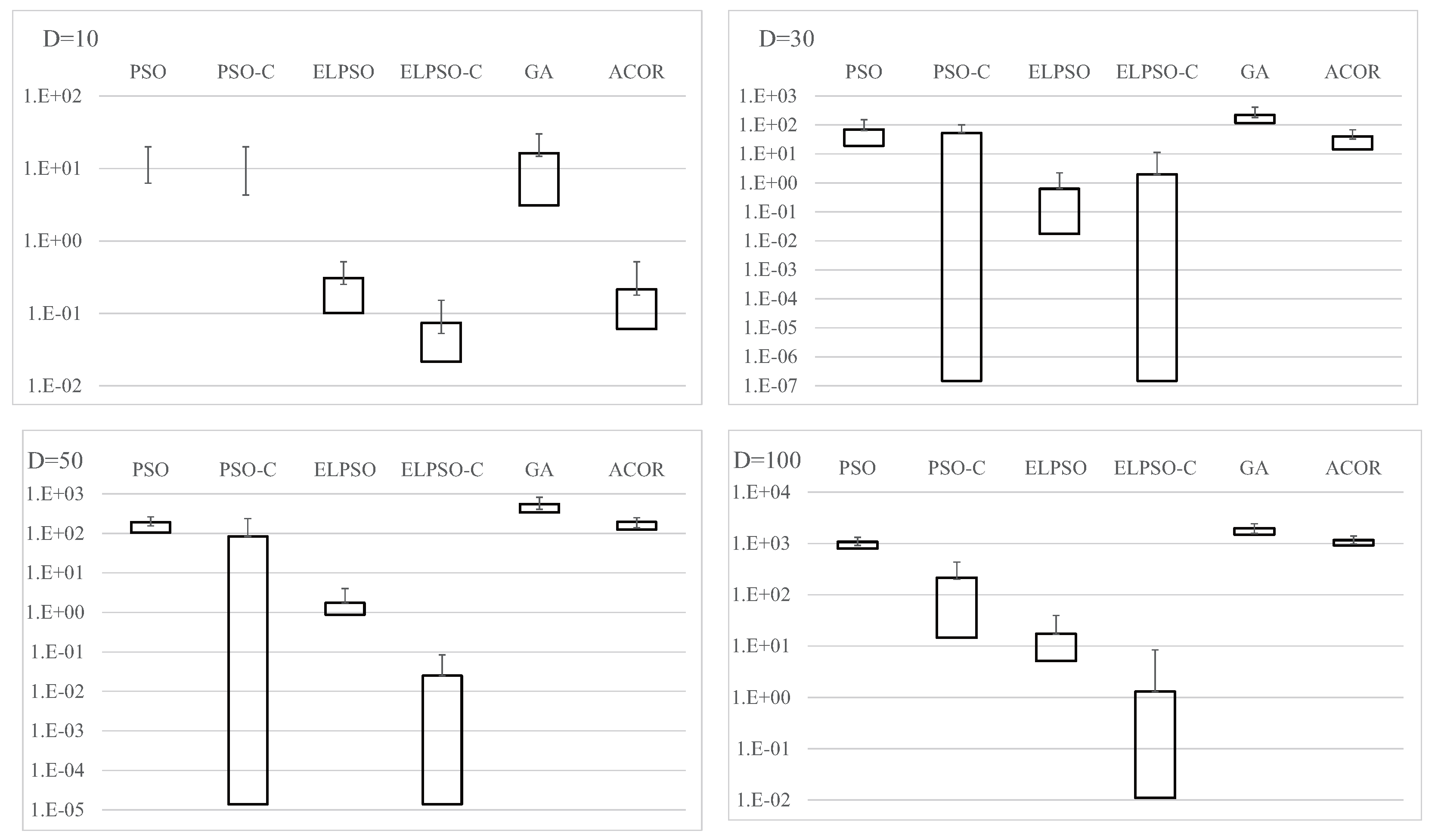
Figure A4.
Results for F4, Schwefel’s problem 2.22, for , 30, 50, and 100 (30 trials).
Figure A4.
Results for F4, Schwefel’s problem 2.22, for , 30, 50, and 100 (30 trials).
Table A6.
Results for F5, Rastrigin’s function, for , 30, 50, and 100 (30 trials).
Table A6.
Results for F5, Rastrigin’s function, for , 30, 50, and 100 (30 trials).
| Metric | PSO | PSO-C | ELPSO | ELPSO-C | GA | ACOR |
|---|
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 59.28 | 157.94 | 31.26 | 73.01 | 19.91 | 648.48 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 91.92 | 315.01 | 55.06 | 165.97 | 40.15 | 1790.37 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 119.01 | 465.44 | 87.80 | 266.02 | 63.56 | 2810.82 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 204.40 | 871.91 | 212.40 | 522.02 | 103.06 | 5959.35 |
Figure A5.
Results for F5, Rastrigin’s function, for , 30, 50, and 100 (30 trials).
Figure A5.
Results for F5, Rastrigin’s function, for , 30, 50, and 100 (30 trials).
Table A7.
Results for F6, Ackley’s function, for , 30, 50, and 100 (30 trials).
Table A7.
Results for F6, Ackley’s function, for , 30, 50, and 100 (30 trials).
| Metric | PSO | PSO-C | ELPSO | ELPSO-C | GA | ACOR |
|---|
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 67.07 | 164.67 | 37.01 | 79.68 | 26.27 | 648.61 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 106.58 | 468.00 | 67.54 | 172.27 | 60.45 | 1798.76 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 139.82 | 468.00 | 110.00 | 288.28 | 79.18 | 2834.69 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 234.20 | 894.19 | 263.98 | 607.14 | 145.99 | 6000.24 |
Figure A6.
Results for F6, Ackley’s function, for , 30, 50, and 100 (30 trials).
Figure A6.
Results for F6, Ackley’s function, for , 30, 50, and 100 (30 trials).
Table A8.
Results for F7, Griewank’s function, for , 30, 50, and 100 (30 trials).
Table A8.
Results for F7, Griewank’s function, for , 30, 50, and 100 (30 trials).
| Metric | PSO | PSO-C | ELPSO | ELPSO-C | GA | ACOR |
|---|
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 61.76 | 156.22 | 31.98 | 74.63 | 21.89 | 598.93 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 92.32 | 343.09 | 57.25 | 175.41 | 40.82 | 1723.48 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 126.57 | 521.18 | 94.94 | 278.49 | 64.13 | 3069.09 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 198.05 | 954.31 | 210.52 | 556.68 | 106.50 | 5522.85 |
Table A9.
Results for F8, Schwefel’s function, for , 30, 50, and 100 (30 trials).
Table A9.
Results for F8, Schwefel’s function, for , 30, 50, and 100 (30 trials).
| Metric | PSO | PSO-C | ELPSO | ELPSO-C | GA | ACOR |
|---|
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 59.94 | 141.47 | 31.18 | 72.24 | 19.60 | 548.77 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 88.01 | 423.91 | 52.58 | 169.98 | 35.97 | 1867.13 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 110.39 | 423.91 | 83.14 | 271.26 | 51.31 | 3045.01 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 186.88 | 804.47 | 188.78 | 554.45 | 98.63 | 5709.09 |
Figure A7.
Results for F7, Griewank’s function, for , 30, 50, and 100 (30 trials).
Figure A7.
Results for F7, Griewank’s function, for , 30, 50, and 100 (30 trials).
Table A10.
Results for F9, the shifted sphere function, for , 30, 50, and 100 (30 trials).
Table A10.
Results for F9, the shifted sphere function, for , 30, 50, and 100 (30 trials).
| Metric | PSO | PSO-C | ELPSO | ELPSO-C | GA | ACOR |
|---|
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 47.65 | 146.66 | 24.03 | 70.87 | 22.32 | 589.62 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 68.76 | 304.05 | 37.00 | 152.84 | 20.79 | 1693.08 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 90.73 | 483.26 | 55.83 | 234.25 | 26.60 | 3024.95 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 126.95 | 852.62 | 104.16 | 405.53 | 46.72 | 5889.74 |
Figure A8.
Results for F8, Schwefel’s function, for , 30, 50, and 100 (30 trials).
Figure A8.
Results for F8, Schwefel’s function, for , 30, 50, and 100 (30 trials).
Figure A9.
Results for F9, the shifted sphere function, for , 30, 50, and 100 (30 trials).
Figure A9.
Results for F9, the shifted sphere function, for , 30, 50, and 100 (30 trials).
Table A11.
Results for F10, the shifted weighted sphere function, for , 30, 50, and 100 (30 trials).
Table A11.
Results for F10, the shifted weighted sphere function, for , 30, 50, and 100 (30 trials).
| Metric | PSO | PSO-C | ELPSO | ELPSO-C | GA | ACOR |
|---|
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 53.40 | 148.42 | 27.40 | 73.84 | 16.23 | 596.94 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 75.54 | 320.95 | 42.63 | 154.29 | 24.04 | 1770.73 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 89.85 | 453.22 | 68.16 | 232.65 | 32.03 | 2786.18 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 141.04 | 819.14 | 124.75 | 409.77 | 54.44 | 5478.88 |
Figure A10.
Results for F10, the shifted weighted sphere function, for , 30, 50, and 100 (30 trials).
Figure A10.
Results for F10, the shifted weighted sphere function, for , 30, 50, and 100 (30 trials).
Table A12.
Results for F11, the shifted Rastrigin function, for , 30, 50, and 100 (30 trials).
Table A12.
Results for F11, the shifted Rastrigin function, for , 30, 50, and 100 (30 trials).
| Metric | PSO | PSO-C | ELPSO | ELPSO-C | GA | ACOR |
|---|
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 60.53 | 155.79 | 31.81 | 73.11 | 23.51 | 598.44 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 93.10 | 298.06 | 55.94 | 162.71 | 39.01 | 1777.60 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 121.00 | 455.80 | 89.63 | 259.88 | 59.30 | 2812.31 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 203.08 | 815.72 | 211.25 | 517.98 | 108.75 | 5490.48 |
Figure A11.
Results for F11, the shifted Rastrigin function, for , 30, 50, and 100 (30 trials).
Figure A11.
Results for F11, the shifted Rastrigin function, for , 30, 50, and 100 (30 trials).
Table A13.
Results for F12, the shifted Griewank function, for , 30, 50, and 100 (30 trials).
Table A13.
Results for F12, the shifted Griewank function, for , 30, 50, and 100 (30 trials).
| Metric | PSO | PSO-C | ELPSO | ELPSO-C | GA | ACOR |
|---|
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 63.34 | 158.73 | 32.34 | 76.54 | 23.78 | 599.34 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 94.91 | 343.29 | 57.25 | 173.29 | 40.05 | 1726.06 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 121.34 | 514.46 | 92.05 | 277.63 | 59.77 | 3069.85 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 208.24 | 899.05 | 221.05 | 544.99 | 111.00 | 5942.49 |
Figure A12.
Results for F12, the shifted Griewank function, for , 30, 50, and 100 (30 trials).
Figure A12.
Results for F12, the shifted Griewank function, for , 30, 50, and 100 (30 trials).
Table A14.
Results for F13, the shifted and rotated Bent Cigar function, for , 30, 50, and 100 (30 trials).
Table A14.
Results for F13, the shifted and rotated Bent Cigar function, for , 30, 50, and 100 (30 trials).
| Metric | PSO | PSO-C | ELPSO | ELPSO-C | GA | ACOR |
|---|
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 52.40 | 143.46 | 27.58 | 68.97 | 18.43 | 636.90 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 72.41 | 425.27 | 40.19 | 149.01 | 22.49 | 1699.99 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 87.18 | 425.27 | 57.36 | 252.92 | 30.69 | 2990.90 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 133.83 | 760.31 | 116.30 | 450.34 | 49.87 | 5925.77 |
Figure A13.
Results for F13, the shifted and rotated Bent Cigar function, for , 30, 50, and 100 (30 trials).
Figure A13.
Results for F13, the shifted and rotated Bent Cigar function, for , 30, 50, and 100 (30 trials).
Figure A14.
Results for F14, the shifted and rotated Zakharov function, for , 30, 50, and 100 (30 trials).
Figure A14.
Results for F14, the shifted and rotated Zakharov function, for , 30, 50, and 100 (30 trials).
Table A15.
Results for F14, the shifted and rotated Zakharov function, for , 30, 50, and 100 (30 trials).
Table A15.
Results for F14, the shifted and rotated Zakharov function, for , 30, 50, and 100 (30 trials).
| Metric | PSO | PSO-C | ELPSO | ELPSO-C | GA | ACOR |
|---|
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 57.79 | 150.52 | 31.68 | 72.00 | 22.49 | 644.34 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 87.23 | 456.02 | 54.99 | 164.45 | 48.28 | 1796.73 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 115.87 | 456.02 | 87.41 | 256.94 | 65.09 | 2920.45 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 194.50 | 837.93 | 207.22 | 543.21 | 112.59 | 5519.70 |
Figure A15.
Results for F15, the shifted and rotated Rosenbrock function, for , 30, 50, and 100 (30 trials).
Figure A15.
Results for F15, the shifted and rotated Rosenbrock function, for , 30, 50, and 100 (30 trials).
Table A16.
Results for F15, the shifted and rotated Rosenbrock function, for , 30, 50, and 100 (30 trials).
Table A16.
Results for F15, the shifted and rotated Rosenbrock function, for , 30, 50, and 100 (30 trials).
| Metric | PSO | PSO-C | ELPSO | ELPSO-C | GA | ACOR |
|---|
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 64.79 | 151.67 | 36.08 | 73.35 | 24.99 | 605.82 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 100.30 | 453.68 | 64.51 | 166.33 | 49.11 | 1729.61 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 130.64 | 453.68 | 102.77 | 264.74 | 69.62 | 2934.19 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 221.13 | 827.10 | 248.35 | 573.31 | 131.72 | 5967.33 |
Figure A16.
Results for F16, the shifted and rotated Rastrigin function, for , 30, 50, and 100 (30 trials).
Figure A16.
Results for F16, the shifted and rotated Rastrigin function, for , 30, 50, and 100 (30 trials).
Table A17.
Results for F16, the shifted and rotated Rastrigin function, for , 30, 50, and 100 (30 trials).
Table A17.
Results for F16, the shifted and rotated Rastrigin function, for , 30, 50, and 100 (30 trials).
| Metric | PSO | PSO-C | ELPSO | ELPSO-C | GA | ACOR |
|---|
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 63.56 | 162.50 | 35.73 | 77.76 | 28.44 | 651.85 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 107.38 | 495.15 | 69.49 | 191.50 | 58.66 | 1798.20 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 145.30 | 495.15 | 118.90 | 293.03 | 89.51 | 2841.95 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 254.33 | 898.74 | 290.62 | 686.85 | 168.89 | 5565.35 |
Figure A17.
Results for F17, the shifted and rotated Schaffer’s F7 function, for , 30, 50, and 100 (30 trials).
Figure A17.
Results for F17, the shifted and rotated Schaffer’s F7 function, for , 30, 50, and 100 (30 trials).
Table A18.
Results for F17, the shifted and rotated Schaffer’s F7 function, for , 30, 50, and 100 (30 trials).
Table A18.
Results for F17, the shifted and rotated Schaffer’s F7 function, for , 30, 50, and 100 (30 trials).
| Metric | PSO | PSO-C | ELPSO | ELPSO-C | GA | ACOR |
|---|
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 73.97 | 161.48 | 41.28 | 79.86 | 36.80 | 610.86 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 143.55 | 362.95 | 101.75 | 212.46 | 86.96 | 1758.90 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 201.67 | 543.62 | 175.18 | 343.40 | 138.55 | 3139.37 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 351.62 | 1024.56 | 437.53 | 759.67 | 274.57 | 6109.19 |
Table A19.
Results for F18, the shifted and rotated Griewank function, for , 30, 50, and 100 (30 trials).
Table A19.
Results for F18, the shifted and rotated Griewank function, for , 30, 50, and 100 (30 trials).
| Metric | PSO | PSO-C | ELPSO | ELPSO-C | GA | ACOR |
|---|
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 208.24 | 154.07 | 221.05 | 544.99 | 22.60 | 592.90 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 93.50 | 465.66 | 55.61 | 180.77 | 40.77 | 1713.71 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 115.48 | 465.66 | 87.72 | 247.32 | 58.49 | 3055.10 |
| − Mean | | | | | | |
| Max | | | | | | |
| Min | | | | | | |
| Var | | | | | | |
| Time [s] | 184.81 | 844.44 | 194.72 | 500.24 | 117.43 | 5940.40 |
Figure A18.
Results for F18, the shifted and rotated Griewank function, for , 30, 50, and 100 (30 trials).
Figure A18.
Results for F18, the shifted and rotated Griewank function, for , 30, 50, and 100 (30 trials).