Spatial Mapping of Genes Implicated in SARS-CoV-2 Neuroinvasion to Dorsolateral Prefrontal Cortex Gray Matter
Abstract
1. Introduction
2. Materials and Methods
2.1. Data Acquisition
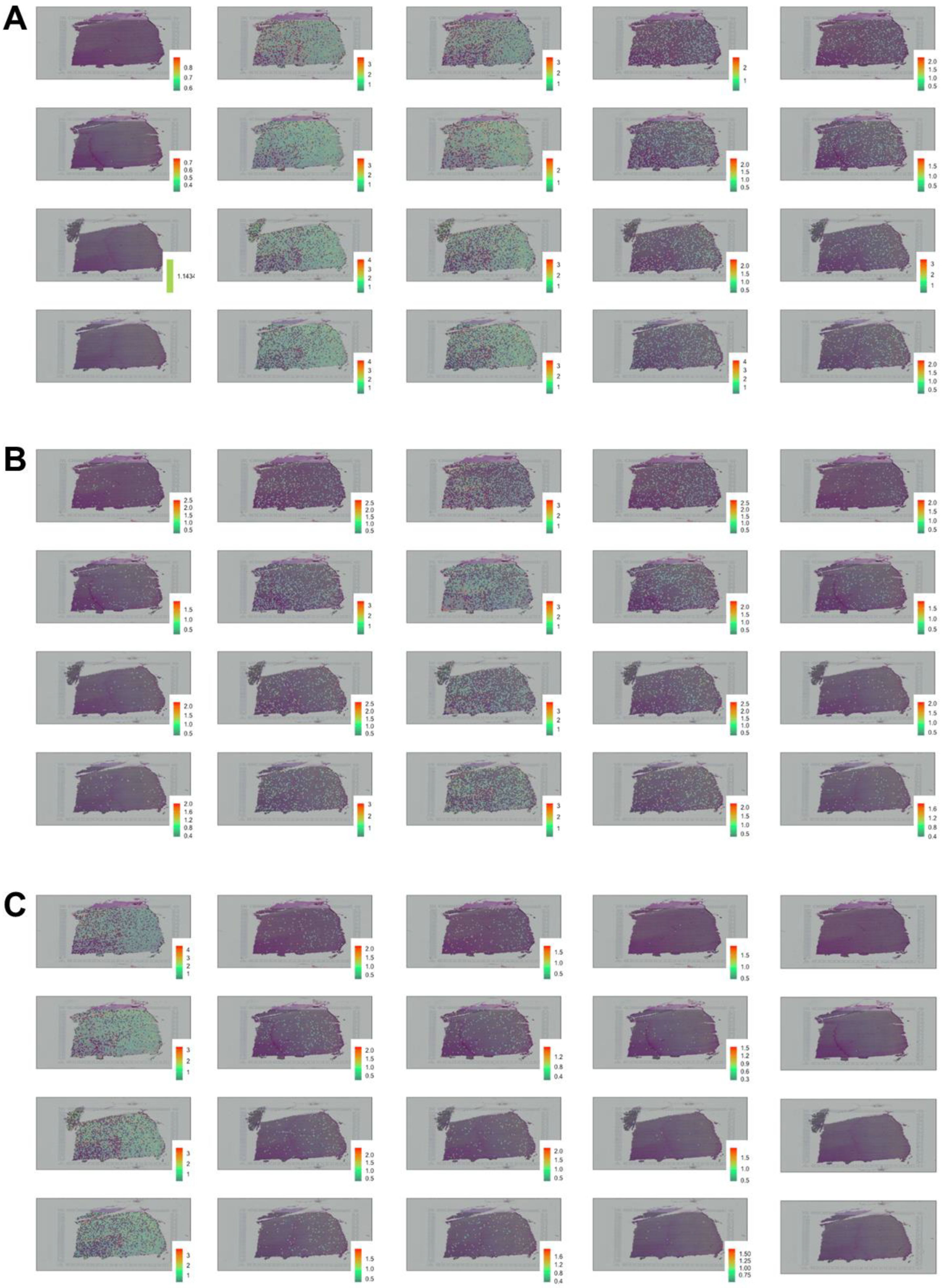
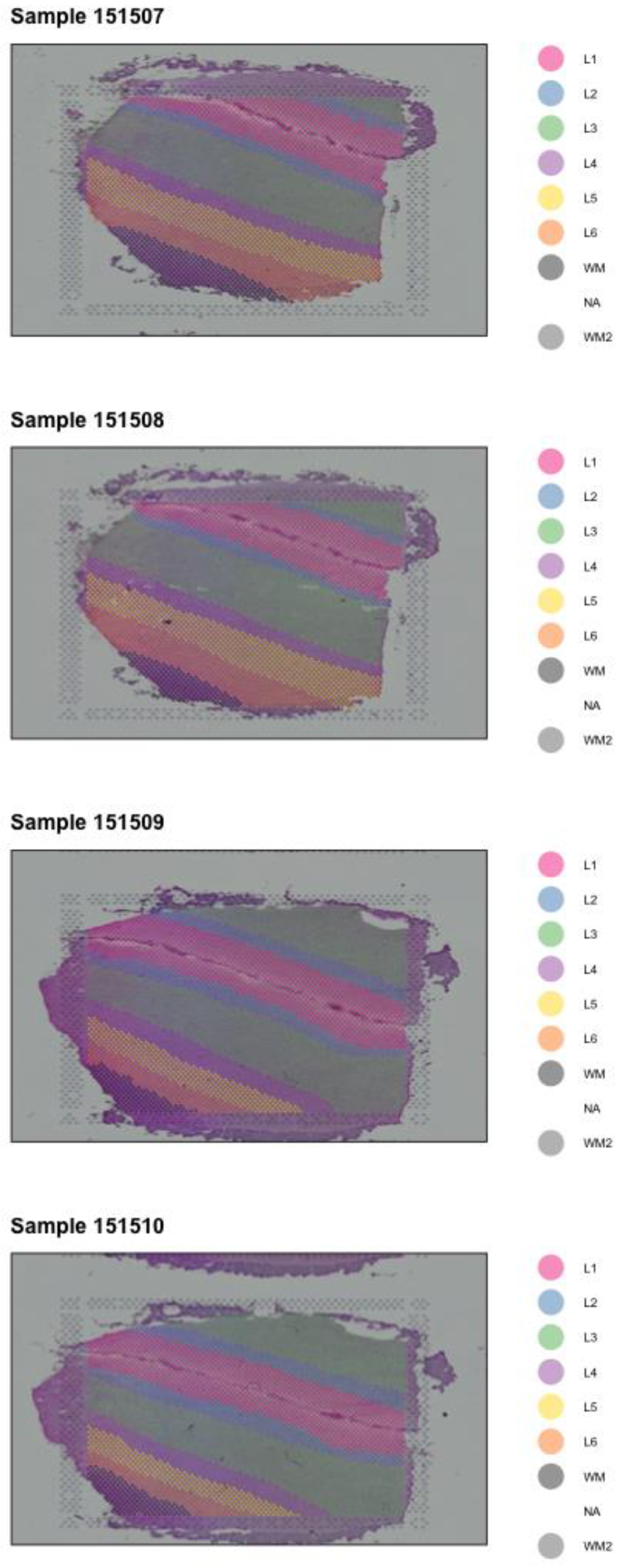
2.2. Spatial Mapping
2.3. Software
3. Results
4. Discussion
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Cascella, M.; Rajnik, M.; Aleem, A.; Dulebohn, S.C.; Di Napoli, R. Features, Evaluation, and Treatment of Coronavirus (COVID-19). In StatPearls; StatPearls Publishing: Treasure Island, FL, USA.
- Ahmad, F.B.; Cisewski, J.A.; Miniño, A.; Anderson, R.N. Provisional Mortality Data—United States, 2020. MMWR. Morb. Mortal. Wkly. Rep. 2021, 70, 519–522. [Google Scholar] [CrossRef]
- Dinakaran, D.; Manjunatha, N.; Naveen Kumar, C.; Suresh, B.M. Neuropsychiatric aspects of COVID-19 pandemic: A selective review. Asian J. Psychiatry 2020, 53, 102188. [Google Scholar] [CrossRef]
- Premraj, L.; Kannapadi, N.V.; Briggs, J.; Seal, S.M.; Battaglini, D.; Fanning, J.; Suen, J.; Robba, C.; Fraser, J.; Cho, S.M. Mid and long-term neurological and neuropsychiatric manifestations of post-COVID-19 syndrome: A meta-analysis. J. Neurol. Sci. 2022, 434, 120162. [Google Scholar] [CrossRef]
- Singh, K.K.; Chaubey, G.; Chen, J.Y.; Suravajhala, P. Decoding SARS-CoV-2 hijacking of host mitochondria in COVID-19 pathogenesis. Am. J. Physiol. Cell Physiol. 2020, 319, C258–C267. [Google Scholar] [CrossRef]
- Najjar, S.; Najjar, A.; Chong, D.J.; Pramanik, B.K.; Kirsch, C.; Kuzniecky, R.I.; Pacia, S.V.; Azhar, S. Central nervous system complications associated with SARS-CoV-2 infection: Integrative concepts of pathophysiology and case reports. J. Neuroinflammation 2020, 17, 231. [Google Scholar] [CrossRef]
- Wu, K.E.; Fazal, F.M.; Parker, K.R.; Zou, J.; Chang, H.Y. RNA-GPS Predicts SARS-CoV-2 RNA Residency to Host Mitochondria and Nucleolus. Cell Syst. 2020, 11, 102–108.e3. [Google Scholar] [CrossRef]
- Generoso, J.S.; Barichello de Quevedo, J.L.; Cattani, M.; Lodetti, B.F.; Sousa, L.; Collodel, A.; Diaz, A.P.; Dal-Pizzol, F. Neurobiology of COVID-19: How can the virus affect the brain? Braz. J. Psychiatry 2021, 43, 650–664. [Google Scholar] [CrossRef]
- Chen, R.; Wang, K.; Yu, J.; Howard, D.; French, L.; Chen, Z.; Wen, C.; Xu, Z. The Spatial and Cell-Type Distribution of SARS-CoV-2 Receptor ACE2 in the Human and Mouse Brains. Front. Neurol. 2021, 11, 573095. [Google Scholar] [CrossRef]
- Bergmann, C.C.; Lane, T.E.; Stohlman, S.A. Coronavirus infection of the central nervous system: Host-virus stand-off. Nat. Rev. Microbiol. 2006, 4, 121–132. [Google Scholar] [CrossRef]
- Iadecola, C.; Anrather, J.; Kamel, H. Effects of COVID-19 on the Nervous System. Cell 2020, 183, 16–27.e1. [Google Scholar] [CrossRef]
- Wang, K.; Chen, W.; Zhang, Z.; Deng, Y.; Lian, J.Q.; Du, P.; Wei, D.; Zhang, Y.; Sun, X.X.; Gong, L.; et al. CD147-spike protein is a novel route for SARS-CoV-2 infection to host cells. Signal Transduct. Target. Ther. 2020, 5, 283. [Google Scholar] [CrossRef] [PubMed]
- Cantuti-Castelvetri, L.; Ojha, R.; Pedro, L.D.; Djannatian, M.; Franz, J.; Kuivanen, S.; van der Meer, F.; Kallio, K.; Kaya, T.; Anastasina, M.; et al. Neuropilin-1 facilitates SARS-CoV-2 cell entry and infectivity. Science 2020, 370, 856–860. [Google Scholar] [CrossRef]
- Maynard, K.R.; Collado-Torres, L.; Weber, L.M.; Uytingco, C.; Barry, B.K.; Williams, S.R.; Catallini, J.L., II; Tran, M.N.; Besich, Z.; Tippani, M.; et al. Transcriptome-scale spatial gene expression in the human dorsolateral prefrontal cortex. Nat. Neurosci. 2021, 24, 425–436. [Google Scholar] [CrossRef]
- Rao, A.; Barkley, D.; França, G.S.; Yanai, I. Exploring tissue architecture using spatial transcriptomics. Nature 2021, 596, 211–220. [Google Scholar] [CrossRef]
- Lee, J.; Yoo, M.; Choi, J. Recent advances in spatially resolved transcriptomics: Challenges and opportunities. BMB Rep. 2022, 55, 113–124. [Google Scholar] [CrossRef]
- Teves, J.M.; Won, K.J. Mapping Cellular Coordinates through Advances in Spatial Transcriptomics Technology. Mol. Cells 2020, 43, 591–599. [Google Scholar] [CrossRef]
- Kapur, A.; Marwah, K.; Alterovitz, G. Gene expression prediction using low-rank matrix completion. BMC Bioinform. 2016, 17, 243. [Google Scholar] [CrossRef]
- Panikratova, Y.R.; Vlasova, R.M.; Akhutina, T.V.; Korneev, A.A.; Sinitsyn, V.E.; Pechenkova, E.V. Functional connectivity of the dorsolateral prefrontal cortex contributes to different components of executive functions. Int. J. Psychophysiol. Off. J. Int. Organ. Psychophysiol. 2020, 151, 70–79. [Google Scholar] [CrossRef]
- Pardo, B.; Spangler, A.; Weber, L.M.; Hicks, S.C.; Jaffe, A.E.; Martinowich, K.; Maynard, K.R.; Collado-Torres, L. spatialLIBD: An R/Bioconductor package to visualize spatially-resolved transcriptomics data. BMC Genom. 2022, 23, 1–5. [Google Scholar] [CrossRef]
- Yang, A.C.; Kern 2022, F.; Losada, P.M.; Agam, M.R.; Maat, C.A.; Schmartz, G.P.; Fehlmann, T.; Stein, J.A.; Schaum, N.; Lee, D.P.; et al. Dysregulation of brain and choroid plexus cell types in severe COVID-19. Nature 2021, 595, 565–571. [Google Scholar] [CrossRef]
- Nejati, V.; Majdi, R.; Salehinejad, M.A.; Nitsche, M.A. The role of dorsolateral and ventromedial prefrontal cortex in the processing of emotional dimensions. Sci. Rep. 2021, 11, 1971. [Google Scholar] [CrossRef]
- Torices, S.; Cabrera, R.; Stangis, M.; Naranjo, O.; Fattakhov, N.; Teglas, T.; Adesse, D.; Toborek, M. Expression of SARS-CoV-2-related receptors in cells of the neurovascular unit: Implications for HIV-1 infection. J. Neuroinflammation 2021, 18, 167. [Google Scholar] [CrossRef] [PubMed]
- Medina-Enríquez, M.M.; Lopez-León, S.; Carlos-Escalante, J.A.; Aponte-Torres, Z.; Cuapio, A.; Wegman-Ostrosky, T. ACE2: The molecular doorway to SARS-CoV-2. Cell Biosci. 2020, 10, 148. [Google Scholar] [CrossRef] [PubMed]
- Wulandari, L.; Hamidah, B.; Pakpahan, C.; Damayanti, N.S.; Kurniati, N.D.; Adiatmaja, C.O.; Wigianita, M.R.; Soedarsono, H.D.; Tinduh, D.; Prakoeswa, C.R.S.; et al. Initial study on TMPRSS2 p.Val160Met genetic variant in COVID-19 patients. Hum. Genom. 2021, 15, 29. [Google Scholar] [CrossRef] [PubMed]
- Callies, L.K.; Tadeo, D.; Simper, J.; Bugge, T.H.; Szabo, R. Iterative, multiplexed CRISPR-mediated gene editing for functional analysis of complex protease gene clusters. J. Biol. Chem. 2019, 294, 15987–15996. [Google Scholar] [CrossRef]
- Shen, W.B.; Logue, J.; Yang, P.; Baracco, L.; Elahi, M.; Reece, E.A.; Wang, B.; Li, L.; Blanchard, T.G.; Han, Z.; et al. SARS-CoV-2 invades cognitive centers of the brain and induces Alzheimer’s-like neuropathology. bioRxiv 2022. [Google Scholar] [CrossRef]
- Krasemann, S.; Haferkamp, U.; Pfefferle, S.; Woo, M.S.; Heinrich, F.; Schweizer, M.; Appelt-Menzel, A.; Cubukova, A.; Barenberg, J.; Leu, J.; et al. The blood-brain barrier is dysregulated in COVID-19 and serves as a CNS entry route for SARS-CoV-2. Stem Cell Rep. 2022, 17, 307–320. [Google Scholar] [CrossRef]
- Roy, S.; Bag, A.K.; Singh, R.K.; Talmadge, J.E.; Batra, S.K.; Datta, K. Multifaceted Role of Neuropilins in the Immune System: Potential Targets for Immunotherapy. Front. Immunol. 2017, 8, 1228. [Google Scholar] [CrossRef]
- Fullard, J.F.; Lee, H.C.; Voloudakis, G.; Suo, S.; Javidfar, B.; Shao, Z.; Peter, C.; Zhang, W.; Jiang, S.; Corvelo, A.; et al. Single-nucleus transcriptome analysis of human brain immune response in patients with severe COVID-19. Genome Med. 2021, 13, 118. [Google Scholar] [CrossRef]
- Khanmohammadi, S.; Rezaei, N.; Khazaei, M.; Shirkani, A. A Case of Autosomal Recessive Interferon Alpha/Beta Receptor Alpha Chain (IFNAR1) Deficiency with Severe COVID-19. J. Clin. Immunol. 2022, 42, 19–24. [Google Scholar] [CrossRef]
- Diamond, M.S.; Farzan, M. The broad-spectrum antiviral functions of IFIT and IFITM proteins. Nature reviews. Immunology 2013, 13, 46–57. [Google Scholar] [CrossRef] [PubMed]
- Łacina, P.; Butrym, A.; Frontkiewicz, D.; Mazur, G.; Bogunia-Kubik, K. Soluble CD147 (BSG) as a Prognostic Marker in Multiple Myeloma. Curr. Issues Mol. Biol. 2022, 44, 350–359. [Google Scholar] [CrossRef]
- Gomes, C.P.; Fernandes, D.E.; Casimiro, F.; da Mata, G.F.; Passos, M.T.; Varela, P.; Mastroianni-Kirsztajn, G.; Pesquero, J.B. Cathepsin L in COVID-19: From Pharmacological Evidences to Genetics. Front. Cell. Infect. Microbiol. 2020, 10, 589505. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Y.; Gao, X.; Bai, X.; Yao, S.; Chang, Y.Z.; Gao, G. The emerging role of furin in neurodegenerative and neuropsychiatric diseases. Transl. Neurodegener. 2022, 11, 39. [Google Scholar] [CrossRef] [PubMed]
- Yang, X.; Yin, H.; Zhang, D.; Peng, L.; Li, K.; Cui, F.; Xia, C.; Li, Z.; Huang, H. Bibliometric Analysis of Cathepsin B Research From 2011 to 2021. Front. Med. 2022, 9, 898455. [Google Scholar] [CrossRef] [PubMed]
- Upadhyay, G. Emerging Role of Lymphocyte Antigen-6 Family of Genes in Cancer and Immune Cells. Front. Immunol. 2019, 10, 819. [Google Scholar] [CrossRef] [PubMed]
- Zhou, Y.; Xu, J.; Hou, Y.; Leverenz, J.B.; Kallianpur, A.; Mehra, R.; Liu, Y.; Yu, H.; Pieper, A.A.; Jehi, L.; et al. Network medicine links SARS-CoV-2/COVID-19 infection to brain microvascular injury and neuroinflammation in dementia-like cognitive impairment. bioRxiv 2021. [CrossRef]
- McQuaid, C.; Brady, M.; Deane, R. SARS-CoV-2: Is there neuroinvasion? Fluids Barriers CNS 2021, 18, 32. [Google Scholar] [CrossRef]
- Yang, Y.; Shi, X.; Liu, W.; Zhou, Q.; Chan Lau, M.; Chun Tatt Lim, J.; Sun, L.; Ng, C.; Yeong, J.; Liu, J. SC-MEB: Spatial clustering with hidden Markov random field using empirical Bayes. Brief. Bioinform. 2022, 23, bbab466. [Google Scholar] [CrossRef]
- Zhang, L.; Chen, D.; Song, D.; Liu, X.; Zhang, Y.; Xu, X.; Wang, X. Clinical and translational values of spatial transcriptomics. Signal Transduct. Target. Ther. 2022, 7, 111. [Google Scholar] [CrossRef]
- Cang, Z.; Ning, X.; Nie, A.; Xu, M.; Zhang, J. SCAN-IT: Domain segmentation of spatial transcriptomics images by graph neural network. BMVC: Proceedings of the British Machine Vision Conference. Br. Mach. Vis. Conf. 2021, 32, 406. [Google Scholar]

| Sample ID | Estimated # of Spots | Mean Reads per Spot | Valid Barcodes | Sequencing Saturation |
|---|---|---|---|---|
| 151507 | 4226 | 76,928 | 96.9% | 95.3% |
| 151508 | 4384 | 60,230 | 96.9% | 95.2% |
| 151509 | 4789 | 58,788 | 96.5% | 94.0% |
| 151510 | 4634 | 58,482 | 96.6% | 94.1% |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Batchu, S.; Diaz, M.J.; Tran, J.T.; Fadil, A.; Taneja, K.; Patel, K.; Lucke-Wold, B. Spatial Mapping of Genes Implicated in SARS-CoV-2 Neuroinvasion to Dorsolateral Prefrontal Cortex Gray Matter. COVID 2023, 3, 82-89. https://doi.org/10.3390/covid3010005
Batchu S, Diaz MJ, Tran JT, Fadil A, Taneja K, Patel K, Lucke-Wold B. Spatial Mapping of Genes Implicated in SARS-CoV-2 Neuroinvasion to Dorsolateral Prefrontal Cortex Gray Matter. COVID. 2023; 3(1):82-89. https://doi.org/10.3390/covid3010005
Chicago/Turabian StyleBatchu, Sai, Michael Joseph Diaz, Jasmine Thuy Tran, Angela Fadil, Kamil Taneja, Karan Patel, and Brandon Lucke-Wold. 2023. "Spatial Mapping of Genes Implicated in SARS-CoV-2 Neuroinvasion to Dorsolateral Prefrontal Cortex Gray Matter" COVID 3, no. 1: 82-89. https://doi.org/10.3390/covid3010005
APA StyleBatchu, S., Diaz, M. J., Tran, J. T., Fadil, A., Taneja, K., Patel, K., & Lucke-Wold, B. (2023). Spatial Mapping of Genes Implicated in SARS-CoV-2 Neuroinvasion to Dorsolateral Prefrontal Cortex Gray Matter. COVID, 3(1), 82-89. https://doi.org/10.3390/covid3010005







