1. Introduction
Findings from recent studies show that the temporal patterns of eating behavior are equally, if not more, important as the nutritional components and the total calories consumed. There is ample evidence that an irregular eating schedule has an adverse effect on body weight regulation and metabolic functions and may significantly increase the risks of metabolic diseases such as obesity, hypertension, diabetes, and stroke [
1]. On the other hand, a recent study shows that proper eating patterns—such as controlling all three meals within certain time window, also known as restricted feeding—help people control body weight [
2]. These findings suggest that
when we eat plays an important role in maintaining health and preventing chronic diseases. Understanding the temporal eating patterns thus can give hints for disease prediction and health intervention.
Traditional methods for studying eating behavior mostly rely on self-report. Several validated questionnaires have been widely used in research studies, including 24 h dietary recall, food frequency questionnaire, and dietary record. These psychometric instruments have the limitation of requiring users to make a log in a food journal every time they eat, which demands strong commitment from users. Another limitation of these methods is the so-called recall bias that may compromise the accuracy and reliability of the collected data.
The goal of our study is to develop and validate a method that can reliably recognize eating activities in a free-living environment using consumer wearable devices. Several automatic eating recognition approaches were previously developed in the literature [
3,
4,
5]. In this study, we propose a new multimodal approach that combines the monitoring of wrist movement, heart rate, and blood glucose in a free-living environment. Despite the drawback of not being able to obtain raw signals from the measurement devices, our method has the advantage of enhanced accessibility as consumer wearable devices are widely used by individual users in daily life. In the meantime, however, we faced the challenge of achieving good accuracy with noisy, imbalanced, and limited input data. The problem of interest was formulated into a binary classification problem. We applied ensemble machine learning algorithms in combination with different resampling strategies.
2. Data Analysis
2.1. Data Collection
We used three non-invasive consumer wearable sensors for collecting biometric data to achieve a better trade-off between accuracy and convenience [
6,
7]: a Mi Band 4, a Fitbit Charge 3, and a FreeStyle Libre 2. In addition to the sensors and their accompanying apps, another app named aTimeLogger was used to log the ground truth of activities. We logged both eating and non-eating activities. Eating activities included main meals and snacking. Non-eating activities included sleeping, tooth brushing, showering, cooking, house cleaning, exercising, walking, indoor working, outdoor working, playing video games, watching videos, browsing the Internet, reading, and smoking.
A total of 16 participants took part in the data collection experiment (nine women, average age: 30 years). Glucose data were not collected from six participants either because they did not provide consent for using FreeStyle Libre sensor or because of sensor malfunctioning (e.g., sensor peeling off). Eventually, 10 participants successfully collected data using all devices (five women, average range: 32 years). The data collection experiment last up to 14 days. Before the data collection experiment started, we held a briefing with each participant individually to explain the objective of this study and to walk them through the data collection process. We also instructed the participants to install the Notify&Fitness app (paired with Mi Band 4), the Fitbit app (paired with Fitbit Charge 3), and the FreeStyle Libre Link app (paired with FreeStyle Libre) and to pair up the corresponding devices through Bluetooth using a given account. During the data collection experiment, participants were asked to wear the Fitbit and Mi Band on their dominant and non-dominant wrist, respectively. The FreeStyle Libre sensor was applied to the back of their non-dominant upper arm on the first day of the experiment. At the end of the experiment, participants were instructed to remove the sensor from their arms.
2.2. Data Export and Cleaning
Participants were asked to sync data with the Fitbit app, the Mi Fit app, and the FreeStyle Libre Link app four times a day: after waking up, before lunch, before dinner, and before bedtime. At the end of the data collection experiment, participants were instructed to export data from the Fitbit web dashboard (because the Fitbit smartphone app does not support data export), the FreeStyle Libre Link app, and the aTimeLogger app. The official app of Mi Band, i.e., the Mi Fit app, did support data export, but the exported files contained no data. We eventually used a third-party app called the Notify&Fitness app to export the data of the Mi Band. Once all the data became available, they emailed us the data using a given email account. These data are heterogeneous in nature and vary in resolution. The time series data were segmented and labelled based on the start and the end of activity events logged using the aTimeLogger app. We not only considered the multimodal sensory data during an event but also involved the data one hour before and after the event, because these data may contain important information for activity recognition. For example, the blood glucose level fluctuates before and after a meal. Data quality has long been an issue with consumer wearable devices. Noise may originate from device sensors and human factors [
8] and is unavoidable when collecting data using consumer wearable devices. The dataset was cleaned according to the following procedure. First, we removed or corrected conflicting data by averaging overall all available values and used the average as the final value for the timestamp. Wrong values of step by the Mi Band were replaced by the average of the previous and the following data points. Missing data were handled using the following rules. It is worth noting that the Rule 2 is meant to deal with very short events. For example, if there is no Fitbit step data during an event, and the event only lasts 3 min, the event will be kept if the maximum time difference between two timestamps in the Fitbit step data is no longer than 30 min.
If an event is longer than a minimum duration and the maximum time difference between two timestamps is longer than a certain threshold during the event or one hour before and after the event, this event will be removed.
If an event is shorter than the minimum duration, the event is kept if the maximum time difference between two timestamps is shorter than the corresponding threshold during the one hour before and after the event.
If an activity lasts longer than 4 h (e.g., sleep) and the missing data in corresponding sensor readings (e.g., step, heart rate, glucose level) is longer than 1.5 h, this activity will be removed.
The second step in data cleaning was to convert the data of different devices into a unified format. The data that we obtained are heterogeneous in nature. The data exported from the Mi Band and the FreeStyle Libre glucose sensor are in the format of csv files, while Fitbit data are in JSON format. We created a python script to parse the JSON data of the Fitbit. In addition, we transformed the step data of the Mi Band from accumulative to interval-wise, so that the format of step data from both wristbands became consistent.
2.3. Feature Construction and Model Training
The tsfresh Python library was applied and computed 3815 features. The missing values in the feature set were replaced by the means of their corresponding features. Random forest was applied to select the features of high discriminative power from the full feature set. Features were selected based on the Gini impurity. Eventually, 1078 features were selected. We observed that the top 20 most important features were all derived from the glucose data. Among the selected features, there were more features related to heart rate than steps. More Fitbit-derived features than Mi Band-derived features were selected; this tendency is especially strong for step data. These observations suggest that the wearable glucose sensor FreeStyle Libre and the Fitbit may be more effective than the Mi Band in detecting eating events. We converted the ground truth of activity events logged using aTimeLogger to two categories: eating events (denoted as 1) and non-eating events (denoted as 0). Eating events include both main meals and snacks, and non-eating events include all other activities such as walking, cooking, teeth brushing, and sleeping.
We applied two tree-based ensemble learning algorithms: random forest (RF) and extreme gradient boosting tree (XGBoost). Since non-eating events significantly outnumber eating events, we applied three resampling techniques—random up sampling, random down sampling, and SMOTE resampling—to balance the two classes. These resampling techniques have been shown to improve classifier accuracy in health-related applications [
9]. We adopted the leave-one-out cross-validation strategy for the evaluation of model performance [
10]. Model performance was evaluated using multiple measures, including accuracy, sensitivity, specificity, precision, F1-score, and Matthew’s correlation coefficient (MCC).
3. Results
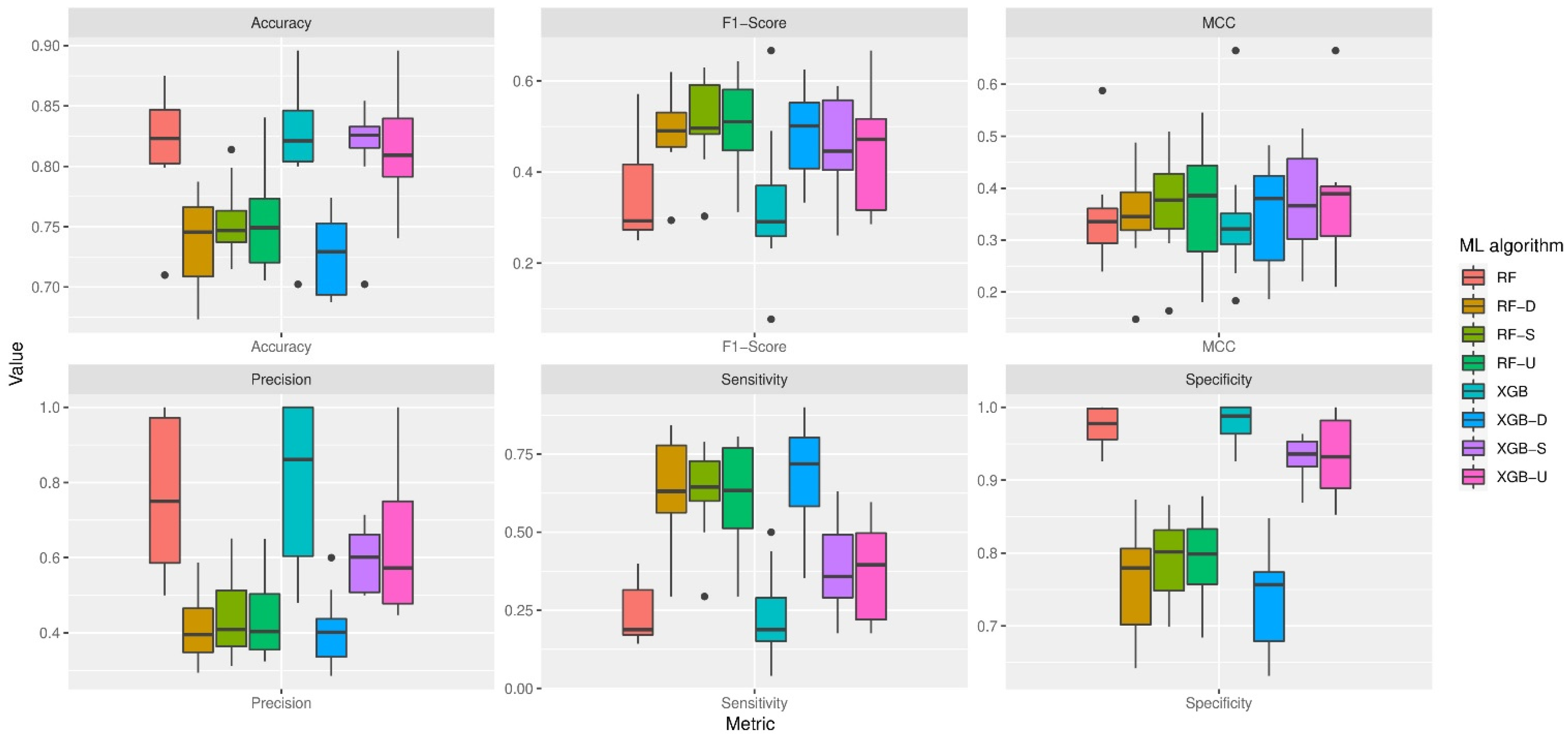
The collected dataset consists of in total 1361 activity events. The number of events from each participant varied from 48 to 342. The combinations of different machine learning algorithms and resampling techniques resulted in eight classification models. The distribution of model performance on each evaluation measure is illustrated in
Figure 1. The results show that without resampling, the performance of XGBoost was statistically similar to random forest. With proper resampling, random forest could achieve slightly better overall performance than XGBoost.
Resampling has a big impact on model performance. In general, resampling techniques significantly improved sensitivity (i.e., the accuracy in detecting eating events) and F1-score, but at the sacrifice of precision and specificity (i.e., the accuracy in detecting non-eating events). On average, XGBoost with down sampling (i.e., XGB-D) achieved the highest sensitivity of 0.67, which is a 180% increase compared to the baseline models (i.e., RF and XGB). Nevertheless, the specificity of XGB-D was reduced by 23% compared to the baseline models. Overall, XGB-D improved F1-score by 42% but did not significantly improve MCC. In contrast, although XGBoost with up sampling or SMOTE sampling (i.e., XGB-U and XGB-S) only achieved a 59% and 66% increase, respectively, in sensitivity, their performance on specificity was statistically similar to that of baseline models. It is also shown that up sampling and SMOTE sampling work better than down sampling for random forest. Distribution-wise, RF-S and RF-U achieved similar sensitivity as XGB-D, and with better specificity, F1-score, and MCC. This indicates that RF-S and RF-U have more balanced performance on both eating events and non-eating events. In addition, the performance of RF-S exhibited smaller variations than that of RF-U.
4. Discussion
We have presented the performance of the eight models with different combinations of machine learning algorithms and resampling techniques. We found that random forest with SMOTE resampling has the best overall performance in all models. Previous studies using sensors exclusively developed for meal detection have reached accuracy between 0.73 and 0.83, precision between 0.78 and 0.91, and sensitivity between 0.74 and 0.93 [
3,
4,
5]. The performance of our method is comparable to previous studies in terms of accuracy and precision but with slightly reduced sensitivity. The possible reasons for the gap could be that (1) many of the previous studies were conducted in a controlled environment, and thus the collected data were less noisy compared to the data collected in the free-living environment in our study, and (2) previous studies used more dedicated devices that allowed the access to raw signals, which were more informative to the models than the processed data that we retrieved from the consumer devices in our study. Despite the slightly compromised performance, our study made a new contribution to the field by demonstrating the feasibility of automatic detection of eating events using widely available consumer devices that are affordable and non-invasive. We also found that random forest is not as sensitive as XGBoost to the type of resampling technique applied. Both machine learning algorithms have the tendency of increasing sensitivity while reducing specificity. That said, XGBoost with up sampling and SMOTE sampling did not sacrifice specificity for enhanced sensitivity, although their improvement in sensitivity were not as significant as the other models. One interpretation is that XGB with up sampling and smote sampling has biased performance over non-eating events (sensitivity = 0.38–0.40, specificity = 0.93), while other models have more balanced performance on both eating and non-eating events (sensitivity = 0.62–0.69, specificity = 0.74–0.79).
The current study has the following limitations. Firstly, we assumed that the start and end of an event was known, whereas in practice that information would not be readily available. The sliding window approach would be more realistic than the event-driven method adopted in this study. Secondly, the current method requires using three sensors to collect multimodal data, which may be too demanding in daily life. It would be interesting to investigate the model performance with reduced modality of input data. We will focus on addressing these limitations in our future work.
Author Contributions
Conceptualization, Z.L.; methodology, Z.L.; software, L.B. and N.C.-M.; validation, L.B. and N.C.-M.; formal analysis, L.B. and N.C.-M.; investigation, Z.L., L.B. and N.C.-M.; resources, Z.L.; data curation, L.B. and N.C.-M.; writing—original draft preparation, Z.L., L.B. and N.C.-M.; writing—review and editing, Z.L.; visualization, Z.L., L.B. and N.C.-M.; supervision, Z.L.; project ad-ministration, Z.L.; funding acquisition, Z.L. All authors have read and agreed to the published version of the manuscript.
Funding
This research was funded by the Japan Society for the Promotion of Science (JSPS) grant numbers 16H07469, 19K20141, and 21K17670.
Institutional Review Board Statement
The study was conducted according to the guidelines of the Declaration of Helsinki and approved by the Ethics Committee of Kyoto University of Advanced Science.
Informed Consent Statement
Written informed consent was obtained from all subjects involved in the study.
Data Availability Statement
The datasets generated during the current study are available from the corresponding author on reasonable request.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Garaulet, M.; Gómez-Abellán, P.; Alburquerque-Béja, J.J.; Lee, Y.C.; Ordovás, J.M. Timing of food intake predicts weight loss effectiveness. Int. J. Obes. 2013, 37, 604–611. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Gabel, K.; Hoddy, K.K.; Haggerty, N.; Song, J.; Kroeger, C.M.; Trepanowski, J.F.; Panda, S.; Varady, K.A. Effects of 8-h time restricted feeding on body weight and metabolic disease risk factors in obese adults: A pilot study. Nutr. Healthy Aging 2018, 4, 345–353. [Google Scholar] [CrossRef] [PubMed]
- Bi, C.; Xing, G.; Hao, T.; Huh, J.; Peng, W.; Ma, M. FamilyLog: Monitoring Family Mealtime Activities by Mobile Devices. IEEE Trans. Mob. Comput. 2020, 19, 1818–1830. [Google Scholar] [CrossRef]
- Bi, S.; Wang, T.; Tobias, N.; Nordrum, J.; Wang, S.; Halvorsen, G.; Sen, S.; Peterson, R.; Odame, K.; Caine, K. Auracle: Detecting Eating Episodes with an Ear-mounted Sensor. Proc. ACM Interact. Mob. Wearable Ubiquitous Technol. 2018, 2, 1–27. [Google Scholar] [CrossRef]
- Papapanagiotou, V.; Diou, C.; Zhou, L.; van den Boer, J.; Mars, M.; Delopoulos, A. A Novel Chewing Detection System Based on PPG, Audio, and Accelerometry. IEEE J. Biomed. Health Inform. 2017, 21, 607–618. [Google Scholar] [CrossRef] [PubMed]
- Liang, Z.; Chapa-Martell, M.A. Accuracy of Fitbit wristbands in measuring sleep stage transitions and the effect of user-specific factors. JMIR Mhealth Uhealth 2019, 7, e13384. [Google Scholar] [CrossRef] [PubMed]
- Liang, Z.; Chapa-Martell, M.A. Validity of consumer activity wristbands and wearable EEG for measuring overall sleep parameters and sleep structure in free-living conditions. J. Healthc. Inf. Res. 2018, 2, 152–178. [Google Scholar] [CrossRef]
- Liang, Z.; Ploderer, B.; Chapa-Martell, M.A. Is fitbit fit for sleep-tracking? Sources of measurement errors and proposed countermeasures. In Proceedings of the 11th EAI International Conference on Pervasive Computing Technologies for Healthcare, Barcelona, Spain, 23–26 May 2017; pp. 476–479. [Google Scholar]
- Liang, Z.; Chapa-Martell, M.A. Achieving accurate ubiquitous sleep sensing with consumer wearable activity wristbands using multi-class imbalanced classification. In Proceedings of the 2019 IEEE Intl Conf on Dependable, Autonomic and Secure Computing, Intl Conf on Pervasive Intelligence and Computing, Intl Conf on Cloud and Big Data Computing, Intl Conf on Cyber Science and Technology Congress (DASC/PiCom/CBDCom/CyberSciTech), Fukuoka, Japan, 5–8 August 2019; pp. 768–775. [Google Scholar] [CrossRef]
- Liang, Z.; Chapa-Martell, M.A. A two-stage imbalanced learning method to sleep stage classification using consumer activity trackers. In Proceedings of the 13th International Conference on Health Informatics, Valletta, Malta, 24–26 February 2020. [Google Scholar]
| Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).