2.1. Critical Analysis of Epidemiological Data at the Regional Level
The Italian regions of Lombardy, Emilia Romagna, Valle d’Aosta, and Veneto have been herein selected since they were among the regions with the largest spread of the contagion. Moreover, they experienced different lockdown measures over time, had different healthcare policies, different initial dates of outbreaks, and are also characterized by different population sizes. Data from the beginning of Covid-19 epidemic in Italy (day 1 corresponds to the 24th of February 2020) till the 30th of April 2020 (last day of phase 1, the so-called Italian lockdown) were considered. Those data are publicly available on the portal of the Italian Civil Protection Centre [
12].
Lombardy has a population size of about 10.4 million inhabitants and was the epicenter of the first major outbreak in Italy. The first lockdown began around the 21st of February 2020, covering eleven municipalities of the province of Lodi that were included in the so-called “red zone” (quarantine zone). On the 8th of March, Italian Prime Minister Conte announced the expansion of the quarantine zone to cover much of northern Italy. With that decree, the initial “red zone” was also abolished, though the municipalities were still within the quarantined area. The locked down area, as of 8 March, covered the entirety of the region of Lombardy, in addition to 14 provinces in Piemonte, Veneto, Emilia-Romagna, and Marche. On the evening of 9 March, the quarantine measures were expanded to the entire country, coming into effect the next day. Conte announced on 11 March that the lockdown would be tightened, with all commercial and retail businesses, except those providing essential services, like grocery stores, food stores, and pharmacies, closed down. Such lockdown measures were prolongated till 4 May. Hence, Lombardy was subject to all the above lockdown measures of increasing intensity.
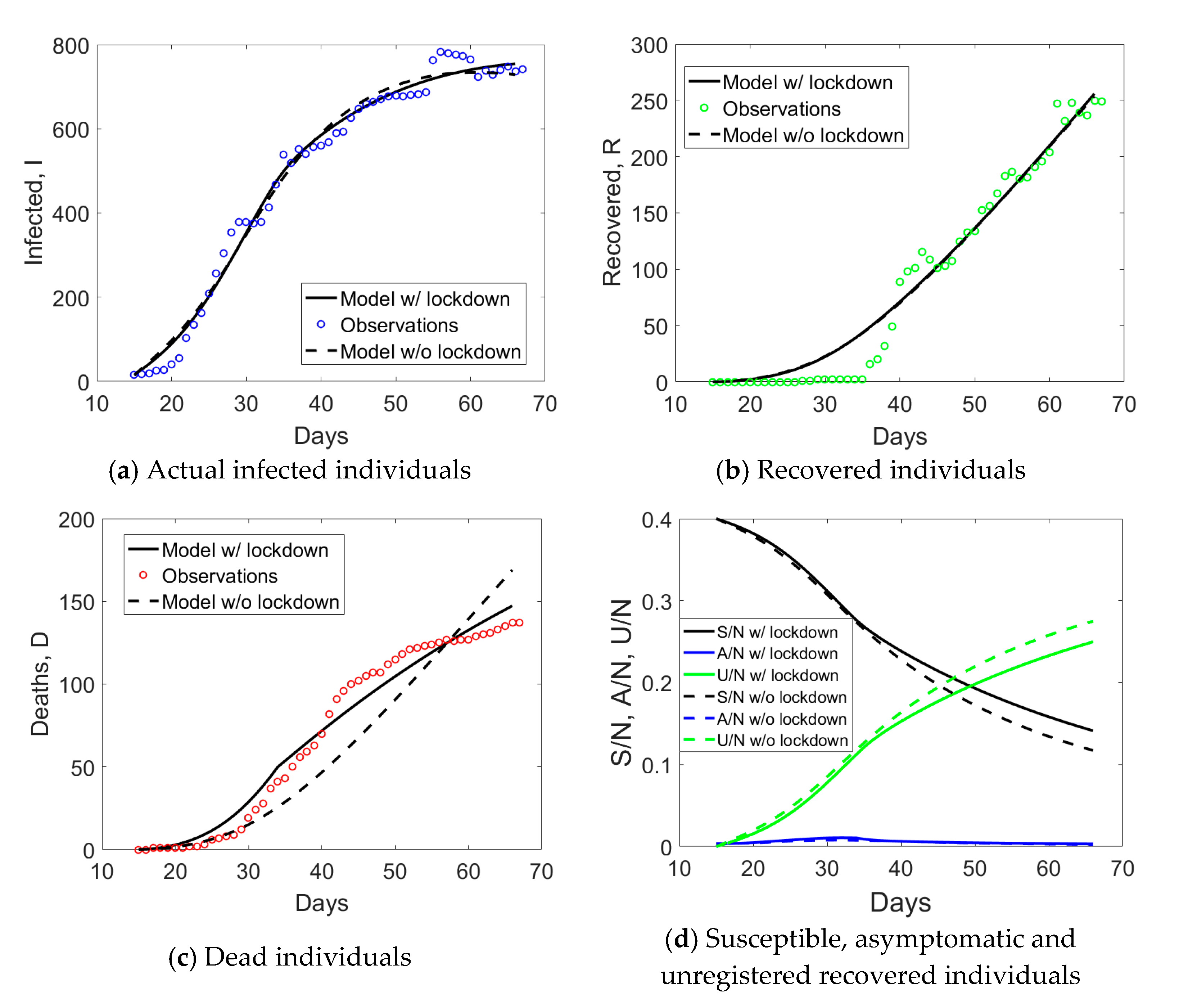
The observed data for the
I (actual infected),
R (recovered), and
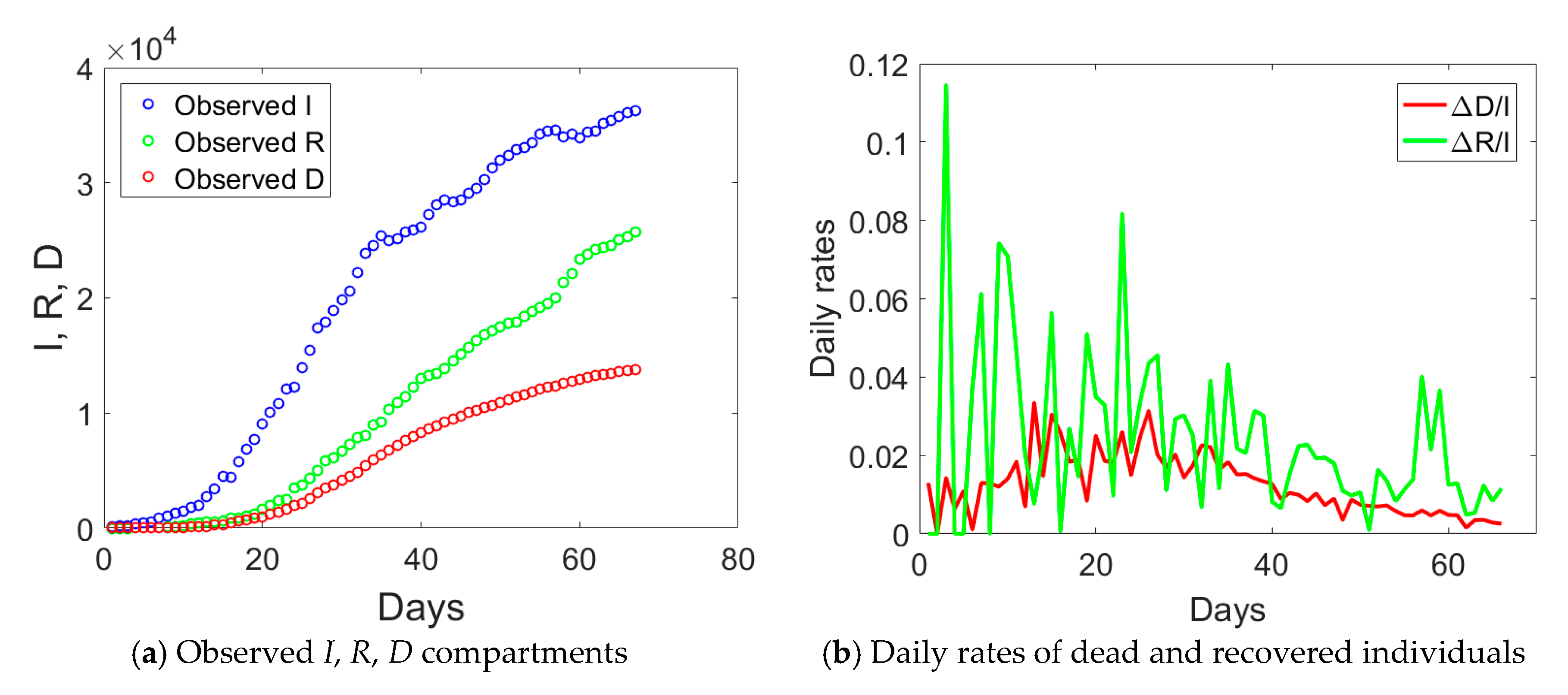
D (dead individuals) compartments for Lombardy are shown in
Figure 1a. The curve corresponding to actual infected individuals (blue circles) had an initial exponential growth, subsequently slowed down after day no. 18, 21 days after the implementation of the first lockdown to the province of Lodi. Another change of slope occurred at day no. 34, again 21 days from the second lockdown that was extending the red zone to the whole Lombardy. At the end of phase 1, the peak of actual infectious individuals was not yet reached. Daily rates Δ
D/
I and Δ
R/
I (Δ stays for the difference between compartmental values over two consecutive days) computed from
Figure 1a are plotted in
Figure 1b and show some distinct trends: an initial increasing one till day no. 18, a plateau, and then a progressive decay over time.
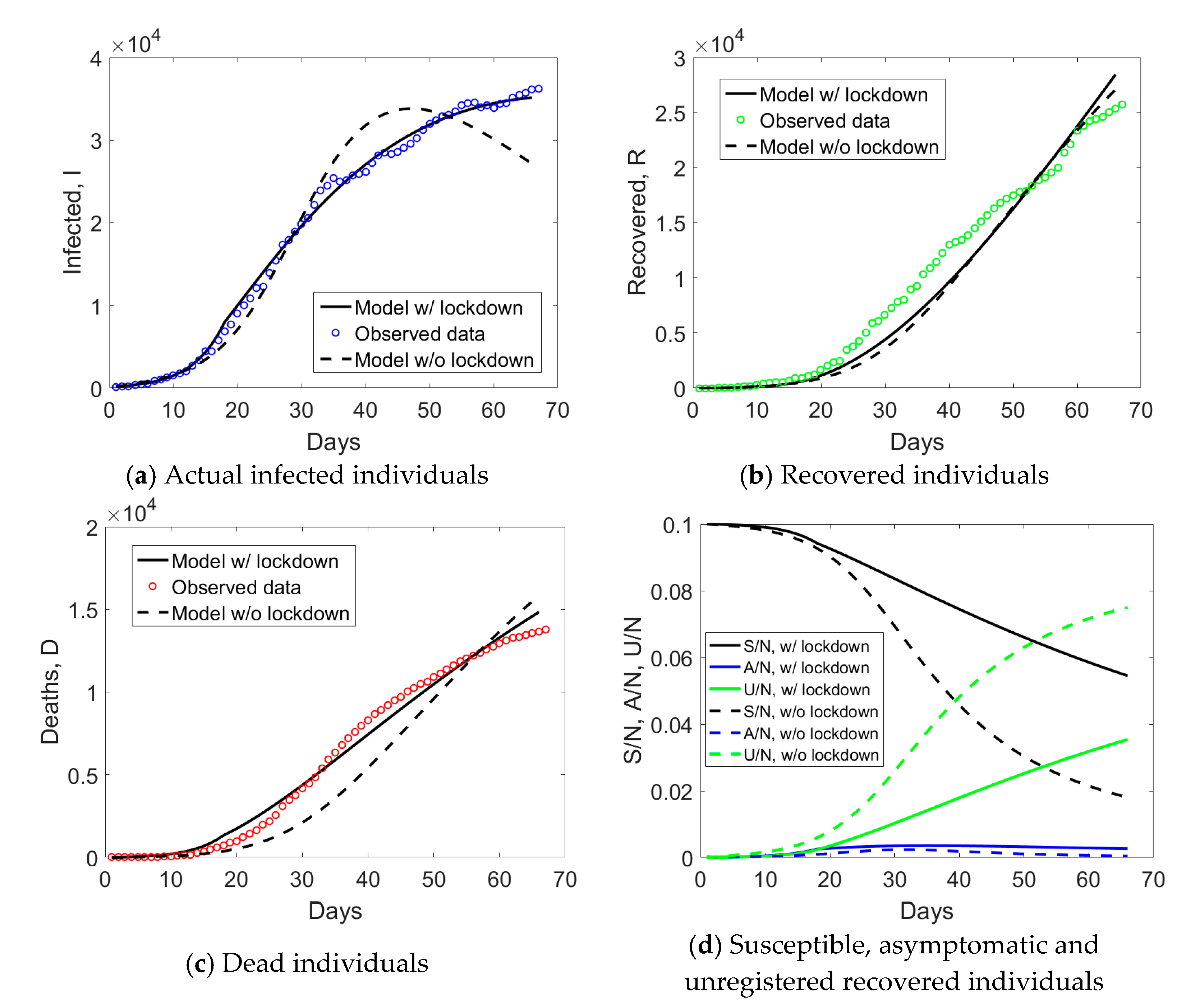
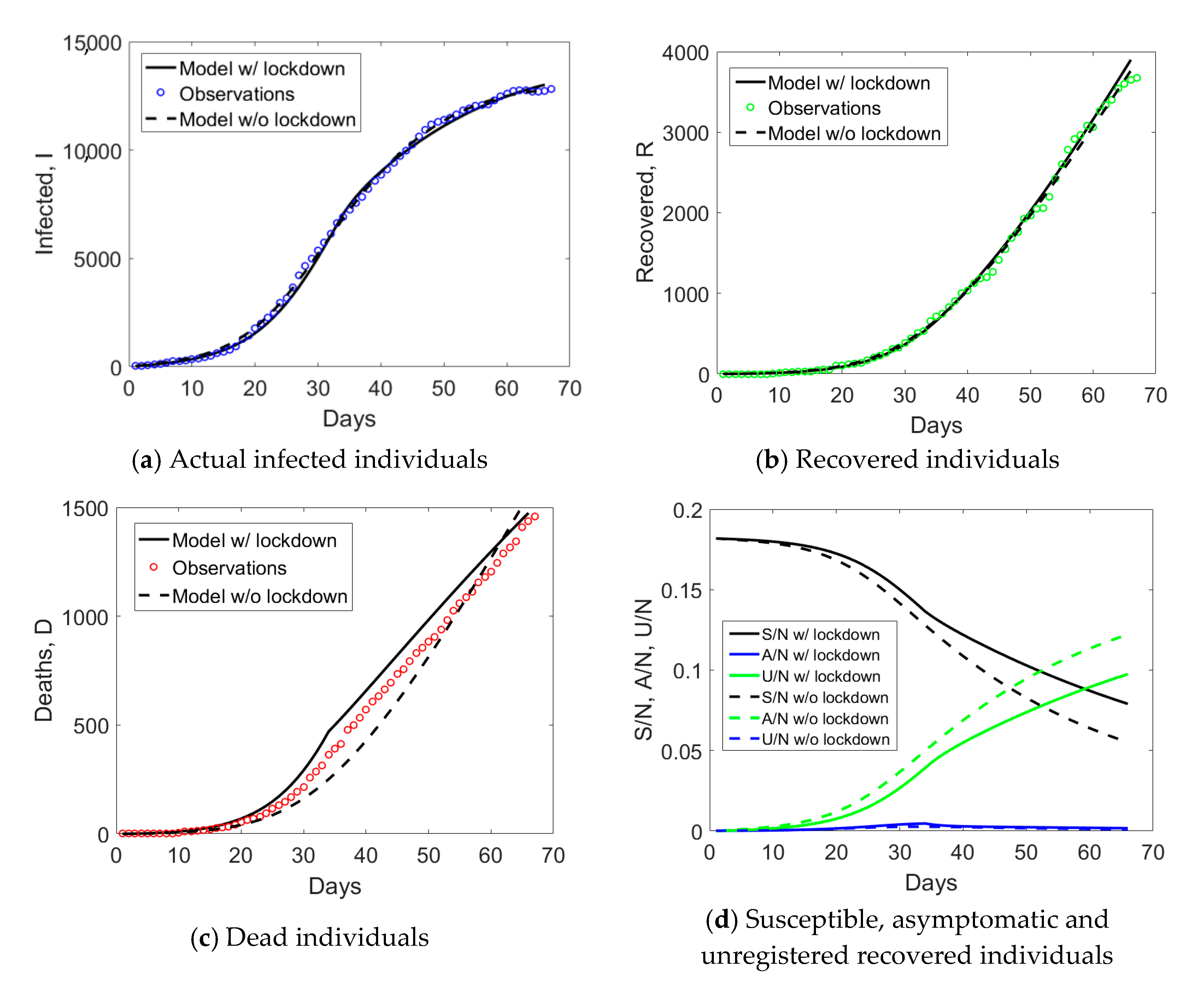
Emilia Romagna has a population size of 4.459 million inhabitants and saw the provinces of Modena, Parma, Piacenza, and Reggio Emilia included in the red zone on the 8th of March. Such an intermediate measure was very short in time, since it was followed by the Italian lockdown on the 9th of March. If we examine the observed
I,
R, and
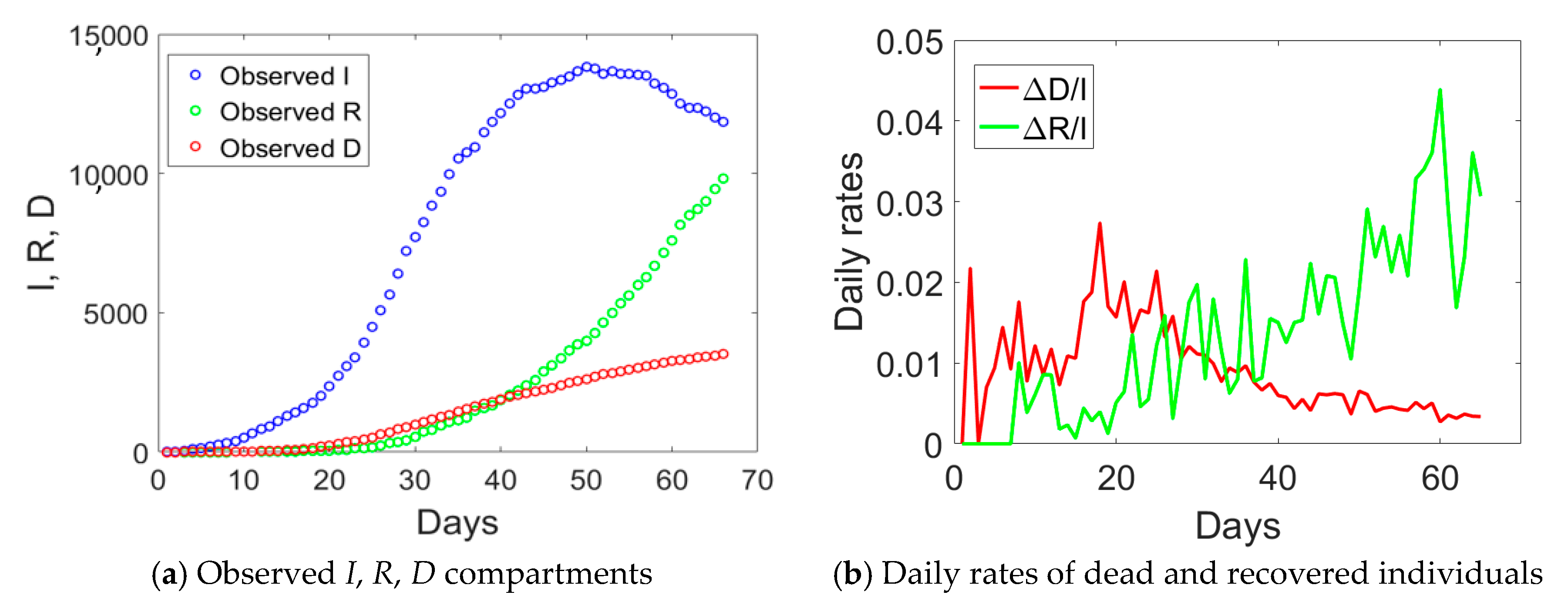
D data for Emilia Romagna in
Figure 2a, one notices an evident change of curvature at day no. 34, about 21 days from the day when lockdown measures took place. The curve corresponding to the actual infected individuals (blue circles) reached the peak at day no. 50. Daily rates Δ
D/
I and Δ
R/
I computed from
Figure 2a are plotted in
Figure 2b. The daily rate of deaths, Δ
D/
I, had an initial increasing trend followed by a decay over time, similarly to Lombardy. The absence of a plateau could be probably related to the fact that Emilia Romagna was subject to two lockdown measures with just one day of time lag between them, rather than 16 days as in Lombardy. The daily rate of recovered individuals shows an increasing trend over time, opposite to the situation observed in Lombardy.
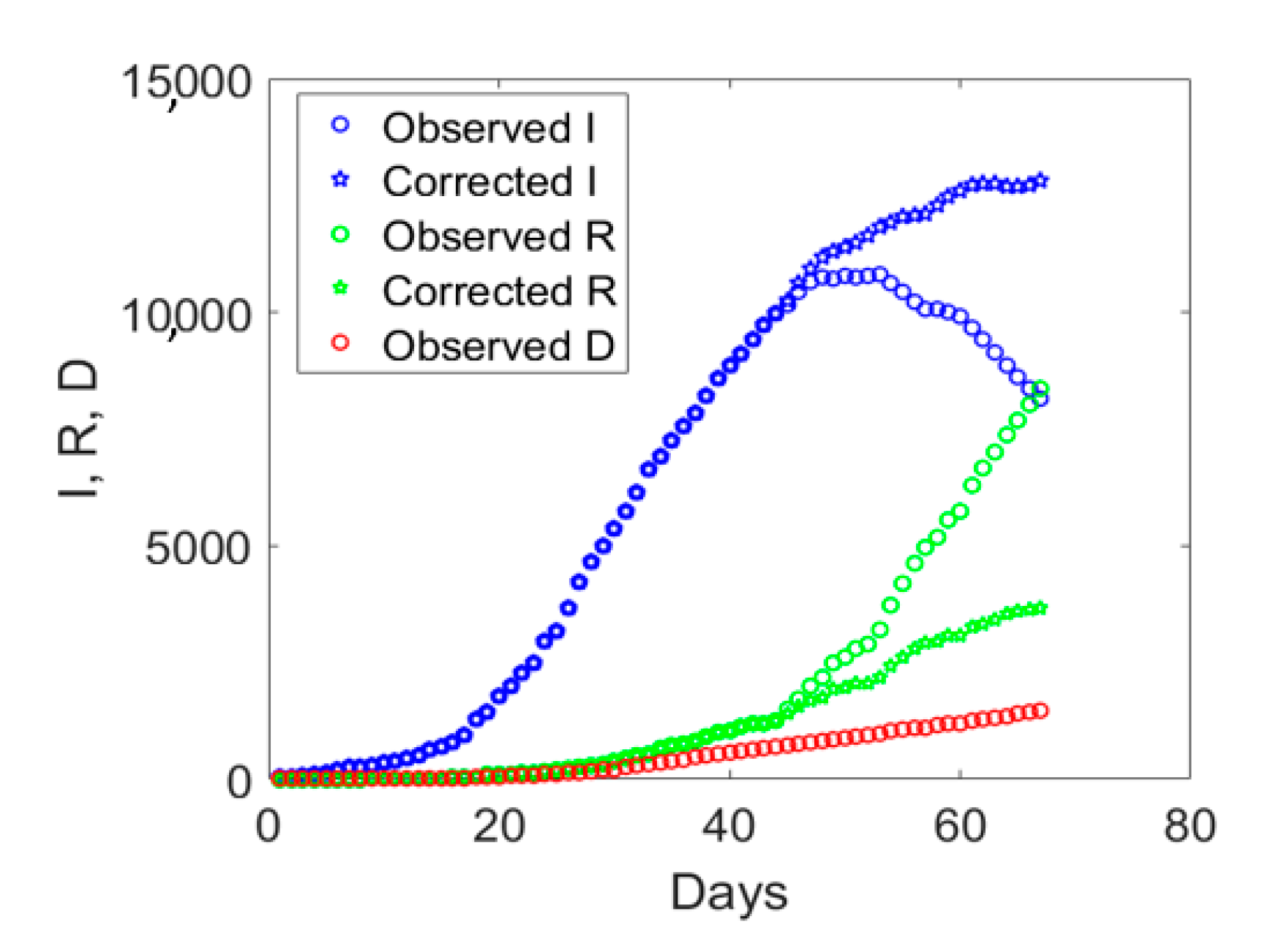
Valle d’Aosta is the smallest Italian region with only 125,666 inhabitants. It was subject to the Italian lockdown taking place on the 9th of March and experienced the largest number of Covid-19 deaths per population.
If we examine the observed
I,
R, and
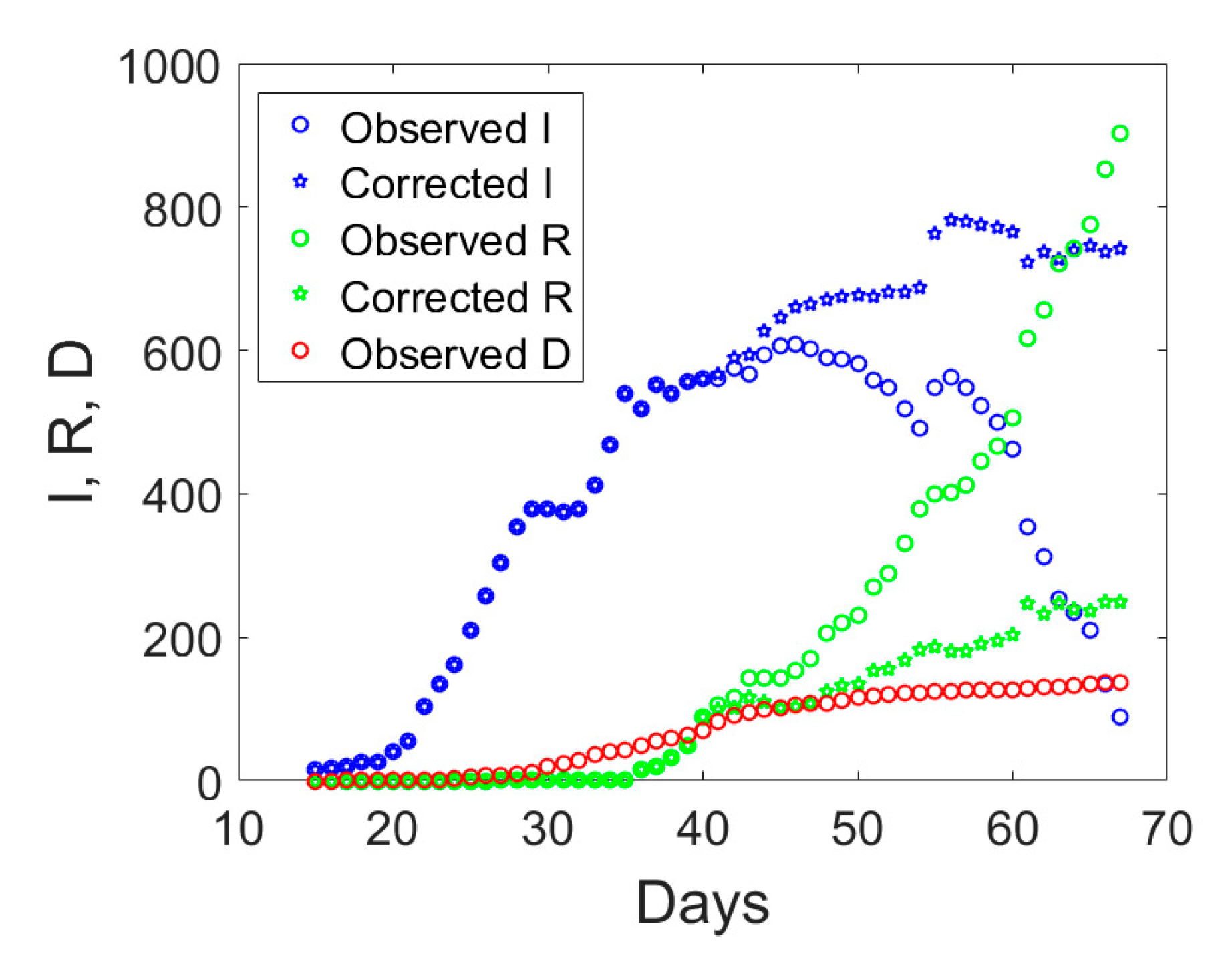
D data displayed with circles in
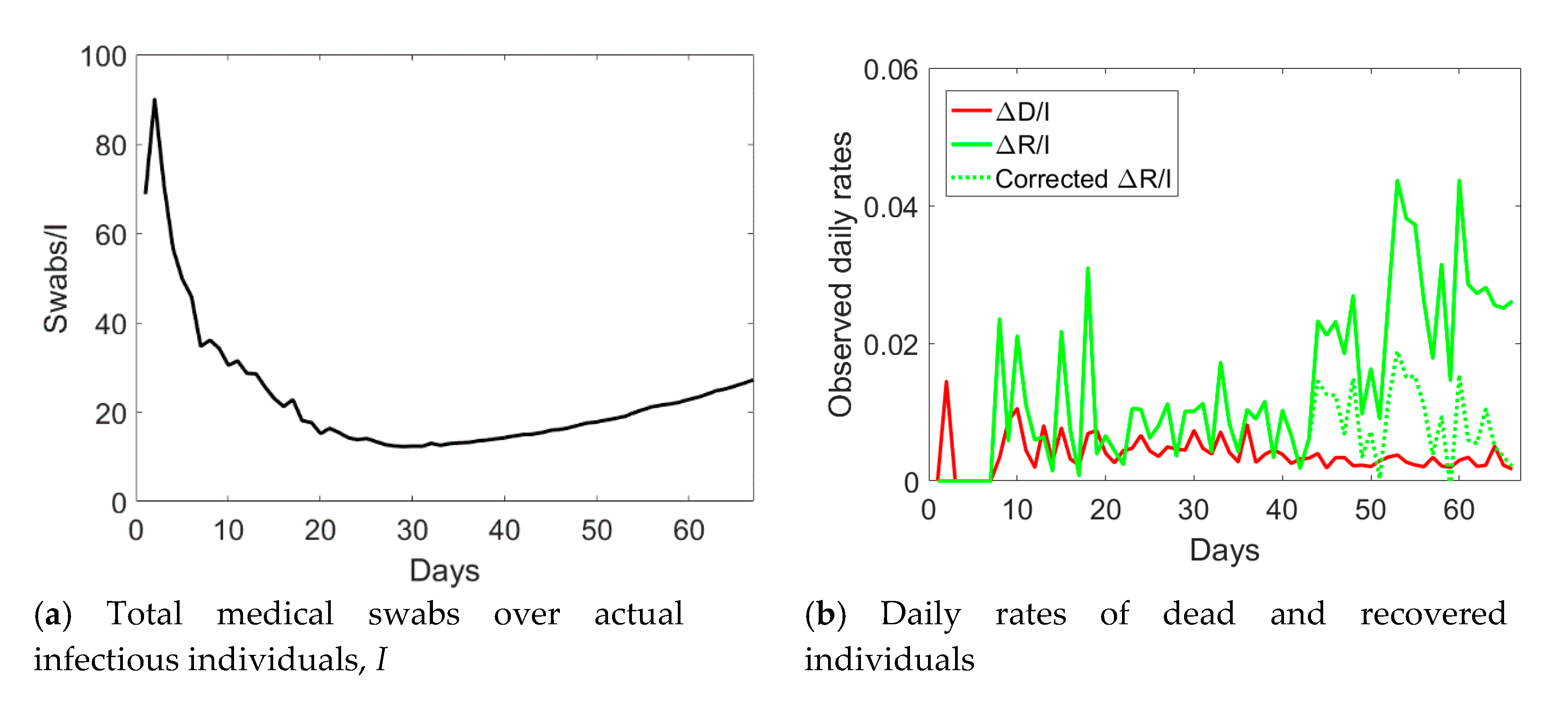
Figure 3, one notices an abrupt sudden increase in recovered individuals after day no. 40. The time series of the actual infected individuals presented also sudden jumps at days no. 40 and 55. To understand the possible reason for such a lack of smoothness in the observed data, one could examine the ratio between the cumulative number of medical swabs and the actual number of infectious individuals,
Sw/
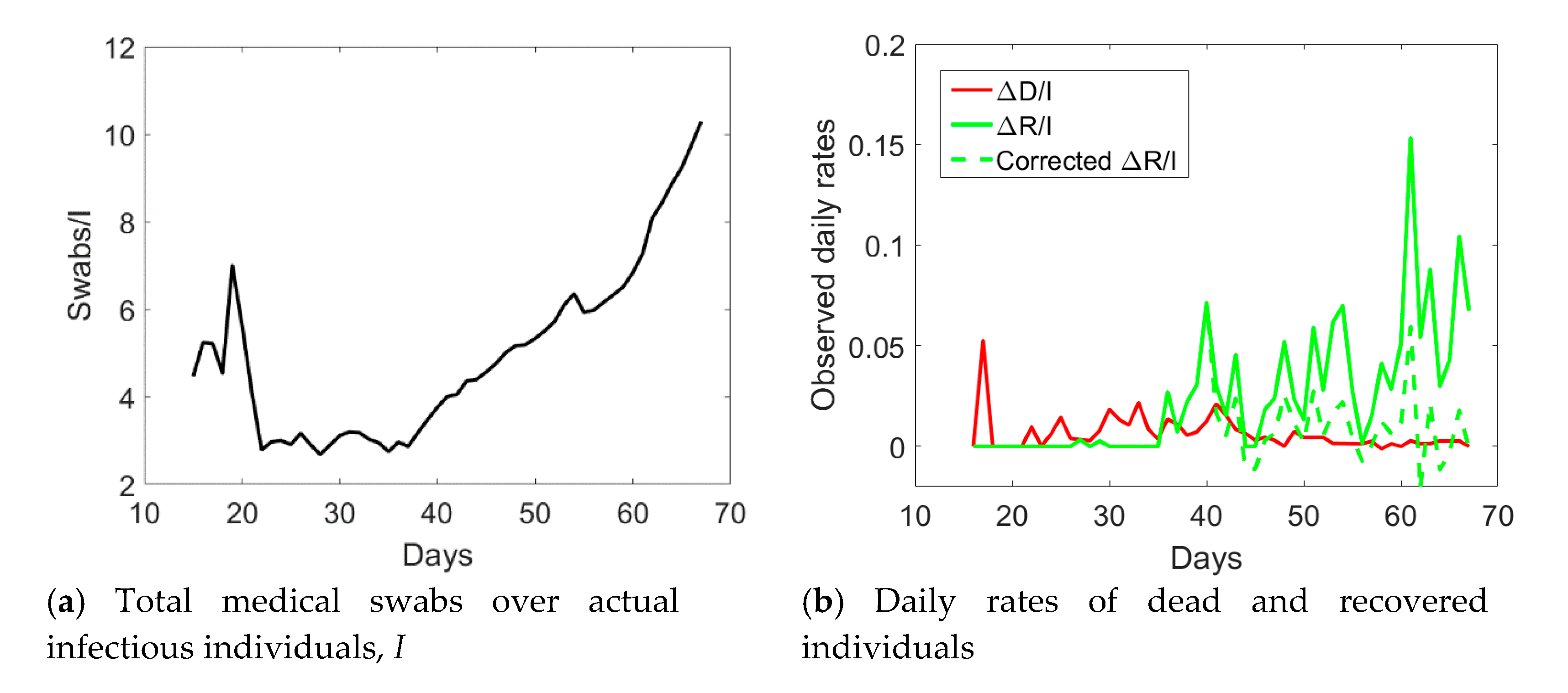
I (see
Figure 4a). Such a ratio was almost constant and reached a minimum at day no. 40. Afterwards, it linearly increased over time with a discontinuity at day no. 55. One could reasonably assume that when the quantity
Sw/
I was at the minimum at the early stage of the epidemic, swab tests were in a number almost strictly necessary to identify infected people claiming Covid-19 symptoms. Later, the increase in the use of medical swabs also to randomly test the population was likely leading to the identification of not only symptomatic, but also asymptomatic infected individuals. Since the open data in the compartment
I include all the confirmed positive cases to Covid-19, regardless of their symptoms, one could expect that the observed data might be biased by the number of medical swabs. Overall, the number of asymptomatic cases is expected to increase as long as medical swabs are used to massively test the population even in absence of reported Covid-19 symptoms. This bias could affect the number of recovered individuals as well, which inevitably sums up the number of symptomatic and asymptomatic recovered patients, which could explain the sudden increase in the daily rate Δ
R/
I after day no. 40.
As a possible empirical correction to the observed data to retain only symptomatic cases in the
I compartment, one could multiply the number of recovered individuals by the ratio between the number of medical swabs divided by the number of medical swabs at day no. 40, when
Sw/
I was minimum in the time series. Since this ratio is less than unity, the corrected number of recovered individuals will diminish as compared to the reported one. The actual number of asymptomatic infected individuals,
I, will be computed as the total number of reported cases minus the number of deaths and the corrected number of recovered individuals. Hence, the actual number of symptomatic infected individuals would increase as compared to the observed data which include also some asymptomatic ones. The application of this correction to the data of Valle d’Aosta leads to the corrected time series shown in
Figure 3 with pentagrams. The daily rate of the corrected recovered individuals shown in
Figure 4b with a dashed green line is now oscillating around a constant average value, without the anomalous fast increasing trend (solid green line) observed from the reported observations.
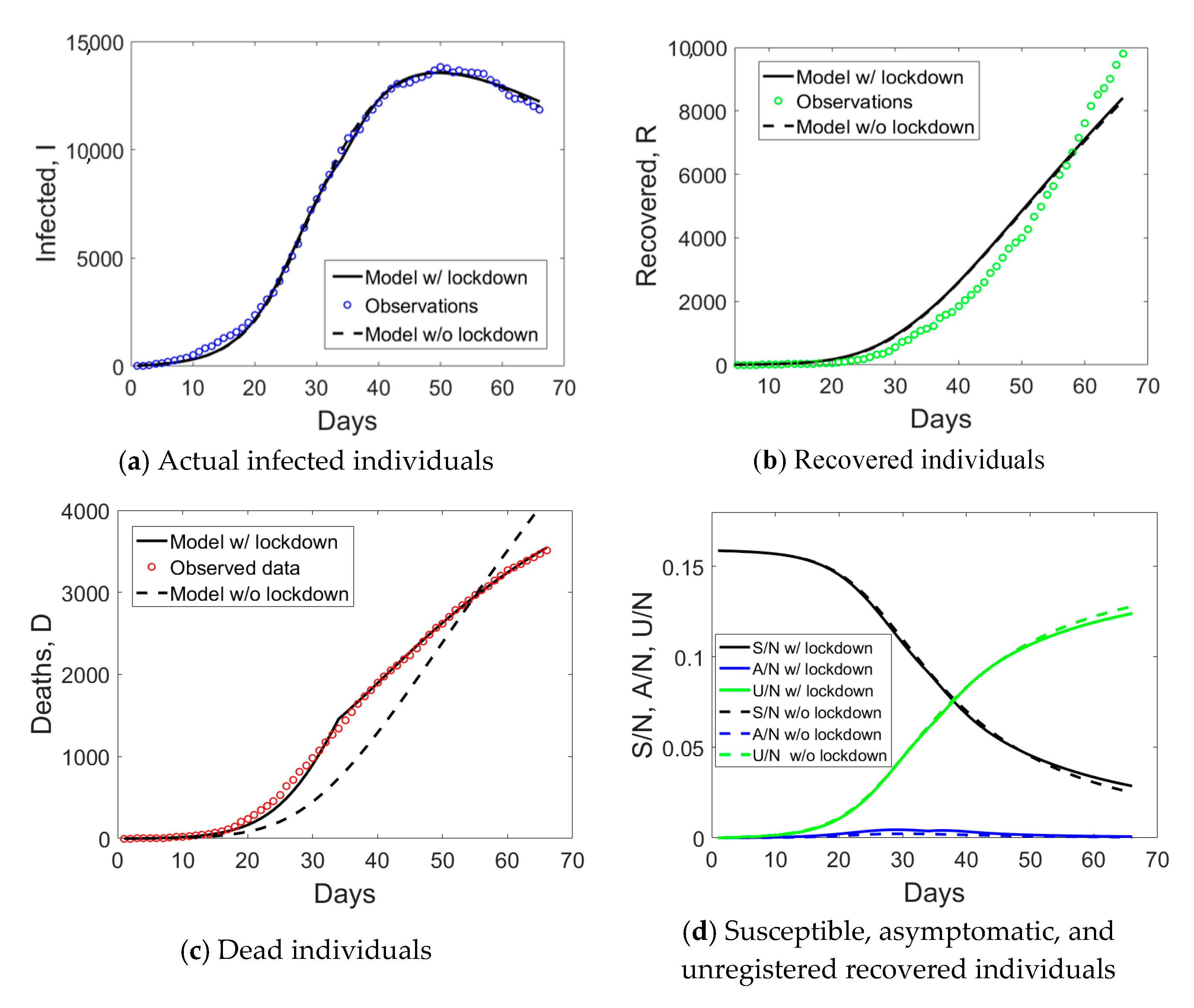
Veneto has a population size of 4.9 million inhabitants. The provinces of Padua, Treviso, and Venice were subject to the lockdown on the 8th of March, followed by the Italian lockdown the day after. As compared to Lombardy, where hospitalization was the major healthcare solution, Veneto largely implemented home treatment. Moreover, Veneto was the only region able to purchase laboratory equipment for automatic testing of up to 9000 medical swabs per day. Such an infrastructure was announced to be operative since the 7th of April 2020, which corresponds to day no. 44 of the time series.
The augmented testing capabilities resulted in an increased number of observed recovered individuals and a fast decay of actual infected people after day no. 44 (see
Figure 5). Correspondingly, the ratio
Sw/
I increased from its minimum value (see
Figure 6a). The application of the correction to the data of Veneto, as previously done for Valle d’Aosta, but considering the number of medical swabs at day no. 44 as a reference, led to the corrected data shown with pentagrams in
Figure 5. Such an empirical correction to the actual infected individuals to remove the asymptomatic ones is noteworthy, since the corrected time series shows that the peak was not yet reached in this region before the start of phase 2.
The corrected daily rate of recovered individuals is shown in
Figure 6b with a green dashed line. As for Valle d’Aosta, the correction leads to a steady rate over time, rather than the anomalous growth reported (green solid line). The rate of dead individuals decreased over time after day no. 34, in line with the trend reported for the other regions.
2.2. An Epidemiological Model for Covid-19 Including Asymptomatic and Lockdown Effects to Interpret Observed Data
SIR epidemiological models have been the subject of intensive research which led to several variants to the basic model including only three compartments. Here, we refer to the model proposed elsewhere [
11] to deal with the large number of asymptomatic individuals for Covid-19 epidemic in Italy. This includes two additional compartments representing asymptomatic infected (
A) and unregistered recovered (
U) individuals (i.e., the recovered asymptomatic patients). In the model, permanent immunity of individuals who were infected and recovered is postulated, which is realistic enough since few cases of repeated infections were reported during phase 1. Moreover, the compartment of dead individuals is also herein included to model living dynamics. As another major variant from the original model [
11], for its application at the regional level, the population size (
N) is split into two parts: one of initial susceptible individuals (
N0), that are involved into the epidemic dynamics, and the other, of numerosity (
N −
N0), composed of people excluded from the Covid-19 epidemic thanks to the effect of the lockdown measures. This is motivated by the fact that many categories, such as public servants, started smart working from the emanation of the earliest lockdown measures, significantly reducing the probability of becoming susceptible to contagion.
Susceptible individuals S, who reduced their mobility but still had contacts justified by their urgent job activities, can evolve in one of the two classes of infected and infective people: symptomatic, which should match the corrected value I of reported infected people, and asymptomatic A. People can move from the compartment of symptomatic individuals into two other compartments, one of registered recovered, which should match the observed R data, and another of dead individuals, that should match the observed D data. People in the compartment of asymptomatic individuals, on the other hand, can be removed and pass to the compartment of unregistered recovered, U, which collects individuals passing unnoticed through the infection.
In formulae, the dynamics is described by the following set of nonlinearly coupled ordinary differential equations (ODEs):
Model parameters entering Equations (1)–(6) are represented by the coefficients β, γ, μ, η, and ξ. The effect of lockdown measures is herein modeled by introducing time-dependent parameters β and μ. A decreasing function of the actual number of infected (symptomatic and asymptomatic) individuals is considered for the coefficient β, in line with Tsiotas and Magafas [
14] and other similar treatments of social distancing measures [
15,
16,
17,
18], and a decreasing function for μ is also introduced as noticed from the observed data (see
Section 2.1):
In Equations (7) and (8), tLD denotes the day when lockdown measures are expected to have an effect on epidemic dynamics, which for Covid-19 is about 21 days after the date when each ministerial decree on lockdown measures is published.
The differential model in Equations (1)–(6) are coupled and nonlinear, and it is solved by integrating the ODEs over time according to an explicit time stepping scheme (rates on the left hand side at time
t + Δ
t, where Δ
t is the time interval, are computed from the quantities evaluated at the previous time step,
t), with the smallest time step possible, Δ
t = 1 day. After evaluating the rates of
S,
I,
A,
R,
U, and
D at time
t + Δ
t, the updated states are incrementally computed from the previous ones. Initial conditions have to be specified and are given by
S0 =
N0,
I0 =
I0,
A0 = ξ/(1 − ξ)
I0, R0 =
D0 =
U0 = 0. Here,
I0 is the initial number of infected individuals reported in the open data, while
A0 is estimated based on the probability ξ of becoming symptomatic, consistent with Gaeta [
11].
The application of such a model to the interpretation of observed data poses several challenges. The first major challenge is related to the fact that only
I,
R, and
D compartments are known and, as discussed in
Section 2.1,
I and
R observed data might be influenced by the number of medical swabs. Few medical swabs might underestimate the number of infectious individuals, while a massive testing of the population might lead to an overestimation of the number of
I and
R, including also individuals belonging to asymptomatic compartments,
A and
U. To overcome this issue, for Valle d’Aosta and Veneto, we consider the empirical correction to the observed data as proposed in
Section 2.1. The second challenge is related to model parameters’ identification. The present simplified model includes five parameters (β
0, γ, μ
0, η, and ξ) plus the value of
N0 that have to be simultaneously identified, if lockdown effects are not included. Otherwise, identification should consider seven parameters (β
0, γ, μ
0, η, ξ, ρ
, ρ
m) plus
N0. 2.3. Model Parameters’ Identification Based on PSO
To overcome the complexity related to model parameters’ identification, a machine learning approach based on PSO is herein proposed to automatically identify the whole set of model parameters. PSO belongs to the wide class of genetic algorithms [
19] and it is an efficient heuristic for global optimization over continuous spaces [
20], overcoming many drawbacks of gradient-based optimization methods [
21]. Its computational performance has been carefully analyzed previously (see [
22]) and in the references therein.
PSO is widely used in physics, to predict collective dynamics, and find optimal solutions originating from highly nonlinear collective phenomena (see e.g., [
23,
24,
25,
26,
27,
28]). Such a computational method is used to minimize the mismatch between model predictions and real observations of the individuals belonging to the
I,
R, and
D compartments by iteratively trying to improve a candidate solution. The method is therefore initialized by a population of candidate solutions in the parameter space, which are called particles, and moving those particles around in the search-space according to simple mathematical formulae over the particle’s position and velocity. Each particle’s movement is influenced by its local best-known position, but it is also guided toward the best-known positions in the search-space, which are updated as better positions are found by other particles. This meta-heuristic gradient-free method is generally expected to move the swarm toward the best solution, and it is particularly effective for nonlinear optimization problems. It has also been efficiently applied to parameters estimation of machine learning algorithms (see [
29] and references therein).
The algorithm specialized to the present problem is sketched in Algorithm 1. The particle has coordinates
xi = (β
0i, γ
i, μ
0i, η
i, ξ
i) if lockdown effects are not modeled (the dimension of the search space is therefore 5), while it is given by
xi = (β
0i, γ
i, μ
0i, η
i, ξ
i, ρ
i, ρ
mi) when lockdown is also considered (the dimension of the search space becomes 7). Usually, the method is already performing well for 30–50 particles. Here we set
n = 100, and a higher number had no effect on the quality of the identified optimal solution. In step 1.1 of Algorithm 1, maximum values are set to be enough wide not to constrain the algorithm. For γ and μ
0, their maximum values can be estimated by the observed daily rates of recovered and dead individuals (see
Section 2.1).
The function
f to be minimized was chosen as the norm of the absolute error between model predictions and observations regarding
I,
R, and
D datasets available from the Italian Civil Protection Centre. The termination criterion was set in terms of maximum iterations achieved,
kmax = 1000. A larger number of iterations had no significant effect on the quality of results. Regarding the PSO algorithm control coefficients
w, φ
p and φ
g, they were set as follows:
w = 0.9 − (0.9 − 0.5)
k/
kmax, φ
p = φ
g = 0.5, where
k is an integer defining the iteration number, in line with recommendations for the use of PSO [
30].
| Algorithm 1. Machine learning algorithm for model parameters’ identification. |
- (1)
for each particle xi, i = 1,…,n, do - (1.1)
Initialize a group of random solutions (particles xi) in the feasible domain. They are scattered over the search space as uniformly as possible, with a uniformly distributed random vector whose entries belong to the range from zero to given maximum values, i.e., xi = rand (0; xi,max). - (1.2)
Initialize the particle’s best known position to its initial position pi ← xi - (1.3)
Solve ODEs and compute S(xi, t), I(xi, t), R(xi, t), D(xi, t), A(xi, t), U(xi, t), for t = 1,…,tmax - (1.4)
Compute f(pi) = norm{I(xi,t) − Iexp(t)} + norm{R(xi, t) − Rexp(t)} + norm{D(xi, t) − Dexp(t)} - (1.5)
Find the swarm’s best known position g: f(g) = mini{ f(pi) } - (1.6)
Initialize the particle’s velocity: vi = rand(−xi,max; xi,max)
- (2)
while a termination criterion is not met do - (2.1)
for each particle i = 1,…,n do - (2.1.1)
for each dimension d = 1,…,5 or 7 do Pick random numbers rp = rand(0, 1), rg = rand(0, 1) Update particle’s velocity: vi,d ← ω vi,d + φp rp (pi,d − xi,d) + φg rg (gd − xi,d)
- (2.1.2)
Update particle’s position: xi ← xi + vi - (2.1.3)
iff(xi) < f(pi) then
|
Regarding the estimation of the initial condition N0, this parameter is not included in the identification algorithm. It has been in fact observed that this parameter has a direct effect upon the scaling of the compartments and on the parameter β0. Therefore, it is manually rescaled in order to match the observed basic reproduction number at the beginning of the epidemic when the SIR model predicts an exponential growth (i.e., in order to have R0 = β0/γ~3).