An Integrated Deep Learning Approach for Poultry Disease Detection and Classification Based on Analysis of Chicken Manure Images
Abstract
1. Introduction
2. Materials and Methods
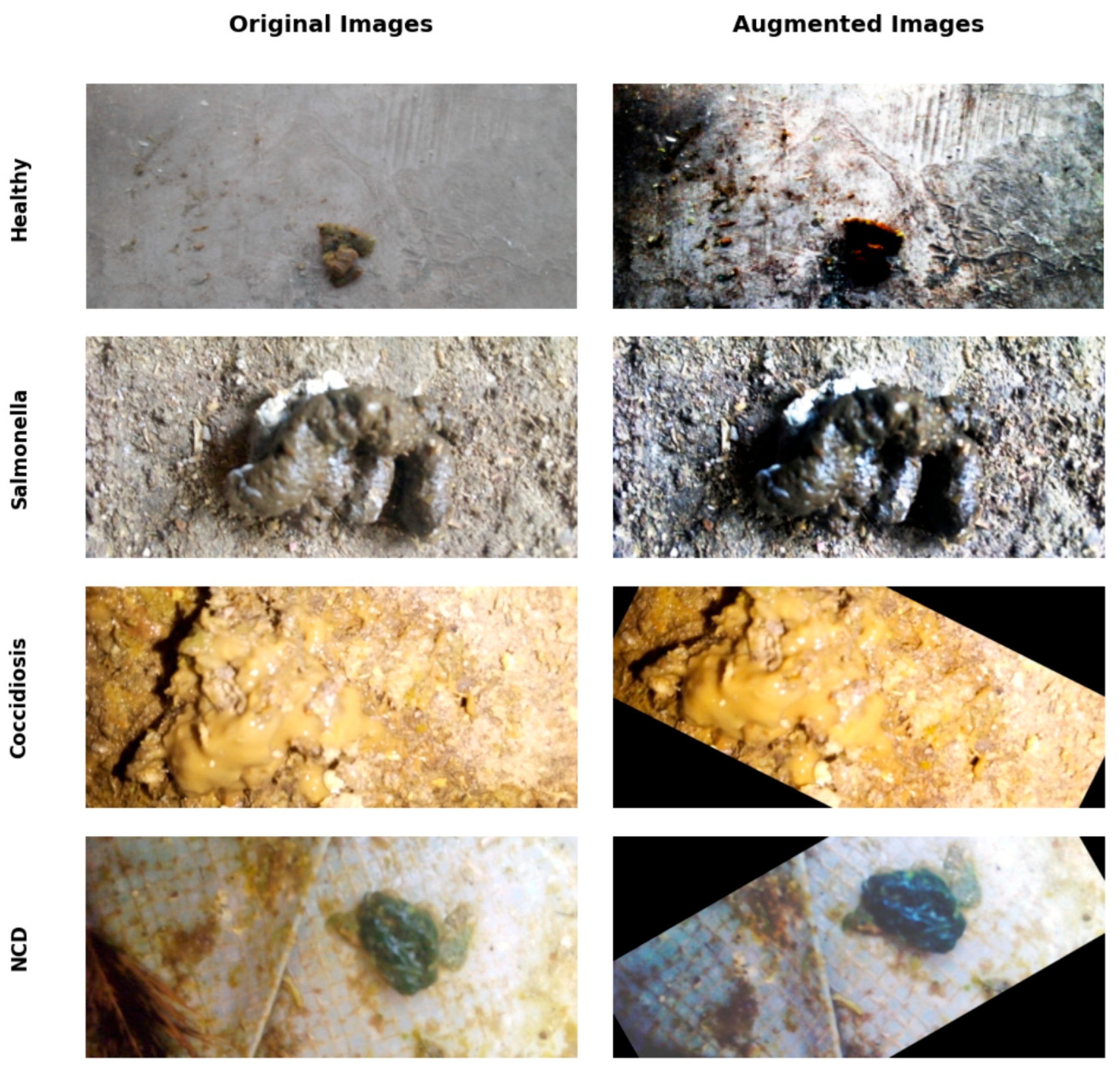
2.1. Dataset and Data Preprocessing
2.2. Model Training and Selection
2.2.1. Detection Model
2.2.2. Classification Model
2.2.3. Model Performance Parameters
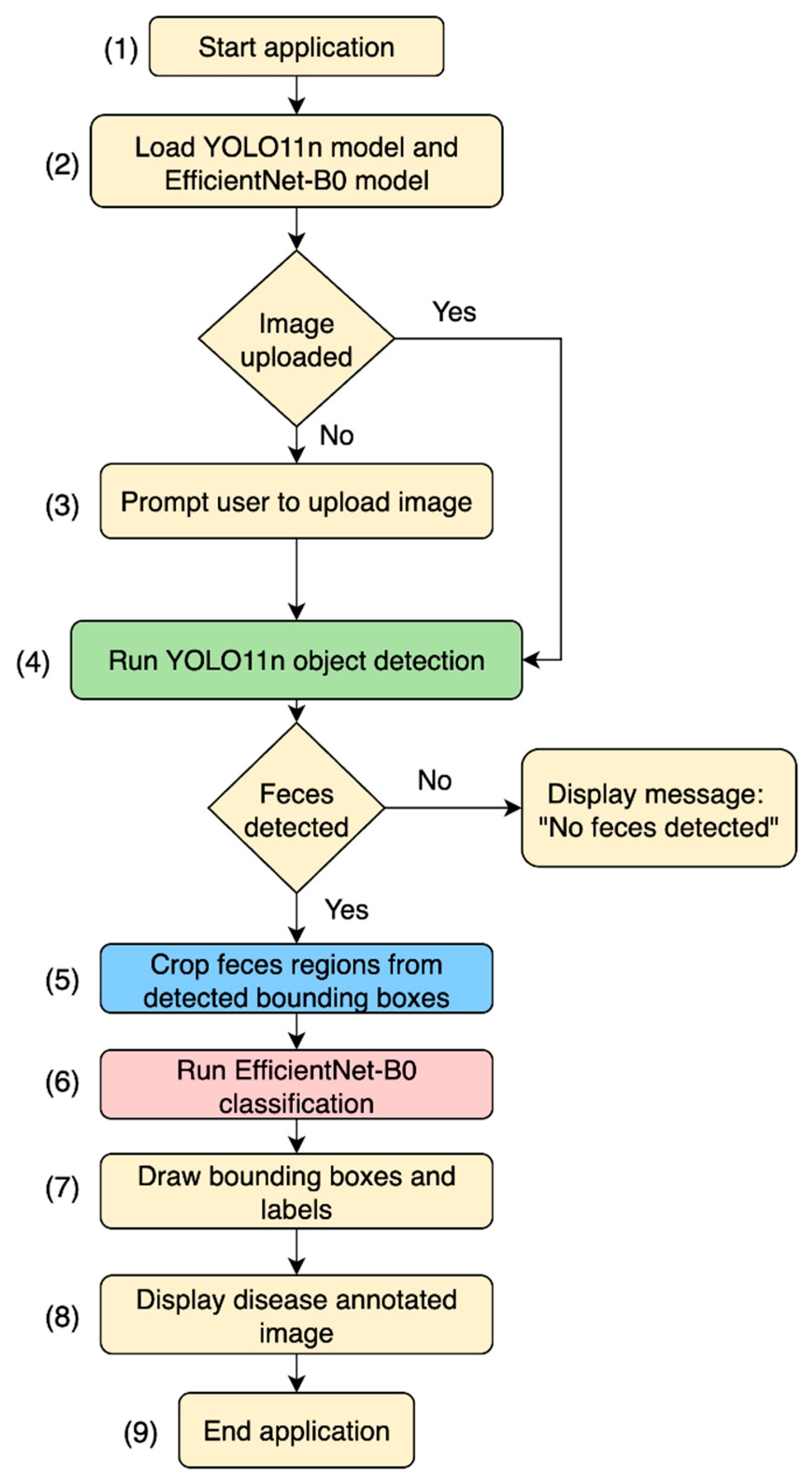
2.3. Integrated Web App
3. Results and Discussion
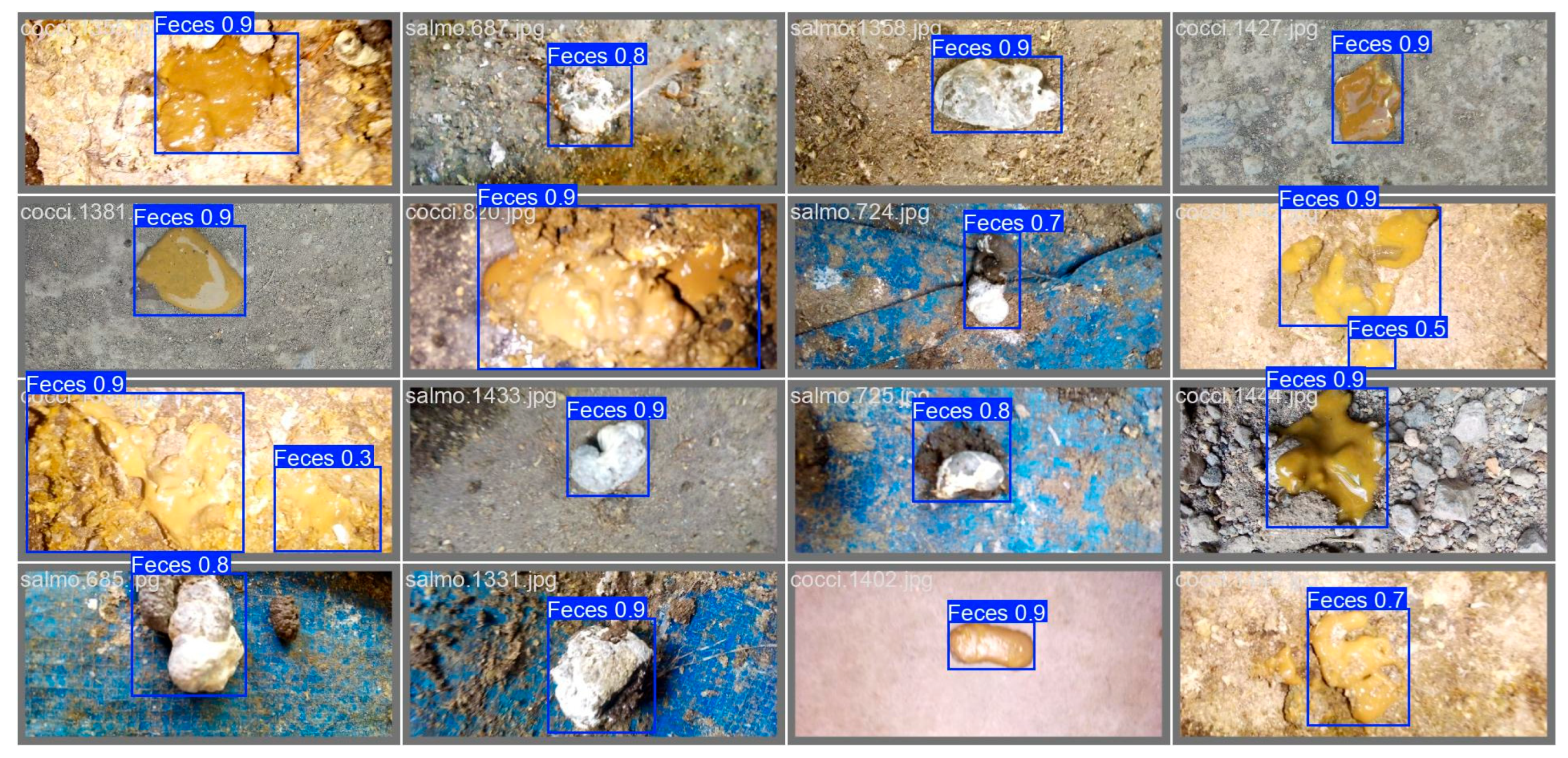
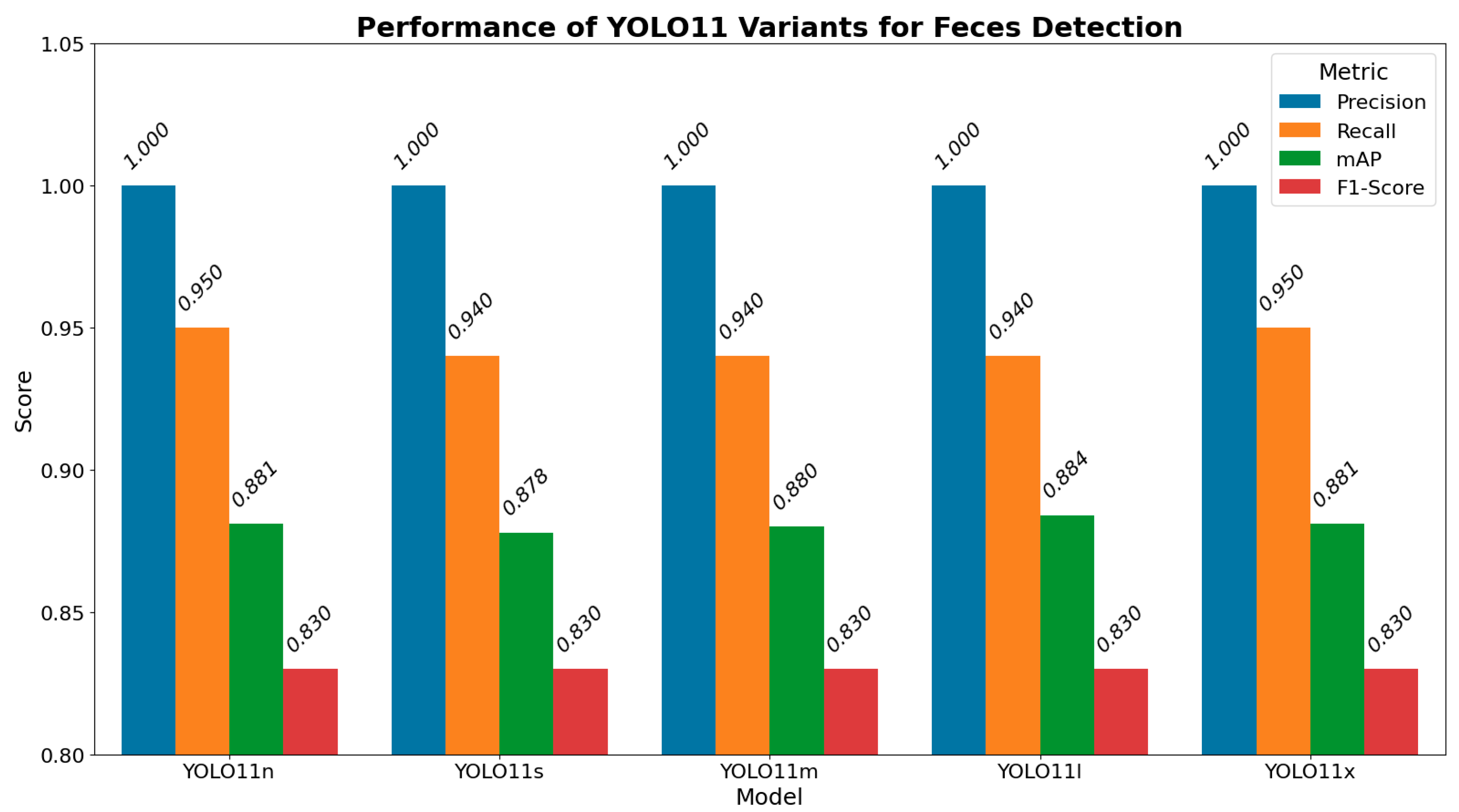
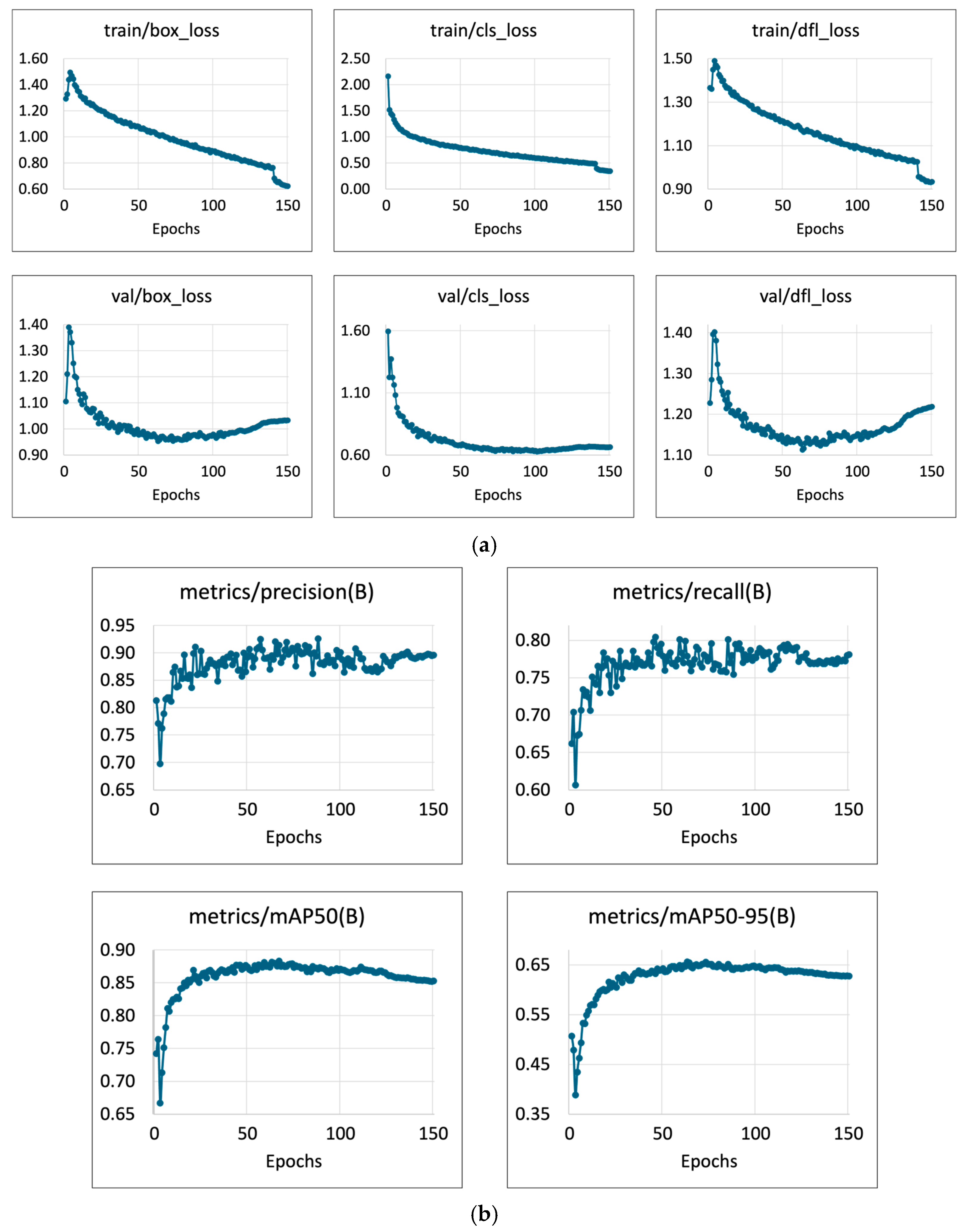
3.1. Feces Detection
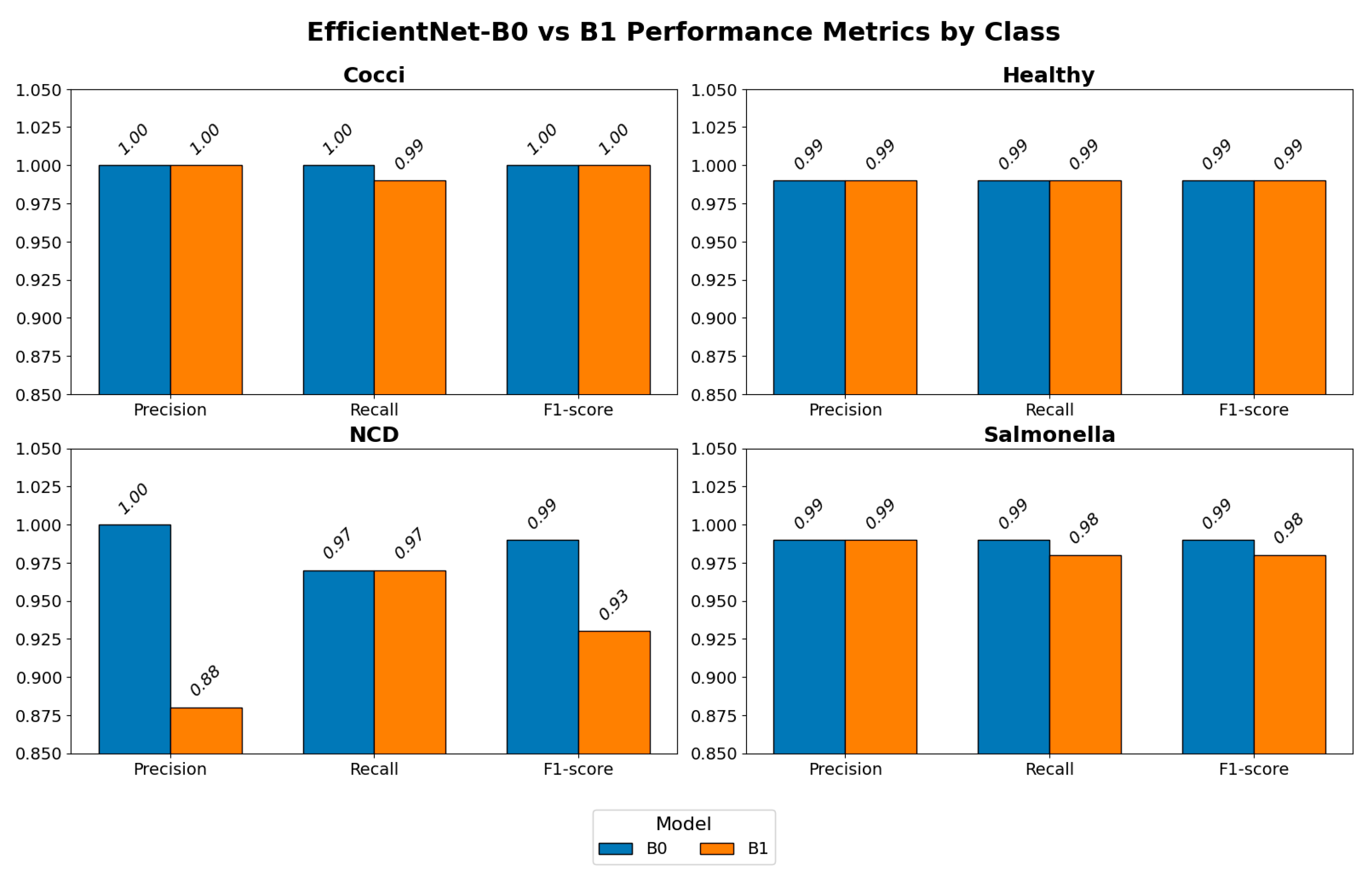
3.2. Disease Classification
3.3. Model Selection
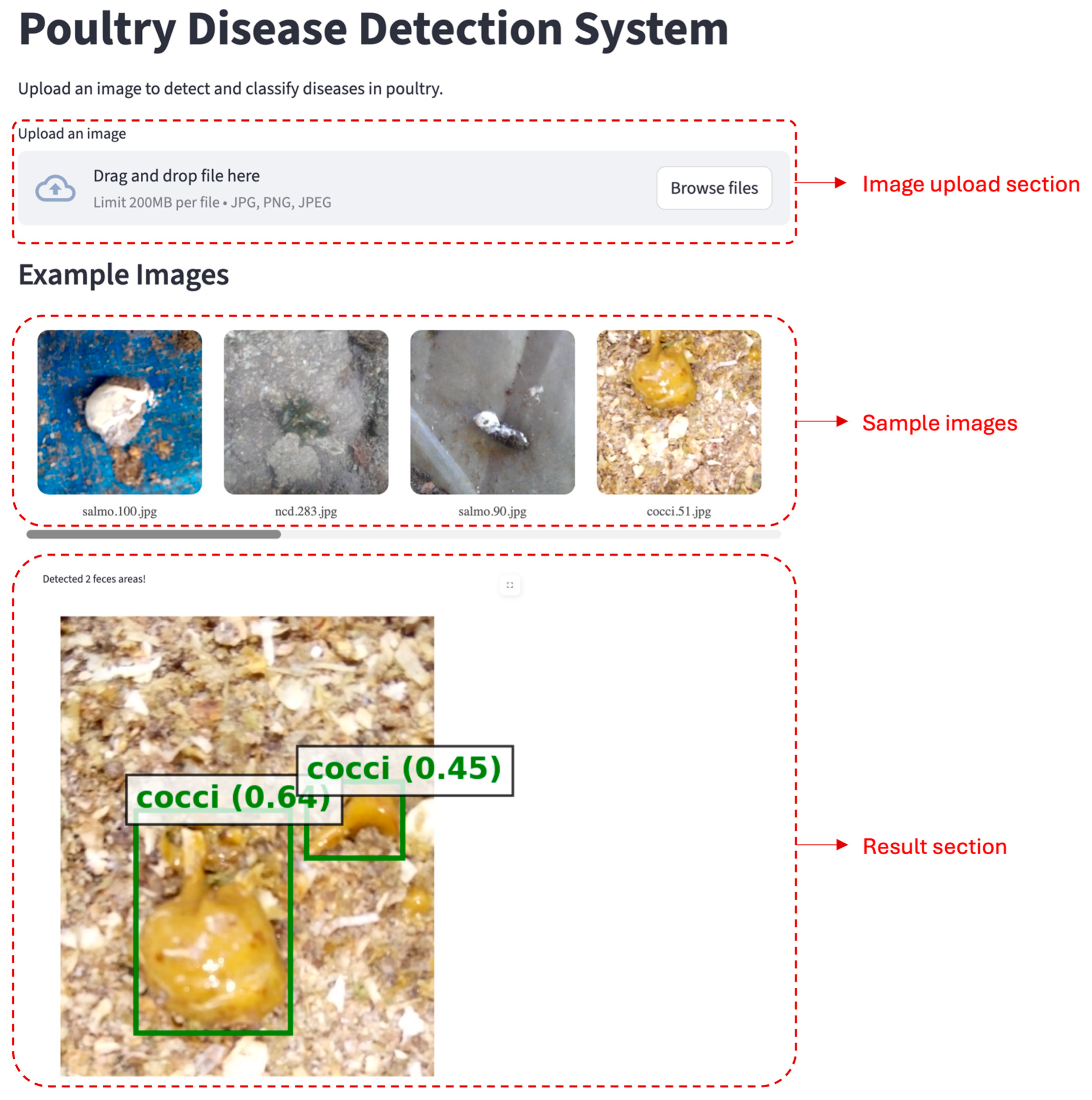
3.4. Web App and User Interface (UI)
3.5. System Performance
4. Conclusions
Author Contributions
Funding
Data Availability Statement
Conflicts of Interest
References
- USDA-NASS. Poultry—Production and Value, 2024 Summary; USDA: Washington, DC, USA, 2025.
- Blake, D.P.; Knox, J.; Dehaeck, B.; Huntington, B.; Rathinam, T.; Ravipati, V.; Ayoade, S.; Gilbert, W.; Adebambo, A.O.; Jatau, I.D. Re-calculating the cost of coccidiosis in chickens. Vet. Res. 2020, 51, 115. [Google Scholar] [CrossRef] [PubMed]
- Gast, R.; Jones, D.; Guraya, R.; Anderson, K.; Karcher, D. Research Note: Horizontal transmission and internal organ colonization by Salmonella Enteritidis and Salmonella Kentucky in experimentally infected laying hens in indoor cage-free housing. Poult. Sci. 2020, 99, 6071–6074. [Google Scholar] [CrossRef]
- Hoorebeke, S.; Immerseel, F.; Haesebrouck, F.; Ducatelle, R.; Dewulf, J. The Influence of the Housing System on Salmonella Infections in Laying Hens: A Review. Zoonoses Public Health 2011, 58, 304–311. [Google Scholar] [CrossRef]
- Hartcher, K.; Jones, B. The Welfare of Layer Hens in Cage and Cage-Free Housing Systems. World S Poult. Sci. J. 2017, 73, 767–782. [Google Scholar] [CrossRef]
- Caputo, V.; Staples, A.J.; Tonsor, G.T.; Lusk, J.L. Egg producer attitudes and expectations regarding the transition to cage-free production: A mixed-methods approach. Poult. Sci. 2023, 102, 103058. [Google Scholar] [CrossRef] [PubMed]
- Lusk, J.L. Consumer preferences for cage-free eggs and impacts of retailer pledges. Agribusiness 2019, 35, 129–148. [Google Scholar] [CrossRef]
- Nezworski, J.; St. Charles, K.; Malladi, S.; Ssematimba, A.; Bonney, P.; Cardona, C.; Halvorson, D.; Culhane, M. A Retrospective Study of Early vs. Late Virus Detection and Depopulation on Egg Laying Chicken Farms Infected with Highly Pathogenic Avian Influenza Virus During the 2015 H5N2 Outbreak in the United States. Avian Dis. 2021, 65, 474–482. [Google Scholar] [CrossRef]
- Wen, J.; Gou, H.; Zhan, Z.; Gao, Y.; Chen, Z.; Bai, J.; Wang, S.; Chen, K.; Lin, Q.; Liao, M.; et al. A Rapid Novel Visualized Loop-Mediated Isothermal Amplification Method for Salmonella Detection Targeting at fimW Gene. Poult. Sci. 2020, 99, 3637–3642. [Google Scholar] [CrossRef]
- Butt, S.L.; Taylor, T.L.; Volkening, J.D.; Dimitrov, K.M.; Williams-Coplin, D.; Lahmers, K.K.; Miller, P.J.; Rana, A.M.; Suarez, D.L.; Afonso, C.L.; et al. Rapid virulence prediction and identification of Newcastle disease virus genotypes using third-generation sequencing. Virol. J. 2018, 15, 179. [Google Scholar] [CrossRef] [PubMed]
- Lee, Y.; Lillehoj, H.S. Development of a new immunodiagnostic tool for poultry coccidiosis using an antigen-capture sandwich assay based on monoclonal antibodies detecting an immunodominant antigen of Eimeria. Poult. Sci. 2023, 102, 102790. [Google Scholar] [CrossRef]
- Mesa-Pineda, C.; Navarro-Ruiz, J.L.; López-Osorio, S.; Chaparro-Gutiérrez, J.J.; Gómez-Osorio, L.M. Chicken Coccidiosis: From the Parasite Lifecycle to Control of the Disease. Front. Vet. Sci. 2021, 8, 787653. [Google Scholar] [CrossRef] [PubMed]
- Damerow, G. The Chicken Health Handbook; Storey Communications: London, UK, 1994. [Google Scholar]
- Morishita, T.Y.; Porter, R.E., Jr. Gastrointestinal and hepatic diseases. In Backyard Poultry Medicine and Surgery: A Guide for Veterinary Practitioners; Wiley-Blackwell: Hoboken, NJ, USA, 2021; pp. 289–316. [Google Scholar]
- Janiesch, C.; Zschech, P.; Heinrich, K. Machine learning and deep learning. Electron. Mark. 2021, 31, 685–695. [Google Scholar] [CrossRef]
- Srivastava, K.; Pandey, P. Deep Learning Based Classification of Poultry Disease. Int. J. Autom. Smart Technol. 2023, 13, 2439. [Google Scholar] [CrossRef]
- Machuve, D.; Nwankwo, E.; Mduma, N.; Mbelwa, J. Poultry diseases diagnostics models using deep learning. Front. Artif. Intell. 2022, 5, 733345. [Google Scholar] [CrossRef]
- Liu, X.; Zhou, Y.; Liu, Y. Poultry Disease Identification Based on Light Weight Deep Neural Networks. In Proceedings of the 2023 IEEE 3rd International Conference on Computer Communication and Artificial Intelligence (CCAI), Taiyuan, China, 26–28 May 2023; pp. 92–96. [Google Scholar]
- Wati, D.F.; Roestam, R. Poultry Disease Detection in Chicken Fecal Images Through Annotated Polymerase Chain Reaction Dataset Using YOLOv7 And Soft-Nms Algorithm. In Proceedings of the 2023 Eighth International Conference on Informatics and Computing (ICIC), Manado, Indonesia, 8–9 December 2023; pp. 1–7. [Google Scholar] [CrossRef]
- Li, G.; Gates, R.S.; Ramirez, B.C. An on-site feces image classifier system for chicken health assessment: A proof of concept. Appl. Eng. Agric. 2023, 39, 417–426. [Google Scholar] [CrossRef]
- Sophia, S.; Gladson, J.J. Human Behaviour and Abnormality Detection using YOLO and Conv2D Net. In Proceedings of the 2024 International Conference on Inventive Computation Technologies (ICICT), Lalitpur, Nepal, 24–26 April 2024; pp. 70–75. [Google Scholar] [CrossRef]
- Hussain, M. YOLOv1 to v8: Unveiling Each Variant–A Comprehensive Review of YOLO. IEEE Access 2024, 12, 42816–42833. [Google Scholar] [CrossRef]
- Redmon, J.; Divvala, S.; Girshick, R.; Farhadi, A. You only look once: Unified, real-time object detection. arXiv 2016, arXiv:1506.02640. [Google Scholar] [CrossRef]
- Tan, M.; Le, Q. Efficientnet: Rethinking model scaling for convolutional neural networks. In Proceedings of the International Conference on Machine Learning, Long Beach, CA, USA, 9–15 June 2019; pp. 6105–6114. [Google Scholar]
- Costa, A.; Da Silva, F.A.; Rios, R. Deep Learning-Based Transfer Learning for Classification of Cassava Disease. arXiv 2025, arXiv:2502.19351. [Google Scholar]
- Sufian, M.A.; Hamzi, W.; Zaman, S.; Alsadder, L.; Hamzi, B.; Varadarajan, J.; Azad, M.A.K. Enhancing Clinical Validation for Early Cardiovascular Disease Prediction through Simulation, AI, and Web Technology. Diagnostics 2024, 14, 1308. [Google Scholar] [CrossRef] [PubMed]
- Warbhe, M.K.; Bore, J.J.; Chaudari, S.N. A Deep Learning Based System to Predict the Plant Disease Using Streamlit. In Proceedings of the 2025 4th International Conference on Sentiment Analysis and Deep Learning (ICSADL), Bhimdatta, Nepal, 18–20 February 2025; pp. 1744–1751. [Google Scholar]
- Depuru, B.K.; Putsala, S.; Mishra, P. Automating poultry farm management with artificial intelligence: Real-time detection and tracking of broiler chickens for enhanced and efficient health monitoring. Trop. Anim. Health Prod. 2024, 56, 75. [Google Scholar] [CrossRef]
- Lu, H.; Yao, C.; An, L.; Song, A.; Ling, F.; Huang, Q.; Cai, Y.; Liu, Y.; Kang, D. Classification and identification of chicken-derived adulteration in pork patties: A multi-dimensional quality profile and machine learning-based approach. Food Control 2025, 176, 111381. [Google Scholar] [CrossRef]
- Machuve, D.; Nwankwo, E.; Lyimo, E.; Maguo, E.; Munisi, C. Machine learning dataset for poultry diseases diagnostics-PCR annotated. Zenodo 2021. Available online: https://zenodo.org/records/5801834 (accessed on 1 May 2025).
- Cubuk, E.; Zoph, B.; Mané, D.; Vasudevan, V.; Le, Q. AutoAugment: Learning Augmentation Strategies From Data. In Proceedings of the 2019 IEEE/CVF Conference on Computer Vision and Pattern Recognition (CVPR), Long Beach, CA, USA, 15–20 June 2019; pp. 113–123. [Google Scholar] [CrossRef]
- Khanam, R.; Hussain, M. Yolov11: An overview of the key architectural enhancements. arXiv 2024, arXiv:2410.17725. [Google Scholar] [CrossRef]
- Afaq, S.; Rao, S. Significance of epochs on training a neural network. Int. J. Sci. Technol. Res 2020, 9, 485–488. [Google Scholar]
- Sandler, M.; Howard, A.; Zhu, M.; Zhmoginov, A.; Chen, L.-C. Mobilenetv2: Inverted residuals and linear bottlenecks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Salt Lake City, UT, USA, 18–23 June 2018; pp. 4510–4520. [Google Scholar]
- Hu, J.; Shen, L.; Sun, G. Squeeze-and-excitation networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Salt Lake City, UT, USA, 18–23 June 2018; pp. 7132–7141. [Google Scholar]
- Hoang, V.-T.; Jo, K. Practical Analysis on Architecture of EfficientNet. In Proceedings of the 2021 14th International Conference on Human System Interaction (HSI), Gdańsk, Poland, 8–10 July 2021; pp. 1–4. [Google Scholar] [CrossRef]
- Hastomo, W.; Karno, A.S.B.; Arif, D.; Wardhana, I.S.K.; Kamilia, N.; Yulianto, R.; Digdoyo, A.; Surawan, T. Brain Tumor Classification Using Four Versions of EfficientNet. Insearch Inf. Syst. Res. J. 2023, 3, 16–23. [Google Scholar] [CrossRef]
- Rosadi, M.I.; Hakim, L. Classification of Coffee Leaf Diseases using the Convolutional Neural Network (CNN) EfficientNet Model. Conf. Ser. 2023, 4, 58–69. [Google Scholar] [CrossRef]
- Cordeiro, L.D.; Nääs, I.D.; Okano, M.T. Smart Postharvest Management of Strawberries: YOLOv8-Driven Detection of Defects, Diseases, and Maturity. AgriEngineering 2025, 7, 246. [Google Scholar] [CrossRef]
- Singh, R.; Sharma, N.; Chauhan, R.; Choudhary, A.; Gupta, R. Precision kidney disease classification using efficientnet-b3 and ct imaging. In Proceedings of the 2023 3rd International Conference on Smart Generation Computing, Communication and Networking (SMART GENCON), Bangalore, India, 29–31 December 2023; pp. 1–6. [Google Scholar]
- Verma, G. Chicken Disease Detection: A Deep Learning Approach. In Proceedings of the 2024 Second International Conference on Intelligent Cyber Physical Systems and Internet of Things (ICoICI), Coimbatore, India, 28–30 August 2024; pp. 725–729. [Google Scholar]
- Kumar, P.; Bhatnagar, R.; Gaur, K.; Bhatnagar, A. Classification of Imbalanced Data: Review of Methods and Applications. IOP Conf. Ser. Mater. Sci. Eng. 2021, 1099, 012077. [Google Scholar] [CrossRef]
- Mbelwa, H.; Mbelwa, J.; Machuve, D. Deep Convolutional Neural Network for Chicken Diseases Detection. Int. J. Adv. Comput. Sci. Appl. 2021, 12, 759–765. [Google Scholar] [CrossRef]
- Mohammed, R.; Rawashdeh, J.; Abdullah, M. Machine Learning with Oversampling and Undersampling Techniques: Overview Study and Experimental Results. In Proceedings of the 2020 11th International Conference on Information and Communication Systems (ICICS), Irbid, Jordan, 7–9 April 2020; pp. 243–248. [Google Scholar]
- Johnson, J.M.; Khoshgoftaar, T.M. Survey on deep learning with class imbalance. J. Big Data 2019, 6, 27. [Google Scholar] [CrossRef]
- Goodfellow, I.J.; Pouget-Abadie, J.; Mirza, M.; Xu, B.; Warde-Farley, D.; Ozair, S.; Courville, A.; Bengio, Y. Generative adversarial nets. Adv. Neural Inf. Process. Syst. 27, NIPS 2014. Available online: https://proceedings.neurips.cc/paper_files/paper/2014/file/f033ed80deb0234979a61f95710dbe25-Paper.pdf (accessed on 1 May 2025).
- Zhang, H.; Cisse, M.; Dauphin, Y.N.; Lopez-Paz, D. mixup: Beyond empirical risk minimization. arXiv 2017, arXiv:1710.09412. [Google Scholar]
- Aworinde, H.O.; Adebayo, S.; Akinwunmi, A.; Alabi, O.M.; Ayandiji, A.; Sakpere, A.; Oyebamiji, A.; Olaide, O.; Kizito, E.; Olawuyi, A. Poultry fecal imagery dataset for health status prediction: A case of South-West Nigeria. Data Brief 2023, 50, 109517. [Google Scholar] [CrossRef] [PubMed]
- Chidziwisano, G.; Samikwa, E.; Daka, C. Deep learning methods for poultry disease prediction using images. Comput. Electron. Agric. 2025, 230, 109765. [Google Scholar] [CrossRef]


Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2025 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dhungana, A.; Yang, X.; Paneru, B.; Dahal, S.; Lu, G.; Chai, L. An Integrated Deep Learning Approach for Poultry Disease Detection and Classification Based on Analysis of Chicken Manure Images. AgriEngineering 2025, 7, 278. https://doi.org/10.3390/agriengineering7090278
Dhungana A, Yang X, Paneru B, Dahal S, Lu G, Chai L. An Integrated Deep Learning Approach for Poultry Disease Detection and Classification Based on Analysis of Chicken Manure Images. AgriEngineering. 2025; 7(9):278. https://doi.org/10.3390/agriengineering7090278
Chicago/Turabian StyleDhungana, Anjan, Xiao Yang, Bidur Paneru, Samin Dahal, Guoyu Lu, and Lilong Chai. 2025. "An Integrated Deep Learning Approach for Poultry Disease Detection and Classification Based on Analysis of Chicken Manure Images" AgriEngineering 7, no. 9: 278. https://doi.org/10.3390/agriengineering7090278
APA StyleDhungana, A., Yang, X., Paneru, B., Dahal, S., Lu, G., & Chai, L. (2025). An Integrated Deep Learning Approach for Poultry Disease Detection and Classification Based on Analysis of Chicken Manure Images. AgriEngineering, 7(9), 278. https://doi.org/10.3390/agriengineering7090278







