The Promise of Hyperspectral Imaging for the Early Detection of Crown Rot in Wheat
Abstract
:1. Introduction
- Through the review, we provide evidence that the crown can change photosynthesis (Section 3.2.2) and water and nutrients uptake (Section 3.2.1) in plants, which are major factors to influence wheat growth and cause yield loss.
- We developed a hypothesis that hyperspectral imaging can detect the changes of photosynthesis, water, and nitrogen uptake before visible symptoms on the upper stems and leaves.
- We conducted an initial experiment to support the hypothesis.
- We point out further research directions of using HSI for crown rot detection.
2. Digital Color Imaging
3. HSI Technologies for Plant Phenotyping
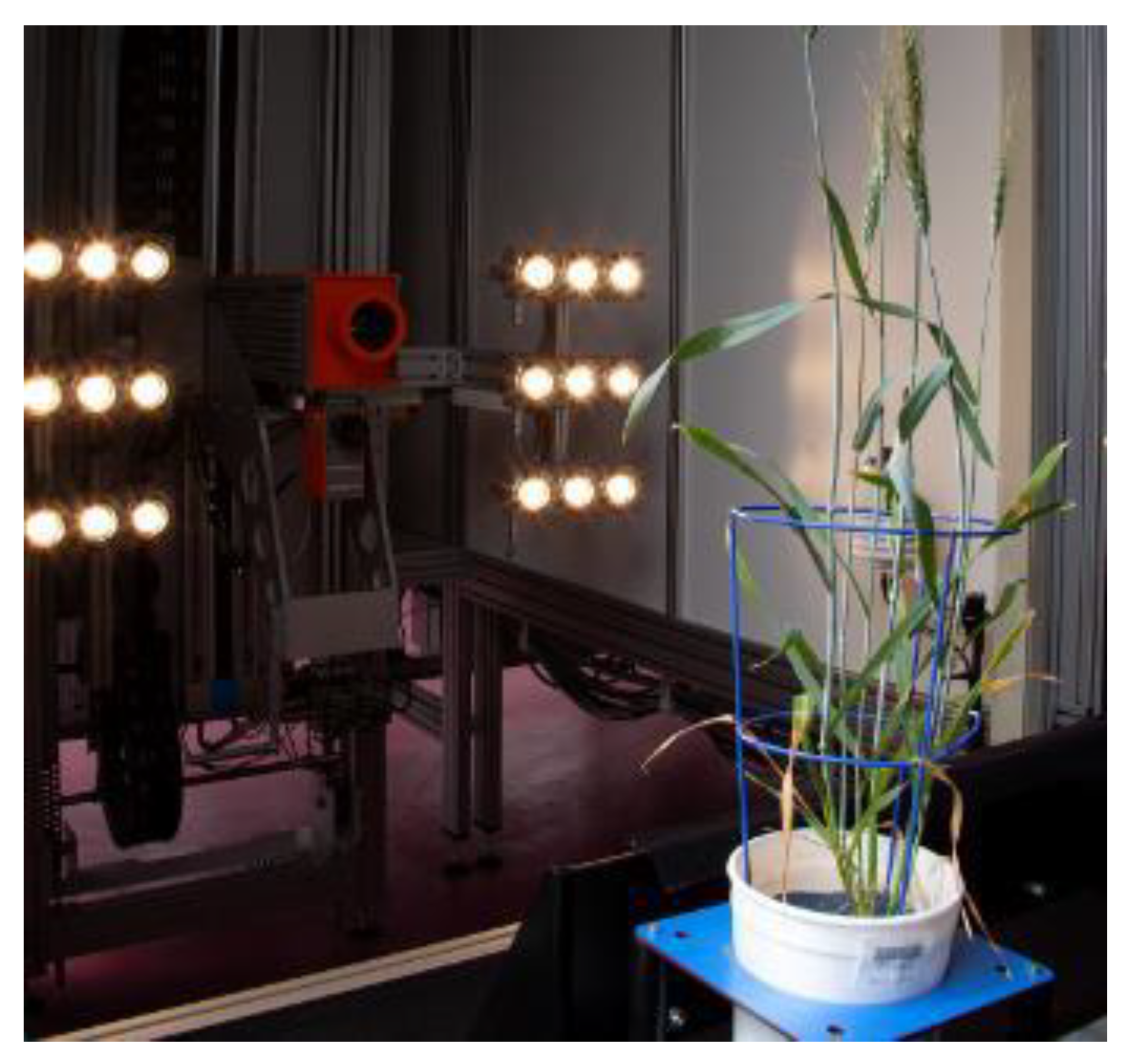
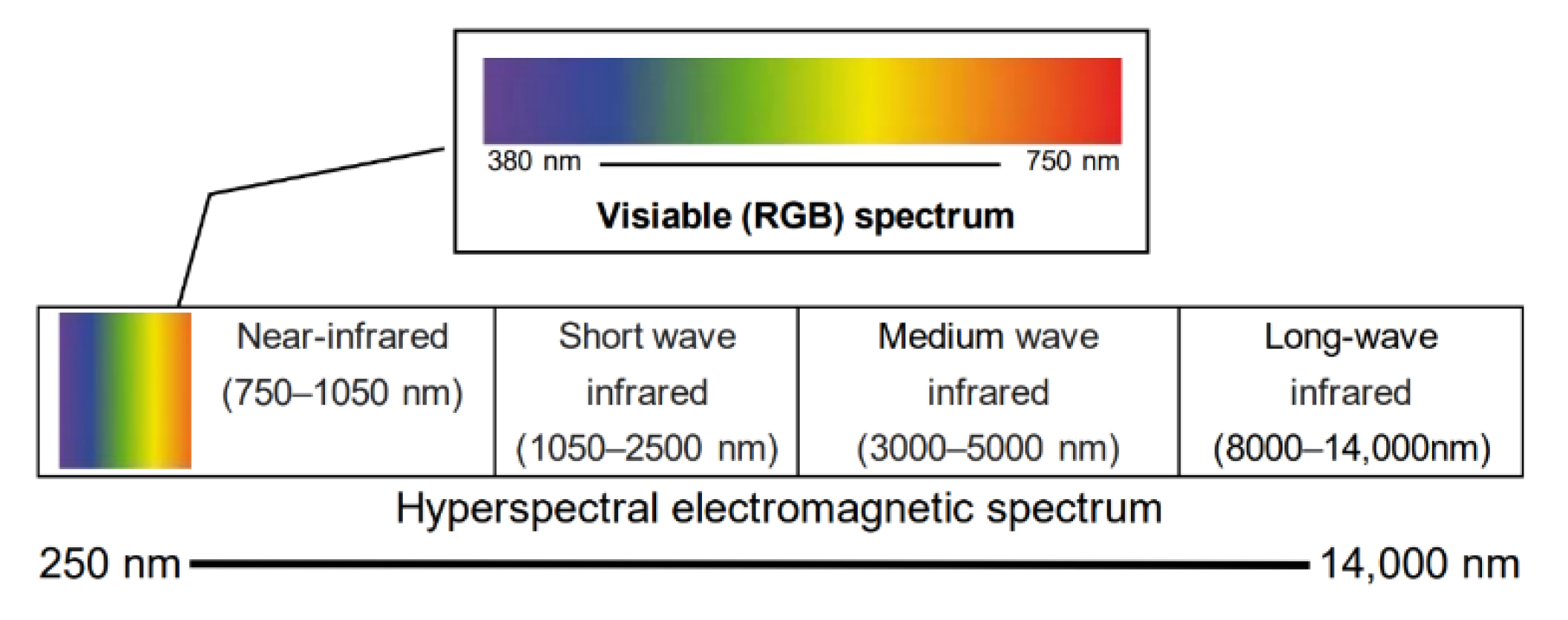
3.1. Hyperspectral Imaging
3.2. HSI and Crown Rot Related Plant Traits
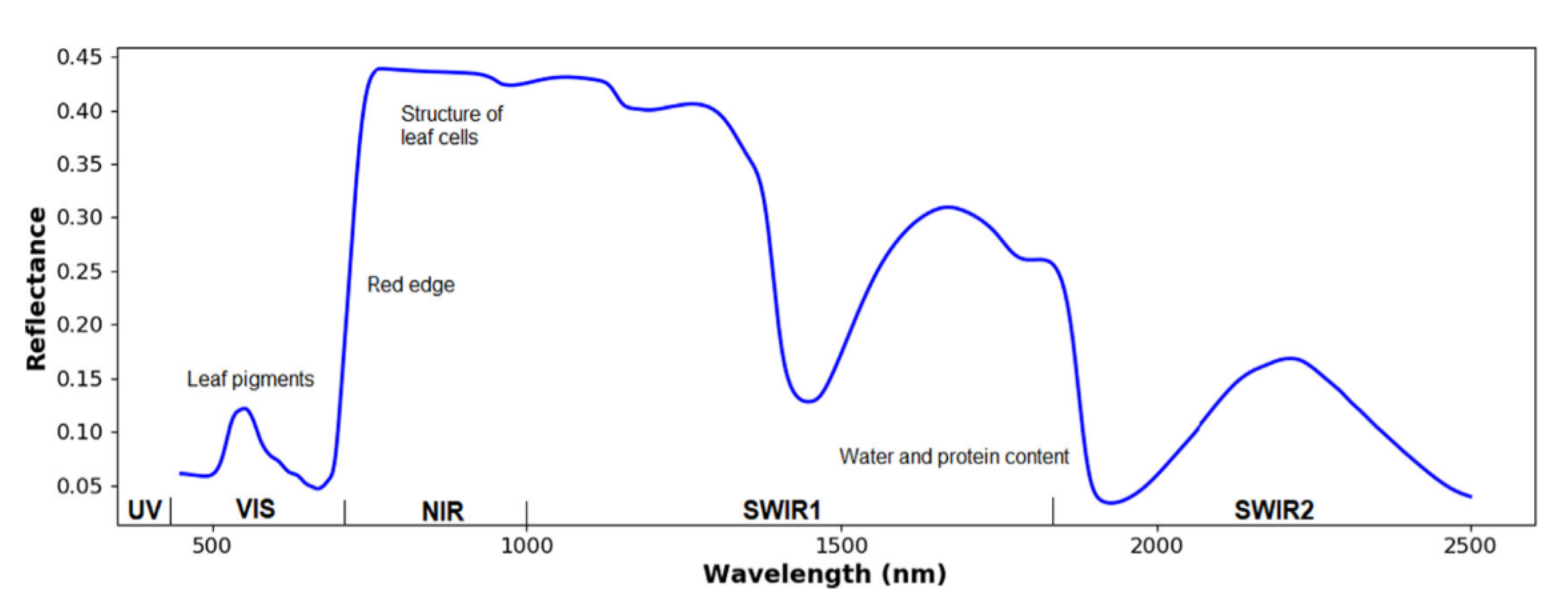
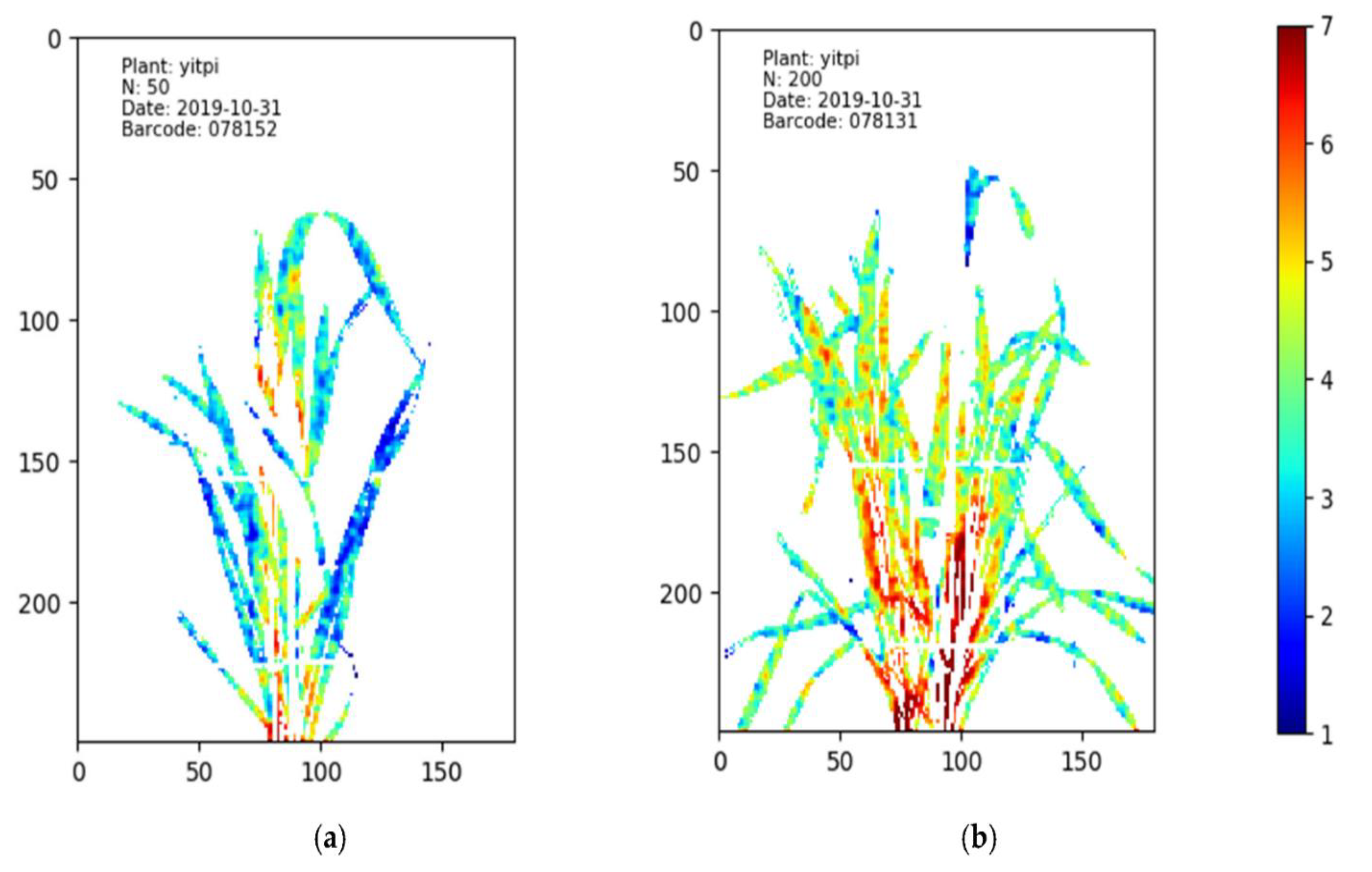
3.2.1. Water and Nutrient Distribution Maps of Wheat
3.2.2. Hyperspectral and Chlorophyll Fluorescence Imaging Interaction in Photosynthesis and DON Screening
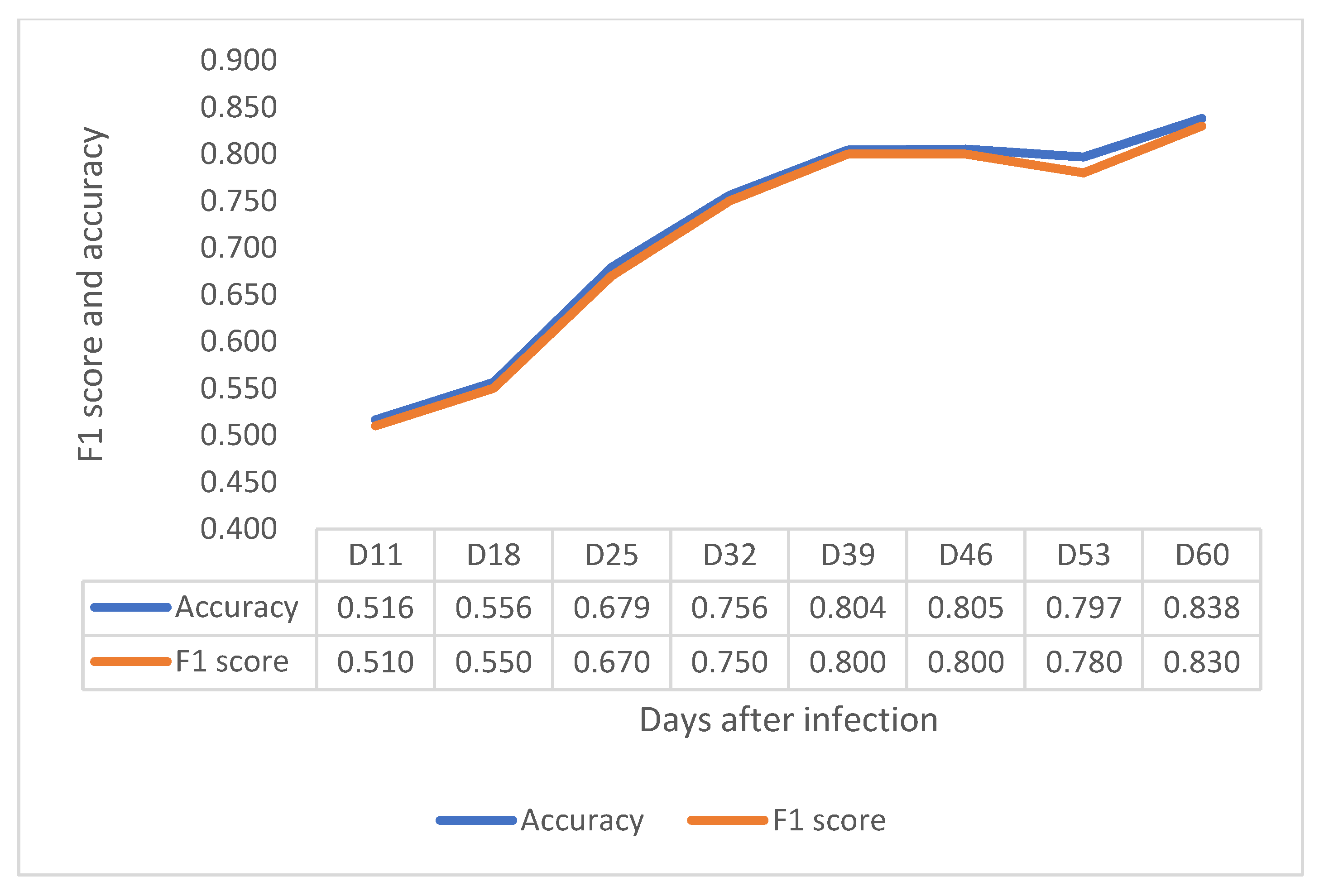
3.3. Preliminary Evaluation of HSI to Detect Crown Rot in Wheat
4. Conclusions
- The wavelengths range of the spectrum mentioned in this study is very wide. The key-wavelengths that play the most important role in the classification need to be further studied. Determining key-wavelengths can guide the design of low-cost, light-weight multispectral sensors for field applications.
- In the preliminary experiment, we used the reflectance data alone as input for SVM classification. Different data types, such as SNV, hyper-hue, or principal components, need to be further studied.
- The preliminary experiment supports the hypothesis that HSI can distinguish the difference between infection and healthy plant, which can provide support for early disease detection. However, further research needs to investigate how to use HSI in disease screening to determine symptom severity level and levels of crown rot resistance in diverse varieties.
- It is important to further analyze how the pathogen affects the transport and distribution of water and nutrients in plants, especially at different growth stages.
- The initial experimental results are limited to side-view imaging only. An experiment of top-view imaging should be undertaken in a further study, since top-view images would be easier to obtain via remote sensing in field trials.
- The initial experiment was limited to the greenhouse environment and further investigation needs to be conducted in field trials.
- We conducted a preliminary experiment to demonstrate the feasibility of using HSI for crown rot disease detection. However, different types of sensors, data collection, data processing, and machine learning algorithms need further intensive study.
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Akinsanmi, O.A.; Mitter, V.; Simpfendorfer, S.; Backhouse, D.; Chakraborty, S. Identity and pathogenicity of Fusarium spp. isolated from wheat fields in Queensland and northern New South Wales. Aust. J. Agric. Res. 2004, 55, 97–107. [Google Scholar] [CrossRef]
- Moya-Elizondo, E.; Arismendi, N.; Castro, M.P.; Doussoulin, H. Distribution and prevalence of crown rot pathogens affecting wheat crops in Southern Chile. Chil. J. Agric. Res. 2015, 75, 78–84. [Google Scholar] [CrossRef] [Green Version]
- Shikur Gebremariam, E.; Sharma-Poudyal, D.; Paulitz, T.C.; Erginbas-Orakci, G.; Karakaya, A.; Dababat, A.A. Identity and pathogenicity of Fusarium species associated with crown rot on wheat (Triticum spp.) in Turkey. Eur. J. Plant Pathol. 2018, 150, 387–399. [Google Scholar] [CrossRef]
- Xu, F.; Yang, G.; Wang, J.; Song, Y.; Liu, L.; Zhao, K.; Li, Y.; Han, Z. Spatial distribution of root and crown rot fungi associated with winter wheat in the north China plain and its relationship with climate variables. Front. Microbiol. 2018, 9, 1054. [Google Scholar] [CrossRef] [PubMed]
- Knight, N.L.; Macdonald, B.; Percy, C.; Sutherland, M.W. Disease responses of hexaploid spring wheat (Triticum aestivum) culms exhibiting premature senescence (dead heads) associated with Fusarium pseudograminearum crown rot. Eur. J. Plant Pathol. 2021, 159, 191–202. [Google Scholar] [CrossRef]
- Chakraborty, S.; Liu, C.J.; Mitter, V.; Scott, J.B.; Akinsanmi, O.A.; Ali, S.; Dill-Macky, R.; Nicol, J.; Backhouse, D.; Simpfendorfer, S. Pathogen population structure and epidemiology are keys to wheat crown rot and Fusarium head blight management. Australas. Plant Pathol. 2006, 35, 643–655. [Google Scholar] [CrossRef]
- Khudhair, M.; Obanor, F.; Kazan, K.; Gardiner, D.M.; Aitken, E.; McKay, A.; Giblot-Ducray, D.; Simpfendorfer, S.; Thatcher, L.F. Genetic diversity of Australian Fusarium pseudograminearum populations causing crown rot in wheat. Eur. J. Plant Pathol. 2021. [Google Scholar] [CrossRef]
- Kazan, K.; Gardiner, D.M. Fusarium crown rot caused by Fusarium pseudograminearum in cereal crops: Recent progress and future prospects. Mol. Plant Pathol. 2018, 19, 1547–1562. [Google Scholar] [CrossRef] [Green Version]
- Murray, G.M.; Brennan, J.P. Estimating disease losses to the Australian wheat industry. Australas. Plant Pathol. 2009, 38, 558–570. [Google Scholar] [CrossRef]
- Moya-Elizondo, E.A.; Jacobsen, B.J. Integrated management of Fusarium crown rot of wheat using fungicide seed treatment, cultivar resistance, and induction of systemic acquired resistance (SAR). Biol. Control 2016, 92, 153–163. [Google Scholar] [CrossRef]
- Simpfendorfer, S. Evaluation of the Seed treatment Rancona Dimension as a Standalone Option for Managing Crown Rot in Wheat–2015; DAN00175; NSW DPI Northern Grains Research Results; Department of Primary Industries NSW: Tamworth, Australia, 2016; pp. 121–124. [Google Scholar]
- Wildermuth, G.; Thomas, G.; Radford, B.; McNamara, R.; Kelly, A. Crown rot and common root rot in wheat grown under different tillage and stubble treatments in southern Queensland, Australia. Soil Tillage Res. 1997, 44, 211–224. [Google Scholar] [CrossRef]
- Backhouse, D. Modelling the behaviour of crown rot in wheat caused by Fusarium pseudograminearum. Australas. Plant Pathol. 2014, 43, 15–23. [Google Scholar] [CrossRef]
- Erginbas-Orakci, G.; Poole, G.; Nicol, J.M.; Paulitz, T.; Dababat, A.A.; Campbell, K. Assessment of inoculation methods to identify resistance to Fusarium crown rot in wheat. J. Plant Dis. Prot. 2016, 123, 19–27. [Google Scholar] [CrossRef]
- Stephens, A.E.; Gardiner, D.M.; White, R.G.; Munn, A.L.; Manners, J.M. Phases of infection and gene expression of Fusarium graminearum during crown rot disease of wheat. Mol. Plant-Microbe Interact. 2008, 21, 1571–1581. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Knight, N.L.; Sutherland, M.W. Histopathological assessment of Fusarium pseudograminearum colonization of cereal culms during crown rot infections. Plant Dis. 2016, 100, 252–259. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Summerell, B.A.; Burgess, L.W.; Backhouse, D.; Bullock, S.; Swan, L.J. Natural occurrence of perithecia of Gibberella coronicola on wheat plants with crown rot in Australia. Australas. Plant Pathol. 2001, 30, 353–356. [Google Scholar] [CrossRef]
- Saad, A.; Macdonald, B.; Martin, A.; Knight, N.L.; Percy, C. Comparison of disease severity caused by four soil-borne pathogens in winter cereal seedlings. Crop Pasture Sci. 2021, 72, 325–334. [Google Scholar] [CrossRef]
- Smiley, R.W. Fusarium Crown Rot Whitehead Symptom as Influenced by Wheat Crop Management and Sampling Date. Plant Dis. 2019, 103, 2612–2623. [Google Scholar] [CrossRef]
- Alahmad, S.; Simpfendorfer, S.; Bentley, A.R.; Hickey, L.T. Crown rot of wheat in Australia: Fusarium pseudograminearum taxonomy, population biology and disease management. Australas. Plant Pathol. 2018, 47, 285–299. [Google Scholar] [CrossRef]
- Alahmad, S. Accelerating the Development of Durum Wheat Adapted to Drought and Crown Rot Conditions. Ph.D. Thesis, The University of Queensland, Brisbane, Australia, 2019. [Google Scholar]
- Hollaway, G.J.; Exell, G.K. Survey of wheat crops for white heads caused by crown rot in Victoria, 1997–2009. Australas. Plant Pathol. 2010, 39, 363–367. [Google Scholar] [CrossRef]
- Moya-Elizondo, E.A. Fusarium crown rot disease: Biology, interactions, management and function as a possible sensor of global climate change. Int. J. Agric. Nat. Resour. 2013, 40, 235–252. [Google Scholar] [CrossRef]
- Pestka, J.J.; Smolinski, A.T. Deoxynivalenol: Toxicology and potential effects on humans. J. Toxicol. Environ. Health Part B 2005, 8, 39–69. [Google Scholar] [CrossRef] [PubMed]
- Sobrova, P.; Adam, V.; Vasatkova, A.; Beklova, M.; Zeman, L.; Kizek, R. Deoxynivalenol and its toxicity. Interdiscip. Toxicol. 2010, 3, 94. [Google Scholar] [CrossRef]
- Mudge, A.M.; Dill-Macky, R.; Dong, Y.; Gardiner, D.M.; White, R.G.; Manners, J.M. A role for the mycotoxin deoxynivalenol in stem colonisation during crown rot disease of wheat caused by Fusarium graminearum and Fusarium pseudograminearum. Physiol. Mol. Plant Pathol. 2006, 69, 73–85. [Google Scholar] [CrossRef]
- Yli-Mattila, T. Ecology and evolution of toxigenic Fusarium species in cereals in northern Europe And Asia. J. Plant Pathol. 2010, 92, 7–18. [Google Scholar]
- Ozdemir, F.; Koc, N.K.; Paulitz, T.; Nicol, J.M.; Schroeder, K.L.; Poole, G. Determination of fusarium crown rot resistance in wheat to Fusarium culmorum and Fusarium pseudogramineaum using real time PCR. Crop Prot. 2020, 135. [Google Scholar] [CrossRef]
- Bauriegel, E.; Giebel, A.; Herppich, W. Hyperspectral and chlorophyll fluorescence imaging to analyse the impact of Fusarium culmorum on the photosynthetic integrity of infected wheat ears. Sensors 2011, 11, 3765–3779. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Davies, P. Resistance and Tolerance Where We Are with Crown Rot Breeding; GRDC, ACT 2604: Kingston, Australia, 2016. [Google Scholar]
- Tapia, M.P.C.; Burrows, R.P.M.; Sepúlveda, B.R.; Concha, M.V.; Palma, C.V.; Moya-Elizondo, E.A. Antagonistic activity of chilean strains of pseudomonas protegens against fungi causing crown and root rot of wheat (Triticum aestivum L.). Front. Plant Sci. 2020, 11, 951. [Google Scholar] [CrossRef] [PubMed]
- Shi, S.; Zhao, J.; Pu, L.; Sun, D.; Han, D.; Li, C.; Feng, X.; Fan, D.; Hu, X. Identification of new sources of resistance to crown rot and Fusarium head blight in Wheat. Plant Dis. 2020, 104, 1979–1985. [Google Scholar] [CrossRef] [PubMed]
- Nassau, K. Colour. Available online: https://www.britannica.com/science/color (accessed on 21 September 2021).
- Moroney, N.; Fairchild, M.D.; Hunt, R.W.; Li, C.; Luo, M.R.; Newman, T. The CIECAM02 color appearance model. In Proceedings of the Color and Imaging Conference, Scottsdale, AZ, USA, 12 November 2002; pp. 23–27. [Google Scholar]
- Ahmed, N.; Asif, H.M.S.; Saleem, G. Leaf Image-based Plant Disease Identification using Color and Texture Features. arXiv 2021, arXiv:2102.04515. [Google Scholar]
- Tian, Y.-W.; LI, C.-h. Color image segmentation method based on statistical pattern recognition for plant disease diagnose. J. Jilin Univ. Technol. (Nat. Sci. Ed.) 2004, 2, 028. [Google Scholar]
- Qiu, R.; Yang, C.; Moghimi, A.; Zhang, M.; Steffenson, B.J.; Hirsch, C.D. Detection of Fusarium Head Blight in wheat using a deep neural network and color imaging. Remote Sens. 2019, 11, 2658. [Google Scholar] [CrossRef] [Green Version]
- Lu, G.; Fei, B. Medical hyperspectral imaging: A review. J. Biomed. Opt. 2014, 19, 10901. [Google Scholar] [CrossRef] [PubMed]
- Bruning, B.; Berger, B.; Lewis, M.; Liu, H.; Garnett, T. Approaches, applications, and future directions for hyperspectral vegetation studies: An emphasis on yield-limiting factors in wheat. Plant Phenome J. 2020, 3, e20007. [Google Scholar] [CrossRef]
- Sahoo, R.N.; Ray, S.S.; Manjunath, K.R. Hyperspectral remote sensing of agriculture. Curr. Sci. 2015, 108, 848–859. [Google Scholar]
- Liu, H.; Bruning, B.; Garnett, T.; Berger, B. Hyperspectral imaging and 3D technologies for plant phenotyping: From satellite to close-range sensing. Comput. Electron. Agric. 2020, 175, 105621. [Google Scholar] [CrossRef]
- Abdulridha, J.; Ampatzidis, Y.; Qureshi, J.; Roberts, P. Laboratory and UAV-based identification and classification of tomato yellow leaf curl, bacterial spot, and target spot diseases in tomato utilizing hyperspectral imaging and machine learning. Remote Sens. 2020, 12, 2732. [Google Scholar] [CrossRef]
- Schut, A.; Ketelaars, J. Monitoring grass swards using imaging spectroscopy. Grass Forage Sci. 2003, 58, 276–286. [Google Scholar] [CrossRef]
- Lowe, A.; Harrison, N.; French, A.P. Hyperspectral image analysis techniques for the detection and classification of the early onset of plant disease and stress. Plant Methods 2017, 13, 80. [Google Scholar] [CrossRef]
- Vigneau, N.; Ecarnot, M.; Rabatel, G.; Roumet, P. Potential of field hyperspectral imaging as a non destructive method to assess leaf nitrogen content in Wheat. Field Crop. Res. 2011, 122, 25–31. [Google Scholar] [CrossRef] [Green Version]
- Liu, H.; Chahl, J.S. A multispectral machine vision system for invertebrate detection on green leaves. Comput. Electron. Agric. 2018, 150, 279–288. [Google Scholar] [CrossRef]
- Liu, H.; Lee, S.H.; Chahl, J.S. Transformation of a high-dimensional color space for material classification. J. Opt. Soc. Am. A Opt. Image Sci. Vis. 2017, 34, 523–532. [Google Scholar] [CrossRef] [PubMed]
- Thennadil, S.; Martens, H.; Kohler, A. Physics-based multiplicative scatter correction approaches for improving the performance of calibration models. Appl. Spectrosc. 2006, 60, 315–321. [Google Scholar] [CrossRef] [PubMed]
- Grisanti, E.; Totska, M.; Huber, S.; Krick Calderon, C.; Hohmann, M.; Lingenfelser, D.; Otto, M. Dynamic Localized SNV, Peak SNV, and Partial Peak SNV: Novel Standardization Methods for Preprocessing of Spectroscopic Data Used in Predictive Modeling. J. Spectrosc. 2018, 2018, 5037572. [Google Scholar] [CrossRef]
- Zhu, H.; Gowen, A.; Feng, H.; Yu, K.; Xu, J.-L. Deep spectral-spatial features of near infrared hyperspectral images for pixel-wise classification of food products. Sensors 2020, 20, 5322. [Google Scholar] [CrossRef]
- Savitzky, A.; Golay, M.J. Smoothing and differentiation of data by simplified least squares procedures. Anal. Chem. 1964, 36, 1627–1639. [Google Scholar] [CrossRef]
- Norris, K.; Williams, P. Optimization of mathematical treatments of raw near-infrared signal in the measurement of protein in hard red spring wheat. I. Influence of particle size. Cereal Chem. 1984, 61, 158–165. [Google Scholar]
- Rinnan, Å.; Van Den Berg, F.; Engelsen, S.B. Review of the most common pre-processing techniques for near-infrared spectra. TrAC Trends Anal. Chem. 2009, 28, 1201–1222. [Google Scholar] [CrossRef]
- Steinier, J.; Termonia, Y.; Deltour, J. Smoothing and differentiation of data by simplified least square procedure. Anal. Chem. 1972, 44, 1906–1909. [Google Scholar] [CrossRef]
- Celestre, R.; Rosenberger, M.; Notni, G. A novel algorithm for bad pixel detection and correction to improve quality and stability of geometric measurements. Proc. J.Phys. Conf.Ser. 2016, 772, 012002. [Google Scholar] [CrossRef] [Green Version]
- Stellacci, A.M.; Castrignanò, A.; Diacono, M.; Troccoli, A.; Ciccarese, A.; Armenise, E.; Gallo, A.; De Vita, P.; Lonigro, A.; Mastro, M.A. Combined approach based on principal component analysis and canonical discriminant analysis for investigating hyperspectral plant response. Ital. J. Agron. 2012, 7, e34. [Google Scholar] [CrossRef] [Green Version]
- Kuhlman, D. A Python Book: Beginning Python, Advanced Python, and Python Exercises; Platypus Global Media: Washington, DC, USA, 2011. [Google Scholar]
- Koza, J.R.; Bennett, F.H.; Andre, D.; Keane, M.A. Automated design of both the topology and sizing of analog electrical circuits using genetic programming. In Artificial Intelligence in Design ’96; Gero, J.S., Sudweeks, F., Eds.; Springer: Dordrecht, The Netherlands, 1996; pp. 151–170. [Google Scholar]
- Pedregosa, F.; Varoquaux, G.; Gramfort, A.; Michel, V.; Thirion, B.; Grisel, O.; Blondel, M.; Prettenhofer, P.; Weiss, R.; Dubourg, V. Scikit-learn: Machine learning in Python. the J. Mach. Learn. Res. 2011, 12, 2825–2830. [Google Scholar]
- Dotto, A.C.; Dalmolin, R.S.D.; ten Caten, A.; Grunwald, S. A systematic study on the application of scatter-corrective and spectral-derivative preprocessing for multivariate prediction of soil organic carbon by Vis-NIR spectra. Geoderma 2018, 314, 262–274. [Google Scholar] [CrossRef]
- Kaur, R.; Kang, S.S. An enhancement in classifier support vector machine to improve plant disease detection. In Proceedings of the 2015 IEEE 3rd International Conference on MOOCs, Innovation and Technology in Education (MITE), Amritsar, India, 1–2 October 2015; pp. 135–140. [Google Scholar]
- Nguyen, C.; Sagan, V.; Maimaitiyiming, M.; Maimaitijiang, M.; Bhadra, S.; Kwasniewski, M.T. Early Detection of Plant Viral Disease Using Hyperspectral Imaging and Deep Learning. Sensors 2021, 21, 742. [Google Scholar] [CrossRef]
- Bruning, B.; Liu, H.; Brien, C.; Berger, B.; Lewis, M.; Garnett, T. The development of hyperspectral distribution maps to predict the content and distribution of nitrogen and water in wheat (Triticum aestivum). Front. Plant Sci. 2019, 10. [Google Scholar] [CrossRef]
- Maity, S.; Sarkar, S.; vinaba Tapadar, A.; Dutta, A.; Biswas, S.; Nayek, S.; Saha, P. Fault area detection in leaf diseases using k-means clustering. In Proceedings of the 2018 2nd International Conference on Trends in Electronics and Informatics (ICOEI), Tirunelveli, India, 11–12 May 2018; pp. 1538–1542. [Google Scholar]
- Jolliffe, I.T.; Cadima, J. Principal component analysis: A review and recent developments. Philos. Trans. R. Soc. A Math. Phys. Eng. Sci. 2016, 374, 20150202. [Google Scholar] [CrossRef] [PubMed]
- Severson, K.A.; Molaro, M.C.; Braatz, R.D. Principal component analysis of process datasets with missing values. Processes 2017, 5, 38. [Google Scholar] [CrossRef] [Green Version]
- Rodarmel, C.; Shan, J. Principal component analysis for hyperspectral image classification. Surv. Land Inf. Sci. 2002, 62, 115–122. [Google Scholar]
- Bock, C.H.; Barbedo, J.G.A.; Del Ponte, E.M.; Bohnenkamp, D.; Mahlein, A.-K. From visual estimates to fully automated sensor-based measurements of plant disease severity: Status and challenges for improving accuracy. Phytopathol. Res. 2020, 2, 9. [Google Scholar] [CrossRef] [Green Version]
- Dong, L.; Na, L.; Dongyan, Z.; Jinling, Z.; Fenfang, L.; Linsheng, H.; Qing, Z.; Yuwan, D. Discrimination of powdery mildew and yellow rust of winter wheat using high resolution hyperspectra and imageries. Infrared Laser Eng. 2017, 46, 50–58. [Google Scholar]
- Guo, A.T.; Huang, W.J.; Ye, H.C.; Dong, Y.Y.; Ma, H.Q.; Ren, Y.; Ruan, C. Identification of wheat yellow rust using spectral and texture features of hyperspectral images. Remote Sens. 2020, 12, 1419. [Google Scholar] [CrossRef]
- Hagerty, C.H.; Irvine, T.; Rivedal, H.M.; Yin, C.; Kroese, D.R. Diagnostic Guide: Fusarium Crown Rot of Winter Wheat. Plant Health Prog. 2021, 22, 176–181. [Google Scholar] [CrossRef]
- Knight, N.L.; Sutherland, M.W. Histopathological assessment of wheat seedling tissues infected by Fusarium pseudograminearum. Plant Pathol. 2013, 62, 679–687. [Google Scholar] [CrossRef]
- Knight, N.L.; Macdonald, B.; Sutherland, M.W. Colonization of durum wheat (Triticum turgidum L. var. durum) culms exhibiting premature senescence (dead heads) associated with Fusarium pseudograminearum crown rot. Plant Dis. 2017, 101, 1788–1794. [Google Scholar] [CrossRef] [Green Version]
- Beccari, G.; Covarelli, L.; Nicholson, P. Infection processes and soft wheat response to root rot and crown rot caused by Fusarium culmorum. Plant Pathol. 2011, 60, 671–684. [Google Scholar] [CrossRef]
- Burgess, L.; Backhouse, D.; Summerell, B.; Swan, L. Crown rot of wheat. In Fusarium: Paul E. Nelson Memorial Symposium; Burgess, L., Backhouse, D., Summerell, B., Swan, L., Eds.; APS Press: St Paul, MN, USA, 2001; pp. 271–294. [Google Scholar]
- Britannica, T.E.o.E. Phloem. Available online: https://www.britannica.com/science/phloem (accessed on 21 August 2021).
- Myburg, A.A.; Sederoff, R.R. Xylem structure and function. eLS 2001. [Google Scholar] [CrossRef]
- Liu, H.; Bruning, B.; Garnett, T.; Berger, B. The performances of hyperspectral sensors for proximal sensing of nitrogen levels in wheat. Sensors 2020, 20, 4550. [Google Scholar] [CrossRef]
- Hogg, A.; Johnston, R.; Dyer, A. Applying real-time quantitative PCR to Fusarium crown rot of wheat. Plant Dis. 2007, 91, 1021–1028. [Google Scholar] [CrossRef] [Green Version]
- Pandey, P.; Ge, Y.; Stoerger, V.; Schnable, J.C. High throughput in vivo analysis of plant leaf chemical properties using hyperspectral imaging. Front. Plant Sci. 2017, 8, 1348. [Google Scholar] [CrossRef] [Green Version]
- Wahabzada, M.; Mahlein, A.-K.; Bauckhage, C.; Steiner, U.; Oerke, E.-C.; Kersting, K. Plant phenotyping using probabilistic topic models: Uncovering the hyperspectral language of plants. Sci. Rep. 2016, 6, 1–11. [Google Scholar] [CrossRef] [Green Version]
- Nguyen, H.D.D.; Pan, V.; Pham, C.; Valdez, R.; Doan, K.; Nansen, C. Night-based hyperspectral imaging to study association of horticultural crop leaf reflectance and nutrient status. Comput. Electron. Agric. 2020, 173, 105458. [Google Scholar] [CrossRef]
- Yu, K.-Q.; Zhao, Y.-R.; Li, X.-L.; Shao, Y.-N.; Liu, F.; He, Y. Hyperspectral imaging for mapping of total nitrogen spatial distribution in pepper plant. PLoS ONE 2014, 9, e116205. [Google Scholar]
- Ansari, M.S.; Young, K.R.; Nicolas, M.E. Determining wavelenth for nitrogen and phosphorus nutrients through hyperspectral remote sensing in wheat (Triticum aestivum L.) plant. Int. J. Bio-Resour. Stress Manag. 2016, 7, 653–662. [Google Scholar]
- Barbagallo, R.P.; Oxborough, K.; Pallett, K.E.; Baker, N.R. Rapid, noninvasive screening for perturbations of metabolism and plant growth using chlorophyll fluorescence imaging. Plant Physiol 2003, 132, 485–493. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Meyer, S.; Saccardy-Adji, K.; Rizza, F.; Genty, B. Inhibition of photosynthesis by Colletotrichum lindemuthianum in bean leaves determined by chlorophyll fluorescence imaging. Plant Cell Environ. 2001, 24, 947–956. [Google Scholar] [CrossRef]
- Bauriegel, E.; Giebel, A.; Geyer, M.; Schmidt, U.; Herppich, W.B. Early detection of Fusarium infection in wheat using hyper-spectral imaging. Comput. Electron. Agric. 2011, 75, 304–312. [Google Scholar] [CrossRef]
- Nogués, S.; Cotxarrera, L.; Alegre, L.; Trillas, M.I. Limitations to photosynthesis in tomato leaves induced by Fusarium wilt. New Phytol. 2002, 461–470. [Google Scholar] [CrossRef]
- Bushnell, W.; Perkins-Veazie, P.; Russo, V.; Collins, J.; Seeland, T. Effects of deoxynivalenol on content of chloroplast pigments in barley leaf tissues. Phytopathology 2010, 100, 33–41. [Google Scholar] [CrossRef] [Green Version]
- Yan, K.; Han, G.; Ren, C.; Zhao, S.; Wu, X.; Bian, T. Fusarium solani infection depressed photosystem performance by inducing foliage wilting in apple seedlings. Front. Plant Sci. 2018, 9, 479. [Google Scholar] [CrossRef] [Green Version]
- Bauriegel, E.; Herppich, W. Hyperspectral and Chlorophyll Fluorescence Imaging for Early Detection of Plant Diseases, with Special Reference to Fusarium spec. Infections on Wheat. Agriculture 2014, 4, 1–26. [Google Scholar] [CrossRef] [Green Version]
- Dammer, K.-H.; Möller, B.; Rodemann, B.; Heppner, D. Detection of head blight (Fusarium ssp.) in winter wheat by color and multispectral image analyses. Crop Prot. 2011, 30, 420–428. [Google Scholar] [CrossRef]
- Kang, Z.; Buchenauer, H. Cytology and ultrastructure of the infection of wheat spikes by Fusarium culmorum. Mycol. Res. 2000, 104, 1083–1093. [Google Scholar] [CrossRef]
- Muhammed, H.H.; Larsolle, A. Feature vector based analysis of hyperspectral crop reflectance data for discrimination and quantification of fungal disease severity in wheat. Biosyst. Eng. 2003, 86, 125–134. [Google Scholar] [CrossRef]
- Baret, F.; Guyot, G.; Major, D. Coupled fluorescence and reflectance measurements to improve crop productivity evaluation. In Applications of Chlorophyll Fluorescence in Photosynthesis Research, Stress Physiology, Hydrobiology and Remote Sensing; Springer: Berlin/Heidelberg, Germany, 1988; pp. 319–324. [Google Scholar]
- Šrobárová, A.; Nadubinská, M.; Čiamporová, M. Relative Efficacy of Fusamotoxins on Young Maize Plants. Cereal Res. Commun. 2004, 32, 241–248. [Google Scholar] [CrossRef]
- Gunupuru, L.R.; Perochon, A.; Doohan, F.M. Deoxynivalenol resistance as a component of FHB resistance. Trop. Plant Pathol. 2017, 42, 175–183. [Google Scholar] [CrossRef] [Green Version]
- Thai, L.H.; Hai, T.S.; Thuy, N.T. Image classification using support vector machine and artificial neural network. Int. J. Inf. Technol. Comput. Sci. 2012, 4, 32–38. [Google Scholar] [CrossRef] [Green Version]
- Humpal, J.; McCarthy, C.; Percy, C.; Thomasson, J.A. Detection of crown rot in wheat utilising near-infrared spectroscopy: Towards remote and robotic sensing. In Proceedings of the SPIE—The International Society for Optical Engineering, Online, 27 April–8 May 2020. [Google Scholar]
- Wallwork, H.; Zwer, P. Cereal Variety Disease Guide 2016; SARDI: Urrbrae, Australia, 2016. [Google Scholar]
- Liu, S.; Tan, X.; Liu, C.Y.; Zhu, C.L.; Li, W.H.; Cui, S.; Du, Y.F.; Huang, D.C.; Xie, F. Recognition of Fusarium Head Blight Wheat Grain Based on Hyperspectral Data Processing Algorithm. Guang Pu Xue Yu Guang Pu Fen Xi/Spectrosc. Spectr. Anal. 2019, 39, 3540–3546. [Google Scholar] [CrossRef]


| Sensor | Wavelength Range or Image Type | Plant | Parameter | Comments |
|---|---|---|---|---|
| Hyperspectral imaging chamber (WIWAM, Ghent, Netherlands) | VNIR (400–1000 nm) and SWIR (1000–2500 nm) | Wheat | Water and nutrient content | 400–2500 nm can predict the water and nutrient content in the plant with high accuracy (validated R2 = 0.69 in water and R2 = 0.66 in nitrogen) [63]. |
| VNIR sensor (FLAME-S-XR1-ES, Ocean Optics, Germany) and SWIR sensor (NQ512-1.7, Ocean Optics, Germany) | VNIR (200–1025 nm) and SWIR (900–1700 nm) | Wheat | Nitrogen content | The N stressed and controled plants were distinguished by the cameras after 30 days of sowing; the wavelengths of 355–515.5 nm, 617–695 nm, and 726–1075 nm played the most important roles [84]. |
| Spectrograph (ImSpector V10E, Spectral Imaging Ltd., Oulu, Finland) and digitally temperature-compensated b/w camera (Pixelfly qe, PCO AG, Kelheim, Germany) | VNIR (400–1000 nm) | Wheat | Water loss, Fusarium culmorum disease, and water content | Fusarium-infected plants and healthy plants were successfully distinguished in 682–733 nm and 927–931 nm wavelengths due to different water loss in plant tissue after growing for 70 days [29]. |
| Charge-coupled device (CCD) camera | Chlorophyll fluorescence imaging | Bean | Colletotrichum lundemuthianum disease | Chlorophyll fluorescence imaging can distinguish healthy and infected plant by analysis of the plant’s photosynthesis rate [86]. |
| Spectroradiometer | VNIR (360–900 nm) | Wheat | Fungal Disease, green reflectance | The fungus-disease-infected plant showed a difference between healthy plants in the 550 and 750 nm wavelength, and the reflectance peak at near-infrared region was decreased [94]. |
| 3-chip CCD camera (red, infrared, and green) and type MS2100 (DuncanTech company, Redlake Inc., San Diego, CA, USA) | red (peak wavelength: 670 nm, and infrared (peak wavelength: 800 nm | Wheat | Fusarium spp. disease, chlorophyll | Fusarium spp. caused a chlorophyll defect in wheat ears and reduced the photosynthesis rate [92]. |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Xie, Y.; Plett, D.; Liu, H. The Promise of Hyperspectral Imaging for the Early Detection of Crown Rot in Wheat. AgriEngineering 2021, 3, 924-941. https://doi.org/10.3390/agriengineering3040058
Xie Y, Plett D, Liu H. The Promise of Hyperspectral Imaging for the Early Detection of Crown Rot in Wheat. AgriEngineering. 2021; 3(4):924-941. https://doi.org/10.3390/agriengineering3040058
Chicago/Turabian StyleXie, Yiting, Darren Plett, and Huajian Liu. 2021. "The Promise of Hyperspectral Imaging for the Early Detection of Crown Rot in Wheat" AgriEngineering 3, no. 4: 924-941. https://doi.org/10.3390/agriengineering3040058
APA StyleXie, Y., Plett, D., & Liu, H. (2021). The Promise of Hyperspectral Imaging for the Early Detection of Crown Rot in Wheat. AgriEngineering, 3(4), 924-941. https://doi.org/10.3390/agriengineering3040058







