EEG-Based Seizure Detection Using Variable-Frequency Complex Demodulation and Convolutional Neural Networks
Abstract
1. Introduction
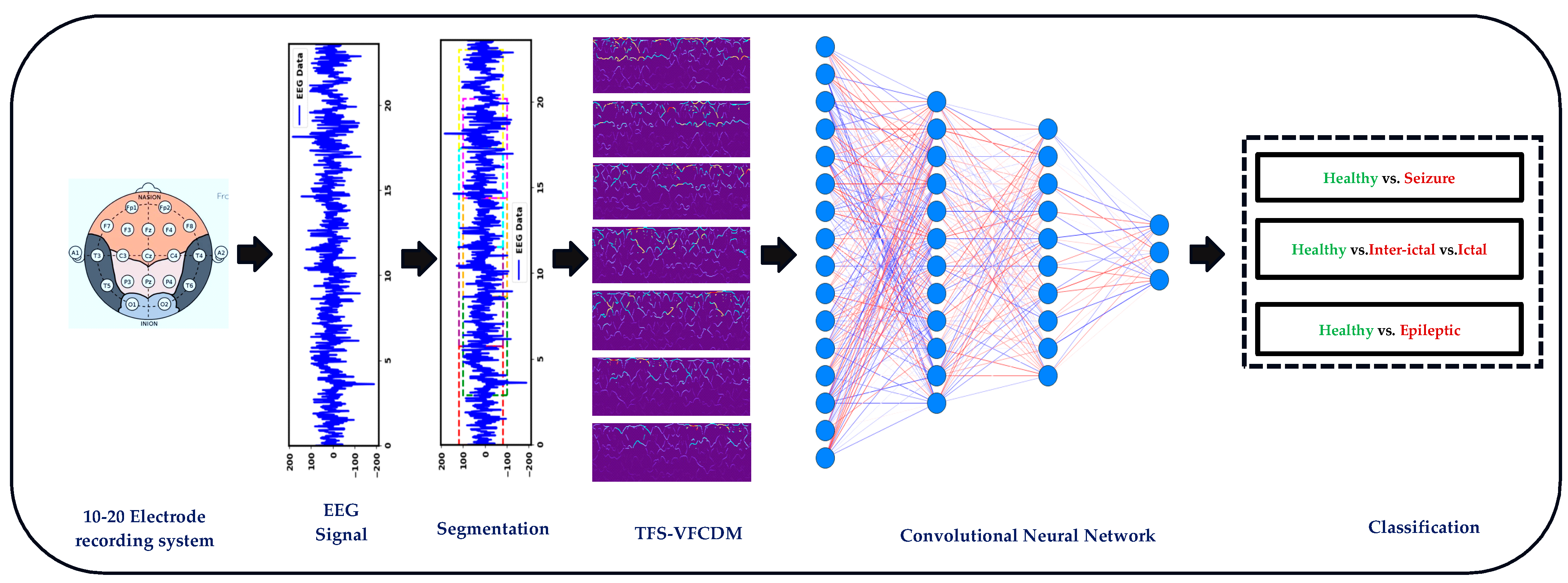
2. Materials and Methods
2.1. Database
2.2. Variable-Frequency Complex Demodulation Algorithm
2.2.1. Complex Demodulation
2.2.2. VFCDM
- 1.
- Filter Design: Design a finite-impulse response LPF with a specified bandwidth ( and filter length (. Set center frequencies as (:
- = Spacing between neighboring center frequencies,
- = Highest signal frequency.
- 2.
- CDM Dominant Frequency Extraction: Employ the CDM technique to identify the dominant frequency within the defined bandwidth and iterate this process by incrementing across the entire frequency band.
- 3.
- Signal Decomposition: Decompose the signal into sinusoidal modulations using the CDM.
- 4.
- Instantaneous Frequency Calculation: Calculate the instantaneous frequencies using Equation (9), based on the phase (as per Equation (5)) and the instantaneous amplitudes (as per Equation (4)) of each sinusoidal modulation component, using the Hilbert transform.
- 5.
- Time–Frequency Representation: Obtain the TFR of the signal by using the estimated instantaneous frequencies and amplitudes, providing a detailed depiction of signal variations across both time and frequency domains. For more details about the VFCDM algorithm, please refer to [21].
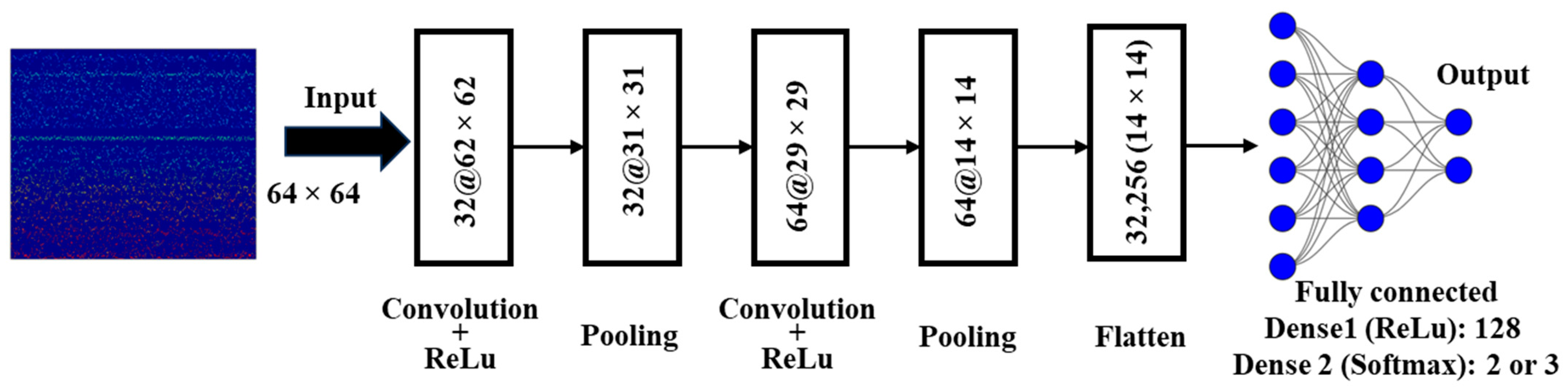
2.3. Convolutional Neural Networks
- (1)
- Convolution Layer (CL)
- (2)
- Pooling Layer (PL)
- (3)
- Fully Connected Layer (FC)
2.4. Performance Evaluation of CNN
3. Results
3.1. Case I: Healthy vs. Ictal State
3.2. Case II: Healthy vs. Epilepsy Subjects
3.3. Case III: Healthy vs. Interictal vs. Ictal States
4. Discussion
| Author | Method | Classifier | Cross-Validation | Performance (%) | |||
|---|---|---|---|---|---|---|---|
| Acc | Pre | Rec | F1 | ||||
| Faust et al. 2010 [38] | Yule–Walker | SVM $ | ˗ | 93.3 | ˗ | ˗ | ˗ |
| Acharya et al. 2009 [39] | Non-linear measures | GMM $ | ˗ | 96.1 | ˗ | ˗ | ˗ |
| Acharya et al. 2013 [40] | DWT Frequency Bands | SVM $ | ˗ | 96.0 | ˗ | ˗ | ˗ |
| Sharma et al. 2017 [47] | Flexible WT/Fractal Dimension | SVM | 10-Fold | 99.2 | ˗ | ˗ | ˗ |
| Acharya et al. 2018 [13] | 1D EEG Features | CNN | 10-Fold | 88.7 | ˗ | ˗ | ˗ |
| Ilias et al. 2023 [14] | Spectrogram/Delta | CNN | 10-Fold | 97.0 | 97.14–97.18 * | 96.00–97.99 * | 96.41–97.52 * |
| Mahfuz et al. 2021 [15] | CWT | CNN | Split 10/90 | 98.46 | ˗ | ˗ | ˗ |
| Hussein et al. 2019 [43] | TD | RNN/LSTM | 3/5/10-Fold | 100 | ˗ | ˗ | ˗ |
| Chanu et al. 2023 [44] | DWT | SONN | Split 30/70 | 99.2 | 98 | 100 | 98.99 |
| Islam et al. 2022 [45] | TD | Epileptic-Net | 10-Fold | 99.95–99.98 * | ˗ | ˗ | ˗ |
| Yuan et al. 2017 [42] | CWT Scalogram | GPCA/SDAE | Split 80/20 | 100 | 100 | 100 | 100 |
| Ullah et al. 2018 [12] | TD | P-1D-CNN | 10-Fold | 97.4–100 * | ˗ | ˗ | ˗ |
| Guo et al. 2011 [49] | Genetic Programming | KNN | Split 60/40 | 93.5 | ˗ | ˗ | ˗ |
| Bhattacharyya et al. 2017 [46] | EWT | RF | 10-Fold | 99.4 | ˗ | ˗ | ˗ |
| Sharmila et al. 2018 [41] | DWT Entropy | SVM | Split 50/50 | 78–100 * | ˗ | ˗ | ˗ |
| Goel et al. 2023 [48] | Recurrence Plots | SVM | - | 98.21 | 99.61 | ˗ | ˗ |
| Abdulhay et al. 2020 [50] | Entropy, non-linear, and spectra | SCANN | 10-Fold | 98.5 | ˗ | ˗ | ˗ |
| Proposed Approach | VFCDM | CNN | LOSO CV | 90–99 * | 96–99 * | 94–99 * | 93–99 * |
5. Conclusions
Author Contributions
Funding
Institutional Review Board Statement
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Epilepsy. Available online: https://www.who.int/news-room/fact-sheets/detail/epilepsy (accessed on 16 October 2023).
- England, M.J.; Liverman, C.T.; Schultz, A.M.; Strawbridge, L.M. Epilepsy across the Spectrum: Promoting Health and Understanding.: A Summary of the Institute of Medicine Report. Epilepsy Behav. 2012, 25, 266–276. [Google Scholar] [CrossRef] [PubMed]
- Abreu, M.; Carmo, A.S.; Franco, A.; Parreira, S.; Vidal, B.; Costa, M.; Peralta, A.R.; da Silva, H.P.; Bentes, C.; Fred, A. Mobile Applications for Epilepsy: Where Are We? Where Should We Go? A Systematic Review. Signals 2022, 3, 40–65. [Google Scholar] [CrossRef]
- Epilepsy and Seizures. Available online: https://www.ninds.nih.gov/health-information/disorders/epilepsy-and-seizures (accessed on 16 October 2023).
- Epilepsy—Seizure Types, Symptoms and Treatment Options. Available online: https://www.aans.org/ (accessed on 16 October 2023).
- Biasiucci, A.; Franceschiello, B.; Murray, M.M. Electroencephalography. Curr. Biol. 2019, 29, R80–R85. [Google Scholar] [CrossRef] [PubMed]
- Noachtar, S.; Rémi, J. The Role of EEG in Epilepsy: A Critical Review. Epilepsy Behav. 2009, 15, 22–33. [Google Scholar] [CrossRef]
- Ramakrishnan, S.; Rayi, A. EEG Localization Related Epilepsies. In StatPearls; StatPearls Publishing: Treasure Island, FL, USA, 2023. [Google Scholar]
- Nigam, V.P.; Graupe, D. A Neural-Network-Based Detection of Epilepsy. Neurol. Res. 2004, 26, 55–60. [Google Scholar] [CrossRef] [PubMed]
- Angus-Leppan, H. Diagnosing Epilepsy in Neurology Clinics: A Prospective Study. Seizure 2008, 17, 431–436. [Google Scholar] [CrossRef]
- Acharya, U.R.; Vinitha Sree, S.; Swapna, G.; Martis, R.J.; Suri, J.S. Automated EEG Analysis of Epilepsy: A Review. Knowl.-Based Syst. 2013, 45, 147–165. [Google Scholar] [CrossRef]
- Ullah, I.; Hussain, M.; Qazi, E.-H.; Aboalsamh, H. An Automated System for Epilepsy Detection Using EEG Brain Signals Based on Deep Learning Approach. Expert Syst. Appl. 2018, 107, 61–71. [Google Scholar] [CrossRef]
- Acharya, U.R.; Oh, S.L.; Hagiwara, Y.; Tan, J.H.; Adeli, H. Deep Convolutional Neural Network for the Automated Detection and Diagnosis of Seizure Using EEG Signals. Comput. Biol. Med. 2018, 100, 270–278. [Google Scholar] [CrossRef]
- Ilias, L.; Askounis, D.; Psarras, J. Multimodal Detection of Epilepsy with Deep Neural Networks. Expert Syst. Appl. 2023, 213, 119010. [Google Scholar] [CrossRef]
- Rashed-Al-Mahfuz, M.; Moni, M.A.; Uddin, S.; Alyami, S.A.; Summers, M.A.; Eapen, V. A Deep Convolutional Neural Network Method to Detect Seizures and Characteristic Frequencies Using Epileptic Electroencephalogram (EEG) Data. IEEE J. Transl. Eng. Health Med. 2021, 9, 2000112. [Google Scholar] [CrossRef]
- Yamashita, R.; Nishio, M.; Do, R.K.G.; Togashi, K. Convolutional Neural Networks: An Overview and Application in Radiology. Insights Imaging 2018, 9, 611–629. [Google Scholar] [CrossRef] [PubMed]
- Ganapathy, N.; Veeranki, Y.R.; Swaminathan, R. Convolutional Neural Network Based Emotion Classification Using Electrodermal Activity Signals and Time-Frequency Features. Expert Syst. Appl. 2020, 159, 113571. [Google Scholar] [CrossRef]
- Craik, A.; He, Y.; Contreras-Vidal, J.L. Deep Learning for Electroencephalogram (EEG) Classification Tasks: A Review. J. Neural Eng. 2019, 16, 031001. [Google Scholar] [CrossRef]
- Jacovi, A.; Shalom, O.S.; Goldberg, Y. Understanding Convolutional Neural Networks for Text Classification. arXiv 2020, arXiv:1809.08037. [Google Scholar]
- Tajbakhsh, N.; Shin, J.Y.; Gurudu, S.R.; Hurst, R.T.; Kendall, C.B.; Gotway, M.B.; Liang, J. Convolutional Neural Networks for Medical Image Analysis: Full Training or Fine Tuning? IEEE Trans. Med. Imaging 2016, 35, 1299–1312. [Google Scholar] [CrossRef]
- Wang, H.; Siu, K.; Ju, K.; Chon, K.H. A High Resolution Approach to Estimating Time-Frequency Spectra and Their Amplitudes. Ann. Biomed. Eng. 2006, 34, 326–338. [Google Scholar] [CrossRef]
- Andrzejak, R.G.; Lehnertz, K.; Mormann, F.; Rieke, C.; David, P.; Elger, C.E. Indications of Nonlinear Deterministic and Finite-Dimensional Structures in Time Series of Brain Electrical Activity: Dependence on Recording Region and Brain State. Phys. Rev. E 2001, 64, 061907. [Google Scholar] [CrossRef]
- Chon, K.H.; Dash, S.; Ju, K. Estimation of Respiratory Rate From Photoplethysmogram Data Using Time–Frequency Spectral Estimation. IEEE Trans. Biomed. Eng. 2009, 56, 2054–2063. [Google Scholar] [CrossRef]
- Siu, K.L.; Sung, B.; Cupples, W.A.; Moore, L.C.; Chon, K.H. Detection of Low-Frequency Oscillations in Renal Blood Flow. Am. J. Physiol.-Ren. Physiol. 2009, 297, F155–F162. [Google Scholar] [CrossRef][Green Version]
- Zhong, Y.; Jan, K.-M.; Chon, K.H. Frequency Modulation between Low- and High-Frequency Components of the Heart Rate Variability Spectrum May Indicate Sympathetic-Parasympathetic Nonlinear Interactions. In Proceedings of the 2006 International Conference of the IEEE Engineering in Medicine and Biology Society, New York, NY, USA, 30 August–3 September 2006; pp. 6438–6441. [Google Scholar]
- Posada-Quintero, H.F.; Florian, J.P.; Orjuela-Cañón, Á.D.; Chon, K.H. Highly Sensitive Index of Sympathetic Activity Based on Time-Frequency Spectral Analysis of Electrodermal Activity. Am. J. Physiol.-Regul. Integr. Comp. Physiol. 2016, 311, R582–R591. [Google Scholar] [CrossRef]
- Veeranki, Y.R.; Ganapathy, N.; Swaminathan, R. Classification of Dichotomous Emotional States Using Electrodermal Activity Signals and Multispectral Analysis. In Challenges of Trustable AI and Added-Value on Health; IOS Press: Amsterdam, The Netherlands, 2022; pp. 941–942. [Google Scholar]
- Monti, A.; Medigue, C.; Mangin, L. Instantaneous Parameter Estimation in Cardiovascular Time Series by Harmonic and Time-Frequency Analysis. IEEE Trans. Biomed. Eng. 2002, 49, 1547–1556. [Google Scholar] [CrossRef]
- Huang, N.E.; Shen, Z.; Long, S.R.; Wu, M.C.; Shih, H.H.; Zheng, Q.; Yen, N.-C.; Tung, C.C.; Liu, H.H. The Empirical Mode Decomposition and the Hilbert Spectrum for Nonlinear and Non-Stationary Time Series Analysis. Proc. R. Soc. London. Ser. A Math. Phys. Eng. Sci. 1998, 454, 903–995. [Google Scholar] [CrossRef]
- Indolia, S.; Goswami, A.K.; Mishra, S.P.; Asopa, P. Conceptual Understanding of Convolutional Neural Network—A Deep Learning Approach. Procedia Comput. Sci. 2018, 132, 679–688. [Google Scholar] [CrossRef]
- Smirnov, E.A.; Timoshenko, D.M.; Andrianov, S.N. Comparison of Regularization Methods for ImageNet Classification with Deep Convolutional Neural Networks. AASRI Procedia 2014, 6, 89–94. [Google Scholar] [CrossRef]
- Wang, J.; Lin, J.; Wang, Z. Efficient Convolution Architectures for Convolutional Neural Network. In Proceedings of the 2016 8th International Conference on Wireless Communications & Signal Processing (WCSP), Yangzhou, China, 13–15 October 2016; pp. 1–5. [Google Scholar]
- Tivive, F.H.C.; Bouzerdoum, A. Efficient Training Algorithms for a Class of Shunting Inhibitory Convolutional Neural Networks. IEEE Trans. Neural Netw. 2005, 16, 541–556. [Google Scholar] [CrossRef] [PubMed]
- Lecun, Y.; Bottou, L.; Bengio, Y.; Haffner, P. Gradient-Based Learning Applied to Document Recognition. Proc. IEEE 1998, 86, 2278–2324. [Google Scholar] [CrossRef]
- Hastie, T.; Tibshirani, R.; Friedman, J. The Elements of Statistical Learning; Springer Series in Statistics; Springer: New York, NY, USA, 2009; ISBN 978-0-387-84857-0. [Google Scholar]
- Veeranki, Y.R.; Kumar, H.; Ganapathy, N.; Natarajan, B.; Swaminathan, R. A Systematic Review of Sensing and Differentiating Dichotomous Emotional States Using Audio-Visual Stimuli. IEEE Access 2021, 9, 124434–124451. [Google Scholar] [CrossRef]
- Veeranki, Y.R.; Ganapathy, N.; Swaminathan, R. Analysis of Fluctuation Patterns in Emotional States Using Electrodermal Activity Signals and Improved Symbolic Aggregate Approximation. Fluct. Noise Lett. 2021, 21, 2250013. [Google Scholar] [CrossRef]
- Faust, O.; Acharya, U.R.; Min, L.C.; Sputh, B.H.C. Automatic Identification of Epileptic and Background Eeg Signals Using Frequency Domain Parameters. Int. J. Neural Syst. 2010, 20, 159–176. [Google Scholar] [CrossRef]
- Acharya, U.R.; Chua, C.K.; Lim, T.-C.; Dorithy; Suri, J.S. Automatic Identification of Epileptic Eeg Signals Using Nonlinear Parameters. J. Mech. Med. Biol. 2009, 09, 539–553. [Google Scholar] [CrossRef]
- Acharya, U.R.; Yanti, R.; Swapna, G.; Sree, V.S.; Martis, R.J.; Suri, J.S. Automated Diagnosis of Epileptic Electroencephalogram Using Independent Component Analysis and Discrete Wavelet Transform for Different Electroencephalogram Durations. Proc. Inst. Mech. Eng. H 2013, 227, 234–244. [Google Scholar] [CrossRef] [PubMed]
- Sharmila, A.; Aman Raj, S.; Shashank, P.; Mahalakshmi, P. Epileptic Seizure Detection Using DWT-Based Approximate Entropy, Shannon Entropy and Support Vector Machine: A Case Study. J. Med. Eng. Technol. 2018, 42, 1–8. [Google Scholar] [CrossRef] [PubMed]
- Yuan, Y.; Xun, G.; Jia, K.; Zhang, A. A Novel Wavelet-Based Model for EEG Epileptic Seizure Detection Using Multi-Context Learning. In Proceedings of the 2017 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), Kansas City, MO, USA, 13–16 November 2017; pp. 694–699. [Google Scholar]
- Hussein, R.; Palangi, H.; Ward, R.K.; Wang, Z.J. Optimized Deep Neural Network Architecture for Robust Detection of Epileptic Seizures Using EEG Signals. Clin. Neurophysiol. 2019, 130, 25–37. [Google Scholar] [CrossRef] [PubMed]
- Chanu, M.M.; Singh, N.H.; Thongam, K. An Automated Epileptic Seizure Detection Using Optimized Neural Network from EEG Signals. Expert Syst. 2023, 40, e13260. [Google Scholar] [CrossRef]
- Islam, M.S.; Thapa, K.; Yang, S.-H. Epileptic-Net: An Improved Epileptic Seizure Detection System Using Dense Convolutional Block with Attention Network from EEG. Sensors 2022, 22, 728. [Google Scholar] [CrossRef] [PubMed]
- Bhattacharyya, A.; Pachori, R.B. A Multivariate Approach for Patient-Specific EEG Seizure Detection Using Empirical Wavelet Transform. IEEE Trans. Biomed. Eng. 2017, 64, 2003–2015. [Google Scholar] [CrossRef] [PubMed]
- Sharma, M.; Pachori, R.B.; Rajendra Acharya, U. A New Approach to Characterize Epileptic Seizures Using Analytic Time-Frequency Flexible Wavelet Transform and Fractal Dimension. Pattern Recognit. Lett. 2017, 94, 172–179. [Google Scholar] [CrossRef]
- Goel, S.; Agrawal, R.; Bharti, R.K. Automated Detection of Epileptic EEG Signals Using Recurrence Plots-Based Feature Extraction with Transfer Learning. Soft Comput. 2023. [Google Scholar] [CrossRef]
- Guo, L.; Rivero, D.; Dorado, J.; Munteanu, C.R.; Pazos, A. Automatic Feature Extraction Using Genetic Programming: An Application to Epileptic EEG Classification. Expert Syst. Appl. 2011, 38, 10425–10436. [Google Scholar] [CrossRef]
- Abdulhay, E.; Elamaran, V.; Chandrasekar, M.; Balaji, V.S.; Narasimhan, K. Automated Diagnosis of Epilepsy from EEG Signals Using Ensemble Learning Approach. Pattern Recognit. Lett. 2020, 139, 174–181. [Google Scholar] [CrossRef]
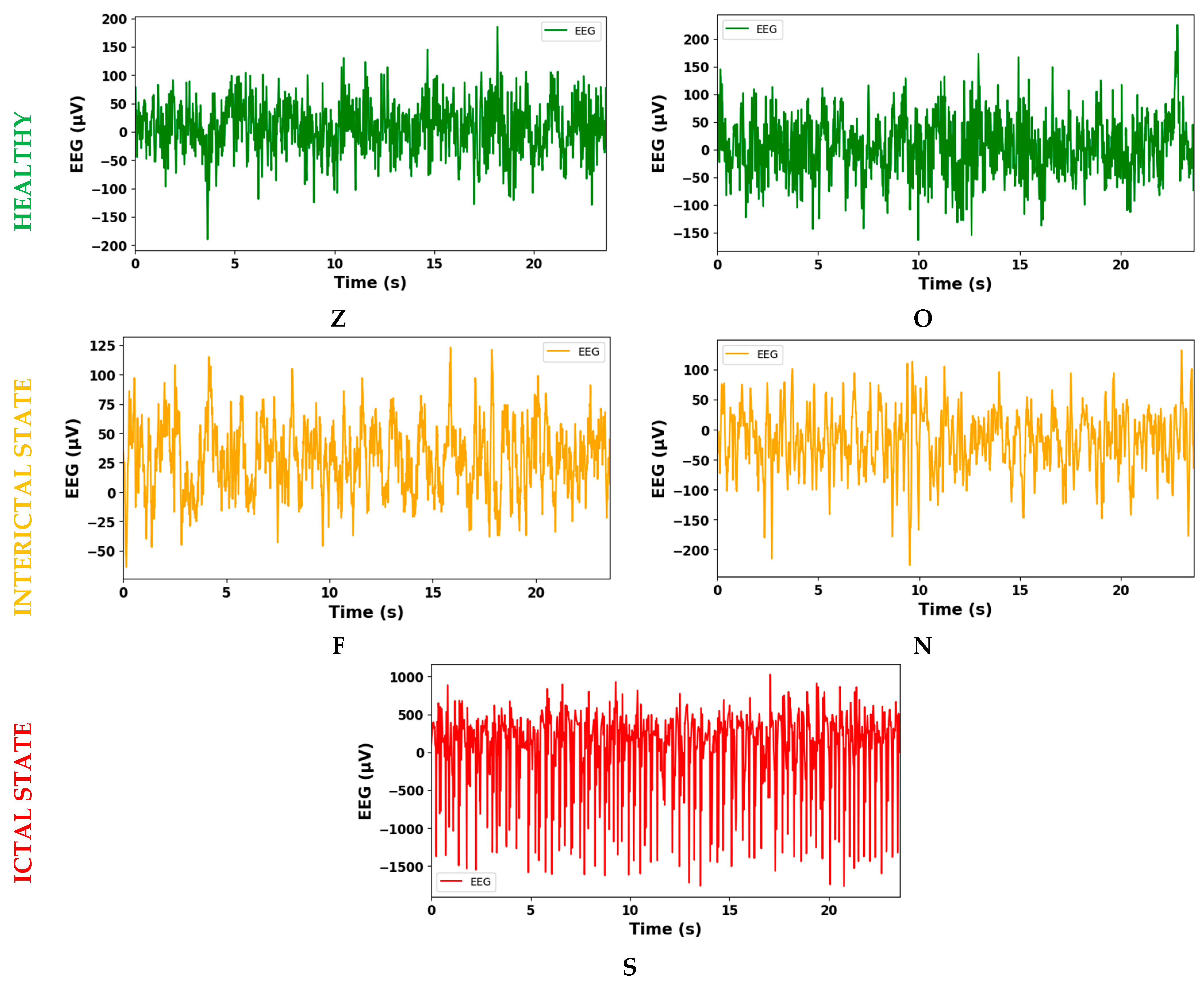








| Healthy Subject | Epilepsy Subject | |||
|---|---|---|---|---|
| Eyes open | Eyes closed | Seizure-free interval (interictal state) | Seizure activity (ictal state) | |
| Hippocampal formation | Hippocampal formation of the opposite hemisphere of the brain | |||
| Z | O | N | F | S |
| Case | Classes | Description |
|---|---|---|
| I |
| Healthy vs. Seizure |
| II |
| Healthy vs. Interictal vs. Ictal |
| III |
| Healthy vs. Epileptic |
| Metric | Description | Expression |
|---|---|---|
| Acc | Measures the proportion of predictions that are correct. | |
| Pre | Quantifies the accuracy of positive predictions. | |
| Rec | Evaluates the model’s ability to identify all actual positives. | |
| F1 | Harmonic means of precision and recall. It provides a balance between precision and recall, considering both false positives and false negatives. |
Disclaimer/Publisher’s Note: The statements, opinions and data contained in all publications are solely those of the individual author(s) and contributor(s) and not of MDPI and/or the editor(s). MDPI and/or the editor(s) disclaim responsibility for any injury to people or property resulting from any ideas, methods, instructions or products referred to in the content. |
© 2023 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Veeranki, Y.R.; McNaboe, R.; Posada-Quintero, H.F. EEG-Based Seizure Detection Using Variable-Frequency Complex Demodulation and Convolutional Neural Networks. Signals 2023, 4, 816-835. https://doi.org/10.3390/signals4040045
Veeranki YR, McNaboe R, Posada-Quintero HF. EEG-Based Seizure Detection Using Variable-Frequency Complex Demodulation and Convolutional Neural Networks. Signals. 2023; 4(4):816-835. https://doi.org/10.3390/signals4040045
Chicago/Turabian StyleVeeranki, Yedukondala Rao, Riley McNaboe, and Hugo F. Posada-Quintero. 2023. "EEG-Based Seizure Detection Using Variable-Frequency Complex Demodulation and Convolutional Neural Networks" Signals 4, no. 4: 816-835. https://doi.org/10.3390/signals4040045
APA StyleVeeranki, Y. R., McNaboe, R., & Posada-Quintero, H. F. (2023). EEG-Based Seizure Detection Using Variable-Frequency Complex Demodulation and Convolutional Neural Networks. Signals, 4(4), 816-835. https://doi.org/10.3390/signals4040045








