1. Introduction
Mode-clustering is a clustering analysis method that partitions the data into groups by the local modes of the underlying density function [
1,
2,
3,
4]. A density local mode is often a signature of a cluster, so mode-clustering leads to clusters that are easy to interpret. In practice, we estimate the density function from the data and perform mode-clustering via the density estimator. When we use a kernel density estimator (KDE), there exists a simple and elegant algorithm called the mean-shift algorithm [
5,
6,
7] that allows us to compute clusters easily. The mean-shift algorithm has made the mode-clustering a numerically friendly problem.
When applied to a scientific problem, we often use a clustering method to gain insight from the data [
8,
9]. Sometimes, finding clusters is not the ultimate goal. The connectivity among clusters may yield valuable information for scientists. To see this, consider the galaxy sample from the Sloan Digital Sky Survey [
10] in
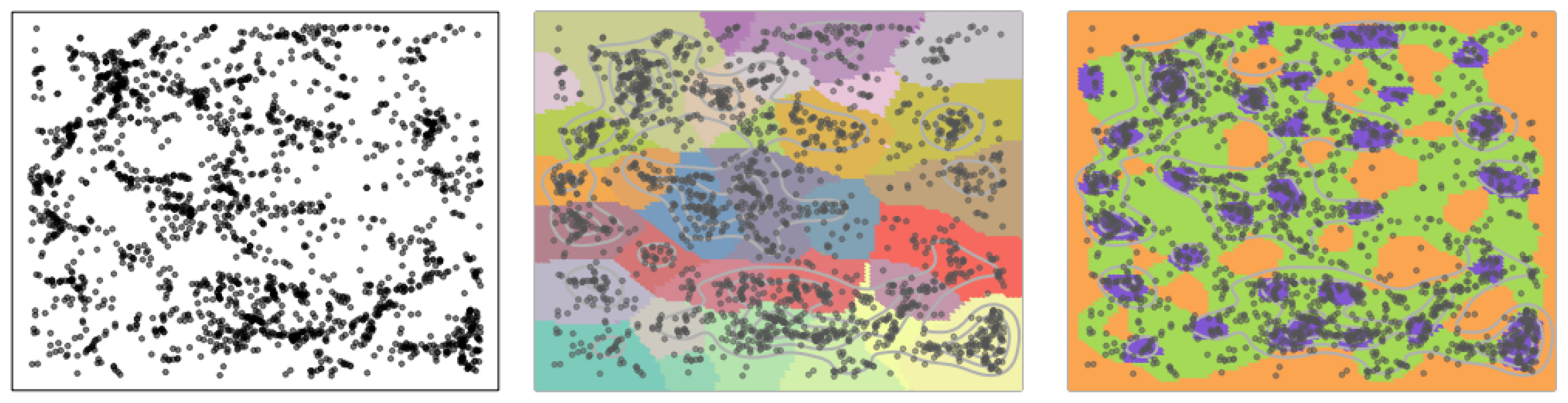
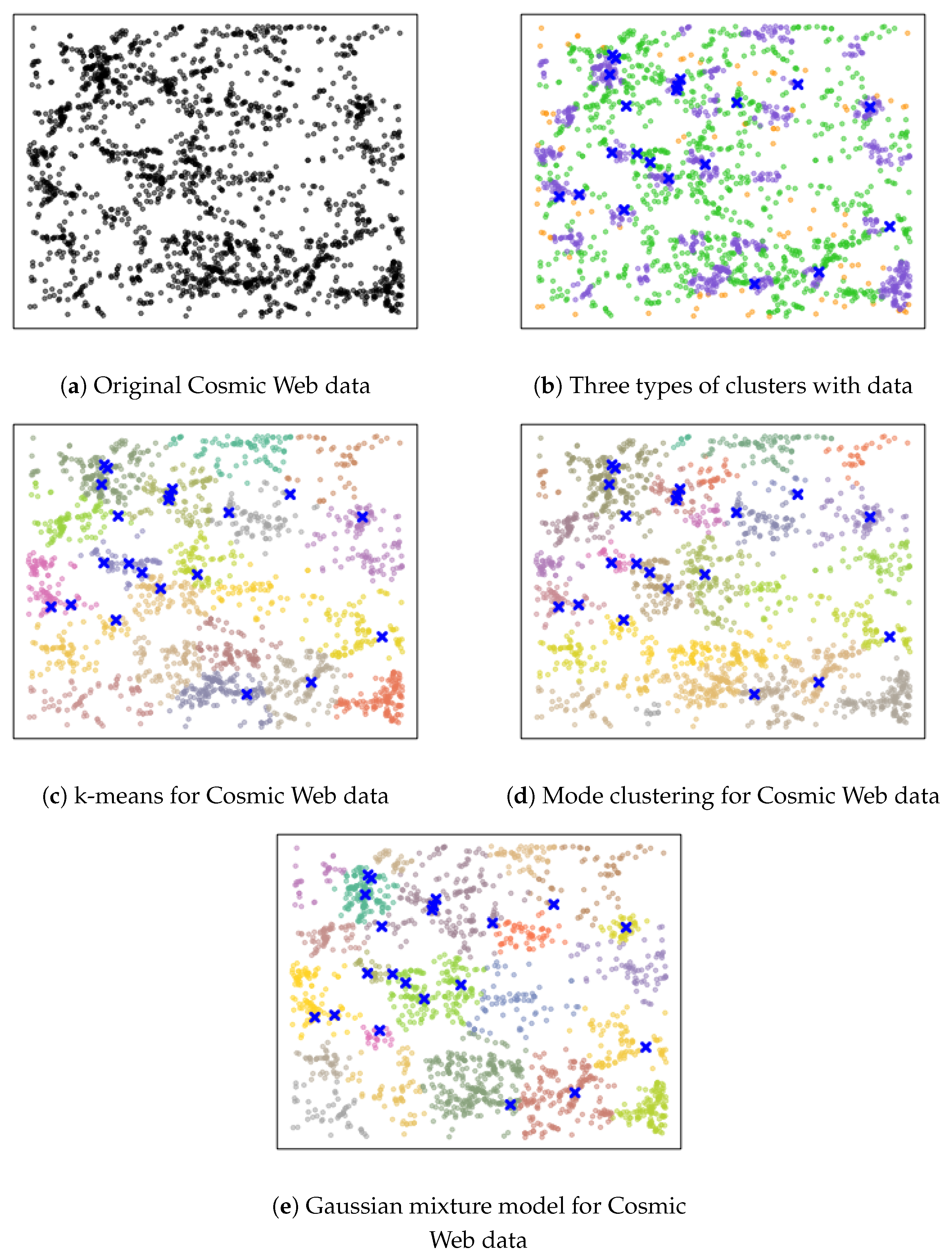
Figure 1. While the original data is 3D, here we use a 2D slice of the original data to illustrate the idea. Each black dot indicates the location of a galaxy at a particular location in the sky. Astronomers seek to find clusters of galaxies and their connectivity, since these quantities (clusters and their connections) are associated with the large-scale structures in the universe. Our method finds the underlying connectivity structures without assuming any parametric form of the underlying distribution. In the middle panel, we display the results by the usual mode-clustering method, which only shows clusters, but not how they connect with each other. On the other hand, our proposed method is given in the right panel, which finds a set of dense clusters (purple regions) along with some regions serving as bridges connecting clusters (green areas) and a set of low-density regions (yellow regions). Thus, our clustering method allows us to better identify the structures of galaxies.
We improve the usual mode-clustering method by (1) adding additional clusters that can further partition the entire sample space, and (2) assigning an attribute label to each cluster. The attribute label will indicate if this cluster is a ‘robust cluster’ (a cluster around a local mode; purple regions in
Figure 1), a ‘boundary cluster’ (a cluster bridging two or more robust clusters; green regions in
Figure 1), or an ‘outlier cluster’ (a cluster representing low-density regions; yellow regions in
Figure 1). With this refined clustering result, we gain further insights into the underlying density function and are able to infer the intricate structure behind the data. Furthermore, we can apply our improved clustering method to the two sample tests. In this case, we can identify the local differences between the two populations and provide a more sensitive result. Note that in the usual case of cluster analysis, adding more clusters is not a preferred idea. However, if our goal is to detect the underlying structures (such as finding the connectivity of high-density regions in the galaxy data in
Figure 1), using more clusters as an intermediate step to find connectivity could be a plausible approach.
To summarize, our main contributions are as follows:
Related work. The idea of using local modes to cluster observations can be dated back to [
5], where the authors used local modes of the KDE to cluster observations and propose the mean-shift algorithm for this purpose [
5,
11]. mode-clustering has been widely studied in statistics and the machine-learning community [
3,
4,
7,
12,
13,
14]. However, the KDE is not the only option for mode-clustering—[
1,
15] proposed a Gaussian mixture model method, and [
16] used a fuzzy clustering algorithm, and [
17] introduced a nearest-neighbor density method.
Outline. The paper is organized as follows. We start with a brief review on mode-clustering in
Section 2 and formally introduce our method in
Section 3. In
Section 4, we combine the two-sample test and our approach to create a local two-sample test. We use simulations to illustrate our method on simple examples in
Section 5. We show the applicability of our approach to three real datasets in
Section 6. Finally, we study both statistical and computational theories of our method in
Section 7.
2. Review of Mode-Clustering
We start with a review of mode-clustering [
2,
4,
12,
18]. The concept of mode-clustering is based on the rationale of associated clusters to the regions around the modes of the density. When the density function is estimated by the kernel density estimator, there is an elegant algorithm called the mean-shift algorithm [
5] that can easily perform the clustering.
In more detail, let
p be a probability density function with a compact support
. Starting at any point
x, mode-clustering creates a gradient ascent flow
such that
Namely, the flow
starts at point
x and moves according to the gradient at the present location. Let
be the destination of the flow
. According to the Morse theory [
19,
20], when the function is smooth (being a Morse function), such a flow converges to a local maximum of
p except for starting points in a set of the Lebesgue measure 0. The mode-clustering partitions the space according to the destination of the gradient flow, that is, for two points
, they will be assigned to the same cluster if
. For a local mode
, we define its basin of attraction as
. The basin of attraction describes the set of points that belongs to the same cluster.
In practice, we do not know
p, so we replace it by a density estimator,
. A common approach to estimate
p as the kernel density estimator, in which
is
where
K is a smooth function (also known, according to the Morse theory, as the kernel function), such as a Gaussian kernel, and
is the smoothing bandwidth that determines the amount of smoothness. Since we used a nonparametric density estimator, we did not need to assume any parametric assumptions on the shape of the distribution.) With this choice, we the define a sample analogue to the flow
as
and partition the space according to the destination of
.
3. Clustering via the Gradient of Slope
3.1. Refining the Clusters by the Gradient of Slope
As is mentioned previously, the mode-clustering has some limitations that the resulting clusters do not provide enough information on the finer structure of the density. To resolve this problem, we introduce a new clustering method by considering gradient descent flows of the ‘slope’ function. Let be the gradient of p. Define the slope function of p as . Namely, the slope function is the squared amplitude of the density gradient.
An interesting property of the slope function is that the minimal points
form the collection of critical points of
p, so it contains local modes of
p as well as other critical points, such as saddle points and local minima. According to the Morse theory [
21,
22], there is a saddle point between two nearby local modes when the function is a Morse function. A Morse function is a smooth function
f, such that all eigenvalues of Hessian Matrix of
f at every critical point are away from 0. This implies that saddle points may be used to bridge connecting regions around two local modes.
With this insight, we propose to create clusters using the gradient ‘descent’ flow of
. Let
be the gradient of the slope function. Given a starting point
, we construct a gradient descent flow as follows:
That is, is a flow starting from x and moving along the direction of . Similar to mode-clustering, we use the destination of gradient flows to cluster the entire sample space.
Note that if the slope function s is a Morse function, the corresponding PDF p will also be a Morse function, as described in the following Lemma.
Lemma 1. If is a Morse function, then is a Morse function.
Throughout this paper, we will assume that the slope function is Morse. Thus, the corresponding PDF will also be a Morse function and all critical points of the PDF will be well-separated.
3.2. Type of Clusters
Recall that is the collection of critical points of density p. Let be the collection of local minima of the slope function . It is easy to see , since any critical point of p has gradient 0, so it is also a local minimum of s.
Thus, the gradient flow in Equation (
1) leads to a partition of the sample space. Specifically, let
be the destination of the gradient flow
. For an element
, let
be its basin of attraction.
We use the sign of eigenvalues of
to assign an additional attribute to each basin, so the set
forms a collection of meaningful disjoint regions. In more detail, for a critical point
such that
for a small threshold
, its
is classified according to
where
is the
l-th ordered eigenvalue of
(
). In the case of
, we always assign it as an outlier cluster. Note that the threshold
was added to stabilize the numerical calculation. In other words, we refer to a basin of attraction in
as a robust cluster if
is a local mode of
p. If
m is a local minimum of
p, then we call its basin of attraction an outlier cluster. The remaining clusters, which are regions connecting robust clusters, are denoted as boundary cluster. Note that the regions outside the support are, by definition, a set of local minima. We assign the same cluster label to those
x whose destination
is outside the support, which is an outlier cluster.
Our classification of
is based on the following observations. Regions around local modes of
p are where we have strong confidence that these points should belong to the cluster represented by their nearby local modes. Regions around local minima of
p are the low-density areas where we should treat them as anomaly points/outliers.
Figure 1 provides a concrete example that our clustering method could lead to more scientific insight–the connectivity among robust clusters may reveal intricate structure of the underlying distribution.
Defining different types of clusters allows us to partition the whole space into meaningful subregions. Given a random sample, to assign the cluster label to each of them, we simply examine which basins of attraction these data points fall in and pass the cluster labels from the regions to the data points. After assigning cluster labels to data points, the cluster categories in Equation (
2) provide additional information about the characteristics of each data point. Those data points in robust clusters are data points that are highly clustered together; points in the outlier clusters are data points in low-density regions, which could be viewed as anomalies; the rest of points are in the boundary clusters, where these points are not well-clustered and are on the connection regions among different robust clusters.
3.3. Estimators
The above procedure is defined when we have access to the true PDF p. In practice, we do not know p, but we have an IID random sample from p with a compact support So we estimate p using and then use the estimated PDF to perform the above clustering task.
While there are many choices of density estimators, we consider the kernel density estimator (KDE) in this paper, since it has a nice form and its derivatives are well-established [
14,
23,
24,
25]. In more detail, the KDE is
where
K is a smooth function (also known as the kernel function) such as a Gaussian kernel, and
is the smoothing bandwidth that determines the amount of smoothness. Note that the bandwidth
h in the KDE could be replaced by
that depends on each observation. This is called the variable bandwidth KDE in Breiman et al. [
26]. However, since the choice of how
depends on each observation is a non-trivial problem, so to simplify the problem, we set all bandwidths to be the same.
Based on
, we first construct a corresponding estimated flow using
:
An appealing feature is that
has an explicit form:
where
and
are the estimated density gradient and Hessian matrix of
p. Thus, to numerically construct the gradient flow
, we update
x by
where
is the learning rate parameter. Algorithm 1 summarizes the gradient descent approach.
| Algorithm 1: Slope minimization via gradient descent. |
1. Input: and a point x.
2. Initialize and iterate the following equation until convergence: ( is a step size that could be set to a constant)
3. Output: . |
With an output from Algorithm 1, we can group observations into different clusters, with each cluster labeled by a local minimum of
. We assign an attribute to each cluster via the rule in Equation (
2). Note that the smoothing bias could cause some biases around the boundary of clusters. However, when
, this bias will asymptotically be negligible.
4. Enhancements in Two-Sample Tests
Our clustering method can be used as a localized two-sample test. An overview of the idea is as follows. Given two random samples, we first merge them and use clustering method to form partitions of the sample space. Under the null hypothesis, the two samples are from the same distribution, so the proportion of each sample within each cluster should be similar. By comparing the difference in proportion, we obtain a localized two-sample test. Algorithm 2 summarizes the procedure.
In more detail, suppose we want to compare two samples and . Let and . The null hypothesis we want to test is against .
Under
, the two samples are from the same distribution, so they have the same PDF
q. We first pull both samples together to form a joint dataset
We then compute the KDE
using
and compute the corresponding estimated slope function
and apply Algorithm 1 to form clusters. Thus, we obtain a partition of
. Under
, the proportion of Sample 1 in each cluster should be roughly the same as the global proportion
. Therefore, we can apply a simple test of the proportion within each cluster to obtain a
p-value. In practice, we often only focus on the robust and boundary clusters and ignore the outlier clusters because of sample size consideration. Let
be the robust and boundary clusters, and
be the global proportion, and
be the observed proportion of cluster
. We use the test statistic
where
is the total number of the pulled sample within cluster
, when
is true and the test statistic
follows from a standard normal distribution asymptotically. Note that since we are conducting multiple tests, we reject the null hypothesis after applying the Bonferroni correction.
| Algorithm 2: Local two-sample test. |
1. Combine two samples ( and ) into one, called and compute from Equation (6).
2. Construct a kernel density estimator using and its slope function and apply Algorithm 1 to form clusters based on the convergent point.
3. Assign an attribute to each cluster according to Equation (2).
4. Let robust clusters and boundary clusters be , where for each j.
5. For each cluster , compute from Equation (7) and construct Z statistic:
Find the corresponding p-value .
6. Reject if for some j under the significance level .
|
We can apply this idea to other clustering algorithms. However, we need to be very careful when implementing it because we are using data twice–first to form clusters, then again to do two-sample tests. This could inflate the Type 1 error. Our approach is asymptotically valid because the clusters from the estimated slope converge to the clusters of the population slope (see
Section 7). Note that our method may not control the Type 1 error in the finite sample situation, but our simulation results in
Section 5.2 show that this procedure still controls the Type 1 error. This might be due to the conservative result of the Bonferroni correction.
The advantage of this new two-sample test is that we are using the local information, so if the two distributions only differ in a small region, this method will be more powerful than a conventional two-sample test. In particular, the robust clusters are often the ones with more power because they have a higher sample size, and the bumps in the pulled sample’s density could be created by a density bump of one sample but not the other, leading to a region with high testing power. In
Section 5, we demonstrate this through some numerical simulations.
4.1. An Approximation Method
The major computational burden of Algorithm 2 comes from Step 2, where we apply Algorithm 1 to ‘every observation’. This may be computationally heavy if the sample size is large. Here we propose a quick approximation to the clustering result.
Instead of applying Algorithm 1 to every observation, we randomly subsample the original data (large dimension) or create a grid (low dimension) of points and only apply Algorithm 1 to this smaller set of points. This gives us an approximated set of local minima of the slope function. We then assign a cluster label of each observation according to the ‘nearest’ local minima.
5. Simulations
In this section, we demonstrate the applicability of our method by applying it to some simulation setups. Note that in practice, we need to choose the smoothing bandwidth
h in the KDE. Silverman’s rule [
27] is one of the most popular methods for bandwidth selection. The idea is to find the optimal bandwidth by minimizing the mean integrated squared error of the estimated density. Silverman [
27] proposed to use the normal density to approximate the second derivative of the true density, and use the interquartile range providing a robust estimation of the sample standard deviation. For the univariate case, it is defined as follows:
where
is the sample standard deviation and
is the interquartile range. As discussed earlier, we choose
where
is a constant. This choice is motivated by theoretical analysis in
Section 7 (Theorem 1). In practice, we do not know
, so we applied a modification of Silverman’s rule [
27]:
where
is the standard deviation of the samples on
kth dimension,
is the interquartile range on
kth dimension, and
. Note that our procedure involves estimating both the gradient and Hessian of the PDF. The optimal bandwidth of the two quantities are different, so one may apply two separated bandwidths for gradient and Hessian estimation. However, our empirical studies show that a single bandwidth (optimal for Hessian estimation) still leads to reliable results. Note that this bandwidth selector tends to oversmooth the data in the sense that some density peaks in Figure 6b were not detected (not in purple color).
5.1. Clustering
Two-Gaussian mixture. We sample points from a mixture of two-dimensional normals and with equal proportions under the following three scenarios:
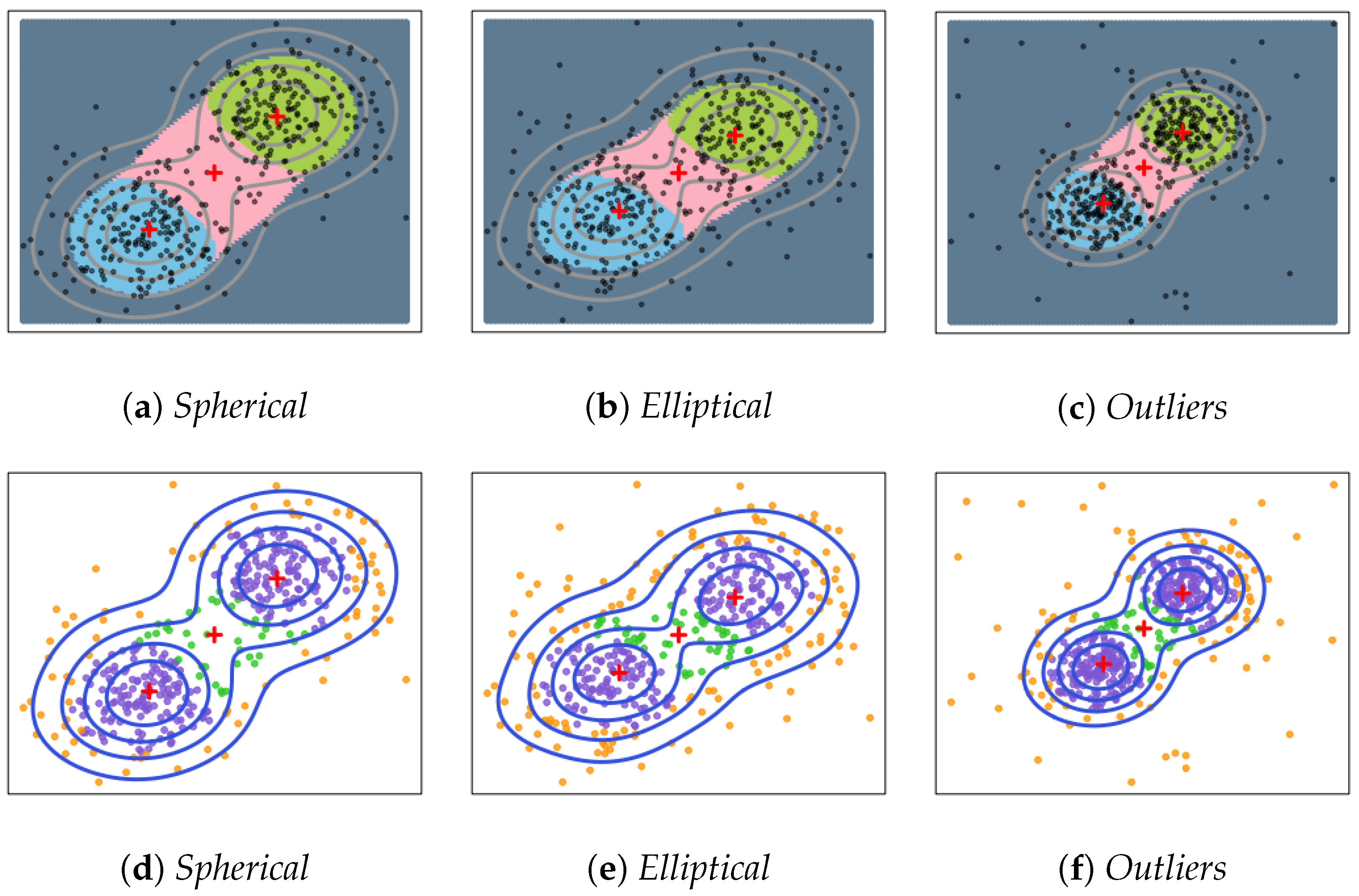
Spherical: = 0, , and .
Elliptical: = 0, , and . (Note that these clusters are elongated in noise directions.)
Outliers: Same construction as Spherical, but with 60 random points (noise) from a uniform distribution over . By design, the outliers differ in such a way that they can only add a little ambiguity.
Note that is the ith standard basis vector, and is the identity matrix. For each scenario, we apply the gradient flow method and draw the contour. If points are outliers, their destinations go to infinity. Thus, we set a threshold to stop them from moving and assign them to outlier clusters.
Figure 2 demonstrates that we identify both two clusters and the boundary of these two clusters. Each colored region is the basin of attraction of a local minimum of
in the picture (a–c). Picture (d–f) provide examples of data points clustering. Given the setting of two equal-sized Gaussian mixture, it is straightforward to verify that the gradient flow algorithm can successfully distinguish points according to their destinations. The purple points represent points that belong to corresponding clusters with strong confidence, while green points represent points in low-density areas that belong to the connection regions among clusters. The yellow points represent points that are not important to any of the clusters. In summary, our proposed method performs well and is not affected by the changes of covariance and outliers.
Four-Gaussian mixture. To show how boundary clusters can serve as bridges among robust clusters, we consider a four-Gaussian mixture. We sample
from a mixture of four two-dimensional normals
,
,
and
with equal proportion. Then we apply our method and display the result in
Figure 3. Each colored region is the basin of attraction of a local minimum of
. The red ‘+’s are the corresponding local minima to each of the basin of attraction. Clearly, we see how robust clusters are connected by the boundary clusters so the additional attributes provide useful information on the connectivity among density modes.
Comparison. To better illustrate the strength of our proposed method, we generate an unbalanced four-Gaussian mixture. We sample
from a mixture of four two-dimensional normals
,
,
and
with proportion
, respectively. Then we apply our method and compare it with the density-based spatial clustering of applications with noise (DBSCAN) [
28] in
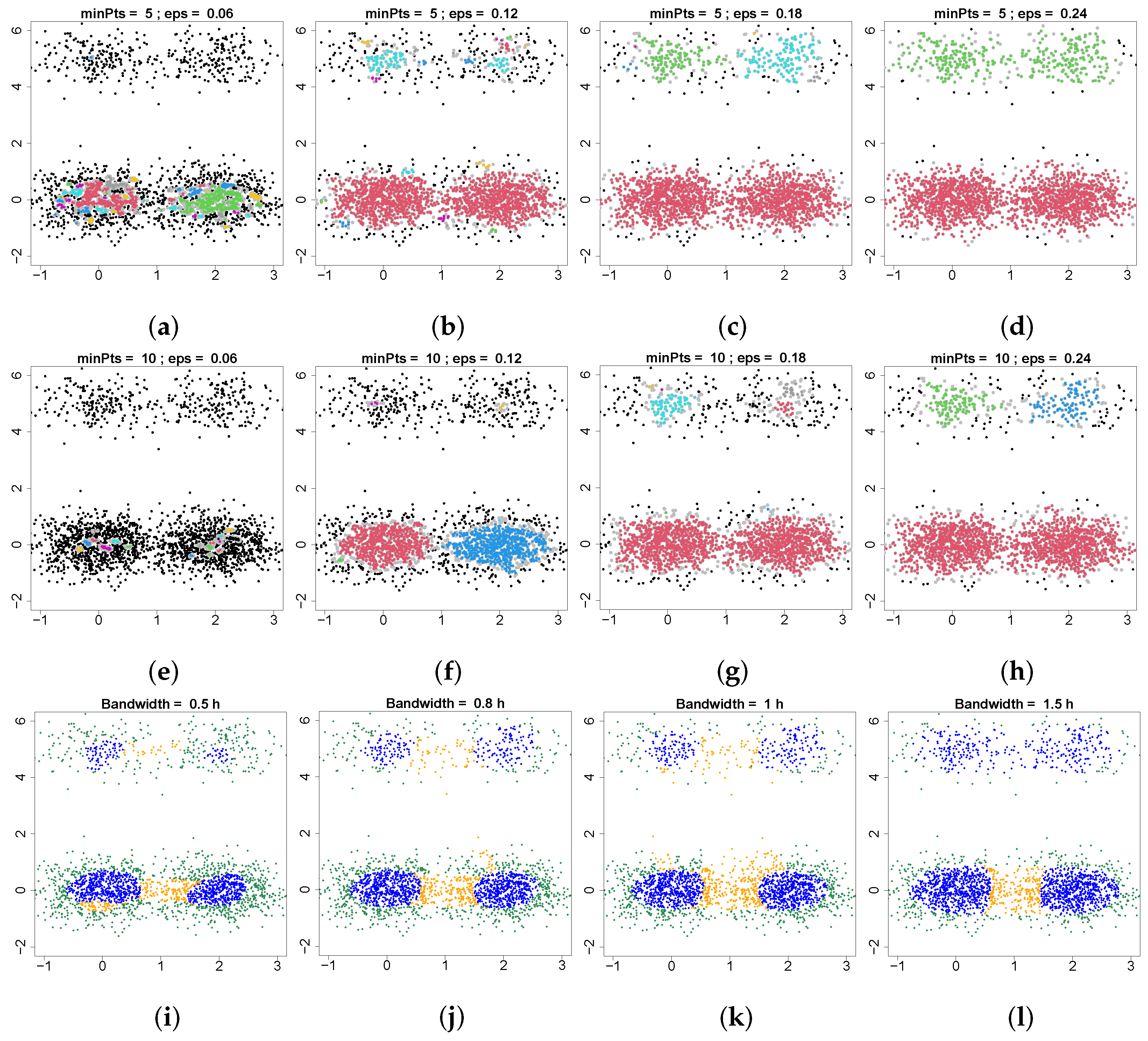
Figure 4. DBSCAN is a classical non-parametric, density-based clustering method that estimates the density around each data point by counting the number of points in a certain neighborhood and applies a threshold minPts to identify core, border and noise points. DBSCAN requires two parameters: the minimum number of nearby points required to form a core point (minPts) and the radius of a neighborhood with respect to a certain point (eps). Two points are connected if they are within the distance of eps. Clusters are the connected components of connected core points. Border points are points connected to a core point, but which do not have enough neighbors to be a core point. Here, we investigate the feasibility of using border points to detect the connectivity of clusters. These two parameters, minPts and eps, are very hard to choose. In the top two rows of
Figure 4, we set minPts equal to 5 and 10 and change the value of eps to see if we can find the connectivity of core points using border points (gray points). Our results show that it is not possible to use border points to find the connectivity of the top two clusters and the bottom two clusters at the same time. When we are able to detect the connectivity of bottom two clusters (panel (f)), we are not able to find the top two clusters. On the other hand, when we can find the connectivity of the top two clusters (panel (c,h)), the bottom two clusters have already merged into a single cluster. The limitation of DBSCAN is that it is based on the density level set, so when the structures involve different density values, DBSCAN will not be applicable. In contrast, our method only requires one parameter, bandwidth, and it has good performance in this case. From
Figure 4i–l, our method detects four robust clusters and their boundaries correctly. In addition, this result also shows that our method is robust to the bandwidth selection.
5.2. Two-Sample Test
In this section, we carry out simulation studies to evaluate the performance of the two-sample test in
Section 4. We compare our method to three other popular approaches: the energy test [
29], the kernel test [
30], and KS [
31] tests based on each of the two variables.
Our simulation is designed as follows. We draw random samples from a two-Gaussian mixture model in Equation (
9):
where
is a cumulative distribution function of normal distribution. For the first group, we choose the parameters as
,
,
,
, and
.
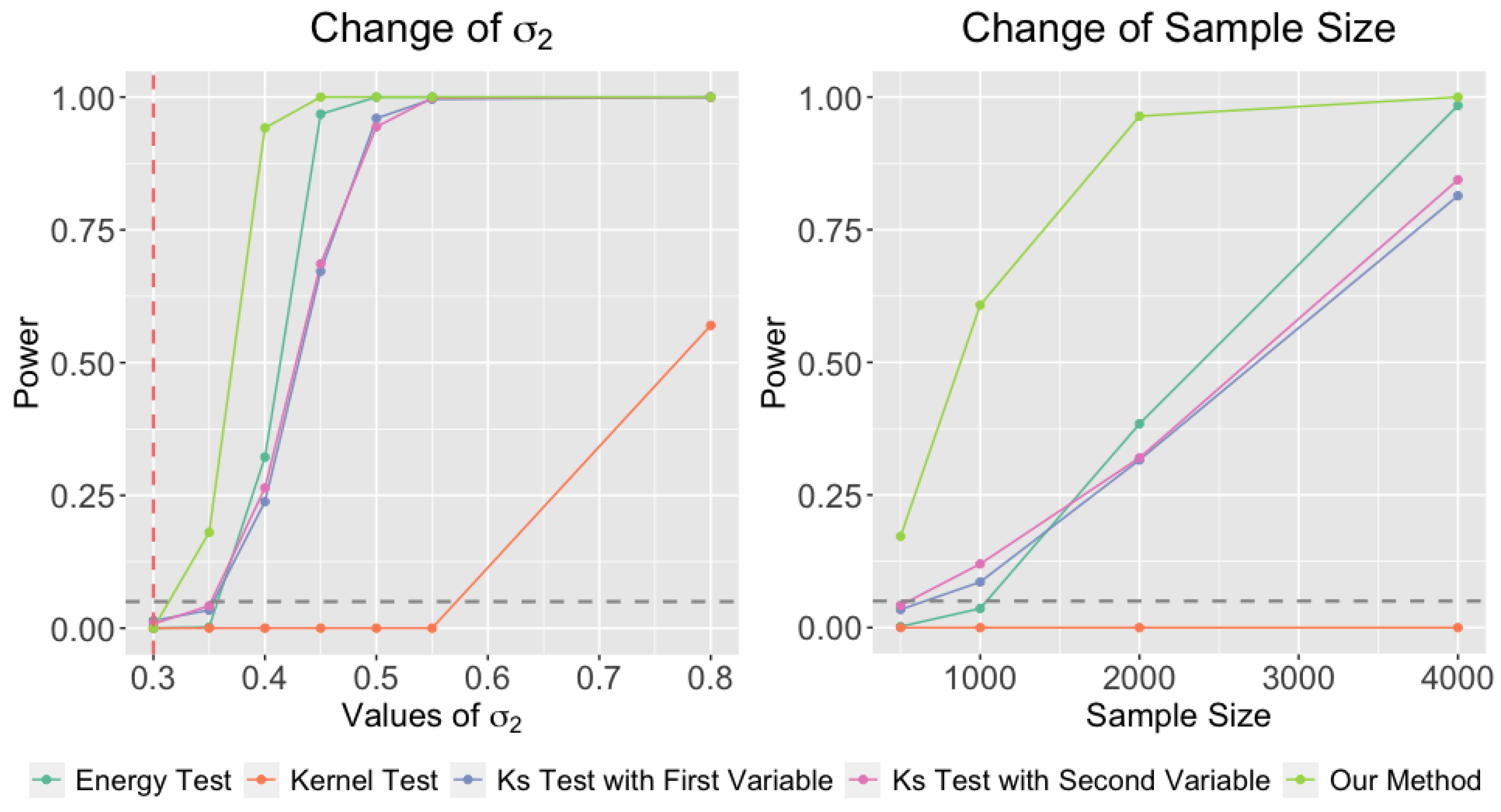
In our first experiment (left panel of
Figure 5), we generate the second sample from a Gaussian mixture with identical setup, except that the second covariance matrix
, and we gradually increase
from 0.3 (
is correct) to 0.8 to see how the power of the test changes. We generate
observations in both samples and repeat the process 500 times to compute the power of the test. This experiment investigates the power as a function of signal strength.
In the second experiment (right panel of
Figure 5), we consider a similar setup except that we fix
and vary the sample size from
to
and examine how the power changes under different sample size. This experiment examines the power as a function of sample size.
In both experiments, all methods control the Type 1 errors. However, our method has better power in both experiments compared to the other alternatives. Our method is more powerful because we utilize the local information from clustering. In this simulation setup, the difference between the two distributions is the width of second Gaussian component. Our method is capable of capturing this local difference and using it as evidence in the hypothesis test.
Finally, we would like to emphasize again that two-sample test after clustering has to be used with caution; we are using data twice, so we may not be able to control the Type 1 error. One needs to theoretically justify that the resulting clusters converge to a population limit and apply numerical analysis to investigate the finite-sample coverage.
7. Theory
In this section, we study both statistical and algorithmic convergence of our method. We start with the convergence of estimated minima to the population minima along with the convergence of the gradient flow. Then we discuss the algorithmic convergence of Algorithm 1.
For a set
, we denote its cardinality by
. For a function
f, we define
to be the
-norm. Let
and
be the gradient and Hessian matrix of
f, respectively. We define
as the element-wise
-norm for
l-th order derivatives of
f. Specifically,
,
for
and
. A twice-differentiable function
f is called Morse [
19,
20,
21] if all eigenvalues of the Hessian matrix of
f at critical points are away from 0.
Recall that our data are random sample from a PDF and . Additionally, , and are the estimated PDF, gradient, and Hessian matrix, respectively. In our analysis, we consider the following assumptions.
Assumptions.
- (P)
The density function is four-times bounded and continuously differentiable.
- (L)
is a Morse function.
- (K)
The kernel K is four-times bounded and continuously differentiable. Moreover, the collection of kernel functions and their partial derivatives up to the third order satisfy the VC-type conditions in Giné and Guillou [
36]. See
Appendix A for more details.
Assumption (P) is slightly stronger than the conventional assumptions for density estimation that we need to be four-times differentiable. This is because we are working with gradient of ‘slope’, which already involves second derivatives. To control the bias, we need additionally two derivatives, leading to a requirement on the fourth-order derivatives. Assumption (L) is slightly stronger than the conventional Morse function assumption on
. We need the slope function to be Morse so that the gradient system is well-behaved. In fact, Assumption (L) implies that
is Morse function due to Lemma 1. Assumption (K) is a common assumption to ensure uniform convergence of a kernel-type estimator; see, for example [
37,
38].
7.1. Estimation Consistency
With the above assumption, we can show that the local minima of converge to the local minima of s.
Theorem 1 (Consistency of local minima of s). Assume (K), (P) and (L). Let be the bound for the partial derivatives of s up to the third order and denote the l-th largest eigenvalues of by (, where d is the dimension). Assume:
- (A1)
There exists such that for any point x with and , we have , where for and .
When is sufficiently small, we have
, and
for every point , there exists a unique element such that
Theorem 1 shows two results. First, asymptotically, there will be a one–one corresponding relationship between a population’s local minimum and an estimated local minimum. The second result shows the rate of convergence, which is the rate of estimating second derivatives. This is reasonable, since the local minima of s is defined through the gradient of , which requires second derivatives of p.
Note that the fourth-order derivative assumption (P) can be relaxed to a smoothed third-order derivative conditions. We use this stronger condition to simplify the derivation, since the global minima of
s are the critical points of
p, the consistency of estimating a global minimum only requires a third-order derivative (or a smooth second-order derivative) assumption; see, for example [
39,
40].
Theorem 1 also implies the rate of the set estimator
in terms of the Hausdorff distance. For given two sets
A,
B, their Hausdorff distance is
where
is the projection distance from point
x to the set
A.
Corollary 1. Assume (K),(P), (L), and (A1). When is sufficiently small, The above results describe the statistical consistency of the convergent points (local minima) of a gradient flow system. In what follows, we show that the gradient flows will also converge under the same set of assumptions.
Theorem 2 (Consistency of gradient flows).
Assume (K), (P) and (L). Then for a fixed point x, when , ,where and are the minimal and maximal eigenvalues of the Hessian matrix of s evaluated at the destination , and . Theorem 2 is mainly inspired by Theorem 2 in Arias-Castro et al. [
3]. It shows that starting at a given point
x, the estimated gradient flow
is a consistent estimator to the population gradient flow
. One may notice that this result shows that the convergence rate is slowed down by the factor
, which comes from the curvature of
s around the local minimum. This is due to the fact that when a flow is close to its convergent point (a local minimum), the speed of flow is decreasing until 0 (when it arrives at a minimum), so the eigenvalues determine the rate of how fast the speed of a flow decreases along a particular direction. When the eigengap (difference between
and
) is large, even a small perturbation could change the orientation of the flow drastically, leading to a slower convergence rate.
Remark. It is possible to obtain the clustering consistency in the sense that the clustering based on
s and
are asymptotically the same [
41]. In [
41], the authors placed conditions on the density function and showed that the mode-clustering of
leads to a consistent partition of the data compared to the mode-clustering of
p. If we generalize their conditions to the slope
s, we will obtain a similar clustering consistency result.
7.2. Algorithmic Consistency
In this section, we study the algorithmic convergence of Algorithm 1. For simplicity, we consider the case where the gradient descent algorithm is applied to
s. The convergence analysis of gradient descent has been well studied in the literature [
42,
43] under convex/concave setups. Our algorithm is a gradient descent algorithm but is applied to a non-convex scenario. Fortunately, if we consider a small ball around each local minimum, the function
s will still be a convex function, so the conventional techniques apply.
Specifically, we need an additional assumption that is slightly stronger than (L).
- (A2)
There are positive numbers such that for all , where and is a ball with center m and radius , all eigenvalues of Hessian matrix are above and .
The assumption (A2) is a local strongly convex condition.
Theorem 3 (Convergence of Algorithm 1).
Assume conditions (P), (K), (A1) and (A2). Let the step size in Algorithm 1 be γ. Recall that is the point at iteration time t and is the initial point. Assume that the step size , where . For any initial point within the ball , there exists a constant such that:Note that is the constant in assumption (A2) and satisfies ; see the proof of this theorem.
Theorem 3 shows that when the initial point is sufficiently close to a local minimum, the algorithm converges linearly [
42,
43] to the local minimum. Additionally, this implies that the ball
is always in the basin of attraction of
m. However, note that the actual basin could be much larger than
.
8. Conclusions
In this paper, we introduced a novel clustering approach based on the gradient of the slope function. The resulting clusters are associated with an attribute label, which provides additional information on each cluster. With this new clustering method, we propose a two-sample test using local information within each cluster, which improves the testing power. Finally, we developed an informative visualization tool that gives the structure of multi-dimensional data.
We studied our improved method’s performance empirically and theoretically. Simulation studies show that our refined clustering method is capable of capturing fine structures within the data. Furthermore, as a two-sample test procedure, our clustering method has better power than conventional approaches. The analysis on Astronomy and GvHD data shows that our method finds meaningful clusters. Finally, we studied both statistical and computational theory of our proposed method. Our proposed method demonstrated good empirical performance and statistical and numerical properties. Finally, we would like to note that while our method works well for the GvHD data (d = 4), it may not be applicable for any higher dimensional data, since our method is a nonparametric procedure involving derivative estimation. The curse of dimensionality prevents us from applying it to data with more dimensions.