A Morphological Post-Processing Approach for Overlapped Segmentation of Bacterial Cell Images
Abstract
:1. Introduction
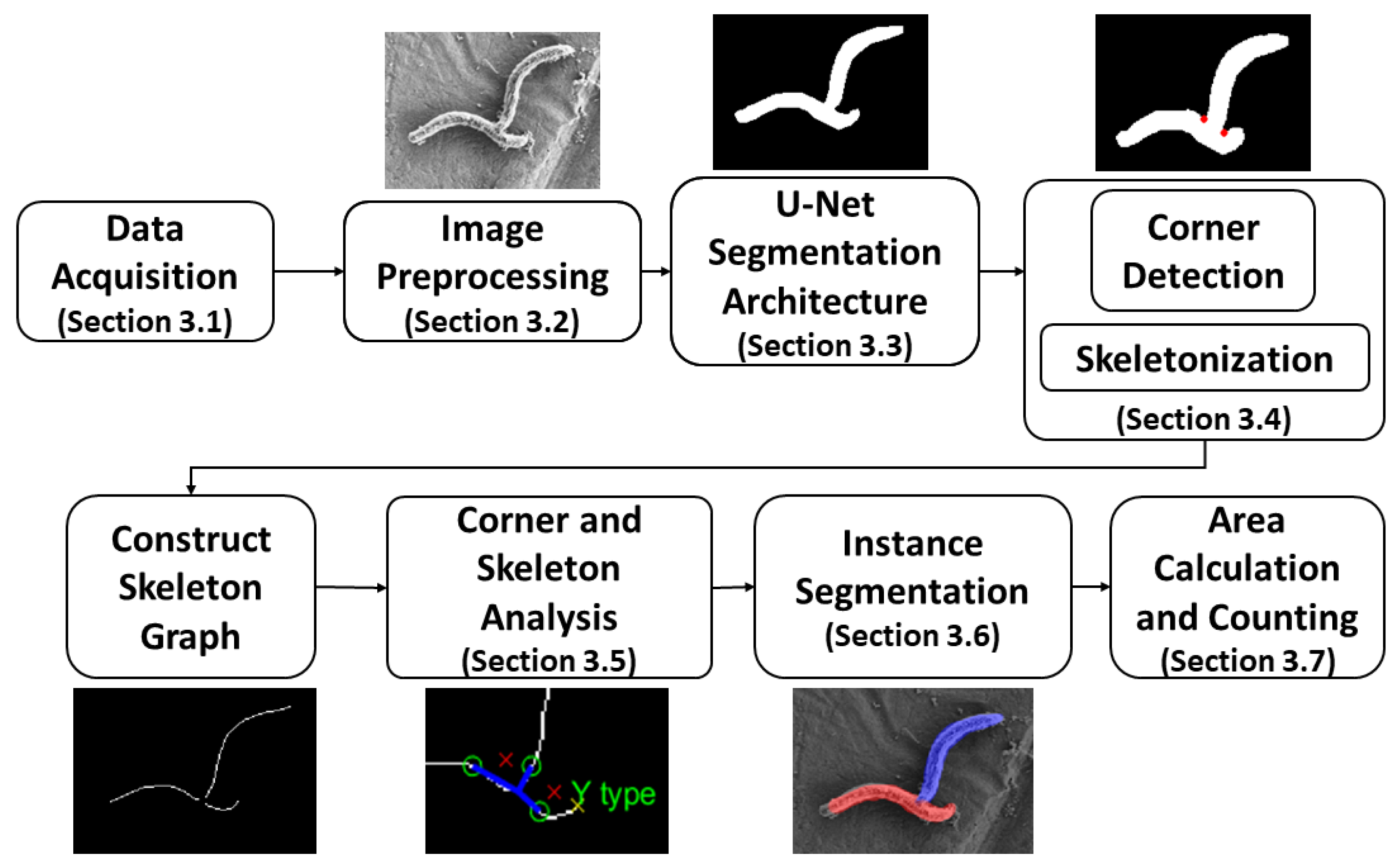
- A bacterial cell segmentation pipeline comprising deep semantic segmentation architecture and morphological post-processing technique is proposed to accommodate the above-mentioned cell extraction complications to retrieve accurate quantitative measures in SEM images.
- Benchmark the segmentation performance compared to the other mature overlapping object segmentation approaches, the proposed method shows a 89.52% Dice similarity score on bacterial segmentation and is validated with a comparison to several cell overlapping object segmentation approaches, in which significant a performance improvement was observed.
2. Related Work
2.1. Traditional Approaches for Overlapping Objects Segmentation
2.2. Contour-Based Methods
2.3. Ellipse-Fitting Methods
2.4. Deep Learning Methods
- Instance segmentation: mask R-CNN [52] is a well-known deep neural network architecture for multi-objects detection, and extends faster R-CNN [14] by adding an extra branch for predicting segmentation masks while simultaneously recognizing the bounding box from the existing branch. Mask R-CNN uses a region proposal-based object detection and uses a high-quality segmentation mask technique to achieve instance segmentation results. However, this method cannot perform well in situations with heavily overlapping object instances or highly close object occurrences due to greedy non-maximum suppression post-processing.
- Semantic segmentation: U-Net [53] deep learning architecture is recognized as another popular semantic segmentation approach that neither employs region proposals nor reuses pooling indices. Instead, it uses encoder–decoder-based neural network architecture to predict a class-based object segmentation output. The U-Net architecture has been successfully used in many overlapping cell segmentation tasks [51,54], especially in the medical community, because of its intrinsic capability to perform down sampling–up sampling. For example, research studies by [55,56] have demonstrated how to use the architecture to accurately segment overlapping cervical cells.
2.5. Our Contribution
3. Methodology
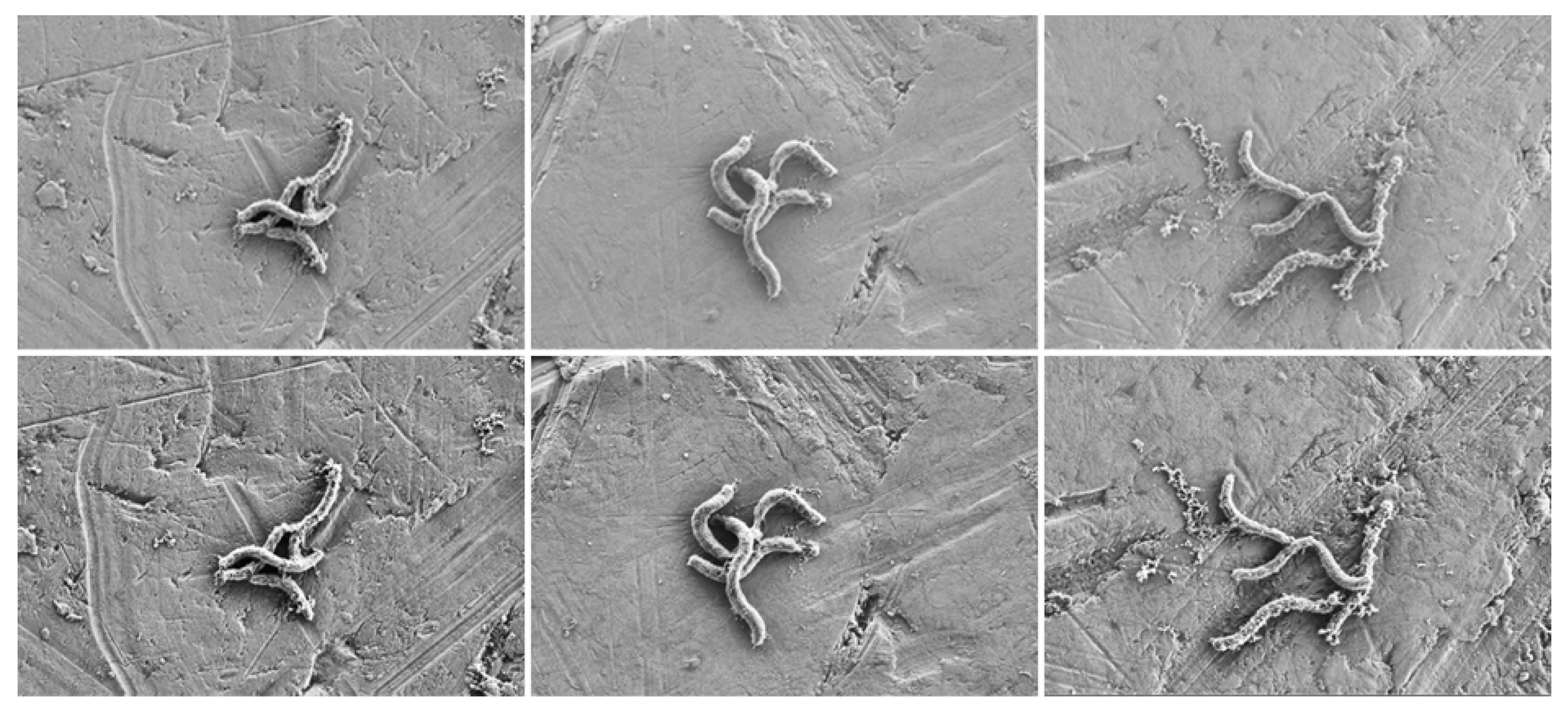
3.1. Data Acquisition
3.2. Images Pre-Processing
3.3. U-Net Segmentation Architecture
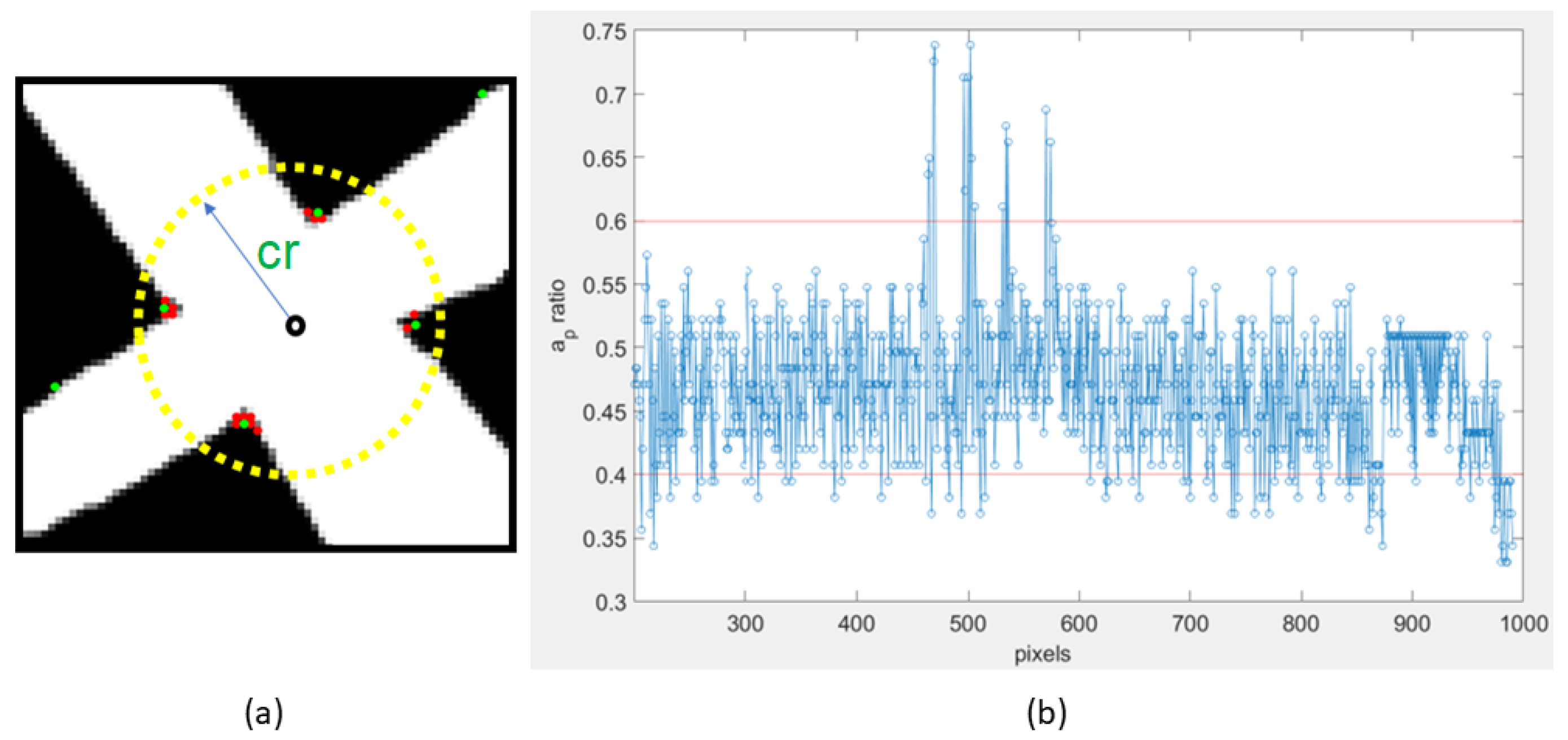
3.4. Corner Detection and Skeletonization
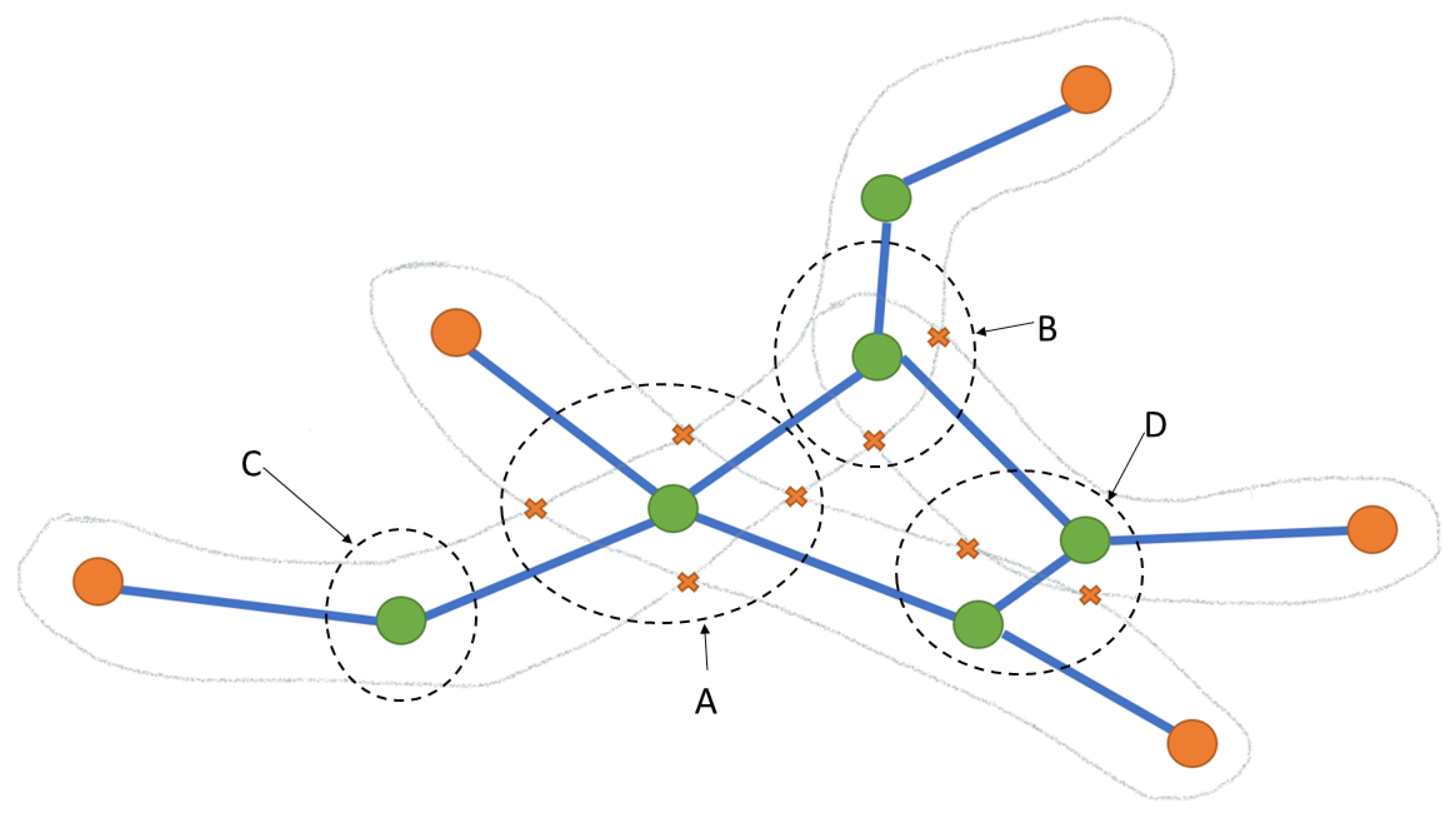
3.5. Corner and Skeleton Analysis
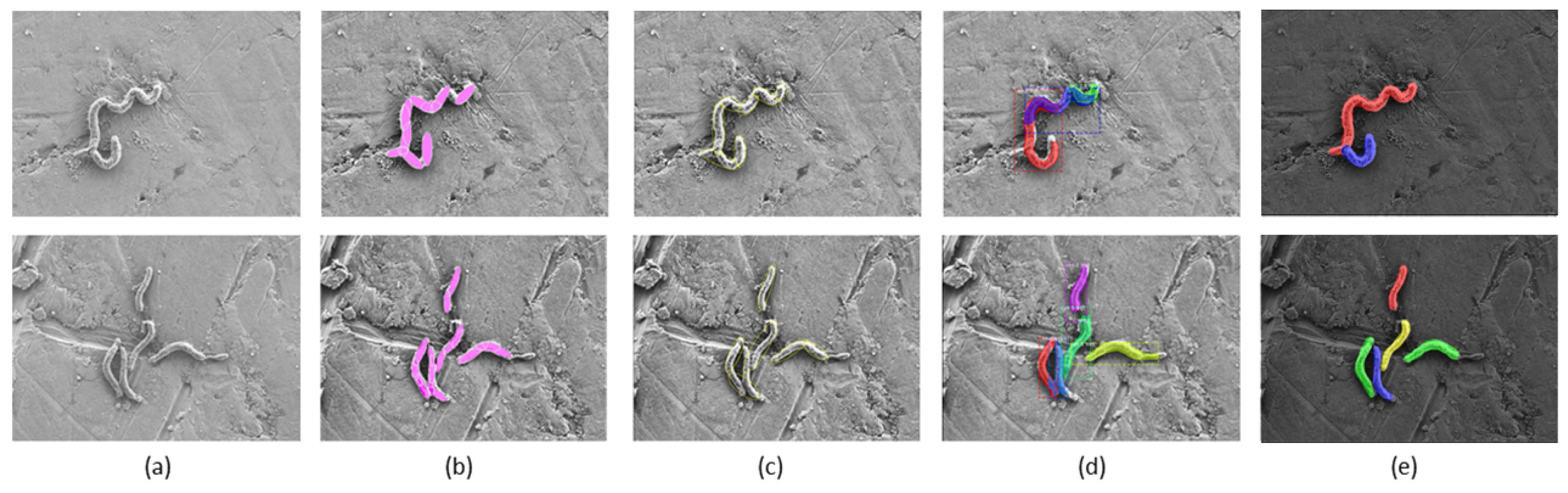
3.6. Instance Segmentation
| Algorithm 1: Bacterial Cell Segmentor |
 |
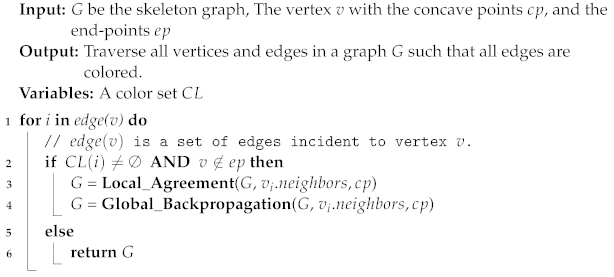
| Algorithm 2: Local Agreement | |||
| Input: G be the skeleton graph, the vertex v with the concave points , and the | |||
| skeleton | |||
| Output: Assign distinguished colors for the edges based on different types | |||
| Variables: radius , and a color set | |||
| 1 | Let , , be the three edges incident to vertex v | ||
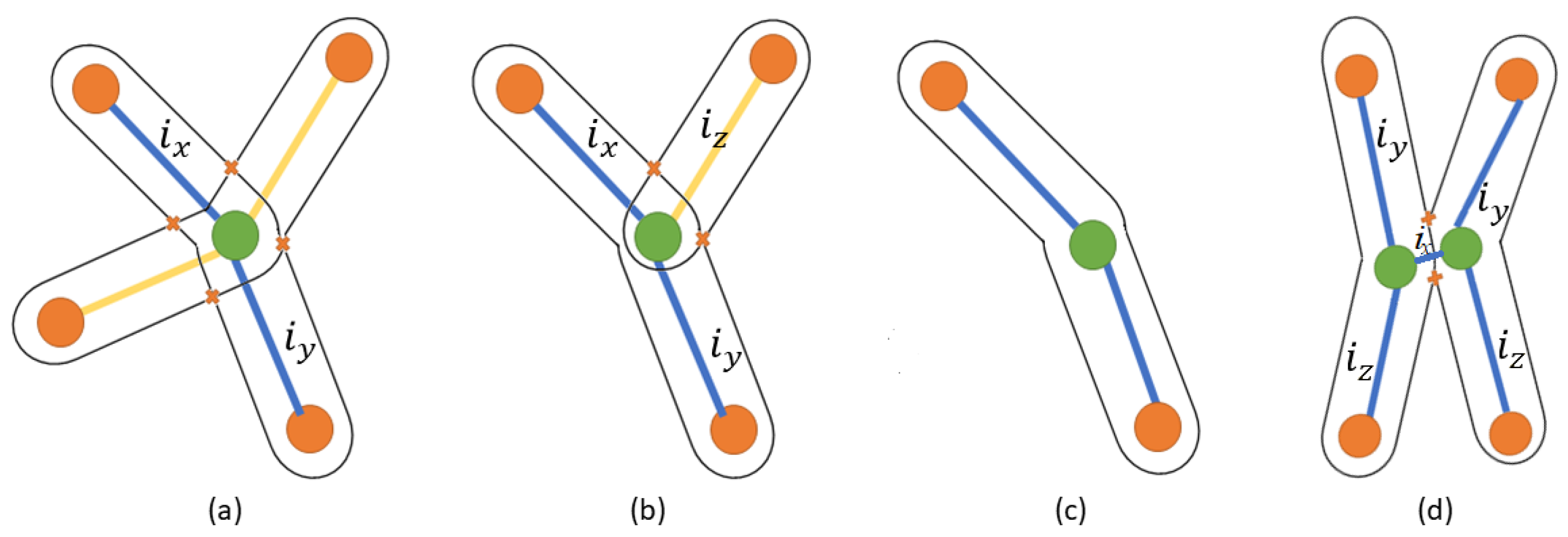
| 2 | ifvinX-typethen | ||
| 3 | Assign a distinguished color such that = , where , | ||
| 4 | AND, are opposite adjacent edges // as shown in Figure 6a | ||
| 5 | else ifvinY-typethen | ||
| 6 | Assign a distinguished color such that = , where , | ||
| 7 | AND, do not share two corner points | ||
| 8 | Assign a distinguished color such that , where | ||
| 9 | AND, share two corner points // as shown in Figure 6b | ||
| 10 | else ifvinH-typethen | ||
| 11 | Remove from | ||
| 12 | AND Remove from E, where // as shown in Figure 6d | ||
| 13 | else | ||
| 14 | Assign the same color ∀ // I-type, as shown in Figure 6c | ||
| 15 | returnG | ||
| Algorithm 3: Global Backpropagation |
 |
3.7. Area Calculation and Counting
4. Experimental Design
4.1. Experimental Setup
4.2. Evaluation Metrics
4.3. Train U-Net Models
4.4. Post-Processing Step
5. Evaluation Results & Discussion
6. Conclusions
Author Contributions
Funding
Data Availability Statement
Acknowledgments
Conflicts of Interest
References
- Sampaio, N.M.V.; Dunlop, M.J. Functional roles of microbial cell-to-cell heterogeneity and emerging technologies for analysis and control. Curr. Opin. Microbiol. 2020, 57, 87–94. [Google Scholar] [CrossRef] [PubMed]
- Keegstra, J.M.; Kamino, K.; Anquez, F.; Lazova, M.D.; Emonet, T.; Shimizu, T.S. Phenotypic diversity and temporal variability in a bacterial signaling network revealed by single-cell FRET. eLife 2017, 6, e27455. [Google Scholar] [CrossRef] [PubMed]
- Westfall, S.; Lomis, N.; Kahouli, I.; Dia, S.Y.; Singh, S.P.; Prakash, S. Microbiome, probiotics and neurodegenerative diseases: Deciphering the gut brain axis. Cell. Mol. Life Sci. 2017, 74, 3769–3787. [Google Scholar] [CrossRef] [PubMed]
- Golding, C.G.; Lamboo, L.L.; Beniac, D.R.; Booth, T.F. The scanning electron microscope in microbiology and diagnosis of infectious disease. Sci. Rep. 2016, 6, 26516. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Brahim Belhaouari, D.; Fontanini, A.; Baudoin, J.P.; Haddad, G.; Le Bideau, M.; Bou Khalil, J.Y.; Raoult, D.; La Scola, B. The strengths of scanning electron microscopy in deciphering SARS-CoV-2 infectious cycle. Front. Microbiol. 2020, 11, 2014. [Google Scholar] [CrossRef]
- Guo, Y.; Liu, Y.; Georgiou, T.; Lew, M.S. A review of semantic segmentation using deep neural networks. Int. J. Multimed. Inf. Retr. 2018, 7, 87–93. [Google Scholar] [CrossRef] [Green Version]
- Zhou, B.; Zhao, H.; Puig, X.; Xiao, T.; Fidler, S.; Barriuso, A.; Torralba, A. Semantic understanding of scenes through the ade20k dataset. Int. J. Comput. Vis. 2019, 127, 302–321. [Google Scholar] [CrossRef] [Green Version]
- Arteta, C.; Lempitsky, V.; Noble, J.A.; Zisserman, A. Learning to detect partially overlapping instances. In Proceedings of the IEEE Conference on Computer Vision And Pattern Recognition, Portland, OR, USA, 23–28 June 2013; pp. 3230–3237. [Google Scholar]
- Yan, L.; Park, C.W.; Lee, S.R.; Lee, C.Y. New separation algorithm for touching grain kernels based on contour segments and ellipse fitting. J. Zhejiang Univ. Sci. 2011, 12, 54–61. [Google Scholar] [CrossRef]
- Zhang, W.; Li, H. Automated segmentation of overlapped nuclei using concave point detection and segment grouping. Pattern Recognit. 2017, 71, 349–360. [Google Scholar] [CrossRef]
- Mosaliganti, K.R.; Noche, R.R.; Xiong, F.; Swinburne, I.A.; Megason, S.G. ACME: Automated cell morphology extractor for comprehensive reconstruction of cell membranes. PLoS Comput. Biol. 2012, 8, e1002780. [Google Scholar] [CrossRef]
- Vyas, N.; Sammons, R.; Addison, O.; Dehghani, H.; Walmsley, A. A quantitative method to measure biofilm removal efficiency from complex biomaterial surfaces using SEM and image analysis. Sci. Rep. 2016, 6, 32694. [Google Scholar] [CrossRef] [Green Version]
- Zou, T.; Pan, T.; Taylor, M.; Stern, H. Recognition of overlapping elliptical objects in a binary image. Pattern Anal. Appl. 2021, 24, 1193–1206. [Google Scholar] [CrossRef]
- Ren, S.; He, K.; Girshick, R.; Sun, J. Faster R-CNN: Towards Real-Time Object Detection with Region Proposal Networks. arXiv 2015, arXiv:cs.CV/1506.01497. [Google Scholar] [CrossRef] [Green Version]
- Litjens, G.; Kooi, T.; Bejnordi, B.E.; Setio, A.A.A.; Ciompi, F.; Ghafoorian, M.; van der Laak, J.A.; van Ginneken, B.; Sánchez, C.I. A survey on deep learning in medical image analysis. Med. Image Anal. 2017, 42, 60–88. [Google Scholar] [CrossRef] [Green Version]
- Nistico, L.; Hall-Stoodley, L.; Stoodley, P. Imaging bacteria and biofilms on hardware and periprosthetic tissue in orthopedic infections. In Microbial Biofilms; Springer: Berlin/Heidelberg, Germany, 2014; pp. 105–126. [Google Scholar]
- Li, J.; Hirota, K.; Goto, T.; Yumoto, H.; Miyake, Y.; Ichikawa, T. Biofilm formation of Candida albicans on implant overdenture materials and its removal. J. Dent. 2012, 40, 686–692. [Google Scholar] [CrossRef]
- Hägi, T.T.; Klemensberger, S.; Bereiter, R.; Nietzsche, S.; Cosgarea, R.; Flury, S.; Lussi, A.; Sculean, A.; Eick, S. A biofilm pocket model to evaluate different non-surgical periodontal treatment modalities in terms of biofilm removal and reformation, surface alterations and attachment of periodontal ligament fibroblasts. PLoS ONE 2015, 10, e0131056. [Google Scholar] [CrossRef] [Green Version]
- Vincent, L. Morphological grayscale reconstruction in image analysis: Applications and efficient algorithms. IEEE Trans. Image Process. 1993, 2, 176–201. [Google Scholar] [CrossRef] [Green Version]
- Cooper, L.A.; Kong, J.; Gutman, D.A.; Wang, F.; Gao, J.; Appin, C.; Cholleti, S.; Pan, T.; Sharma, A.; Scarpace, L.; et al. Integrated morphologic analysis for the identification and characterization of disease subtypes. J. Am. Med. Inform. Assoc. 2012, 19, 317–323. [Google Scholar] [CrossRef] [Green Version]
- Vincent, L.; Soille, P. Watersheds in digital spaces: An efficient algorithm based on immersion simulations. IEEE Comput. Archit. Lett. 1991, 13, 583–598. [Google Scholar] [CrossRef] [Green Version]
- Nath, S.K.; Palaniappan, K.; Bunyak, F. Cell Segmentation Using Coupled Level Sets and Graph-Vertex Coloring. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention 2006, Copenhagen, Denmark, 1–6 October 2006; Larsen, R., Nielsen, M., Sporring, J., Eds.; Springer: Berlin/Heidelberg, Germany, 2006; pp. 101–108. [Google Scholar]
- Dzyubachyk, O.; Niessen, W.; Meijering, E. Advanced level-set based multiple-cell segmentation and tracking in time-lapse fluorescence microscopy images. In Proceedings of the 2008 5th IEEE International Symposium on Biomedical Imaging: From Nano to Macro, Paris, France, 14–17 May 2008; pp. 185–188. [Google Scholar] [CrossRef]
- Chang, H.; Han, J.; Borowsky, A.; Loss, L.; Gray, J.W.; Spellman, P.T.; Parvin, B. Invariant delineation of nuclear architecture in glioblastoma multiforme for clinical and molecular association. IEEE Trans. Med. Imaging 2012, 32, 670–682. [Google Scholar] [CrossRef]
- Al-Kofahi, Y.; Lassoued, W.; Lee, W.; Roysam, B. Improved automatic detection and segmentation of cell nuclei in histopathology images. IEEE Trans. Biomed. Eng. 2009, 57, 841–852. [Google Scholar] [CrossRef] [PubMed]
- Kumar, N.; Verma, R.; Sharma, S.; Bhargava, S.; Vahadane, A.; Sethi, A. A dataset and a technique for generalized nuclear segmentation for computational pathology. IEEE Trans. Med. Imaging 2017, 36, 1550–1560. [Google Scholar] [CrossRef] [PubMed]
- Jeulin, D. Morphological Models of Random Structures; Springer: Berlin/Heidelberg, Germany, 2021. [Google Scholar]
- Yin, Z.; Bise, R.; Chen, M.; Kanade, T. Cell segmentation in microscopy imagery using a bag of local Bayesian classifiers. In Proceedings of the 2010 IEEE International Symposium on Biomedical Imaging: From Nano to Macro, Rotterdam, The Netherlands, 14–17 April 2010; pp. 125–128. [Google Scholar] [CrossRef] [Green Version]
- Su, H.; Yin, Z.; Huh, S.; Kanade, T. Cell segmentation in phase contrast microscopy images via semi-supervised classification over optics-related features. Med. Image Anal. 2013, 17, 746–765. [Google Scholar] [CrossRef] [PubMed]
- Wang, W.; Taft, D.A.; Chen, Y.J.; Zhang, J.; Wallace, C.T.; Xu, M.; Watkins, S.C.; Xing, J. Learn to segment single cells with deep distance estimator and deep cell detector. Comput. Biol. Med. 2019, 108, 133–141. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Al-Rifaie, M.M.; Aber, A.; Hemanth, D.J. Deploying swarm intelligence in medical imaging identifying metastasis, micro-calcifications and brain image segmentation. IET Syst. Biol. 2015, 9, 234–244. [Google Scholar] [CrossRef]
- Brezočnik, L.; Fister, I., Jr.; Podgorelec, V. Swarm intelligence algorithms for feature selection: A review. Appl. Sci. 2018, 8, 1521. [Google Scholar] [CrossRef] [Green Version]
- Li, C.; Xu, C.; Gui, C.; Fox, M.D. Distance regularized level set evolution and its application to image segmentation. IEEE Trans. Image Process. 2010, 19, 3243–3254. [Google Scholar] [CrossRef]
- Zhang, K.; Song, H.; Zhang, L. Active contours driven by local image fitting energy. Pattern Recognit. 2010, 43, 1199–1206. [Google Scholar] [CrossRef]
- Niu, S.; Chen, Q.; De Sisternes, L.; Ji, Z.; Zhou, Z.; Rubin, D.L. Robust noise region-based active contour model via local similarity factor for image segmentation. Pattern Recognit. 2017, 61, 104–119. [Google Scholar] [CrossRef] [Green Version]
- Zhang, J.; Lu, Z.; Li, M. Active contour-based method for finger-vein image segmentation. IEEE Trans. Instrum. Meas. 2020, 69, 8656–8665. [Google Scholar] [CrossRef]
- Fernandez, G.; Kunt, M.; Zryd, J.P. A new plant cell image segmentation algorithm. In Proceedings of the International Conference on Image Analysis and Processing, Washington, DC, USA, 23–26 October 1995; Springer: Berlin/Heidelberg, Germany, 1995; pp. 229–234. [Google Scholar]
- He, Y.; Meng, Y.; Gong, H.; Chen, S.; Zhang, B.; Ding, W.; Luo, Q.; Li, A. An automated three-dimensional detection and segmentation method for touching cells by integrating concave points clustering and random walker algorithm. PLoS ONE 2014, 9, e104437. [Google Scholar] [CrossRef] [Green Version]
- Wang, H.; Zhang, H.; Ray, N. Clump splitting via bottleneck detection and shape classification. Pattern Recognit. 2012, 45, 2780–2787. [Google Scholar] [CrossRef]
- Xing, F.; Yang, L. Chapter 4—Machine learning and its application in microscopic image analysis. In Machine Learning and Medical Imaging; Wu, G., Shen, D., Sabuncu, M.R., Eds.; The Elsevier and MICCAI Society Book Series; Academic Press: Cambridge, MA, USA, 2016; pp. 97–127. [Google Scholar] [CrossRef]
- Pavlidis, T. Algorithms for shape analysis of contours and waveforms. IEEE Trans. Pattern Anal. Mach. Intell. 1980, 4, 301–312. [Google Scholar] [CrossRef]
- Harris, C.G.; Stephens, M. A combined corner and edge detector. In Proceedings of the Alvey Vision Conference, Manchester, UK, 9–10 September 1988; Citeseer: University Park, PA, USA, 1988; Volume 15, pp. 10–5244. [Google Scholar]
- Miró-Nicolau, M.; Moyà-Alcover, B.; González-Hidalgo, M.; Jaume-i Capó, A. Segmenting overlapped objects in images. A study to support the diagnosis of sickle cell disease. arXiv 2020, arXiv:2008.00997. [Google Scholar]
- Zafari, S.; Eerola, T.; Sampo, J.; Kälviäinen, H.; Haario, H. Segmentation of overlapping elliptical objects in silhouette images. IEEE Trans. Image Process. 2015, 24, 5942–5952. [Google Scholar] [CrossRef]
- Zhang, G.; Jayas, D.S.; White, N.D. Separation of Touching Grain Kernels in an Image by Ellipse Fitting Algorithm. Biosyst. Eng. 2005, 92, 135–142. [Google Scholar] [CrossRef]
- Panagiotakis, C.; Argyros, A.A. Cell Segmentation Via Region-Based Ellipse Fitting. In Proceedings of the 2018 25th IEEE International Conference on Image Processing (ICIP), Athens, Greece, 7–10 October 2018; pp. 2426–2430. [Google Scholar] [CrossRef]
- Panagiotakis, C.; Argyros, A. Parameter-Free Modelling of 2D Shapes with Ellipses. Pattern Recogn. 2016, 53, 259–275. [Google Scholar] [CrossRef]
- Panagiotakis, C.; Argyros, A. Region-based Fitting of Overlapping Ellipses and its application to cells segmentation. Image Vis. Comput. 2020, 93, 103810. [Google Scholar] [CrossRef]
- Abeyrathna, D.; Life, T.; Rauniyar, S.; Ragi, S.; Sani, R.; Chundi, P. Segmentation of Bacterial Cells in Biofilms Using an Overlapped Ellipse Fitting Technique. In Proceedings of the 2021 IEEE International Conference on Bioinformatics and Biomedicine (BIBM), Houston, TX, USA, 9–12 December 2021; pp. 3548–3554. [Google Scholar] [CrossRef]
- Yang, L.; Zhang, Y.; Guldner, I.H.; Zhang, S.; Chen, D.Z. 3D segmentation of glial cells using fully convolutional networks and k-terminal cut. In Proceedings of the International Conference on Medical Image Computing and Computer-Assisted Intervention, Athens, Greece, 17–21 October 2016; Springer: Berlin/Heidelberg, Germany, 2016; pp. 658–666. [Google Scholar]
- Saleh, H.M.; Saad, N.H.; Isa, N.A.M. Overlapping chromosome segmentation using U-net: Convolutional networks with test time augmentation. Procedia Comput. Sci. 2019, 159, 524–533. [Google Scholar] [CrossRef]
- He, K.; Gkioxari, G.; Dollár, P.; Girshick, R.B. Mask R-CNN. In Proceedings of the 2017 IEEE International Conference on Computer Vision (ICCV), Venice, Italy, 22–29 October 2017; pp. 2980–2988. [Google Scholar]
- Ronneberger, O.; Fischer, P.; Brox, T. U-Net: Convolutional Networks for Biomedical Image Segmentation. In Proceedings of the Medical Image Computing and Computer-Assisted Intervention—MICCAI 2015, Munich, Germany, 5–9 October 2015; pp. 234–241. [Google Scholar] [CrossRef] [Green Version]
- Hu, R.L.; Karnowski, J.; Fadely, R.; Pommier, J.P. Image segmentation to distinguish between overlapping human chromosomes. arXiv 2017, arXiv:1712.07639. [Google Scholar]
- Kurnianingsih.; Allehaibi, K.H.S.; Nugroho, L.E.; Widyawan.; Lazuardi, L.; Prabuwono, A.S.; Mantoro, T. Segmentation and Classification of Cervical Cells Using Deep Learning. IEEE Access 2019, 7, 116925–116941. [Google Scholar] [CrossRef]
- Lu, Z.; Carneiro, G.; Bradley, A.P. An Improved Joint Optimization of Multiple Level Set Functions for the Segmentation of Overlapping Cervical Cells. IEEE Trans. Image Process. 2015, 24, 1261–1272. [Google Scholar] [CrossRef]
- Carlson, C.; Singh, N.K.; Bibra, M.; Sani, R.K.; Venkateswaran, K. Pervasiveness of UVC254-resistant Geobacillus strains in extreme environments. Appl. Microbiol. Biotechnol. 2018, 102, 1869–1887. [Google Scholar] [CrossRef] [PubMed]
- Lam, C.; Yi, D.; Guo, M. Automated Detection of Diabetic Retinopathy using Deep Learning. Amia Jt. Summits Transl. Sci. Proc. 2018, 2018, 147–155. [Google Scholar]
- Dutta, A.; Zisserman, A. The VIA Annotation Software for Images, Audio and Video. In Proceedings of the 27th ACM International Conference on Multimedia, MM’19, Nice, France, 21–25 October 2019; ACM: New York, NY, USA, 2019. [Google Scholar] [CrossRef] [Green Version]
- Krizhevsky, A.; Sutskever, I.; Hinton, G.E. Imagenet classification with deep convolutional neural networks. Adv. Neural Inf. Process. Syst. 2012, 25, 84–90. [Google Scholar] [CrossRef] [Green Version]
- Tan, S.; Ma, X.; Mai, Z.; Qi, L.; Wang, Y. Segmentation and counting algorithm for touching hybrid rice grains. Comput. Electron. Agric. 2019, 162, 493–504. [Google Scholar] [CrossRef]
- Maragos, P.; Schafer, R. Morphological skeleton representation and coding of binary images. IEEE Trans. Acoust. Speech Signal Process. 1986, 34, 1228–1244. [Google Scholar] [CrossRef]
- Yao, Q.; Zhou, Y.; Wang, J. An automatic segmentation algorithm for touching rice grains images. In Proceedings of the 2010 International Conference on Audio, Language and Image Processing, Shanghai, China, 23–25 November 2010; pp. 802–805. [Google Scholar] [CrossRef]
- Rana, D.S. Segmentation of Overlapping Wheat Grains for Quality Detection. 2018. Available online: https://www.semanticscholar.org/paper/Segmentation-of-Overlapping-Wheat-Grains-for-Rana/5fffb195cb6d3bf329310f40e0e9b71be7db6377 (accessed on 1 August 2022).
- Zhou, C.; Lin, K.; Xu, D.; Liu, J.; Zhang, S.; Sun, C.; Yang, X. Method for segmentation of overlapping fish images in aquaculture. Int. J. Agric. Biol. Eng. 2019, 12, 135–142. [Google Scholar] [CrossRef]
- Dice-Score, L.R. Measures of the Amount of Ecologic Association Between Species. Ecology 1945, 26, 297–302. [Google Scholar] [CrossRef]
- Kingma, D.P.; Ba, J. Adam: A method for stochastic optimization. arXiv 2014, arXiv:1412.6980. [Google Scholar]
- Abdulla, W. Mask R-CNN for Object Detection and Instance Segmentation on Keras and TensorFlow. 2017. Available online: https://github.com/matterport/Mask_RCNN (accessed on 1 August 2022).
- Foresti, R.; Rossi, S.; Pinelli, S.; Alinovi, R.; Barozzi, M.; Sciancalepore, C.; Galetti, M.; Caffarra, C.; Lagonegro, P.; Scavia, G.; et al. Highly-defined bioprinting of long-term vascularized scaffolds with Bio-Trap: Complex geometry functionalization and process parameters with computer aided tissue engineering. Materialia 2020, 9, 100560. [Google Scholar] [CrossRef]


| Method | SR | SERR (%) | SEFR (%) | Dice-Score (%) | Avg Time (s) |
|---|---|---|---|---|---|
| U-Net + [13] | 1.60 | 74.69 | 52.63 | 79.10 ± 0.04 | 15.2 |
| U-Net + [49] | 0.79 | 23.11 | 77.02 | 80.60 ± 5.08 | 32.6 |
| Mask R-CNN | 0.82 | 27.08 | 68.25 | 83.44 ± 3.62 | 20.1 |
| In the Present Study | 1.13 | 14.71 | 88.28 | 89.52 ± 1.31 | 22.9 |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Abeyrathna, D.; Rauniyar, S.; Sani, R.K.; Huang, P.-C. A Morphological Post-Processing Approach for Overlapped Segmentation of Bacterial Cell Images. Mach. Learn. Knowl. Extr. 2022, 4, 1024-1041. https://doi.org/10.3390/make4040052
Abeyrathna D, Rauniyar S, Sani RK, Huang P-C. A Morphological Post-Processing Approach for Overlapped Segmentation of Bacterial Cell Images. Machine Learning and Knowledge Extraction. 2022; 4(4):1024-1041. https://doi.org/10.3390/make4040052
Chicago/Turabian StyleAbeyrathna, Dilanga, Shailabh Rauniyar, Rajesh K. Sani, and Pei-Chi Huang. 2022. "A Morphological Post-Processing Approach for Overlapped Segmentation of Bacterial Cell Images" Machine Learning and Knowledge Extraction 4, no. 4: 1024-1041. https://doi.org/10.3390/make4040052






