Abstract
The outbreak of COVID-19 has caused more than 200,000 deaths so far in the USA alone, which instigates the necessity of initial screening to control the spread of the onset of COVID-19. However, screening for the disease becomes laborious with the available testing kits as the number of patients increases rapidly. Therefore, to reduce the dependency on the limited test kits, many studies suggested a computed tomography (CT) scan or chest radiograph (X-ray) based screening system as an alternative approach. Thereby, to reinforce these approaches, models using both CT scan and chest X-ray images need to develop to conduct a large number of tests simultaneously to detect patients with COVID-19 symptoms. In this work, patients with COVID-19 symptoms have been detected using eight distinct deep learning techniques, which are VGG16, InceptionResNetV2, ResNet50, DenseNet201, VGG19, MobilenetV2, NasNetMobile, and ResNet15V2, using two datasets: one dataset includes 400 CT scan and another 400 chest X-ray images. Results show that NasNetMobile outperformed all other models by achieving an accuracy of 82.94% in CT scan and 93.94% in chest X-ray datasets. Besides, Local Interpretable Model-agnostic Explanations (LIME) is used. Results demonstrate that the proposed models can identify the infectious regions and top features; ultimately, it provides a potential opportunity to distinguish between COVID-19 patients with others.
Keywords:
chest X-ray; COVID-19; CT scan; deep learning; explainable AI; image processing; radiography; SARS-CoV-2; small data 1. Introduction
The novel coronavirus, also known as COVID-19, created a global health crisis early in 2020. The disease originates from the virus known as a severe acute respiratory syndrome or coronavirus 2, also called SARS-CoV-2 [1], a socially transmitted disease and can infect individuals because of close contact to the infected patients. The number of infected individuals from COVID-19 cases surpasses 30 million, and death raises close to one million as of 09 September 2020 [2]. While many COVID-19 cases exhibit mild symptoms, a small percentage suffers from severe or critical conditions [3]. In increasingly genuine cases, the contamination can cause pneumonia, extreme intense respiratory discomfort, multi-organ failure, and death [4]. The health systems have been overwhelmed among developed countries such as the USA, UK, and Italy due to the expanding demand for intensive care units, as those units filled with COVID-19 patients with severe medical conditions [5].
Until no fruitful vaccine is developed, expanding the screening and isolating the COVID-19 patients from the mass population is the only solution to reduce the transmissions on a large scale. Currently, all over the world, reverse transcription-polymerase chain reaction (RT-PCR) has been used as a standard gold test to detect COVID-19 patients. However, the test results often produce false alarms, and the current success rate is merely 70% [4]. Additionally, test results take time to acquire, leaving behind a higher risk and possibility of spreading the disease among other peoples by the patients.
Therefore, to limit the dependency on limited test kits and control the exponential growth of COVID-19 patients, many studies suggested chest radiograph (X-ray) based screening procedures at the early stages of this pandemic and demonstrated satisfactory results by achieving higher accuracy than the RT-PCR test. However, since the disease outbreak in 2020, most of the studies had to deal with limited data and reported their result with those available resources. For example, Ghoshal et al. (2020) [6] experimented on a dataset comprises of 70 COVID-19 images from one source [7] and non-COVID-19 images from another sources [8]. Their proposed Bayesian CNN model improves the detection rate from 85.7% to 92.9% along with the VGG16 model [9]. Similarly, Narin et al. (2020) [10] used only 100 images to conduct that experiment, and the dataset consist, 50 chest X-rays of COVID-19 and 50 normal chest X-ray of non-COVID-19 patients. Additionally, Zhang et al. (2020) presented the ResNet model, using 70 COVID-19 and 1008 non-COVID-19 pneumonia patients from different data sources. The evaluation result showed 96% sensitivity, 70.7% specificity and 0.952 of AUC [11]. Wang et al. (2020) introduced a deep CNN based model known as COVID-Net, which attained 83.5% accuracy to detect COVID-19 patients from a dataset of 5941 images which includes 1203 healthy, 931 bacterial pneumonia, 660 viral pneumonia, and 45 individuals with COVID-19 cases [12].
Apart from chest X-ray, some literature suggested chest computed tomography (CT) based screening to distinguish between COVID-19 and non-COVID-19 patients [11,13,14,15,16,17]. For instance, Chen et al. (2020) used UNet++ to classify COVID-19 and non-COVID-19 patients considering 132 sample images using 51 COVID-19, 55 non-COVID-19, 16 viral pneumonia, and 11 non-pneumonia patients and revealed that artificial intelligence reduces the reading time of radiologists up to 65% [13]. Zhang et al. (2020) [11] used the UNet model for lung segmentation, considering 540—included 313 COVID-19 and 229 non-COVID-19 patients’ CT scan—images, and reported the result with 90.7% sensitivity and 91.1% specificity score. Besides, several studies employed the ResNet model to detect COVID-19 patients from the chest CT [14,15,16]. Jin et al. (2020) [14] proposed the ResNet15V2 model for detecting COVID-19 patients with the dataset of 1881 images (496 COVID-19 and 1385 non-COVID-19) and study result shows 94.1% of sensitivity, and 95.5% of specificity [14]. Song et al. (2020) and Li et al. (2020) implemented ResNet50 to detect COVID-19 patients from chest CT scan and achieved 86% and 96% accuracy, respectively [15,17].
Since most of the early studies used limited data, therefore questions raised regarding their models’ stability. Thereby, a better approach to present such limited data result is to provide the result with confidence intervals, which are missing almost in every study. The existing proposed models either demonstrated their potential on chest radiography or CT scan-based datasets, not in both scenarios. Therefore, a model developed with mixed data—chest radiography and CT scan—might provide that answer.
Recently, explanatory artificial intelligence (EAI) gained much popularity in medical image analysis as it helps to understand, visualize, and interpret any machine learning models used for disease prediction [18,19]. Ribeiro et al. (2016) proposed Local Interpretable Model-agnostic Explanations (LIME), a novel approach that explains any classifier’s performance in an interpretable manner [20]. LIME drew much attention by showing superiority in explaining how Google’s pre-trained network predicts by merely analyzing some random images [21]. Holzinger et al. (2017) consent that explainable-AI might be the next future in medical domains when health professionals rely on AI to understand the patients’ conditions [22]. Thus, an AI that explains the X-ray or CT scan images’ infectious regions might help the general practitioners— doctors, nurses, and health professionals, especially in rural areas— to detect between COVID-19 patients with others.
In general, this investigation found that a large portion of the study either considered chest X-ray or CT scan image analysis with a couple of deep learning models because of the time constraints. However, in accordance with the recent literature and extending the current work one step further, this research contributes as follows: (1) Proposed and tested eight individual convolutional neural network-based models—VGG16 [23], InceptionResNetV2 [24], ResNet50 [25], DenseNet201 [26], VGG19 [23], MobileNetV2 [27], NasNetMobile [28], and ResNet15V2 [29]—to detect COVID-19 patients using CT scan and chest X-ray images; (2) analyzed between existing models with the proposed ones in terms of accuracy, precision, recall, and f1-score; and (3) finally, applied LIME to explain features that help the model to identify COVID-19 patients from others.
The rest of the paper is organized as follows: Section 2 discusses the research methodology of this study. This is followed by the results of the proposed research for detecting COVID-19 from chest X-rays and CT scan in Section 3. Section 4 then provides detailed discussion and insight by analyzing the results in terms of models’ overall performances, comparing with the previous studies, and others. Finally, Section 5 concludes the article summarizing our findings, with an identification of opportunities for future work.
2. Methodology
This research uses two distinct datasets containing X-ray and CT scan images collected from the open-source Kaggle datasets repository [8]. At the time of the study, one dataset contains 400 chest X-ray images, and another includes 400 CT scan images. Table 1 summarizes the datasets used in this study. As commonly adopted in data mining techniques, this study used 80% data for training, whereas the remaining 20% was used for testing. The experiment was repeated two times and the model’s performance was evaluated by averaging those two outcomes.

Table 1.
Chest CT and X-ray datasets used in this study.
A transfer learning technique was implemented to develop the models [30] by acquiring model’s weight from pre-trained models (i.e., ImageNet [31]). The primary model’s architecture contains three components—pre-trained network, modified head, and prediction class (inspired from [11]). The pre-trained networks are employed to extract the high-level features and connected to the modified network and classification head, respectively. Figure 1 illustrates one of the modified proposed models’ architecture. The architecture contains 16 [32] CNN layers with different filter numbers, sizes, and stride values.
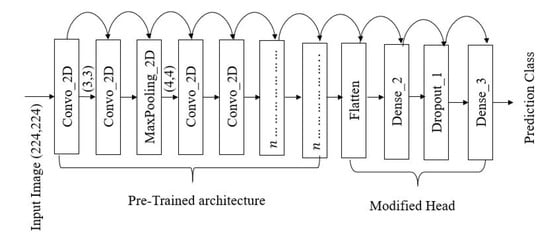
Figure 1.
VGG16 architecture implemented during this experiment [23].
The proposed models are constructed using three basic layers:
If is considered as the input layer, then the proposed models layout for VGG16 may be expressed as:
A robust model also relays on proper feature extraction techniques as well [33]. Let the letter,
Then the two-dimensional convolutional operation can be expressed as follows [34]:
where * represents the discrete convolution operation [34]. Kernel, K slides over the images with the stride parameters. The Rectified Linear unit (ReLu) is used as an activation function in the dense layer. ReLu function can be calculated with the following equations [34]:
During this experiment convolution filter with pool size is used for feature extraction [23]. An illustration of input images flows from the convolutional layer to the Maxpooling layer is given in Figure 2.
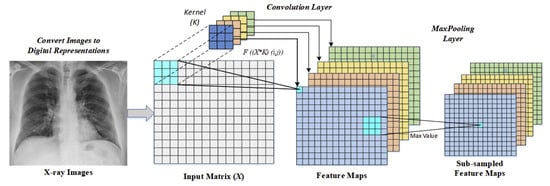
Figure 2.
An illustration of convolutional and maxpooling layer operations [34].
As a part of parameter tuning, initially, the batch size, the number of epochs and learning rate are considered [35]. Following parameters are randomly selected at the beginning of the experiment:
Finally, using grid search methods the optimal parameters are found as follows:
During the training phase, an optimization algorithm requires to set to optimize the model [36]. Some of the most popular optimization algorithms includes- adaptive learning rate optimization algorithm (Adam) [37], stochastic gradient descent (Sgd) [38], and root means square propagation (Rmsprop) [39]. To optimize the model, Adam is used due to its effectiveness in binary image classification [40,41].
Finally, the overall result was statistically analyzed based on accuracy, precision, recall, and f1-score [42]:
where True Positive ()= COVID-19 patient classified as patient, False Positive ()= Healthy people classified as patient, True Negative ()=Healthy people classified as healthy, and False Negative ()= COVID-19 patient classified as healthy. Figure 3 shows the overall flow diagram of the experiment. The best model was selected based on the statistical analysis on CT scan and chest X-ray image datasets.
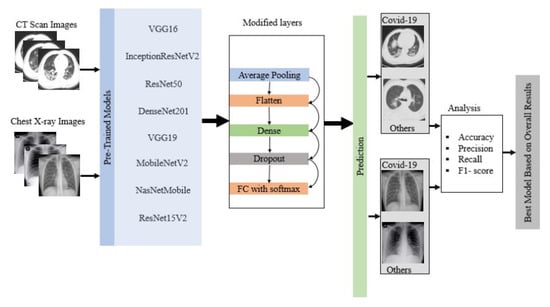
Figure 3.
Flow diagram of the overall experiment.
3. Results
During this experiment overall accuracy, precision, recall, and f1-score were measured for eight different deep learning approaches considering CT scan and X-ray image using Equations (3)–(6).
3.1. CT Scan
Table 2 summarizes the average accuracy, precision, recall, and f1-score for eight pre-trained deep learning models used in this study on the train set. Among all the models, MobileNetV2 performed best in terms of accuracy (99%), precision (99%), recall (99%), and f1-score (99%), and ResNet50 performed worst across all measures.

Table 2.
Overall model’s performance on CT scan train set.
In contrast, on the test set, NasNetMobile showed the best and ResNet50 showed the worst performance across all measures, as shown in Table 3.

Table 3.
Overall model’s performance on CT scan test set.
3.2. X-ray Image
Table 4 summarizes the overall model’s performance amid this experiment on train sets for X-ray images. From the result- most of the model performed well on the train set except ResNet50. 100% accuracy, precision, recall, and f1-score were achieved for VGG16, DenseNet201, MobileNetV2, NasNetMobile, and ResNet15V2.

Table 4.
Overall model’s performance on X-ray train set.
Table 5 summarizes the model’s performance on the test set. 100% accuracy, precision, recall and f1-score were measured for NasNetMobile. On the other hand, ResNet50 appeared to have lowest performance compared to any other models by achieving 64% accuracy, 79% precision, 64% recall, and 58% f-1 score.

Table 5.
Overall model’s performance on X-ray test set.
3.3. Confusion Matrix
The confusion matrix was calculated on the test set to simplify the understanding of the model’s performance. Figure 4 displays the confusion matrix for different models on given CT scan images. In CT scan, the test set contained 80 patients, where 40 were COVID-19 and 40 were non-COVID-19. In this case, NasNetMobile demonstrated the best result by correctly classifying 72 samples, while ResNet50 showed the worst performance by classifying 44 samples out of 80.

Figure 4.
Confusion matrix of different deep learning model for CT scan image dataset.
On chest X-ray test set, NasNetMobile outperformed all other models by correctly classifying all the samples, while ResNet50 displayed the worst performance by misclassifying 29 out of 80 samples, as shown in Figure 5.
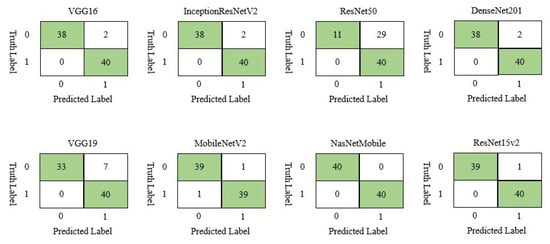
Figure 5.
Confusion matrix of different deep learning model for chest X-ray image dataset.
3.4. Confidence Interval
The confidence interval (CI) was calculated using two standard methods: the Wilson score [43] and Bayesian interval [44]. Both approaches are widely used and suitable to measure the performance on a small dataset [45]. Table 6 delineates 95% CI for model accuracy on the test set for CT scan and chest X-ray datasets. For instance, On the CT scan image dataset, ResNet50 has the lowest accuracy, ranges from 0.441 to 0.654 and 0.441 to 0.656; in contrast, NasNetmobile has the highest accuracy set out from 0.815 to 0.948 and 0.820 to 0.952 respectively.

Table 6.
Confidence Interval () of CT scan and chest X-ray in terms of accuracy on test set. Sample size, n = 80 for both studies.
Additionally, On the Chest X-ray test set, accuracy for VGG16, InceptionResNetV2, DenseNet201, and MobileNetV2 was achieved between 0.913 and 0.993, 0.922 and 0.995 using Wilson score and Bayesian interval, respectively. However, among all the models, higher accuracy was measured for NasNetMobile, and lower accuracy was acquired for ResNet50.
4. Discussion
MobileNetV2 and NasNetMobile outperformed all other models in terms of accuracy, precision, recall, and f1-score on train set and test set, respectively as shown in Table 7. Additionally, the misclassification difference between MobileNetV2 and NasNetMobile is just one which indicates that both models almost equally performed on the CT scan image dataset.

Table 7.
Overall summary of the best model found considering various factor on CT scan image dataset.
Similarly, on chest X-ray data set, Table 8 showed that NasNetMobile outperformed all of the models taking into account the statistical measurement as such accuracy (100%), precision (100%), recall (100%), f1-score (100%), confusing matrix (100%), and loss calculation.

Table 8.
Overall summary of the best model found considering various factors on chest X-ray image dataset.
To find out the best model among MobileNetV2 and NasNetMobile, a comparison was made between those two models. The models’ performance was calculated by averaging the overall performance both on the train and test set. From Table 9, MobileNetV2 outperformed NasNetMobile in terms of accuracy, precision, recall, and f1-score. However, misclassification rate for MobileNetV2 (11.25%) is slightly higher than NasNetMobile (10%). Since the dataset is small, the error rate may not be significant, yet, for a larger dataset, the misclassification rate may significantly impact.

Table 9.
Comparison between MobileNetV2 and NasNetMobile on both dataset.
Average accuracy was calculated by averaging the training and testing accuracy of all the models. Table 10 shows the average accuracy for CT scan and chest X-ray image dataset. Results show that almost all models performed better on the X-ray image data set compared to the CT scan. The average accuracy for all the models on CT scan and X-ray image dataset is 82.94% and 93.94%, respectively.

Table 10.
Models average accuracy on both datasets.
4.1. Feature Territory Highlighted by the Model on Different Layer
In this work, we tried to understand how each layer dealt with the actual image. Figure 6 demonstrated CT scan images during different layers. Note that, just a few of the layers from VGG16 were addressed here.
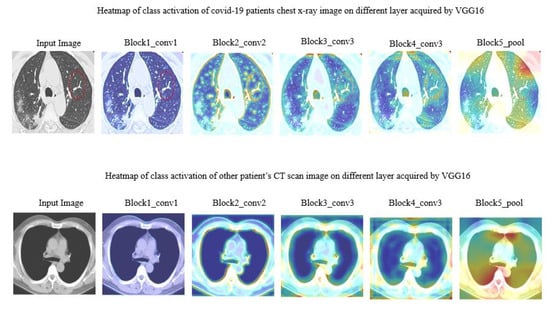
Figure 6.
Heat map of class activation of CT scan image on different layer acquired by VGG16.
Figure 7 manifested the different layer’s activity of model ResNet50 on chest X-ray images. The region spotted by model ResNet50 was highlighted with a heatmap.
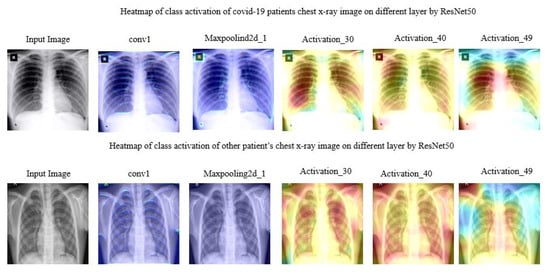
Figure 7.
Heat map of class activation of chest X-ray image on different layer acquired by ResNet50.
4.2. Models Interpretability with LIME
To identify which specific features help the deep learning model (MobileNetV2, NasNetMobile) to differentiate between COVID-19 and non-COVID-19 patients, Local Interpretable Model-agnostic Explanations (LIME) was used. LIME is a procedure that helps to understand how the input features of a deep learning model affect its predictions. For example, for image classification, LIME finds the set of super-pixels with the most grounded relationship with a prediction label [46]. LIME makes clarifications by creating another dataset of random perturbations (with their separate forecasts) around the occasion being clarified and afterward fitting a weighted neighborhood proxy model. This neighborhood model is usually a more straightforward model with natural interpretability, such as a linear regression model. LIME creates perturbations by turning on and off a portion of the super-pixels in the image. A quick shift strategy was utilized with the following parameters in order to calculate the super pixel, as shown in Table 11:

Table 11.
Parameter used to calculate maximum pixels.
Figure 8 is the output after computing the super-pixels on a sample chest CT scan image.
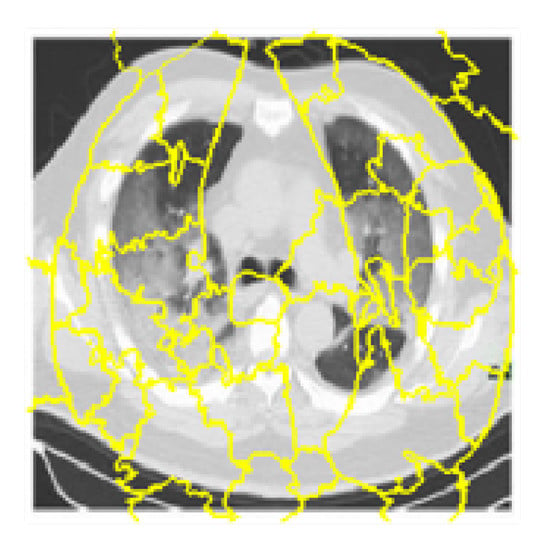
Figure 8.
Super-pixels on a sample chest CT scan images.
Additionally, Figure 9 shows different image conditions considering perturbation vectors and perturbed images. To predict the class, during this experiment 150 perturbations were used.
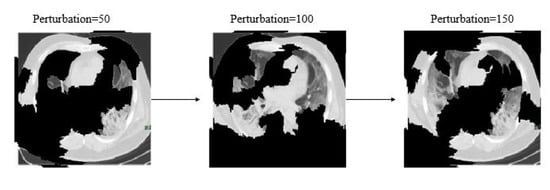
Figure 9.
Examples of perturbation vectors and perturbed images.
The distance metric was utilized to assess how far each perturbation is from the original image. Cosine metrics were used with kernel width as 1/4 to measure the original image’s distance and perturbed images. A weighted linear model was used to explain the overall model. A coefficient was found for every superpixel in the picture that represents how solid the superpixels impact in the prediction of COVID-19 patients. Finally, top features were sorted in order to determine what are the most important superpixel. Here, Figure 10 demonstrates the top four critical features.

Figure 10.
Top four features (a) on COVID-19 patients CT scan image (b) on other patients CT scan image.
Here Figure 11, depicts the overall interpretability for image classification with LIME on chest X-ray images considering each step. The prediction was conducted using NasNetMobile.
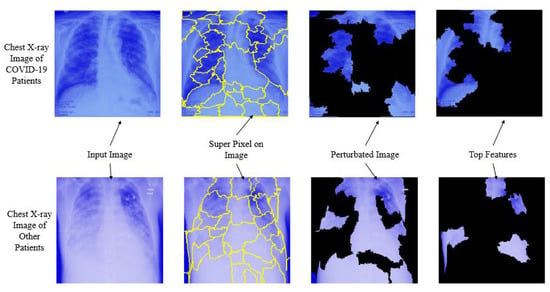

Figure 11.
Overall prediction analysis using Local Interpretable Model-agnostic Explanations.
In brief, based on the overall experiment, this study found that, among all eight deep learning models, MobileNetv2 and NasNetMobile performed better both on CT scan and chest X-ray image datasets. Additionally, all deep learning models performed well on the chest X-ray image dataset compared to CT scan images with an average 8% higher accuracy. This research addressed that existing deep learning approaches could be used along with RT-PCR testing as an alternative approach for detecting COVID-19 patients on a mixed datasets. The study results also revealed that, NasNetMobile can be used to identify COVID-19 patients both CT scan and chest X-ray images. Additionally, the proposed models detect the top features along with the predictions, which might assist the general practitioners to understand about the virus and the infectious regions.
5. Conclusions
In this study, in the CT scan image dataset, MobileNetV2 and NasNetMobile outperformed all other models, while NasNetMobile is the best model on the chest X-ray image dataset alone. NasNetMobile outperformed all other models—except MobileNetV2—with 95% CI on CT scan datasets, and accuracy ranges from 81.5% to 95.2%, and on chest X-ray image dataset, the accuracy varies 95.4% to 100%. Additionally, top features that differentiate between COVID-19 and other patients were analyzed using LIME. With this short time and pandemic situations, we expect this study will give some insights to researchers and developers who are actively seeking the alter screening procedures by using both CT scan and chest X-ray image datasets for COVID-19 patients. Additionally, the experimental result may imitate other current studies, reflecting the possibility of developing a COVID-19 screening system using a deep-learning approach. Further analysis includes but is not limited to- understanding deep learning models performance with highly imbalanced data, model performance with a larger dataset, Check for data bias [47], parameter tuning, and developing a decision support system.
Author Contributions
Conceptualization, M.M.A. and K.D.G.; methodology, M.M.A. and K.D.G.; software, M.M.A.; validation, K.D.G.; formal analysis, S.S., M.M.I. and M.L.R.; investigation, K.D.G.; writing—original draft preparation, M.M.A., M.S.H. and K.D.G.; writing—review and editing, S.S., M.M.R., M.L.R. and M.S.H.; visualization, M.M.I., M.L.R. and M.S.H. All authors have read and agreed to the published version of the manuscript.
Funding
This research received no external funding.
Conflicts of Interest
The authors declare no conflict of interest.
References
- Stoecklin, S.B.; Rolland, P.; Silue, Y.; Mailles, A.; Campese, C.; Simondon, A.; Mechain, M.; Meurice, L.; Nguyen, M.; Bassi, C.; et al. First cases of coronavirus disease 2019 (COVID-19) in France: Surveillance, investigations and control measures, January 2020. Eurosurveillance 2020, 25, 2000094. [Google Scholar]
- Dashbord. Covid-19 WorldMeter, September 2020. Available online: https://www.worldometers.info/coronavirus/ (accessed on 9 September 2020).
- McKeever, A. Here’s what coronavirus does to the body. Natl. Geogr. 2020. Available online: https://www.freedomsphoenix.com/Media/Media-Files/Heres-what-coronavirus-does-to-the-body.pdf (accessed on 12 March 2020).
- Mahase, E. Coronavirus: Covid-19 Has Killed More People than SARS and MERS Combined, Despite Lower Case Fatality Rate; BMJ: London, UK, 2020. [Google Scholar]
- Tanne, J.H.; Hayasaki, E.; Zastrow, M.; Pulla, P.; Smith, P.; Rada, A.G. Covid-19: How doctors and healthcare systems are tackling coronavirus worldwide. BMJ 2020, 368. [Google Scholar] [CrossRef]
- Ghoshal, B.; Tucker, A. Estimating uncertainty and interpretability in deep learning for coronavirus (COVID-19) detection. arXiv 2020, arXiv:2003.10769. [Google Scholar]
- Cohen, J.P.; Morrison, P.; Dao, L. COVID-19 image data collection. arXiv 2020, arXiv:2003.11597. [Google Scholar]
- Chest X-ray Images (Pneumonia). Available online: https://www.kaggle.com/paultimothymooney/chest-xray-pneumonia (accessed on 10 March 2020).
- Shi, F.; Wang, J.; Shi, J.; Wu, Z.; Wang, Q.; Tang, Z.; He, K.; Shi, Y.; Shen, D. Review of artificial intelligence techniques in imaging data acquisition, segmentation and diagnosis for covid-19. IEEE Rev. Biomed. Eng. 2020. [Google Scholar] [CrossRef] [PubMed]
- Narin, A.; Kaya, C.; Pamuk, Z. Automatic detection of coronavirus disease (covid-19) using x-ray images and deep convolutional neural networks. arXiv 2020, arXiv:2003.10849. [Google Scholar]
- Zhang, J.; Xie, Y.; Li, Y.; Shen, C.; Xia, Y. Covid-19 screening on chest x-ray images using deep learning based anomaly detection. arXiv 2020, arXiv:2003.12338. [Google Scholar]
- Wang, L.; Wong, A. COVID-Net: A Tailored Deep Convolutional Neural Network Design for Detection of COVID-19 Cases from Chest X-Ray Images. arXiv 2020, arXiv:2003.09871. [Google Scholar]
- Chen, J.; Wu, L.; Zhang, J.; Zhang, L.; Gong, D.; Zhao, Y.; Hu, S.; Wang, Y.; Hu, X.; Zheng, B.; et al. Deep learning-based model for detecting 2019 novel coronavirus pneumonia on high-resolution computed tomography: A prospective study. medRxiv. 2020. Available online: https://www.medrxiv.org/content/10.1101/2020.02.25.20021568v2.full.pdf (accessed on 12 March 2020).
- Jin, C.; Chen, W.; Cao, Y.; Xu, Z.; Zhang, X.; Deng, L.; Zheng, C.; Zhou, J.; Shi, H.; Feng, J. Development and Evaluation of an AI System for COVID-19 Diagnosis. medRxiv 2020. [Google Scholar] [CrossRef]
- Song, Y.; Zheng, S.; Li, L.; Zhang, X.; Zhang, X.; Huang, Z.; Chen, J.; Zhao, H.; Jie, Y.; Wang, R.; et al. Deep learning enables accurate diagnosis of novel coronavirus (COVID-19) with CT images. medRxiv 2020. [Google Scholar] [CrossRef]
- Butt, C.; Gill, J.; Chun, D.; Babu, B.A. Deep learning system to screen coronavirus disease 2019 pneumonia. Appl. Intell. 2020, 1. [Google Scholar] [CrossRef]
- Li, L.; Qin, L.; Xu, Z.; Yin, Y.; Wang, X.; Kong, B.; Bai, J.; Lu, Y.; Fang, Z.; Song, Q.; et al. Artificial intelligence distinguishes COVID-19 from community acquired pneumonia on chest CT. Radiology 2020, 200905. [Google Scholar] [CrossRef] [PubMed]
- Samek, W.; Wiegand, T.; Müller, K.R. Explainable artificial intelligence: Understanding, visualizing and interpreting deep learning models. arXiv 2017, arXiv:1708.08296. [Google Scholar]
- Gilpin, L.H.; Bau, D.; Yuan, B.Z.; Bajwa, A.; Specter, M.; Kagal, L. Explaining explanations: An overview of interpretability of machine learning. In Proceedings of the 2018 IEEE 5th International Conference on Data Science and Advanced Analytics (DSAA), Turin, Italy, 1–3 October 2018; pp. 80–89. [Google Scholar]
- Ribeiro, M.T.; Singh, S.; Guestrin, C. “Why should i trust you?” Explaining the predictions of any classifier. In Proceedings of the 22nd ACM SIGKDD International Conference on Knowledge Discovery and Data Mining, San Francisco, CA, USA, 13 August 2016; pp. 1135–1144. [Google Scholar]
- Guidotti, R.; Monreale, A.; Ruggieri, S.; Turini, F.; Giannotti, F.; Pedreschi, D. A survey of methods for explaining black box models. ACM Comput. Surv. (CSUR) 2018, 51, 1–42. [Google Scholar] [CrossRef]
- Holzinger, A.; Biemann, C.; Pattichis, C.S.; Kell, D.B. What do we need to build explainable AI systems for the medical domain? arXiv 2017, arXiv:1712.09923. [Google Scholar]
- Simonyan, K.; Zisserman, A. Very deep convolutional networks for large-scale image recognition. arXiv 2014, arXiv:1409.1556. [Google Scholar]
- Längkvist, M.; Karlsson, L.; Loutfi, A. A review of unsupervised feature learning and deep learning for time-series modeling. Pattern Recognit. Lett. 2014, 42, 11–24. [Google Scholar] [CrossRef]
- Akiba, T.; Suzuki, S.; Fukuda, K. Extremely large minibatch sgd: Training resnet-50 on imagenet in 15 minutes. arXiv 2017, arXiv:1711.04325. [Google Scholar]
- Huang, G.; Liu, Z.; Van Der Maaten, L.; Weinberger, K.Q. Densely connected convolutional networks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Honolulu, HI, USA, 21–26 July 2017; pp. 4700–4708. [Google Scholar]
- Sandler, M.; Howard, A.; Zhu, M.; Zhmoginov, A.; Chen, L.C. Mobilenetv2: Inverted residuals and linear bottlenecks. In Proceedings of the IEEE Conference on Computer Vision and Pattern Recognition, Salt Lake City, UT, USA, 18–23 June 2018; pp. 4510–4520. [Google Scholar]
- da Nóbrega, R.V.M.; Peixoto, S.A.; da Silva, S.P.P.; Rebouças Filho, P.P. Lung nodule classification via deep transfer learning in CT lung images. In Proceedings of the 2018 IEEE 31st International Symposium on Computer-Based Medical Systems (CBMS), Karlstad, Sweden, 18–21 June 2018; pp. 244–249. [Google Scholar]
- Varatharasan, V.; Shin, H.S.; Tsourdos, A.; Colosimo, N. Improving Learning Effectiveness For Object Detection and Classification in Cluttered Backgrounds. In Proceedings of the 2019 Workshop on Research, Education and Development of Unmanned Aerial Systems (RED UAS), Cranfield, UK, 25–27 November 2019; pp. 78–85. [Google Scholar]
- Pan, S.J.; Yang, Q. A survey on transfer learning. IEEE Trans. Knowl. Data Eng. 2009, 22, 1345–1359. [Google Scholar] [CrossRef]
- Deng, J.; Dong, W.; Socher, R.; Li, L.J.; Li, K.; Fei-Fei, L. Imagenet: A large-scale hierarchical image database. In Proceedings of the 2009 IEEE Conference on Computer Vision and Pattern Recognition, Miami, FL, USA, 20–25 June 2009; pp. 248–255. [Google Scholar]
- Yu, W.; Yang, K.; Bai, Y.; Xiao, T.; Yao, H.; Rui, Y. Visualizing and comparing AlexNet and VGG using deconvolutional layers. In Proceedings of the 33 rd International Conference on Machine Learning, New York City, NY, USA, 19–24 June 2016. [Google Scholar]
- Gupta, K.D.; Ahsan, M.; Andrei, S.; Alam, K.M.R. A Robust Approach of Facial Orientation Recognition from Facial Features. BRAIN. Broad Res. Artif. Intell. Neurosci. 2017, 8, 5–12. [Google Scholar]
- Ozturk, T.; Talo, M.; Yildirim, E.A.; Baloglu, U.B.; Yildirim, O.; Acharya, U.R. Automated detection of COVID-19 cases using deep neural networks with X-ray images. Comput. Biol. Med. 2020, 103792. [Google Scholar] [CrossRef] [PubMed]
- Denil, M.; Shakibi, B.; Dinh, L.; Ranzato, M.; De Freitas, N. Predicting parameters in deep learning. In Proceedings of the 26th International Conference on Neural Information Processing Systems—Volume 2, Lake Tahoe, NV USA, 5–10 December 2013; pp. 2148–2156. [Google Scholar]
- Sutskever, I.; Martens, J.; Dahl, G.; Hinton, G. On the importance of initialization and momentum in deep learning. In Proceedings of the 30th International Conference on Machine Learning, Atlanta, GA, USA; 2013; pp. 1139–1147. Available online: http://proceedings.mlr.press/v28/sutskever13.pdf (accessed on 12 March 2020).
- Kingma, D.P.; Ba, J. Adam: A method for stochastic optimization. arXiv 2014, arXiv:1412.6980. [Google Scholar]
- Zhang, C.; Liao, Q.; Rakhlin, A.; Miranda, B.; Golowich, N.; Poggio, T. Theory of deep learning IIb: Optimization properties of SGD. arXiv 2018, arXiv:1801.02254. [Google Scholar]
- Bengio, Y. Rmsprop and equilibrated adaptive learning rates for nonconvex optimization. arXiv 2015, arXiv:1502.04390v1. [Google Scholar]
- Perez, L.; Wang, J. The effectiveness of data augmentation in image classification using deep learning. arXiv 2017, arXiv:1712.04621. [Google Scholar]
- Filipczuk, P.; Fevens, T.; Krzyżak, A.; Monczak, R. Computer-aided breast cancer diagnosis based on the analysis of cytological images of fine needle biopsies. IEEE Trans. Med. Imaging 2013, 32, 2169–2178. [Google Scholar] [CrossRef]
- Ahsan, M.M. Real Time Face Recognition in Unconstrained Environment; Lamar University-Beaumont: Beaumont, TX, USA, 2018. [Google Scholar]
- Wilson, E.B. Probable inference, the law of succession, and statistical inference. J. Am. Stat. Assoc. 1927, 22, 209–212. [Google Scholar] [CrossRef]
- Edwards, W.; Lindman, H.; Savage, L.J. Bayesian statistical inference for psychological research. Psychol. Rev. 1963, 70, 193. [Google Scholar] [CrossRef]
- Brownlee, J. Machine Learning Mastery. 2014. Available online: http://machinelearningmastery.com/discover-feature-engineering-howtoengineer-features-and-how-to-getgood-at-it (accessed on 12 March 2020).
- Khan, A.; Gupta, K.D.; Kumar, N.; Venugopal, D. CIDMP: Completely Interpretable Detection of Malaria Parasite in Red Blood Cells using Lower-dimensional Feature Space. In Proceedings of the 2020 International Joint Conference on Neural Networks (IJCNN 2020), Glasgow, UK, 19–24 July 2020. [Google Scholar]
- Sen, S.; Dasgupta, D.; Gupta, K.D. An Empirical Study on Algorithmic Bias. In Proceedings of the 2020 IEEE 44th Annual Computers, Software, and Applications Conference (COMPSAC), Madrid, Spain, 13–17 July 2020. [Google Scholar]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2020 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).