Methanolysis of 2-Cyanopyridine in the Coordination Sphere of Manganese(II). The Structure of Mn4L6Cl2 cluster (L = Methyl Picolinimidate) †
Abstract
:1. Introduction
2. Materials and Methods
3. Results and Discussion
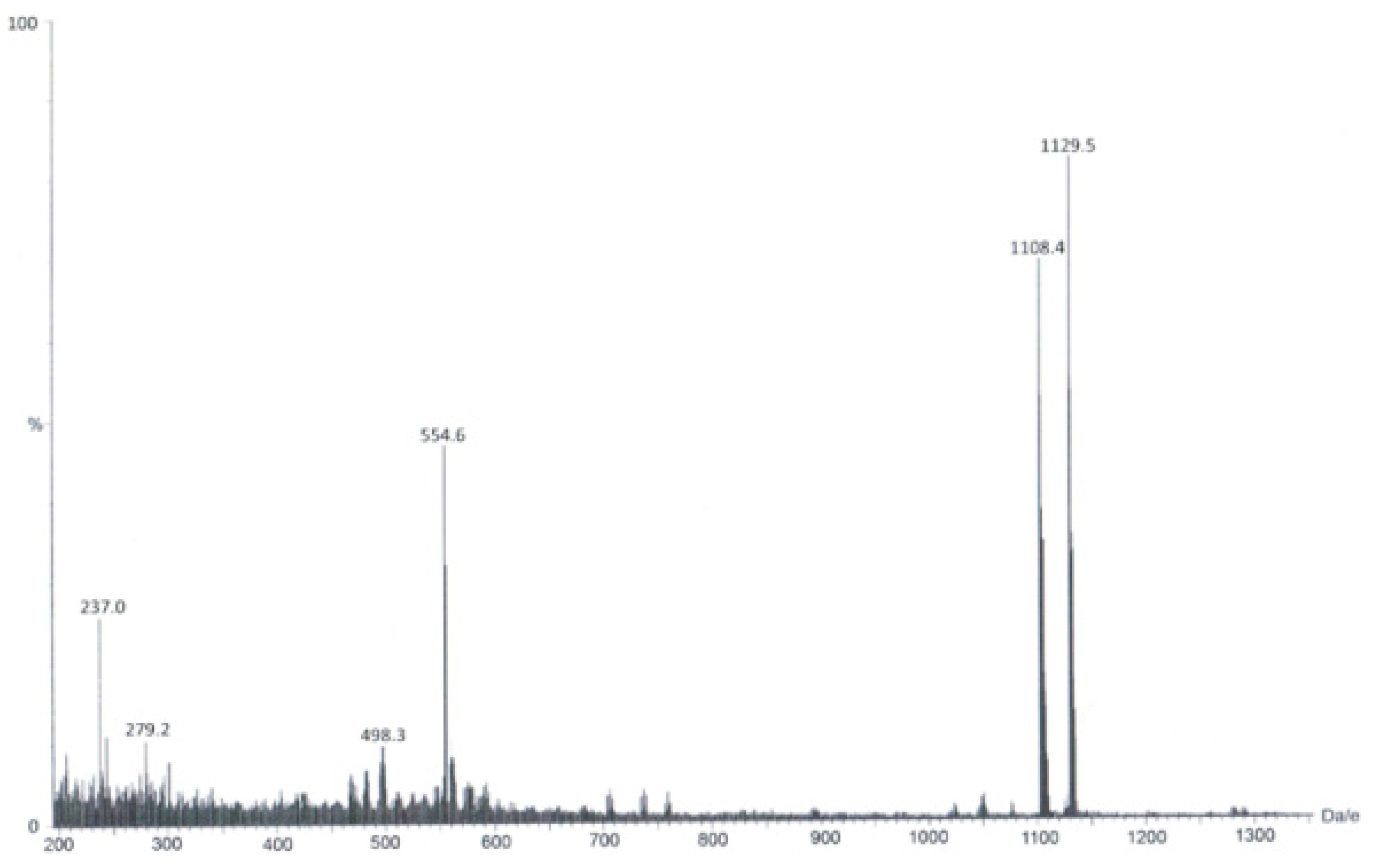
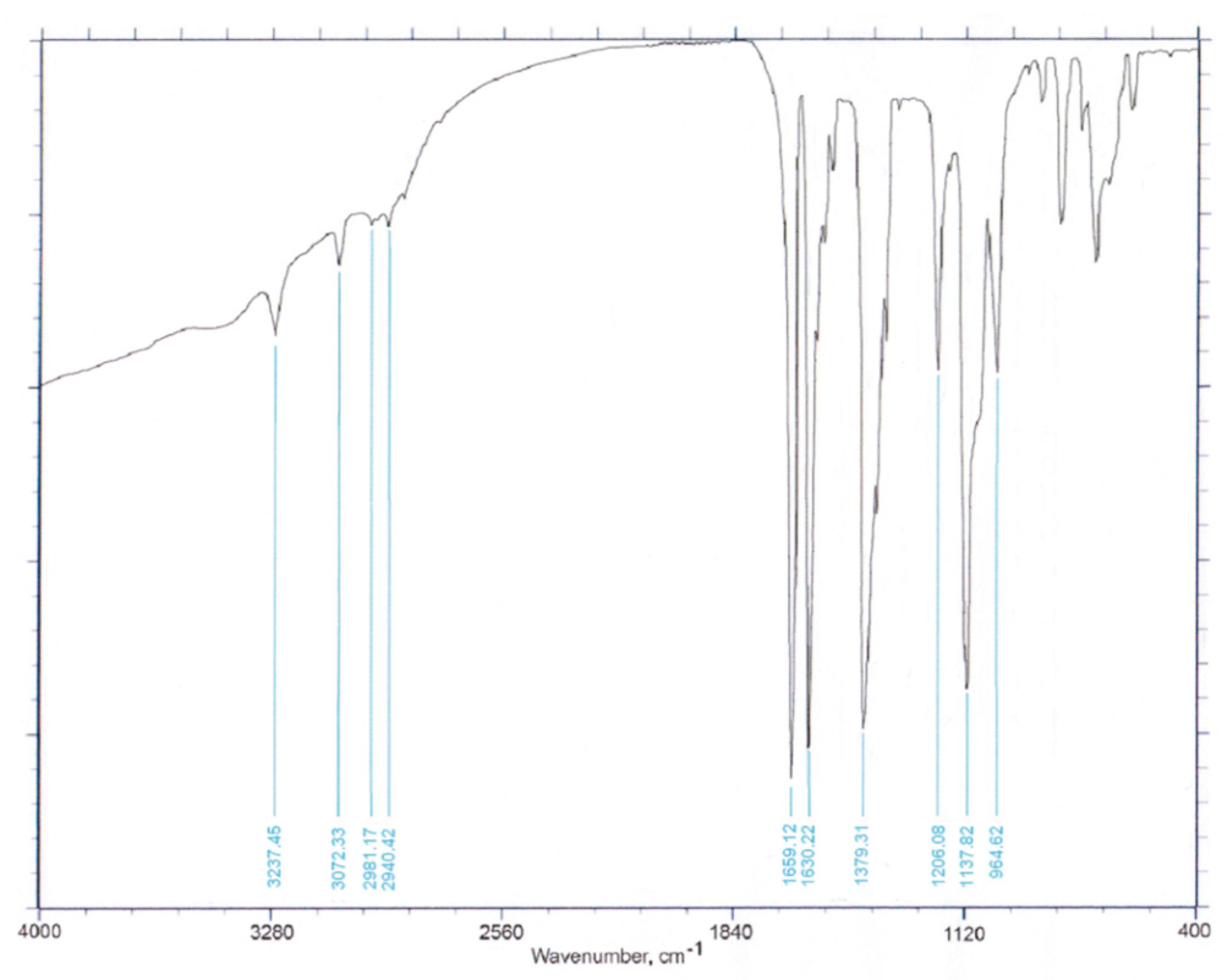
3.1. Synthesis and Spectroscopic Characterization
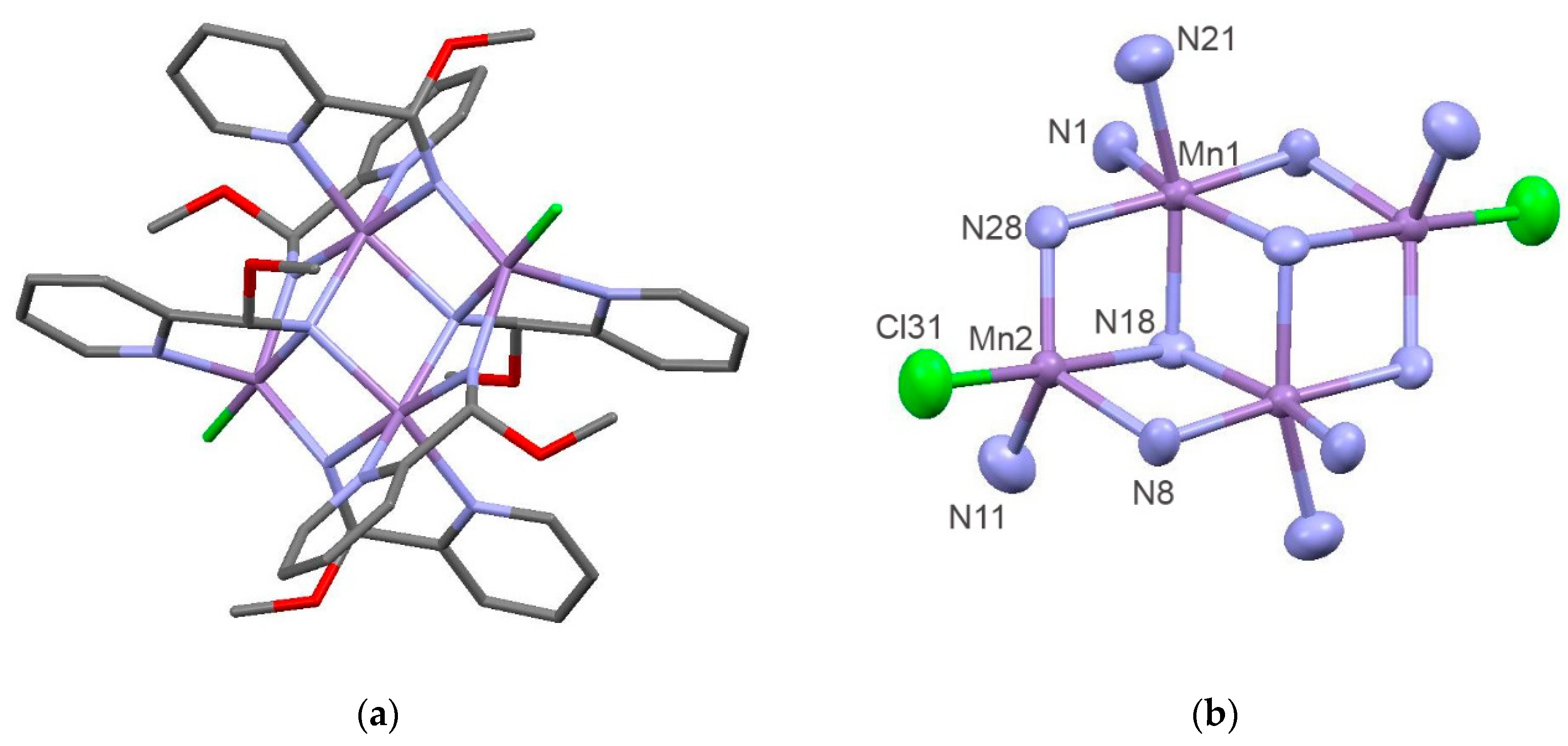
3.2. Crystallographic Studies
4. Conclusions
Acknowledgments
Conflicts of Interest
References
- Soucek, M.D.; Khattab, T.; Wu, J. Review of autoxidation and driers. Prog. Org. Coat. 2012, 73, 435–454. [Google Scholar] [CrossRef]
- Rajic, Z.; Tovmasyan, A.; de Santana, O.L.; Peixoto, I.N.; Spasojevic, I.; Do Monte, S.A.; Ventura, E.; Reboucas, J.S.; Batinic-Haberle, I. Challenges encountered during development of Mn porphyrin-based, potent redox-active drug and superoxide dismutase mimic, MnTnBuOE-2-PyP5+, and its alkoxyalkyl analogues. J. Inorg. Biochem. 2017, 169, 50–60. [Google Scholar] [CrossRef] [PubMed]
- Vázquez-Fernández, A.; Bermejo, M.R.; Fernández-García, M.I.; González-Riopedre, G.; Rodríguez-Doutón, M.J.; Maneiro, M. Influence of the geometry around the manganese ion on the peroxidase and catalase activities of Mn(III)-Schiff base complexes. J. Inorg. Biochem. 2011, 105, 1538–1547. [Google Scholar] [CrossRef] [PubMed]
- Forgacs, A.; Pujales-Paradela, R.; Regueiro-Figueroa, M.; Valencia, L.; Esteban-Gomez, D.; Botta, M.; Platas-Iglesias, C. Developing the family of picolinate ligands for Mn2+ complexation. Dalton Trans. 2017, 46, 1546–1558. [Google Scholar] [CrossRef] [PubMed]
- Dorkov, P.; Pantcheva, I.N.; Sheldrick, W.S.; Mayer-Figge, H.; Petrova, R.; Mitewa, M. Synthesis, structure and antimicrobial activity of manganese(II) and cobalt(II) complexes of the polyether ionophore antibiotic Sodium Monensin A. J. Inorg. Biochem. 2008, 102, 26–32. [Google Scholar] [CrossRef] [PubMed]
- Belaid, S.; Landreau, A.; Djebbar, S.; Benali-Baitich, O.; Bouet, G.; Bouchara, J.-P. Synthesis, characterization and antifungal activity of a series of manganese(II) and copper(II) complexes with ligands derived from reduced N,N′-O-phenylenebis(salicylideneimine). J. Inorg. Biochem. 2008, 102, 63–69. [Google Scholar] [CrossRef] [PubMed]
- Barraclough, C.G.; Gregson, A.K.; Mitra, S. Interpretation of the magnetic properties of manganese(II) phthalocyanine. J. Chem. Phys. 1974, 60, 962–968. [Google Scholar] [CrossRef]
- Bruker. SAINT, Siemens Area Detector Integration Software; Bruker AXS Inc.: Madison, WI, USA, 2003. [Google Scholar]
- Sheldrick, G.M. SADABS, Program for Scaling and Correction of Area Detector Data; University of Göttingen: Göttingen, Germany, 1996. [Google Scholar]
- Altomare, A.; Burla, M.C.; Camalli, M.; Cascarano, G.L.; Giacovazzo, C.; Guagliardi, A.; Moliterni, A.G.G.; Polidori, G.; Spagna, R. SIR97: A new tool for crystal structure determination and refinement. J. Appl. Crystallogr. 1999, 32, 115–119. [Google Scholar] [CrossRef]
- Sheldrick, G.M. A short history of SHELX. Acta Cryst. 2008, A64, 112. [Google Scholar] [CrossRef] [PubMed]
- Jamnicky, M.; Segla, P.; Koman, M. Methanolysis of pyridine-2-carbonitrile in the coordination sphere of copper(II), cobalt(II) and nickel(II). The structure of [Ni(O-methylpyridine-2-carboximidate)3]Br2·4H2O. Polyhedron 1995, 14, 1837–1847. [Google Scholar] [CrossRef]
- Garduno, J.A.; García, J.J. Synthesis of annidines and benzoxazoles from activated nitriles with Ni(0) catalysts. ACS Catal. 2015, 5, 3470–3477. [Google Scholar] [CrossRef]
- Devi, S.P.; Devi, R.B.; Devi, N.S.; Singh, L.J.; Singh, R.H. Structural and spectroscopic investigations on bis (1-amidino-O-2-alkoxyethylurea) copper (II) perchlorate complexes (alkoxy= methoxy, ethoxy or butoxy). Polyhedron 2012, 47, 1–8. [Google Scholar] [CrossRef]
- Pedrido, R.; Bermejo, M.R.; García-Deibe, A.M.; González-Noya, A.M.; Maneiro, M.; Vázquez, M. Metal complexes of a novel achiral symmetric pentadentate ligand—Crystal structures of monohelical zinc(II) and cadmium(II) complexes. Eur. J. Inorg. Chem. 2003, 3193–3200. [Google Scholar] [CrossRef]
- Addison, A.; Nageswara, R.T.; Reedijk, J.; Van Rijn, J.; Verschoor, G.C. Synthesis, structure, and spectroscopic properties of copper(II) compounds containing nitrogen-sulphur donor ligands; the crystal and molecular structure of aqua[1,7-bis(N-methylbenzimidazol-2’-yl)-2,6-dithiaheptane]copper(II) perchlorate. J. Chem. Soc. Dalton Trans. 1984, 1349–1356. [Google Scholar] [CrossRef]
| Compound | 2 |
|---|---|
| Empirical formula | C21H24ClMn2N6O3 |
| Formula weight | 553.79 |
| Temperature [K] | 293(2) |
| Wavelength [Å] | 0.71069 |
| Crystal system | Monoclinic |
| Space group | P21/n |
| a [Å] | 11.953(5) |
| b [Å] | 11.256(5) |
| c [Å] | 17.889(5) |
| α [°] | 90 |
| β [°] | 99.051(5) |
| γ [°] | 90 |
| Volume [Å3] | 2376.9(16) |
| Z | 4 |
| Dcalcd. [g cm−3] | 1.548 |
| μ [mm−1] | 1.21 |
| F(000) | 1132 |
| θmin/max[°] | 1.92/24.73 |
| Goodness-of-fit on F2 | 1.067 |
| Total data | 4038 |
| Unique data | 4038 |
| Data/restrains/parameters | 4038/0/299 |
| Final R indices [I > 2σ(I)] | R1 = 0.0894; wR2 = 0.2658 |
| R indices (all data) | R1 = 0.1223; wR2 =0.2812 |
| Mn(1)–N(28) | 2.048(8) | Mn(1)–Mn(2)#1 | 3.206(2) |
| Mn(1)–N(8) | 2.051(8) | Mn(2)–N(28)#1 | 1.939(8) |
| Mn(1)–N(18)#1 | 2.120(7) | Mn(2)–N(8) | 1.971(8) |
| Mn(1)–N(1) | 2.123(9) | Mn(2)–N(11) | 2.081(11) |
| Mn(1)–N(18) | 2.141(8) | Mn(2)–Cl(31) | 2.305(4) |
| Mn(1)–N(21) | 2.143(10) | Mn(2)–N(18) | 2.353(8) |
| Mn(1)–Mn(2) | 3.203(2) | Mn(2)–Mn(1)#1 | 3.206(2) |
| N(28)–Mn(1)–N(8) | 175.2(3) | N(28)#1–Mn(2)–N(8) | 124.9(4) |
| N(28)–Mn(1)–N(18)#1 | 80.3(3) | N(28)#1–Mn(2)–N(11) | 118.5(4) |
| N(8)–Mn(1)–N(18)#1 | 102.2(3) | N(8)–Mn(2)–N(11) | 101.3(4) |
| N(28)–Mn(1)–N(1) | 98.5(3) | N(28)#1–Mn(2)–Cl(31) | 101.6(2) |
| N(8)–Mn(1)–N(1) | 77.1(3) | N(8)–Mn(2)–Cl(31) | 102.9(2) |
| N(18)#1–Mn(1)–N(1) | 98.9(3) | N(11)–Mn(2)–Cl(31) | 105.2(3) |
| N(28)–Mn(1)–N(18) | 102.7(3) | N(28)#1–Mn(2)–N(18) | 76.8(3) |
| N(8)–Mn(1)–N(18) | 81.9(3) | N(8)–Mn(2)–N(18) | 78.4(3) |
| N(18)#1–Mn(1)–N(18) | 78.2(3) | N(11)–Mn(2)–N(18) | 75.4(4) |
| N(1)–Mn(1)–N(18) | 157.7(3) | Cl(31)–Mn(2)–N(18) | 178.4(2) |
| N(28)–Mn(1)–N(21) | 77.9(4) | ||
| N(8)–Mn(1)–N(21) | 100.4(4) | ||
| N(18)#1–Mn(1)–N(21) | 155.7(4) | ||
| N(1)–Mn(1)–N(21) | 94.8(4) | ||
| N(18)–Mn(1)–N(21) | 96.2(4) |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rouco, L.; Pedrido, R.; Fernández-García, M.I.; González-Noya, A.M.; Maneiro, M. Methanolysis of 2-Cyanopyridine in the Coordination Sphere of Manganese(II). The Structure of Mn4L6Cl2 cluster (L = Methyl Picolinimidate). Proceedings 2019, 9, 60. https://doi.org/10.3390/ecsoc-22-05659
Rouco L, Pedrido R, Fernández-García MI, González-Noya AM, Maneiro M. Methanolysis of 2-Cyanopyridine in the Coordination Sphere of Manganese(II). The Structure of Mn4L6Cl2 cluster (L = Methyl Picolinimidate). Proceedings. 2019; 9(1):60. https://doi.org/10.3390/ecsoc-22-05659
Chicago/Turabian StyleRouco, Lara, Rosa Pedrido, M. Isabel Fernández-García, Ana M. González-Noya, and Marcelino Maneiro. 2019. "Methanolysis of 2-Cyanopyridine in the Coordination Sphere of Manganese(II). The Structure of Mn4L6Cl2 cluster (L = Methyl Picolinimidate)" Proceedings 9, no. 1: 60. https://doi.org/10.3390/ecsoc-22-05659
APA StyleRouco, L., Pedrido, R., Fernández-García, M. I., González-Noya, A. M., & Maneiro, M. (2019). Methanolysis of 2-Cyanopyridine in the Coordination Sphere of Manganese(II). The Structure of Mn4L6Cl2 cluster (L = Methyl Picolinimidate). Proceedings, 9(1), 60. https://doi.org/10.3390/ecsoc-22-05659






