Artificial Neural Networks for Automated Cell Quantification in Lensless LED Imaging Systems †
Abstract
:1. Introduction
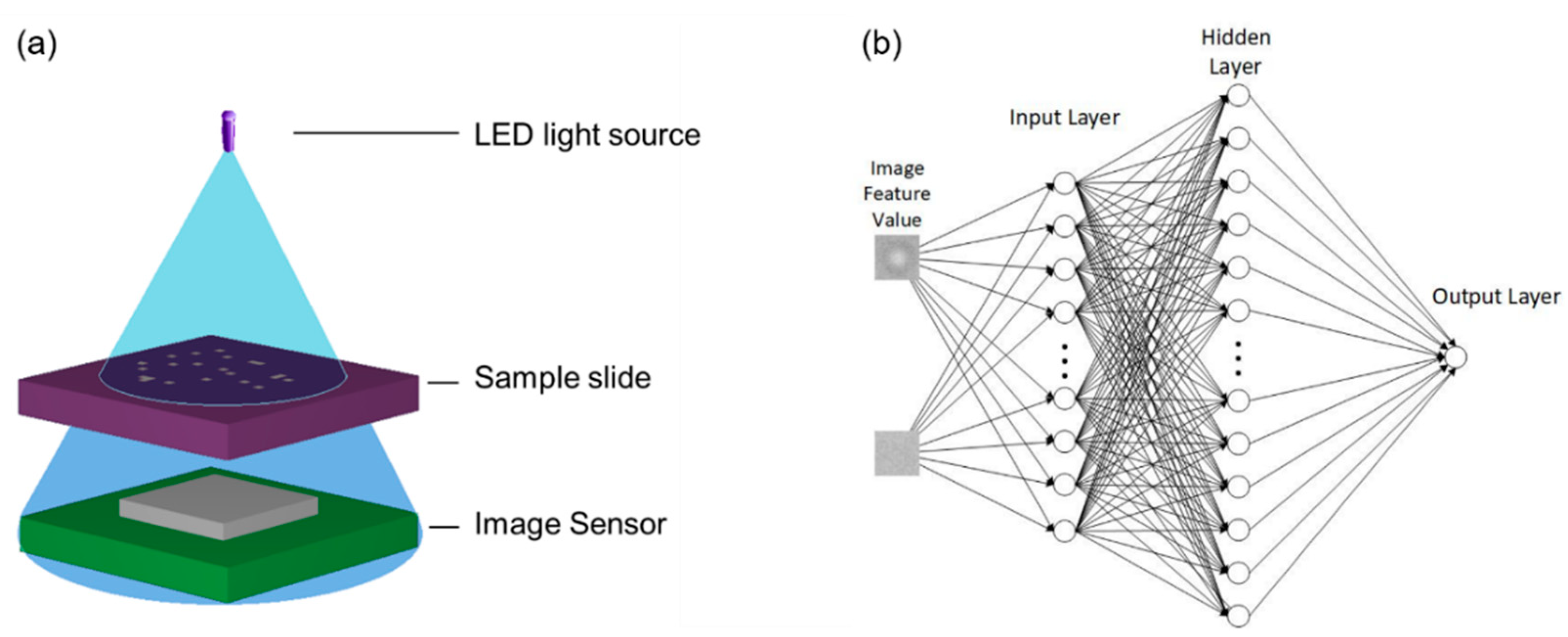
2. Cell Counting Method
3. Experimental Details
4. Results
5. Conclusions
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Wu, Y.; Ozcan, A. Lensless digital holographic microscopy and its applications in biomedicine and environmental monitoring. Methods 2018, 136, 4–16. [Google Scholar] [CrossRef] [PubMed]
- Zhang, Z. Artificial Neural Network. In Multivariate Time Series Analysis in Climate and Environmental Research; Springer: Cham, Switzerland, 2018; ISBN 978-3-319-67340-0. [Google Scholar]
- Rivenson, Y.; Zhang, Y.; Günaydın, H.; Teng, D.; Ozcan, A. Phase recovery and holographic image reconstruction using deep learning in neural networks. Light Sci. Appl. 2018, 7, 17141. [Google Scholar] [CrossRef] [PubMed]
- Cardot, H.; Degras, D. Online Principal Component Analysis in High Dimension: Which Algorithm to Choose? Int. Stat. Rev. 2017, 86, 29–50. [Google Scholar] [CrossRef]
- Scholz, G.; Xu, Q.; Schulze, T.; Boht, H.; Mattern, K.; Hartmann, J.; Dietzel, A.; Scherneck, S.; Rustenbeck, I.; Prades, J.D.; et al. LED-based tomographic imaging for live-cell monitoring of pancreatic islets in microfluidic channels. Proceedings 2017, 1, 552. [Google Scholar] [CrossRef]
- Poon, T.C.; Liu, J.P. Introduction to Modern Digital Holography; Cambridge University Press: Cambridge, UK, 2014. [Google Scholar]

Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2018 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Dharmawan, A.B.; Scholz, G.; Mariana, S.; Hörmann, P.; Ardiyanto, I.; Wibirama, S.; Hartmann, J.; Prades, J.D.; Hiller, K.; Waag, A.; et al. Artificial Neural Networks for Automated Cell Quantification in Lensless LED Imaging Systems. Proceedings 2018, 2, 989. https://doi.org/10.3390/proceedings2130989
Dharmawan AB, Scholz G, Mariana S, Hörmann P, Ardiyanto I, Wibirama S, Hartmann J, Prades JD, Hiller K, Waag A, et al. Artificial Neural Networks for Automated Cell Quantification in Lensless LED Imaging Systems. Proceedings. 2018; 2(13):989. https://doi.org/10.3390/proceedings2130989
Chicago/Turabian StyleDharmawan, Agus Budi, Gregor Scholz, Shinta Mariana, Philipp Hörmann, Igi Ardiyanto, Sunu Wibirama, Jana Hartmann, Joan Daniel Prades, Karsten Hiller, Andreas Waag, and et al. 2018. "Artificial Neural Networks for Automated Cell Quantification in Lensless LED Imaging Systems" Proceedings 2, no. 13: 989. https://doi.org/10.3390/proceedings2130989
APA StyleDharmawan, A. B., Scholz, G., Mariana, S., Hörmann, P., Ardiyanto, I., Wibirama, S., Hartmann, J., Prades, J. D., Hiller, K., Waag, A., & Wasisto, H. S. (2018). Artificial Neural Networks for Automated Cell Quantification in Lensless LED Imaging Systems. Proceedings, 2(13), 989. https://doi.org/10.3390/proceedings2130989






