DNA-Origami-Aided Lithography for Sub-10 Nanometer Pattern Printing †
Abstract
:1. Introduction
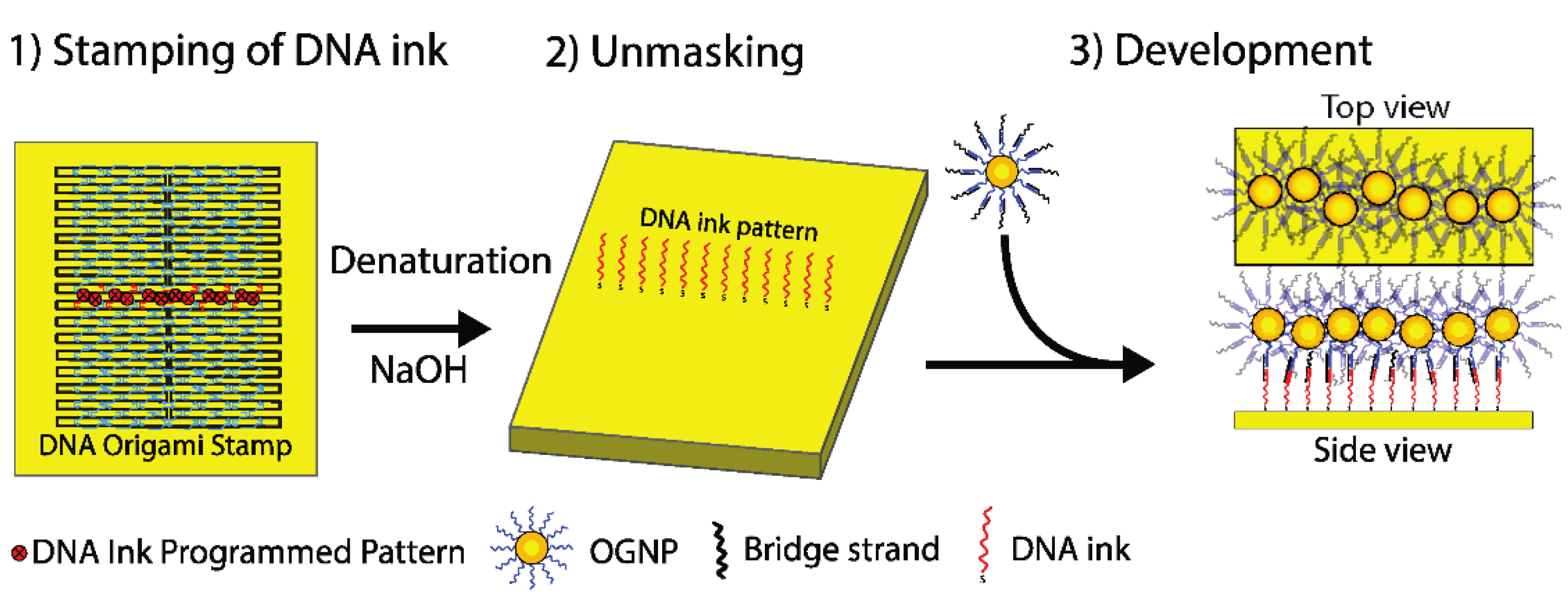
2. The Method
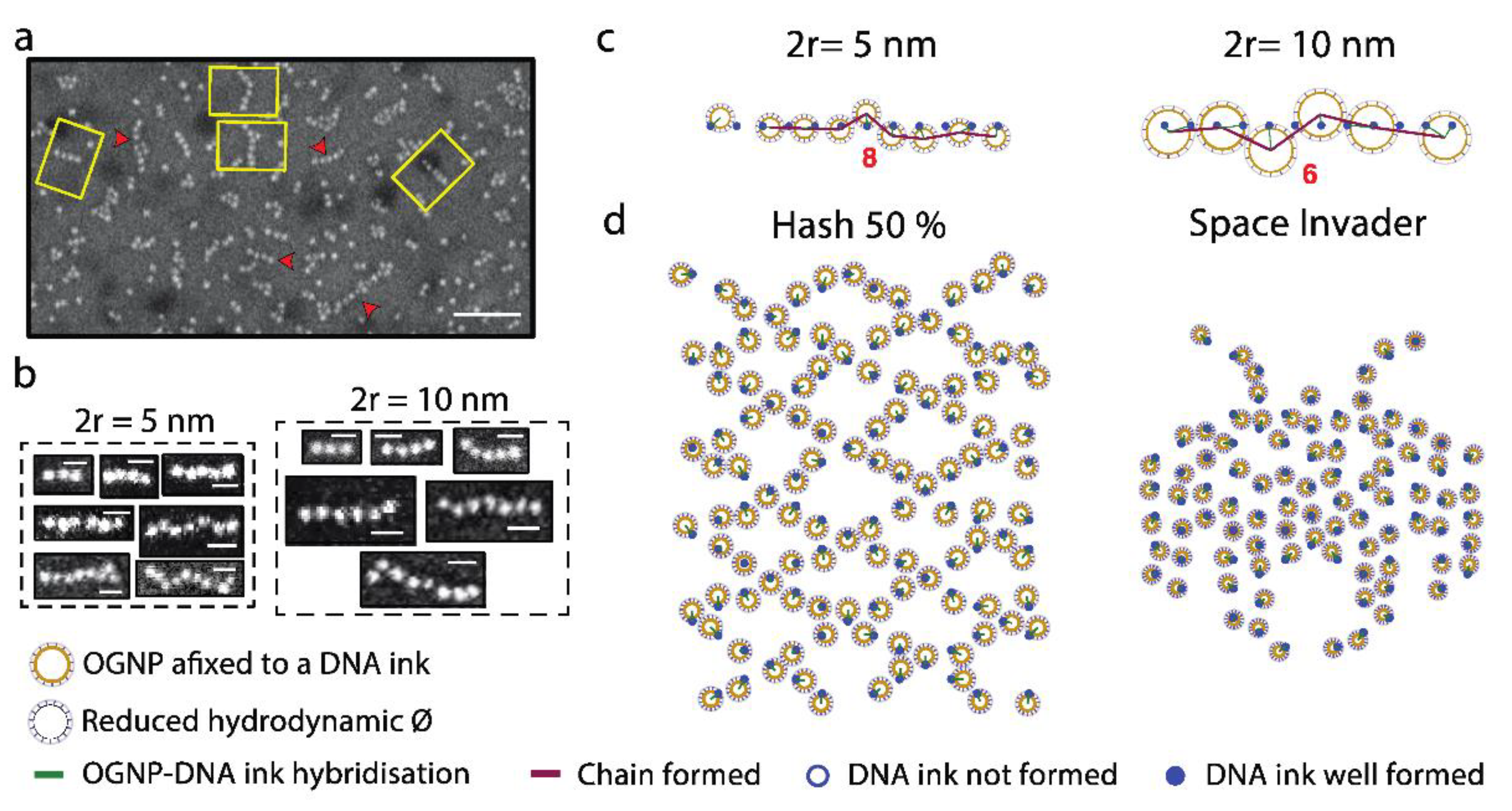
3. Results and Discussion
4. Conclusions
Acknowledgments
Conflicts of Interest
References
- Seeman, N.C. Nanomaterials Based on DNA. Annu. Rev. Biochem. 2010, 79, 65–87. [Google Scholar] [CrossRef] [PubMed]
- Rothemund, P.W. Folding DNA to create nanoscale shapes and patterns. Nature 2006, 440, 297–302. [Google Scholar] [CrossRef] [PubMed]
- Kershner, R.J.; Bozano, L.D.; Micheel, C.M.; Hung, A.M.; Fornof, A.R.; Cha, J.N.; Rettner, C.T.; Bersani, M.; Frommer, J.; Rothemund, P.W.K.; et al. Placement and orientation of individual DNA shapes on lithographically patterned surfaces. Nat. Nanotechnol. 2009, 4, 557–561. [Google Scholar] [CrossRef] [PubMed]
- Scheible, M.B.; Pardatscher, G.; Kuzyk, A.; Simmel, F.C. Single Molecule Characterization of DNA Binding and Strand Displacement Reactions on Lithographic DNA Origami Microarrays. Nano Lett. 2014, 14, 1627–1633. [Google Scholar] [CrossRef]
- Gállego, I.; Manning, B.; Prades, J.D.; Mir, M.; Samitier, J.; Eritja, R. DNA-Origami-Driven Lithography for Patterning on Gold Surfaces with Sub-10 nm Resolution. Adv. Mater. 2017, 29, 1–7. [Google Scholar] [CrossRef] [PubMed]
- Hsieh, K.; Ferguson, B.S.; Eisenstein, M.; Plaxco, K.W.; Soh, H.T. Integrated electrochemical microsystems for genetic detection of pathogens at the point of care. Acc. Chem. Res. 2015, 48, 911–920. [Google Scholar] [CrossRef] [PubMed]
- Kildishev, A.V.; Boltasseva, A.; Shalaev, V.M. Planar Photonics with Metasurfaces. Science 2013, 339, 1232009. [Google Scholar] [CrossRef] [PubMed]
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2017 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Gállego, I.; Manning, B.; Prades, J.D.; Mir, M.; Samitier, J.; Eritja, R. DNA-Origami-Aided Lithography for Sub-10 Nanometer Pattern Printing. Proceedings 2017, 1, 325. https://doi.org/10.3390/proceedings1040325
Gállego I, Manning B, Prades JD, Mir M, Samitier J, Eritja R. DNA-Origami-Aided Lithography for Sub-10 Nanometer Pattern Printing. Proceedings. 2017; 1(4):325. https://doi.org/10.3390/proceedings1040325
Chicago/Turabian StyleGállego, Isaac, Brendan Manning, Joan Daniel Prades, Mònica Mir, Josep Samitier, and Ramon Eritja. 2017. "DNA-Origami-Aided Lithography for Sub-10 Nanometer Pattern Printing" Proceedings 1, no. 4: 325. https://doi.org/10.3390/proceedings1040325
APA StyleGállego, I., Manning, B., Prades, J. D., Mir, M., Samitier, J., & Eritja, R. (2017). DNA-Origami-Aided Lithography for Sub-10 Nanometer Pattern Printing. Proceedings, 1(4), 325. https://doi.org/10.3390/proceedings1040325





