RIAM: A Universal Accessible Protocol for the Isolation of High Purity DNA from Various Soils and Other Humic Substances
Abstract
1. Introduction
2. Experimental Design
2.1. Important Materials
2.2. Materials
- GMA 80 Mesh Garnet Abrasive Media (GMA Group, Melbourne, Australia);
- MOLYKOTE Vacuum grease (DuPont, Wilmington, DE, USA).
2.3. Equipment
- Precellys Evolution Homogenizer (Bertin Technologies SAS, Montigny-le-Bretonneux, France);
- Microcentrifuge.
3. Procedure
- Place a sample of soil (50–300 mg) in a 2 mL test tube with a screw cap, add 300 µL of Sα solution to the tube and suspend the soil by vigorous shaking on a shaker for 5–10 min.
- Add to the suspension 1 g of Garnet abrasive (fraction 80 Mesh, 0.18–0.3 mm) and 500 µL of a mixture of phenol/chloroform taken in a ratio of 30/70 parts, and tightly tighten the caps;
- Place the tubes in a bead homogenizer (Precellys Evolution, France) and shake at 6000 rpm 2 times for 30 s;
- Centrifuge the homogenate at a maximum speed of (16,000–20,000 g) for 2 min.
- Add 500 μL of Sβ solution to the same tube, close the tube, mix the contents by turning the tube upside down 8–10 times, and centrifuge at maximum speed for 5 min. (To speed up the isolation process at this stage, after adding the Sβ solution, approximately 1 g of silicone grease (Dow Corning High Vacuum Grease, Molykote) can be added to the tube, which after centrifugation forms a tight plug between the aqueous and phenol-chloroform phases, which allows to separate the aqueous phase simply by pouring from test tube to test tube without tedious pipetting. Before use, the lubricant is preferably autoclaved for 30 min at 1 atmosphere);
- Transfer the aqueous phase into a new 1.5 mL microcentrifuge tube, add an equal volume of Sγ solution (volume determination accuracy ± 10%), mix thoroughly, incubate at room temperature for 5 min and precipitate DNA-CTAB complexes by centrifugation for 5 min at maximum speed. (Volume of the aqueous phase may vary depending on the amount of clay minerals in the soil);
- Remove the supernatant by inverting the tube, remove the remaining drops by placing the neck of the tube on sterile (autoclaved) filter paper. Complete removal of the supernatant is not necessary. Add 500 μL of Sδ solution to the precipitate, incubate for 5 min at room temperature with vigorous shaking, then precipitate by centrifugation for 5 min at maximum speed;
- Remove the supernatant as in step 7. Add 400 µL of Sε solution to the pellet and incubate for 5 min at room temperature with vigorous shaking. Then add an equal volume of a mixture of chloroform-isoamyl alcohol (24:1) to the solution and, after vigorous shaking, separate the aqueous and organic phases by centrifugation at maximum speed for 5 min. (For some soils incubation of test tubes at protocol steps 6, 7, and 8 at 65 °C instead room temperature can increase the DNA yield);
- Transfer the aqueous phase into a new 1.5 mL centrifuge tube, add 2.5 volumes (1 mL) of chilled 96% ethanol, mix well and precipitate the nucleic acids by centrifugation at maximum speed for 10 min;
- Remove the supernatant, wash the precipitate by adding 400 µL of cold 70% ethanol solution, followed by centrifugation for 5 min at maximum speed;
- Carefully remove the supernatant (preferably with an aspirator), dry the precipitate for 1 min at 65 °C, and dissolve in the desired volume of autoclaved water or 5 mM Tris-HCl, pH 8.0. To speed up the dissolution, the tubes can be incubated for 5 min at 65 °C.
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Guerra, V.; Beule, L.; Lehtsaar, E.; Liao, H.L.; Karlovsky, P. Improved protocol for DNA extraction from subsoils using phosphate lysis buffer. Microorganisms 2020, 8, 532. [Google Scholar] [CrossRef] [PubMed]
- Saidi-Mehrabad, A.; Neuberger, P.; Cavaco, M.; Froese, D.; Lanoil, B. Optimization of subsampling, decontamination, and DNA extraction of difficult peat and silt permafrost samples. Sci. Rep. 2020, 10, 14295. [Google Scholar] [CrossRef] [PubMed]
- Echeverría-Beirute, F.; Varela-Benavides, I.; Jiménez-Madrigal, J.P.; Carvajal-Chacon, M.; Guzmán-Hernández, T. EDNA Extraction protocol for metagenomic studies in tropical soils. Biotechniques 2021, 71, 581–586. [Google Scholar] [CrossRef] [PubMed]
- Sakai, Y. Improvements in extraction methods of high-molecular-weight DNA from soils by modifying cell lysis conditions and reducing adsorption of DNA onto soil particles. Microbes Environ. 2021, 36, ME21017. [Google Scholar] [CrossRef] [PubMed]
- Cai, P.; Huang, Q.; Zhang, X.; Chen, H. Adsorption of DNA on Clay Minerals and Various Colloidal Particles from an Alfisol. Soil. Biol. Biochem. 2006, 38, 471–476. [Google Scholar] [CrossRef]
- Franchi, M.; Ferris, J.P.; Gallori, E. Cations as mediators of the adsorption of nucleic acids on clay surfaces in prebiotic environments. Orig. Life Evol. Biosph. 2003, 33, 1–16. [Google Scholar] [CrossRef]
- Beall, G.W.; Sowersby, D.S.; Roberts, R.D.; Robson, M.H.; Lewis, L.K. Analysis of oligonucleotide DNA binding and sedimentation properties of montmorillonite clay using ultraviolet light spectroscopy. Biomacromolecules 2009, 10, 105–112. [Google Scholar] [CrossRef]
- Direito, S.O.L.; Marees, A.; Röling, W.F.M. sensitive life detection strategies for low-biomass environments: Optimizing extraction of nucleic acids adsorbing to terrestrial and Mars analogue minerals. FEMS Microbiol. Ecol. 2012, 81, 111–123. [Google Scholar] [CrossRef]
- Ikeda, S.; Tsurumaru, H.; Wakai, S.; Noritake, C.; Fujishiro, K.; Akasaka, M.; Ando, K. Evaluation of the effects of different additives in improving the DNA extraction yield and quality from Andosol. Microbes Environ. 2008, 23, 159–166. [Google Scholar] [CrossRef]
- Lever, M.A.; Torti, A.; Eickenbusch, P.; Michaud, A.B.; Šantl-Temkiv, T.; Jørgensen, B.B. A modular method for the extraction of DNA and RNA, and the separation of DNA pools from diverse environmental sample types. Front. Microbiol. 2015, 6, 476. [Google Scholar] [CrossRef]
- Jacobsen, C.S.; Nielsen, T.K.; Vester, J.K.; Stougaard, P.; Nielsen, J.L.; Voriskova, J.; Winding, A.; Baldrian, P.; Liu, B.; Frostegård, Å.; et al. Inter-laboratory testing of the effect of DNA blocking reagent G2 on DNA extraction from low-biomass clay samples. Sci. Rep. 2018, 8, 5711. [Google Scholar] [CrossRef] [PubMed]
- Tanaka, T.; Sakai, R.; Kobayashi, R.; Hatakeyama, K.; Matsunaga, T. Contributions of phosphate to DNA adsorption/desorption behaviors on Aminosilane-modified magnetic nanoparticles. Langmuir 2009, 25, 2956–2961. [Google Scholar] [CrossRef] [PubMed]
- Hurt, R.A.; Robeson, M.S.; Shakya, M.; Moberly, J.G.; Vishnivetskaya, T.A.; Gu, B.; Elias, D.A. Improved yield of high molecular weight DNA coincides with increased microbial diversity access from iron oxide cemented sub-surface clay environments. PLoS ONE 2014, 9, e102826. [Google Scholar] [CrossRef]
- Engel, K.; Coyotzi, S.; Vachon, M.A.; McKelvie, J.R.; Neufeld, J.D. Validating DNA extraction protocols for bentonite clay. mSphere 2019, 4, e00334-19. [Google Scholar] [CrossRef]
- Phosphate Buffer. Cold Spring Harb. Protoc. 2006, pdb.rec8543. [CrossRef]
- Jones, A.S. Use of Alkyltrimethylammonium Bromides for the isolation of ribo- and deoxribo-nucleic acids. Nature 1963, 199, 280–282. [Google Scholar] [CrossRef]
- Cho, J.-C.; Lee, D.-H.; Cheol, C.-Y.; Cho, J.-C.; Kim, S.-J. Direct extraction of DNA from soil for amplification of 16S RRNA gene sequences by polymerase chain reaction. J. Microbiol. 1996, 34, 229–235. [Google Scholar]
- Zhou, J.; Bruns, M.A.; Tiedje, J.M. DNA recovery from soils of diverse composition. Appl. Environ. Microbiol. 1996, 62, 316–322. [Google Scholar] [CrossRef]
- Kachiprath, B.; Jayanath, G.; Solomon, S.; Sarasan, M. CTAB influenced differential elution of metagenomic DNA from saltpan and marine sediments. 3 Biotech 2018, 8, 44. [Google Scholar] [CrossRef]
- Pattarkine, M.V.; Ganesh, K.N. DNA-surfactant interactions: Coupled cooperativity in ligand binding leads to duplex stabilization. Biochem. Biophys. Res. Commun. 1999, 263, 41–46. [Google Scholar] [CrossRef]
- Frostegård, Å.; Courtois, S.; Ramisse, V.; Clerc, S.; Bernillon, D.; le Gall, F.; Jeannin, P.; Nesme, X.; Simonet, P. Quantification of bias related to the extraction of DNA directly from soils. Appl. Environ. Microbiol. 1999, 65, 5409–5420. [Google Scholar] [CrossRef]
- Jackson, C.R.; Harper, J.P.; Willoughby, D.; Roden, E.E.; Churchill, P.F. A simple, efficient method for the separation of Humic substances and DNA from environmental samples. Appl. Environ. Microbiol. 1997, 63, 4993–4995. [Google Scholar] [CrossRef]
- Anderson, M. A Lab-Made Method for extracting DNA from soils. Soil Res. 2018, 56, 560–567. [Google Scholar] [CrossRef]
- Miller, D.N.; Bryant, J.E.; Madsen, E.L.; Ghiorse, W.C. Evaluation and optimization of DNA extraction and purification procedures for soil and sediment samples. Appl. Environ. Microbiol. 1999, 65, 4715–4724. [Google Scholar] [CrossRef]
- Sarma, P.V.S.R.N.; Srinivas, V.; Anil, K.; Podile, A.R. Isolation and purification of microbial community DNA from soil naturally enriched for chitin. Biologia 2012, 67, 644–648. [Google Scholar] [CrossRef]
- Devi, S.G.; Fathima, A.A.; Radha, S.; Arunraj, R.; Curtis, W.R.; Ramya, M. A rapid and economical method for efficient DNA extraction from diverse soils suitable for metagenomic applications. PLoS ONE 2015, 10, e0132441. [Google Scholar] [CrossRef]
- Arbeli, Z.; Fuentes, C.L. Improved purification and PCR amplification of DNA from environmental samples. FEMS Microbiol. Lett. 2007, 272, 269–275. [Google Scholar] [CrossRef]
- Braid, M.D.; Daniels, L.M.; Kitts, C.L. Removal of PCR Inhibitors from Soil DNA by Chemical Flocculation. J. Microbiol. Methods 2003, 52, 389–393. [Google Scholar] [CrossRef]
- Zaveri, P.; Patel, R.; Patel, M.; Sarodia, D.; Munshi, N.S. Modification of extraction method for community DNA isolation from salt affected compact wasteland soil samples. MethodsX 2017, 4, 63–67. [Google Scholar] [CrossRef]
- Alvarez, R.; Evans, L.A.; Milham, P.J.; Wilson, M.A. Effects of humic material on the precipitation of calcium phosphate. Geoderma 2004, 118, 245–260. [Google Scholar] [CrossRef]
- Arkhipchenko, I.A.; Salkinoja-Salonen, M.S.; Karyakina, J.N.; Tsitko, I. Study of three fertilizers produced from farm waste. Appl. Soil. Ecol. 2005, 30, 126–132. [Google Scholar] [CrossRef]
- Bates, S.T.; Berg-Lyons, D.; Caporaso, J.G.; Walters, W.A.; Knight, R.; Fierer, N. Examining the global distribution of dominant archaeal populations in soil. ISME J. 2011, 5, 908–917. [Google Scholar] [CrossRef]
- Callahan, B.J.; McMurdie, P.J.; Rosen, M.J.; Han, A.W.; Johnson, A.J.A.; Holmes, S.P. DADA2: High-resolution sample inference from Illumina amplicon data. Nat. Methods 2016, 13, 581–583. [Google Scholar] [CrossRef]
- Wright, E.S. Using DECIPHER v2.0 to analyze big biological sequence data in R. R J. 2016, 8, 352–359. [Google Scholar] [CrossRef]
- Quast, C.; Pruesse, E.; Yilmaz, P.; Gerken, J.; Schweer, T.; Yarza, P.; Peplies, J.; Glöckner, F.O. The SILVA ribosomal RNA gene database project: Improved data processing and web-based tools. Nucleic Acids Res. 2013, 41, D590–D596. [Google Scholar] [CrossRef]
- Janssen, S.; McDonald, D.; Gonzalez, A.; Navas-Molina, J.A.; Jiang, L.; Xu, Z.Z.; Winker, K.; Kado, D.M.; Orwoll, E.; Manary, M.; et al. Phylogenetic placement of exact amplicon sequences improves associations with clinical information. mSystems 2018, 3, e00021-18. [Google Scholar] [CrossRef]
- Bolyen, E.; Rideout, J.R.; Dillon, M.R.; Bokulich, N.A.; Abnet, C.C.; Al-Ghalith, G.A.; Alexander, H.; Alm, E.J.; Arumugam, M.; Asnicar, F.; et al. Reproducible, interactive, scalable and extensible microbiome data science using QIIME 2. Nat. Biotechnol. 2019, 37, 852–857. [Google Scholar] [CrossRef]
- Voyron, S.; Marisa, Í.; Victorino, M.; Caser, M.; Demasi, S.; Scariot, V.; Bianciotto, V.; Ghignone, S.; Lumini, E. Truth or lie: Does the DNA extraction procedure really affect the insight in composition and diversity of microbial communities in saffron cultivated soils? Appl. Microbiol. 2022, 2, 492–501. [Google Scholar] [CrossRef]
- Shaffer, J.P.; Carpenter, C.S.; Martino, C.; Salido, R.A.; Minich, J.J.; Bryant, M.; Sanders, K.; Schwartz, T.; Humphrey, G.; Swafford, A.D.; et al. A comparison of six DNA extraction protocols for 16S, ITS and shotgun metagenomic sequencing of microbial communities. Biotechniques 2022, 73, 34–46. [Google Scholar] [CrossRef]
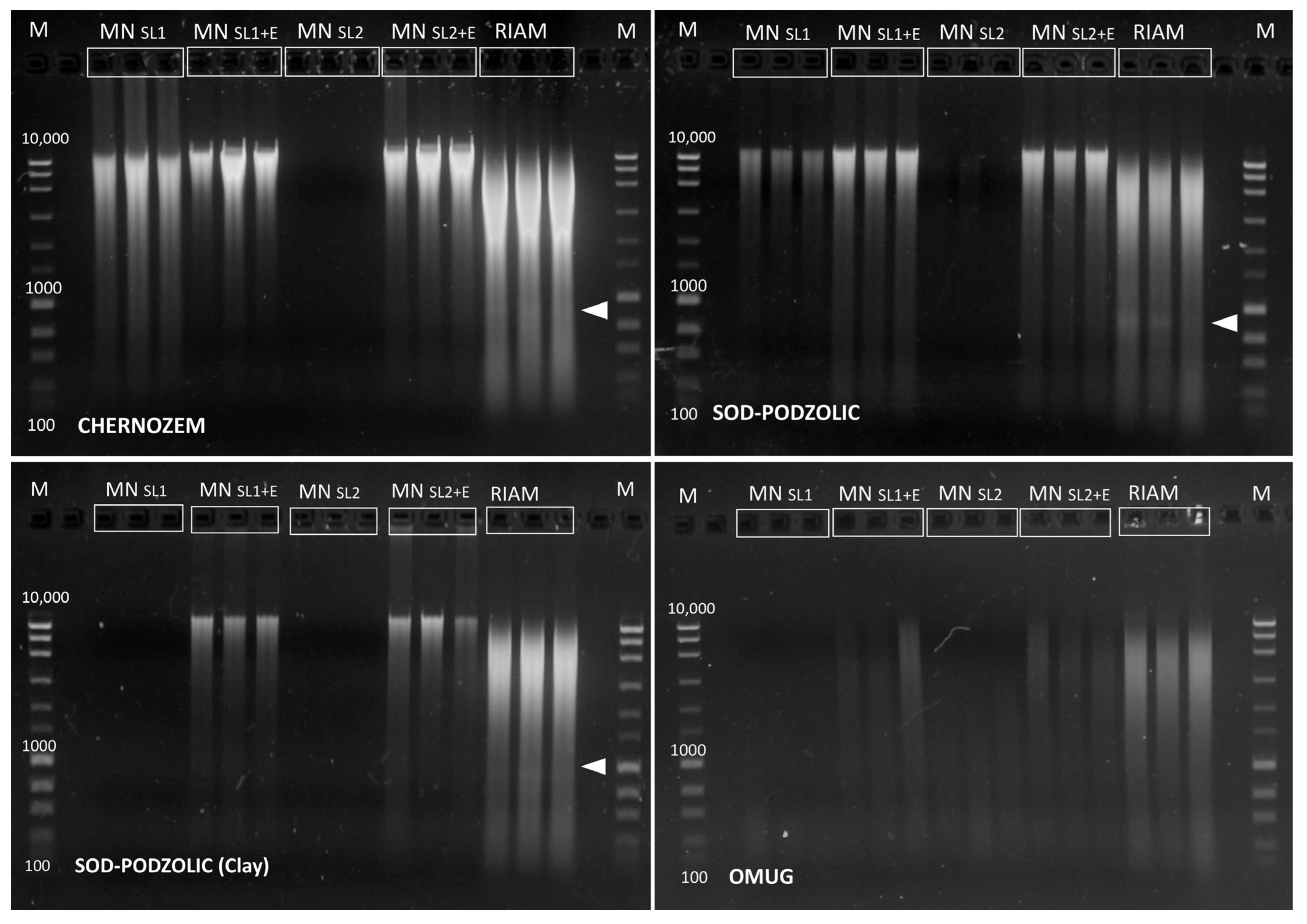
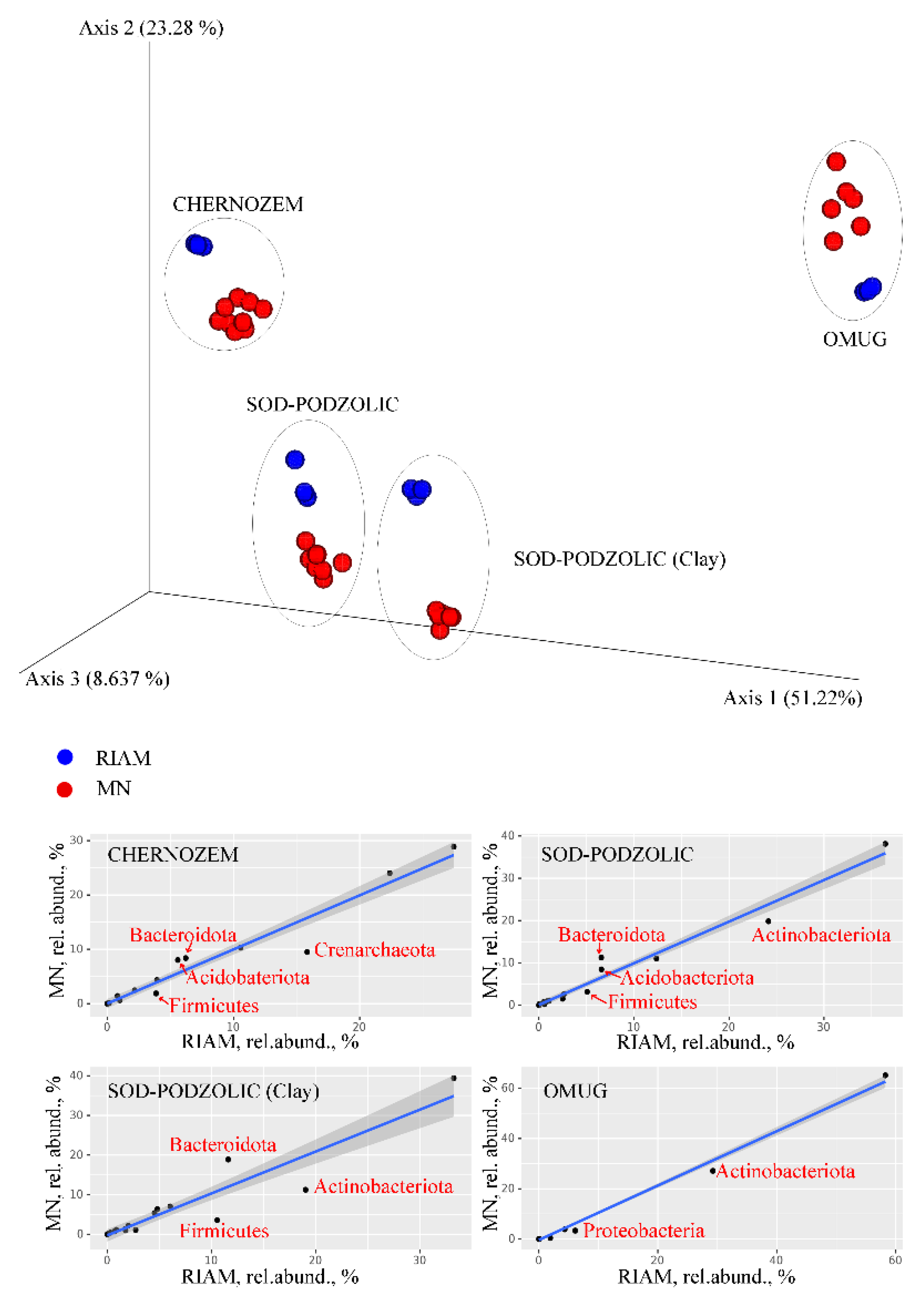
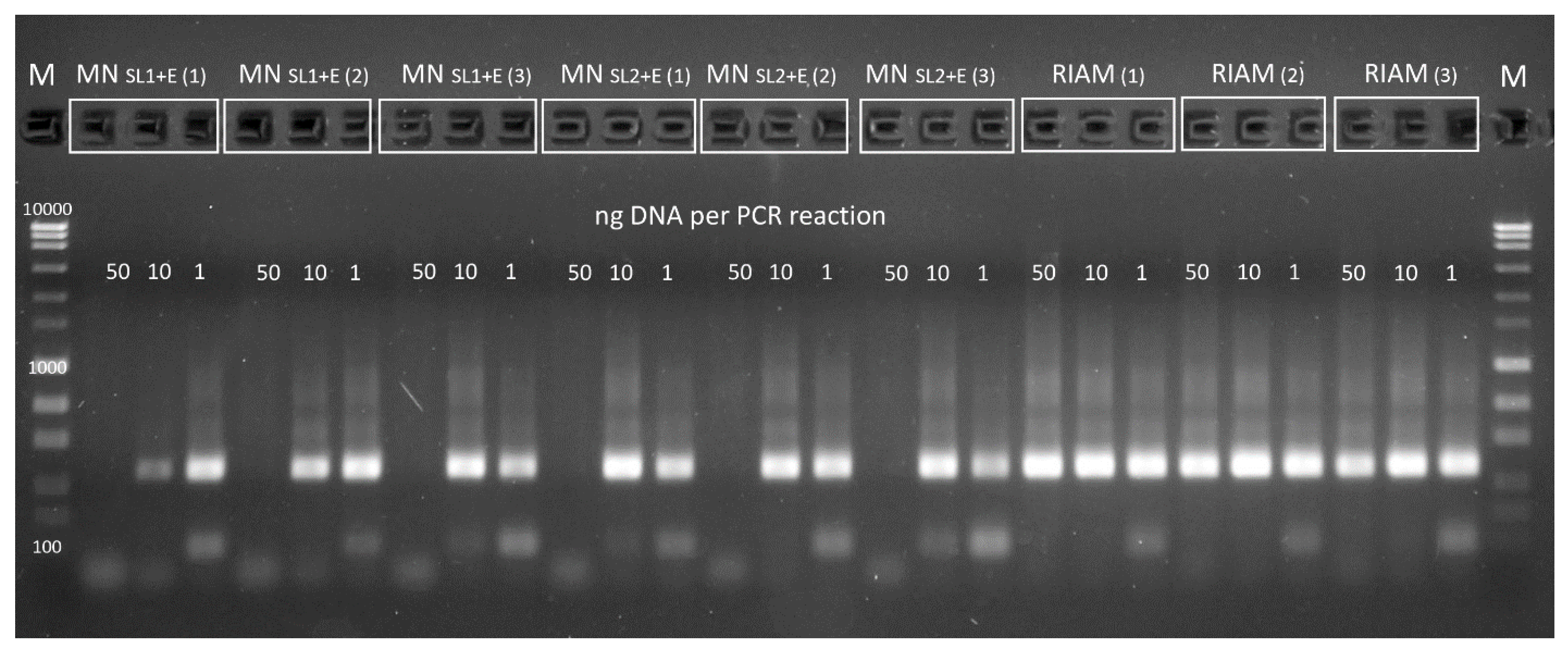
| Soil | MN SL1 | MN SL1+E | MN SL2 | MN SL2+E | RIAM | |
|---|---|---|---|---|---|---|
| conc. ng/µL | chernozem | 49.37 ± 5.7 | 57.93 ± 4.7 | nd 1 | 62.90 ± 16.1 | 168.67 ± 5.7 * |
| sod-podzolic | 6.25 ± 0.5 | 25.20 ± 2.0 | nd | 19.08 ± 3.9 | 32.73 ± 1.5 * | |
| sod-podzolic (clay) | nd | 8.30 ± 0.9 | 8.18 ± 1.8 | 3.43 ± 0.9 | 42.00 ± 3.5 * | |
| OMUG | nd | 3.34 ± 0.9 | nd | 3.35 ± 0.4 | 12.81 ± 1.47 * | |
| A 260/280 | chernozem | 1.92 ± 0.02 | 1.91 ± 0.01 | nd | 1.89 ± 0.01 | 1.92 ± 0.0 |
| sod-podzolic | 1.68 ± 0.03 | 1.81 ± 0.06 | nd | 1.75 ± 0.03 | 1.86 ± 0.01 | |
| sod-podzolic (clay) | nd | 1.87 ± 0.23 | 2.23 ± 0.82 | 2.07 ± 0.49 | 1.91 ± 0.01 | |
| OMUG | nd | 1.87 ± 0.26 | nd | 1.87 ± 0.28 | 1.87 ± 0.0 | |
| A 260/230 | chernozem | 0.35 ± 0.16 | 0.33 ± 0.08 | nd | 0.80 ± 0.22 | 2.07 ± 0.10 |
| sod-podzolic | 0.34 ± 0.13 | 0.50 ± 0.30 | nd | 0.76 ± 0.25 | 2.07 ± 0.11 | |
| sod-podzolic (clay) | nd | 0.34 ± 0.21 | 0.27 ± 0.15 | 0.23 ± 0.19 | 2.07 ± 0.12 | |
| OMUG | nd | 0.20 ± 0.17 | nd | 0.06 ± 0.03 | 2.07 ± 0.13 | |
| PCR inhibition with 50 ng of DNA | OMUG 2 | nd | yes | nd | yes | no |
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2022 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Pinaev, A.G.; Kichko, A.A.; Aksenova, T.S.; Safronova, V.I.; Kozhenkova, E.V.; Andronov, E.E. RIAM: A Universal Accessible Protocol for the Isolation of High Purity DNA from Various Soils and Other Humic Substances. Methods Protoc. 2022, 5, 99. https://doi.org/10.3390/mps5060099
Pinaev AG, Kichko AA, Aksenova TS, Safronova VI, Kozhenkova EV, Andronov EE. RIAM: A Universal Accessible Protocol for the Isolation of High Purity DNA from Various Soils and Other Humic Substances. Methods and Protocols. 2022; 5(6):99. https://doi.org/10.3390/mps5060099
Chicago/Turabian StylePinaev, Alexander G., Arina A. Kichko, Tatiana S. Aksenova, Vera I. Safronova, Elena V. Kozhenkova, and Evgeny E. Andronov. 2022. "RIAM: A Universal Accessible Protocol for the Isolation of High Purity DNA from Various Soils and Other Humic Substances" Methods and Protocols 5, no. 6: 99. https://doi.org/10.3390/mps5060099
APA StylePinaev, A. G., Kichko, A. A., Aksenova, T. S., Safronova, V. I., Kozhenkova, E. V., & Andronov, E. E. (2022). RIAM: A Universal Accessible Protocol for the Isolation of High Purity DNA from Various Soils and Other Humic Substances. Methods and Protocols, 5(6), 99. https://doi.org/10.3390/mps5060099






