Optimized Nuclear Pellet Method for Extracting Next-Generation Sequencing Quality Genomic DNA from Fresh Leaf Tissue
Abstract
:1. Introduction
2. Materials and Methods
2.1. Plant Materials
2.2. Genomic DNA Extraction
2.2.1. CTAB-Based Method
2.2.2. Qiagen Method
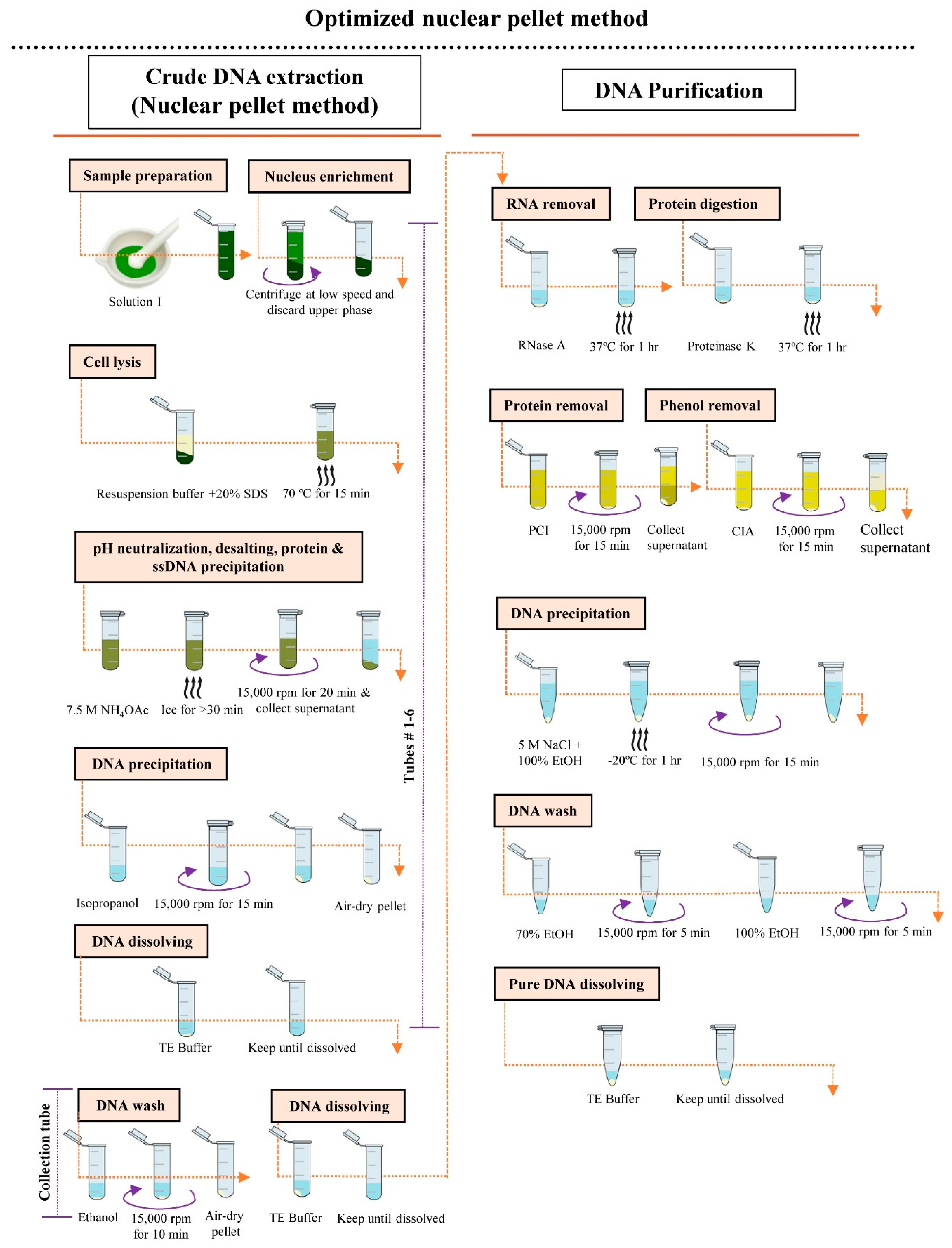
2.2.3. Optimization of Nuclear Pellet Method
Reagents
- Solution I: 15% sucrose, 50 mM Tris-HCl (pH 8.0), 50 mM EDTA (pH 8.0), and 500 mM NaCl
- Resuspension buffer: 20 mM Tris-HCl (pH 8.0) and 10 mM EDTA (pH 8.0)
- 20% sodium dodecyl sulfate (SDS)
- 7.5 M ammonium acetate
- 5 M NaCl
- Isopropanol
- TE buffer: 10 mM Tris-HCl (pH 7.6) and 0.1 mM EDTA
- RNase A: a working solution of 1 µg/µL (final concentration ≈ 10 ng/µL)
- Proteinase K: a working solution of 2 µg/µL (final concentration ≈ 80 ng/µL)
- PCI: phenol/chloroform/isoamyl alcohol (25:24:1)
- CIA: chloroform/isoamyl alcohol (24:1)
- 100% ethanol (v/v)
- 70% ethanol (v/v)
- QubitTM dsDNA HS Assay Kit (Thermo Fisher Scientific Inc., Waltham, MA, USA)
Equipment
- Mortar and pestle
- Dry block heater
- Table-top high-speed microcentrifuge
- Freezer (−20 °C)
- Gel electrophoresis system
- Eppendorf BioSpectrometer® fluorescence (Eppendorf AG, Hamburg, Germany)
Protocol
- Weigh 1 g of young fresh leaf tissue and cut into small pieces using clean and sharp scissors. Place the cut tissue in liquid nitrogen and grind thoroughly with a mortar and pestle. Add 12 mL of Solution I to the fine tissue powder and suspend well. Transfer the suspended tissue (2 mL in each) into six individual 2 mL microcentrifuge tubes.
- Centrifuge the tubes containing the suspended tissue at 500 rpm for 3 min at 4 °C. Discard the upper phase carefully. Centrifuge at 1000 rpm for 1 min at 4 °C and discard the upper phase.
- Add 450 µL of resuspension buffer and 30 µL of 20% SDS into each tube containing tissue, shake briefly, and incubate at 70 °C for 15 min.
- Add 230 µL of 7.5 M NH4OAc into each tube, shake vigorously, and incubate the reaction mixture on ice for at least 30 min.
- Centrifuge the reaction mixture at 15,000 rpm for 20 min at 4 °C and divide equal amounts of the cleared supernatant into six individual microcentrifuge tubes, labeled #1–6. Repeat this step (once).
- Add 1 volume of isopropanol to the supernatant of each tube, gently mix by inverting, and centrifuge at 15,000 rpm for 15 min at 25 °C. Discard the supernatant and air-dry pellet for 5–10 min (do not excess dry).
- Add 50 µL of TE buffer individually to tubes #1–5 and wait until the pellet has dissolved. Transfer the dissolved DNA from tubes #1–5 into tube #6. Add 50 µL TE buffer to the original tube #1, allow to dissolve, then transfer it to tube #2. Repeat this transfer and dissolving process until tube #5, then transfer the DNA to tube #6. The final volume of the DNA sample in tube #6 will be 300 µL.
- Add 600 µL of 100% ethanol to the sample in tube #6, centrifuge at 15,000 rpm for 10 min at 25 °C, and decant the supernatant. Again, add 600 µL of 100% ethanol to the DNA pellet in tube #6, centrifuge at 15,000 rpm for 1 min, and decant the supernatant.
- Air-dry the pellet for 5–10 min. Resuspend the pellet in 50 µL of TE buffer (do not vortex).
- Add 0.5 µL of RNase A to the solution and incubate at 37 °C for 1 h.
- Add 2.2 µL of proteinase K to the solution and incubate at 37 °C for 1 h.
- Check gDNA quality by electrophoresis using a 0.7% agarose gel (optional).
- Add 400 µL of TE buffer to the digested sample containing the DNA and mix gently by pipetting.
- Add 450 µL of PCI to the DNA solution, gently mix, and centrifuge at 15,000 rpm for 15 min at 4 °C. Transfer the cleared supernatant to a new 1.5 mL collection tube. Repeat this step (once).
- Add 450 µL of CIA to the DNA solution, mix gently, and centrifuge at 15,000 rpm for 15 min at 4 °C. Transfer the cleared supernatant to a new 1.5 mL collection tube. Repeat this step (once).
- Add 27 µL of 5 M NaCl and 1 mL of 100% ethanol to the solution, gently mix, and incubate at −20 °C for 1 h.
- Centrifuge at 15,000 rpm for 15 min at 4 °C and discard the supernatant.
- Wash the pellet with 100 µL of 70% ethanol, centrifuge at 15,000 rpm for 5 min, and discard the ethanol.
- Wash the pellet with 100 µL of 100% ethanol, centrifuge at 15,000 rpm for 5 min, and discard the supernatant. Again, add 100 µL of 100% ethanol to the pellet and wash and discard the ethanol.
- Air-dry the pellet for 5–10 min. Resuspend the pellet in 11 µL of TE buffer.
- Dilute the DNA with TE buffer (as required) for downstream analysis.
2.3. DNA Quantification
2.4. DNA Quality Assessment
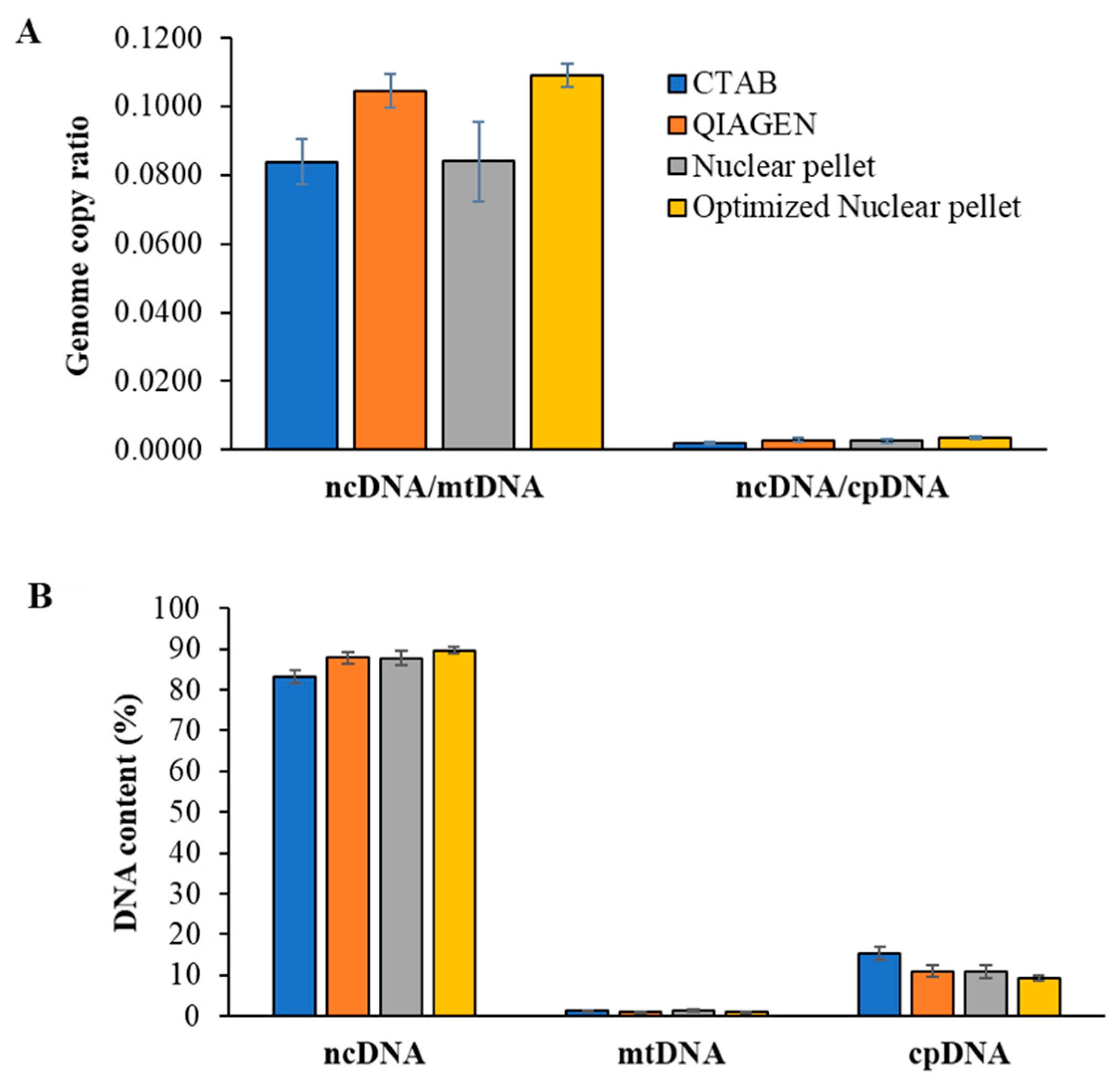
2.5. Quantification of Nuclear, Chloroplast, and Mitochondrial DNAs by qPCR
3. Results and Discussion
4. Conclusions
Supplementary Materials
Author Contributions
Funding
Acknowledgments
Conflicts of Interest
References
- Egan, A.N.; Schlueter, J.; Spooner, D.M. Applications of next-generation sequencing in plant biology. Am. J. Bot. 2012, 99, 175–185. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Healey, A.; Furtado, A.; Cooper, T.; Henry, R.J. Protocol: A simple method for extracting next-generation sequencing quality genomic DNA from recalcitrant plant species. Plant Methods 2014, 10, 21. [Google Scholar] [CrossRef] [PubMed]
- Anderson, C.B.; Franzmayr, B.K.; Hong, S.W.; Larking, A.C.; van Stijn, T.C.; Tan, R.; Moraga, R.; Faville, M.J.; Griffiths, A.G. Protocol: A versatile, inexpensive, high-throughput plant genomic DNA extraction method suitable for genotyping-by-sequencing. Plant Methods 2018, 14, 75. [Google Scholar] [CrossRef] [PubMed]
- Kasem, S.; Rice, N.; Henry, R. DNA extraction from plant tissue. In Plant Genotyping II SNP Technol; Henry, R.J., Ed.; CAB International: Wallingford, Oxfordshire, UK, 2008; pp. 219–271. [Google Scholar]
- Abdel-Latif, A.; Osman, G. Comparison of three genomic DNA extraction methods to obtain high DNA quality from maize. Plant Methods 2017, 13, 1. [Google Scholar] [CrossRef] [PubMed]
- Rhoads, A.; Au, K.F. PacBio Sequencing and Its Applications. Genomics. Proteomics Bioinf. 2015, 13, 278–289. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Mayjonade, B.; Gouzy, J.; Donnadieu, C.; Pouilly, N.; Marande, W.; Callot, C.; Langlade, N.; Muños, S. Extraction of high-molecular-weight genomic DNA for long-read sequencing of single molecules. Biotechniques 2016, 61, 2003–2005. [Google Scholar] [CrossRef] [PubMed]
- Doyle, J.J.; Doyle, J.L. A rapid DNA isolation procedure for small quantities of fresh leaf tissue. Phytochem. Bull. 1987, 19, 11–15. [Google Scholar]
- Xu, X.; Kawasaki, S.; Fujimura, T.; Wang, C. A protocol for high-throughput extraction of DNA from rice leaves. Plant Mol. Biol. Report. 2005, 23, 291–295. [Google Scholar] [CrossRef]
- Whitlock, R.; Hipperson, H.; Mannarelli, M.; Burke, T. A high-throughput protocol for extracting high-purity genomic DNA from plants and animals. Mol. Ecol. Resour. 2008, 8, 736–741. [Google Scholar] [CrossRef]
- Pereira, J.C.; Chaves, R.; Bastos, E.; Leitão, A.; Guedes-Pinto, H. An efficient method for genomic DNA extraction from different molluscs species. Int. J. Mol. Sci. 2011, 12, 8086–8095. [Google Scholar] [CrossRef]
- Telfer, E.; Graham, N.; Stanbra, L.; Manley, T.; Wilcox, P. Extraction of high purity genomic DNA from pine for use in a high-throughput Genotyping Platform. New Zeal. J. For. Sci. 2013, 43, 3. [Google Scholar] [CrossRef] [Green Version]
- Werth, S.; Reynisdóttir, S.; Gudmundsson, H.; Andrésson, Ó.S. A Fast and Inexpensive High-Throughput Protocol for Isolating High Molecular Weight Genomic DNA from Lichens. Herzogia 2016, 29, 610–616. [Google Scholar] [CrossRef]
- Minas, K.; McEwan, N.R.; Newbold, C.J.; Scott, K.P. Optimization of a high-throughput CTAB-based protocol for the extraction of qPCR-grade DNA from rumen fluid, plant and bacterial pure cultures. FEMS Microbiol. Lett. 2011, 325, 162–169. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Simbolo, M.; Gottardi, M.; Corbo, V.; Fassan, M.; Mafficini, A.; Malpeli, G.; Lawlor, R.T.; Scarpa, A. DNA Qualification Workflow for Next Generation Sequencing of Histopathological Samples. PLoS ONE 2013, 8. [Google Scholar] [CrossRef] [PubMed]
- Walbot, V.; Warren, C. Regulation of Mu element copy number in maize lines with an active or inactive Mutator transposable element system. Mol. Gen. Genet. 1988, 211, 27–34. [Google Scholar] [CrossRef]
- Shimamoto, K.; Terada, R.; Izawa, T.; Fujimoto, H. Fertile transgenic rice plants regenerated from transformed protoplasts. Nature 1989, 338, 274–276. [Google Scholar] [CrossRef]
- Takamatsu, T.; Baslam, M.; Inomata, T.; Oikawa, K.; Itoh, K.; Ohnishi, T.; Kinoshita, T.; Mitsui, T. Optimized Method of Extracting Rice Chloroplast DNA for High-Quality Plastome Resequencing and de Novo Assembly. Front. Plant Sci. 2018, 9, 266. [Google Scholar] [CrossRef]
- Rana, M.M.; Takamatsu, T.; Baslam, M.; Kaneko, K.; Itoh, K.; Harada, N.; Sugiyama, T.; Ohnishi, T.; Kinoshita, T.; Takagi, H.; et al. Salt Tolerance Improvement in Rice through Efficient SNP Marker-Assisted Selection Coupled with Speed-Breeding. Int. J. Mol. Sci. 2019, 20, 2585. [Google Scholar] [CrossRef]
- Bolger, A.M.; Lohse, M.; Usadel, B. Trimmomatic: A flexible trimmer for Illumina sequence data. Bioinformatics 2014, 30, 2114–2120. [Google Scholar] [CrossRef]
- Jain, M.; Koren, S.; Miga, K.H.; Quick, J.; Rand, A.C.; Sasani, T.A.; Tyson, J.R.; Beggs, A.D.; Dilthey, A.T.; Fiddes, I.T.; et al. Nanopore sequencing and assembly of a human genome with ultra-long reads. Nat. Biotechnol. 2018, 36, 338–345. [Google Scholar] [CrossRef] [Green Version]
- Seo, J.-S.; Rhie, A.; Kim, J.; Lee, S.; Sohn, M.-H.; Kim, C.-U.; Hastie, A.; Cao, H.; Yun, J.-Y.; Kim, J.; et al. De novo assembly and phasing of a Korean human genome. Nature 2016, 538, 243–247. [Google Scholar] [CrossRef] [PubMed] [Green Version]
- Huddleston, J.; Chaisson, M.J.P.; Steinberg, K.M.; Warren, W.; Hoekzema, K.; Gordon, D.; Graves-Lindsay, T.A.; Munson, K.M.; Kronenberg, Z.N.; Vives, L.; et al. Discovery and genotyping of structural variation from long-read haploid genome sequence data. Genome Res. 2017, 27, 677–685. [Google Scholar] [CrossRef] [PubMed]
- Mitsuhashi, S.; Nakagawa, S.; Takahashi Ueda, M.; Imanishi, T.; Frith, M.C.; Mitsuhashi, H. Nanopore-based single molecule sequencing of the D4Z4 array responsible for facioscapulohumeral muscular dystrophy. Sci. Rep. 2017, 7, 14789. [Google Scholar] [CrossRef] [PubMed]
- Colle, M.; Leisner, C.P.; Wai, C.M.; Ou, S.; Bird, K.A.; Wang, J.; Wisecaver, J.H.; Yocca, A.E.; Alger, E.I.; Tang, H.; et al. Haplotype-phased genome and evolution of phytonutrient pathways of tetraploid blueberry. Gigascience 2019, 8. [Google Scholar] [CrossRef] [PubMed] [Green Version]
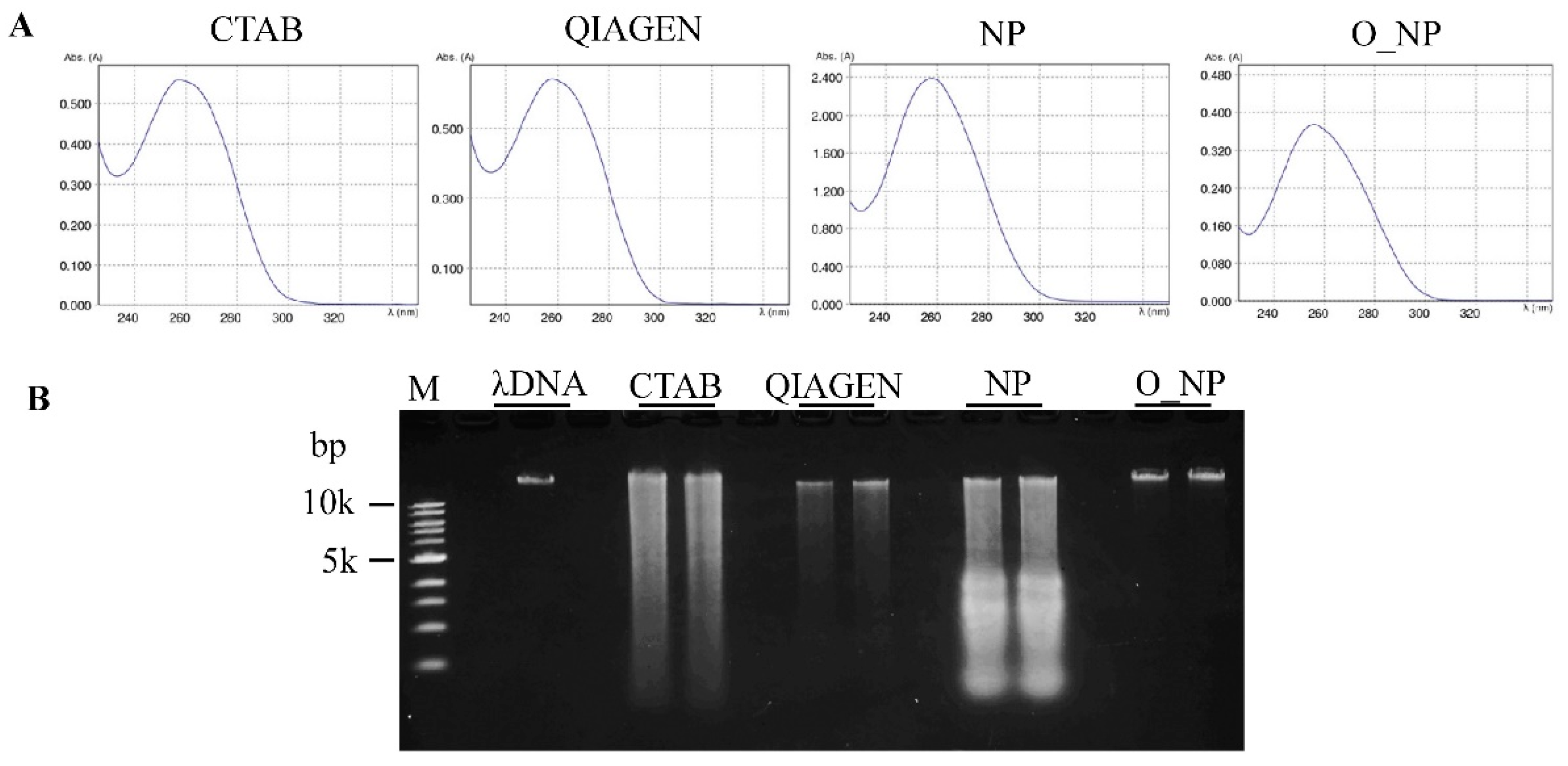
| Experimental Method | A260/A280 Ratio | A260/A230 Ratio | DNA Yield (µg/g Fresh Tissue) |
|---|---|---|---|
| CTAB-based | 2.06 ± 0.10 | 2.39 ± 0.04 | 6.59 ± 0.80 c |
| QIAGEN DNeasy Plant Mini Kit | 1.91 ± 0.03 | 2.31 ± 0.03 | 9.03 ± 1.50 b |
| Nuclear pellet | 2.08 ± 0.01 | 2.46 ± 0.02 | 11.78 ± 1.38 a |
| Optimized nuclear pellet | 1.87 ± 0.02 | 2.26 ± 0.05 | 10.40 ± 1.32 ab |
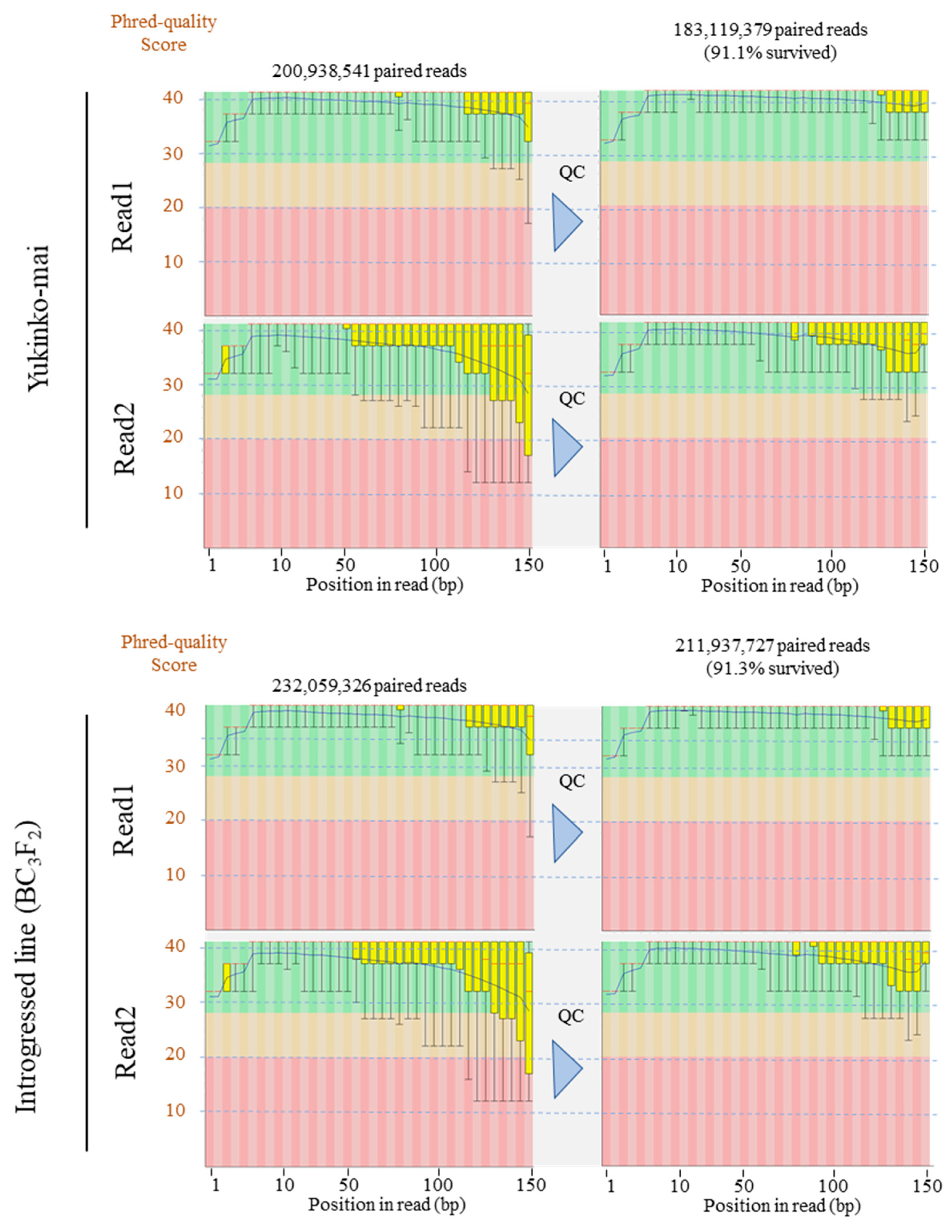
| Sample Name | DNA Submitted (ng/µL TE Buffer) | Total DNA Submitted (µg) | NGS Quality Control Results |
|---|---|---|---|
| Yukinko-mai | 330.7 | 5.29 | Passed |
| Introgressed line (BC3F2) | 256.8 | 4.11 | Passed |
© 2019 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (http://creativecommons.org/licenses/by/4.0/).
Share and Cite
Rana, M.M.; Aycan, M.; Takamatsu, T.; Kaneko, K.; Mitsui, T.; Itoh, K. Optimized Nuclear Pellet Method for Extracting Next-Generation Sequencing Quality Genomic DNA from Fresh Leaf Tissue. Methods Protoc. 2019, 2, 54. https://doi.org/10.3390/mps2020054
Rana MM, Aycan M, Takamatsu T, Kaneko K, Mitsui T, Itoh K. Optimized Nuclear Pellet Method for Extracting Next-Generation Sequencing Quality Genomic DNA from Fresh Leaf Tissue. Methods and Protocols. 2019; 2(2):54. https://doi.org/10.3390/mps2020054
Chicago/Turabian StyleRana, Md Masud, Murat Aycan, Takeshi Takamatsu, Kentaro Kaneko, Toshiaki Mitsui, and Kimiko Itoh. 2019. "Optimized Nuclear Pellet Method for Extracting Next-Generation Sequencing Quality Genomic DNA from Fresh Leaf Tissue" Methods and Protocols 2, no. 2: 54. https://doi.org/10.3390/mps2020054
APA StyleRana, M. M., Aycan, M., Takamatsu, T., Kaneko, K., Mitsui, T., & Itoh, K. (2019). Optimized Nuclear Pellet Method for Extracting Next-Generation Sequencing Quality Genomic DNA from Fresh Leaf Tissue. Methods and Protocols, 2(2), 54. https://doi.org/10.3390/mps2020054






