MRI Imaging Characteristics of Glioblastoma with Concurrent Gain of Chromosomes 19 and 20
Abstract
1. Introduction
2. Materials and Methods
2.1. Patient Population
2.2. Image Analysis
2.3. Statistical Analysis
3. Results
3.1. Patient Population
3.2. Tumor Location
3.3. Tumor Size
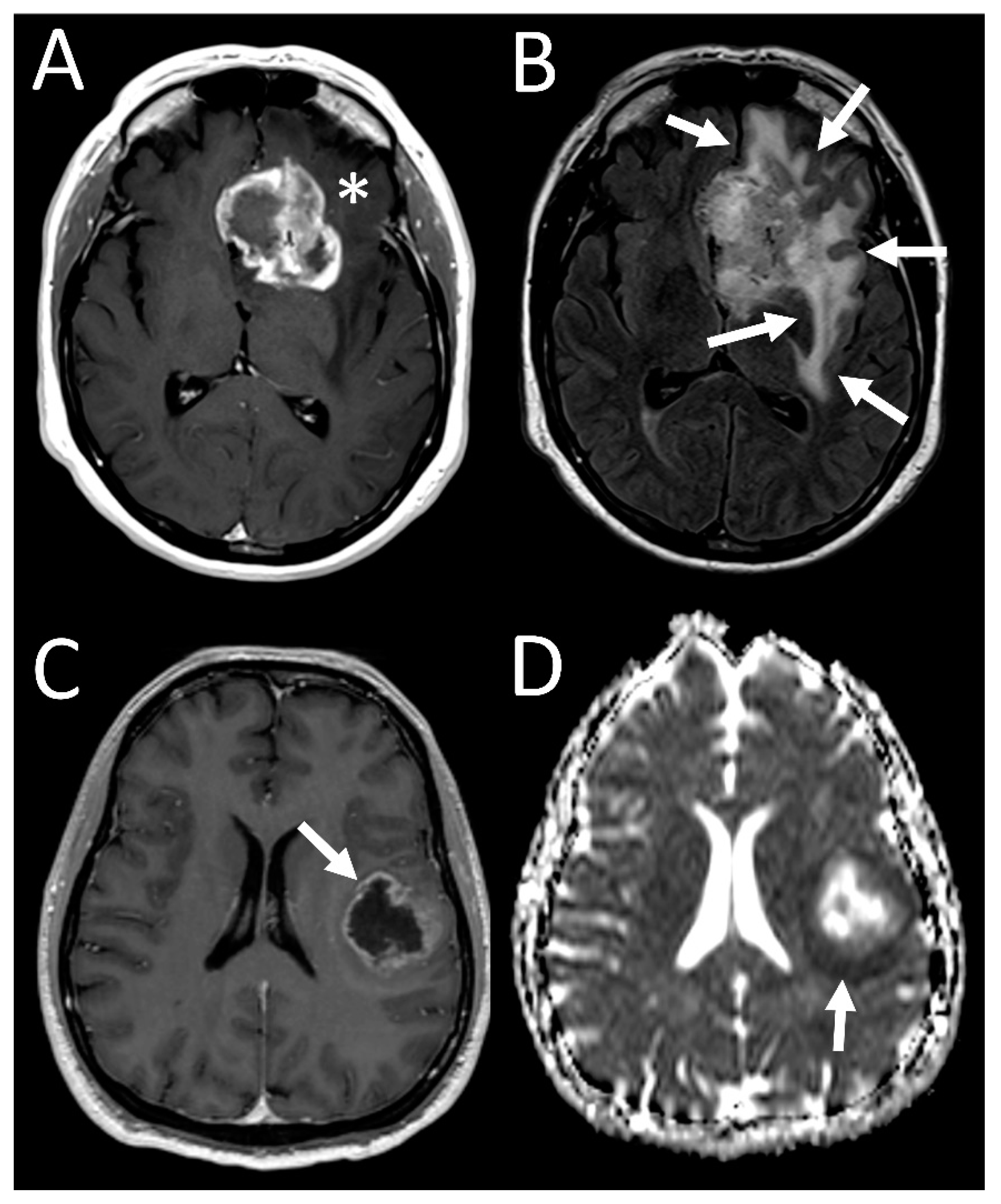
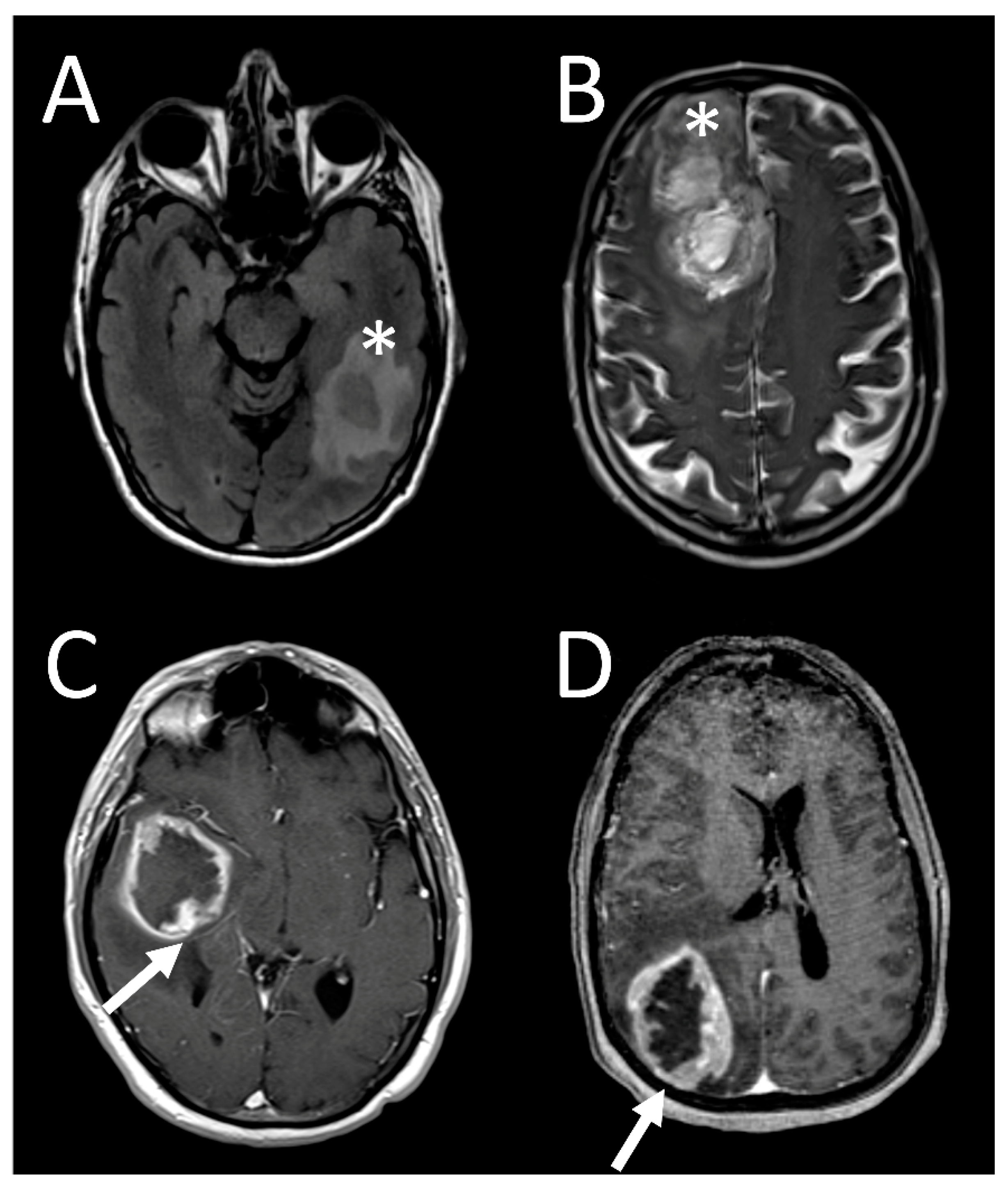
3.4. Tumor Characteristics
3.5. Overall Survival
4. Discussion
5. Conclusions
Supplementary Materials
Author Contributions
Funding
Institutional Review Board Statement
Informed Consent Statement
Data Availability Statement
Conflicts of Interest
References
- Ostrom, Q.T.; Patil, N.; Cioffi, G.; Waite, K.; Kruchko, C.; Barnholtz-Sloan, J.S. CBTRUS Statistical Report: Primary Brain and Other Central Nervous System Tumors Diagnosed in the United States in 2013–2017. Neuro-Oncology 2020, 22, iv1–iv96. [Google Scholar] [CrossRef]
- Marenco-Hillembrand, L.; Wijesekera, O.; Suarez-Meade, P.; Mampre, D.; Jackson, C.; Peterson, J.; Trifiletti, D.; Hammack, J.; Ortiz, K.; Lesser, E.; et al. Trends in glioblastoma: Outcomes over time and type of intervention: A systematic evidence based analysis. J. Neuro-Oncol. 2020, 147, 297–307. [Google Scholar] [CrossRef]
- Stupp, R.; Mason, W.P.; van den Bent, M.J.; Weller, M.; Fisher, B.; Taphoorn, M.J.; Belanger, K.; Brandes, A.A.; Marosi, C.; Bogdahn, U.; et al. Radiotherapy plus concomitant and adjuvant temozolomide for glioblastoma. N. Engl. J. Med. 2005, 352, 987–996. [Google Scholar] [CrossRef] [PubMed]
- Phillips, H.S.; Kharbanda, S.; Chen, R.; Forrest, W.F.; Soriano, R.H.; Wu, T.D.; Misra, A.; Nigro, J.M.; Colman, H.; Soroceanu, L.; et al. Molecular subclasses of high-grade glioma predict prognosis, delineate a pattern of disease progression, and resemble stages in neurogenesis. Cancer Cell 2006, 9, 157–173. [Google Scholar] [CrossRef] [PubMed]
- Verhaak, R.G.; Hoadley, K.A.; Purdom, E.; Wang, V.; Qi, Y.; Wilkerson, M.D.; Miller, C.R.; Ding, L.; Golub, T.; Mesirov, J.P.; et al. Integrated genomic analysis identifies clinically relevant subtypes of glioblastoma characterized by abnormalities in PDGFRA, IDH1, EGFR, and NF1. Cancer Cell 2010, 17, 98–110. [Google Scholar] [CrossRef]
- Hegi, M.E.; Diserens, A.C.; Gorlia, T.; Hamou, M.F.; de Tribolet, N.; Weller, M.; Kros, J.M.; Hainfellner, J.A.; Mason, W.; Mariani, L.; et al. MGMT gene silencing and benefit from temozolomide in glioblastoma. N. Engl. J. Med. 2005, 352, 997–1003. [Google Scholar] [CrossRef] [PubMed]
- Yan, H.; Parsons, D.W.; Jin, G.; Mclendon, R.; Rasheed, B.A.; Yuan, W.; Kos, I.; Batinic-Haberle, I.; Jones, S.; Riggins, G.J.; et al. IDH1andIDH2Mutations in Gliomas. N. Engl. J. Med. 2009, 360, 765–773. [Google Scholar] [CrossRef] [PubMed]
- Hartmann, C.; Hentschel, B.; Simon, M.; Westphal, M.; Schackert, G.; Tonn, J.C.; Loeffler, M.; Reifenberger, G.; Pietsch, T.; von Deimling, A.; et al. Long-term survival in primary glioblastoma with versus without isocitrate dehydrogenase mutations. Clin. Cancer Res. 2013, 19, 5146–5157. [Google Scholar] [CrossRef] [PubMed]
- Louis, D.N.; Perry, A.; Reifenberger, G.; Von Deimling, A.; Figarella-Branger, D.; Cavenee, W.K.; Ohgaki, H.; Wiestler, O.D.; Kleihues, P.; Ellison, D.W. The 2016 World Health Organization Classification of Tumors of the Central Nervous System: A summary. Acta Neuropathol. 2016, 131, 803–820. [Google Scholar] [CrossRef]
- Ohgaki, H.; Kleihues, P. The Definition of Primary and Secondary Glioblastoma. Clin. Cancer Res. 2013, 19, 764–772. [Google Scholar] [CrossRef]
- Turcan, S.; Rohle, D.; Goenka, A.; Walsh, L.A.; Fang, F.; Yilmaz, E.; Campos, C.; Fabius, A.W.M.; Lu, C.; Ward, P.S.; et al. IDH1 mutation is sufficient to establish the glioma hypermethylator phenotype. Nature 2012, 483, 479–483. [Google Scholar] [CrossRef]
- Brat, D.J.; Aldape, K.; Colman, H.; Holland, E.C.; Louis, D.N.; Jenkins, R.B.; Kleinschmidt-DeMasters, B.K.; Perry, A.; Reifenberger, G.; Stupp, R.; et al. cIMPACT-NOW update 3: Recommended diagnostic criteria for Diffuse astrocytic glioma, IDH-wildtype, with molecular features of glioblastoma, WHO grade IV. Acta Neuropathol. 2018, 136, 805–810. [Google Scholar] [CrossRef]
- Chow, D.; Chang, P.; Weinberg, B.D.; Bota, D.A.; Grinband, J.; Filippi, C.G. Imaging Genetic Heterogeneity in Glioblastoma and Other Glial Tumors: Review of Current Methods and Future Directions. Am. J. Roentgenol. 2018, 210, 30–38. [Google Scholar] [CrossRef]
- Fathi Kazerooni, A.; Bakas, S.; Saligheh Rad, H.; Davatzikos, C. Imaging signatures of glioblastoma molecular characteristics: A radiogenomics review. J. Magn. Reson. Imaging 2020, 52, 54–69. [Google Scholar] [CrossRef] [PubMed]
- Bae, S.; Choi, Y.S.; Ahn, S.S.; Chang, J.H.; Kang, S.G.; Kim, E.H.; Kim, S.H.; Lee, S.K. Radiomic MRI Phenotyping of Glioblastoma: Improving Survival Prediction. Radiology 2018, 289, 797–806. [Google Scholar] [CrossRef] [PubMed]
- Ellingson, B.M.; Lai, A.; Harris, R.J.; Selfridge, J.M.; Yong, W.H.; Das, K.; Pope, W.B.; Nghiemphu, P.L.; Vinters, H.V.; Liau, L.M.; et al. Probabilistic radiographic atlas of glioblastoma phenotypes. Am. J. Neuroradiol. 2013, 34, 533–540. [Google Scholar] [CrossRef]
- Altieri, R.; Zenga, F.; Ducati, A.; Melcarne, A.; Cofano, F.; Mammi, M.; Di Perna, G.; Savastano, R.; Garbossa, D. Tumor location and patient age predict biological signatures of high-grade gliomas. Neurosurg. Rev. 2018, 41, 599–604. [Google Scholar] [CrossRef]
- Chang, P.; Grinband, J.; Weinberg, B.D.; Bardis, M.; Khy, M.; Cadena, G.; Su, M.-Y.; Cha, S.; Filippi, C.G.; Bota, D.; et al. Deep-Learning Convolutional Neural Networks Accurately Classify Genetic Mutations in Gliomas. Am. J. Neuroradiol. 2018, 39, 1201–1207. [Google Scholar] [CrossRef] [PubMed]
- Geisenberger, C.; Mock, A.; Warta, R.; Rapp, C.; Schwager, C.; Korshunov, A.; Nied, A.K.; Capper, D.; Brors, B.; Jungk, C.; et al. Molecular profiling of long-term survivors identifies a subgroup of glioblastoma characterized by chromosome 19/20 co-gain. Acta Neuropathol. 2015, 130, 419–434. [Google Scholar] [CrossRef] [PubMed]
- Gevaert, O.; Mitchell, L.A.; Achrol, A.S.; Xu, J.; Echegaray, S.; Steinberg, G.K.; Cheshier, S.H.; Napel, S.; Zaharchuk, G.; Plevritis, S.K. Glioblastoma multiforme: Exploratory radiogenomic analysis by using quantitative image features. Radiology 2014, 273, 168–174. [Google Scholar] [CrossRef] [PubMed]
- Abedalthagafi, M.; Barakeh, D.; Foshay, K.M. Immunogenetics of glioblastoma: The future of personalized patient management. NPJ Precis. Oncol. 2018, 2. [Google Scholar] [CrossRef]
- Qazi, M.A.; Vora, P.; Venugopal, C.; Sidhu, S.S.; Moffat, J.; Swanton, C.; Singh, S.K. Intratumoral heterogeneity: Pathways to treatment resistance and relapse in human glioblastoma. Ann. Oncol. 2017, 28, 1448–1456. [Google Scholar] [CrossRef]
- Comprehensive genomic characterization defines human glioblastoma genes and core pathways. Nature 2008, 455, 1061–1068. [CrossRef]
- Brennan, C.W.; Verhaak, R.G.; Mckenna, A.; Campos, B.; Noushmehr, H.; Salama, S.R.; Zheng, S.; Chakravarty, D.; Sanborn, J.Z.; Berman, S.H.; et al. The Somatic Genomic Landscape of Glioblastoma. Cell 2013, 155, 462–477. [Google Scholar] [CrossRef] [PubMed]
- Muscat, A.M.; Wong, N.C.; Drummond, K.J.; Algar, E.M.; Khasraw, M.; Verhaak, R.; Field, K.; Rosenthal, M.A.; Ashley, D.M. The evolutionary pattern of mutations in glioblastoma reveals therapy-mediated selection. Oncotarget 2018, 9, 7844–7858. [Google Scholar] [CrossRef] [PubMed]
- Georgescu, M.-M.; Olar, A. Genetic and histologic spatiotemporal evolution of recurrent, multifocal, multicentric and metastatic glioblastoma. Acta Neuropathol. Commun. 2020, 8. [Google Scholar] [CrossRef] [PubMed]
- Carrillo, J.A.; Lai, A.; Nghiemphu, P.L.; Kim, H.J.; Phillips, H.S.; Kharbanda, S.; Moftakhar, P.; Lalaezari, S.; Yong, W.; Ellingson, B.M.; et al. Relationship between Tumor Enhancement, Edema, IDH1 Mutational Status, MGMT Promoter Methylation, and Survival in Glioblastoma. Am. J. Neuroradiol. 2012, 33, 1349–1355. [Google Scholar] [CrossRef]
- Yamashita, K.; Hiwatashi, A.; Togao, O.; Kikuchi, K.; Hatae, R.; Yoshimoto, K.; Mizoguchi, M.; Suzuki, S.O.; Yoshiura, T.; Honda, H. MR Imaging–Based Analysis of Glioblastoma Multiforme: Estimation ofIDH1Mutation Status. Am. J. Neuroradiol. 2016, 37, 58–65. [Google Scholar] [CrossRef] [PubMed]
- Hong, E.K.; Choi, S.H.; Shin, D.J.; Jo, S.W.; Yoo, R.-E.; Kang, K.M.; Yun, T.J.; Kim, J.-H.; Sohn, C.-H.; Park, S.-H.; et al. Radiogenomics correlation between MR imaging features and major genetic profiles in glioblastoma. Eur. Radiol. 2018, 28, 4350–4361. [Google Scholar] [CrossRef]
- Ellingson, B.M.; Cloughesy, T.F.; Pope, W.B.; Zaw, T.M.; Phillips, H.; Lalezari, S.; Nghiemphu, P.L.; Ibrahim, H.; Naeini, K.M.; Harris, R.J.; et al. Anatomic localization of O6-methylguanine DNA methyltransferase (MGMT) promoter methylated and unmethylated tumors: A radiographic study in 358 de novo human glioblastomas. NeuroImage 2012, 59, 908–916. [Google Scholar] [CrossRef]
- Rundle-Thiele, D.; Day, B.; Stringer, B.; Fay, M.; Martin, J.; Jeffree, R.L.; Thomas, P.; Bell, C.; Salvado, O.; Gal, Y.; et al. Using the apparent diffusion coefficient to identifying MGMT promoter methylation status early in glioblastoma: Importance of analytical method. J. Med. Radiat. Sci. 2015, 62, 92–98. [Google Scholar] [CrossRef]
- Reeves, G.I.; Marks, J.E. Prognostic Significance of Lesion Size for Glioblastoma Multiforme. Radiology 1979, 132, 469–472. [Google Scholar] [CrossRef] [PubMed]
- Wood, J.R.; Green, S.B.; Shapiro, W.R. The prognostic importance of tumor size in malignant gliomas: A computed tomographic scan study by the Brain Tumor Cooperative Group. J. Clin. Oncol. 1988, 6, 338–343. [Google Scholar] [CrossRef]
- Hammoud, M.A.; Sawaya, R.; Shi, W.; Thall, P.F.; Leeds, N.E. Prognostic significance of preoperative MRI scans in glioblastoma multiforme. J. Neuro-Oncol. 1996, 27, 65–73. [Google Scholar] [CrossRef] [PubMed]
- Li, B.; Senbabaoglu, Y.; Peng, W.; Yang, M.-L.; Xu, J.; Li, J.Z. Genomic Estimates of Aneuploid Content in Glioblastoma Multiforme and Improved Classification. Clin. Cancer Res. 2012, 18, 5595–5605. [Google Scholar] [CrossRef] [PubMed]
| Control Group | 19/20 Co-Gain Group | |
|---|---|---|
| Total patients | 19 | 18 |
| Average age in years | 61.8 ± 2.4 | 61.7 ± 2.2 |
| Male/female | 13/6 | 12/6 |
| IDH | Wildtype | Wildtype |
| MGMT methylated/unmethylated | 7/12 | 7/11 |
| Control Group | Frontal | Parietal | Occipital | Temporal | Sum | |
|---|---|---|---|---|---|---|
| MGMT methylated | Right | 0 | 0 | 0 | 0 | 0 |
| Left | 0 | 1 | 2 | 4 | 7 | |
| MGMT unmethylated | Right | 2 | 3 | 2 | 2 | 9 |
| Left | 0 | 0 | 1 | 1 | 2 | |
| Sum | 2 | 4 | 5 | 7 | 18 * | |
| 19/20 co-gain group | Frontal | Parietal | Occipital | Temporal | Sum | |
| MGMT methylated | Right | 1 | 2 | 1 | 0 | 4 |
| Left | 2 | 0 | 0 | 1 | 3 | |
| MGMT unmethylated | Right | 1 | 0 | 3 | 1 | 5 |
| Left | 1 | 3 | 0 | 2 | 6 | |
| Sum | 5 | 5 | 4 | 4 | 18 | |
| Control Group | 19/20 Co-Gain Group | p-Value (t-Test) | |||||
|---|---|---|---|---|---|---|---|
| Edema | 7.3 ± 0.5 | 7.1 ± 0.5 | 0.84 | ||||
| ENH tumor | 4.7 ± 0.3 | 4.5 ± 0.3 | 0.64 | ||||
| MGMT+ | MGMT− | p-value (t-test) | MGMT+ | MGMT− | p-value (t-test) | ||
| Edema | 6.5 ± 1.0 | 7.7 ± 0.4 | 0.21 | 7.4 ± 1.4 | 7.0 ± 0.6 | 0.72 | |
| ENH tumor | 3.6 ± 0.3 | 5.4 +/− 0.4 | 0.0065 | 4.4 ± 0.7 | 4.5 ± 0.5 | 0.92 | |
| Control Group | 19/20 Co-Gain Group | p-Value (Fisher’s Exact Test) | |
|---|---|---|---|
| T1 to FLAIR ratio (expansive/mixed or infiltrative) | 4/15 | 4/14 | 1.00 |
| Hemorrhage (present/absent) | 12/7 | 11/7 | 1.00 |
| Ependymal extension (present/absent) | 13/6 | 11/7 | 0.74 |
| Multifocal or multicentric (yes/no) | 5/14 | 6/12 | 0.73 |
| Satellites (present/absent) | 7/12 | 5/13 | 0.73 |
| Diffusion restriction (present/absent) | 12/7 | 9/9 | 0.51 |
| Pial invasion (present/absent) | 13/6 | 6/12 | 0.05 |
| Control Group | 19/20 Co-Gain Group | |||
|---|---|---|---|---|
| MGMT Methylated | MGMT Unmethylated | MGMT Methylated | MGMT Unmethylated | |
| Present | 2 | 11 | 5 | 6 |
| Absent | 5 | 1 | 2 | 5 |
| p-value (Fisher’s exact test) | 0.01 | 0.64 | ||
Publisher’s Note: MDPI stays neutral with regard to jurisdictional claims in published maps and institutional affiliations. |
© 2021 by the authors. Licensee MDPI, Basel, Switzerland. This article is an open access article distributed under the terms and conditions of the Creative Commons Attribution (CC BY) license (https://creativecommons.org/licenses/by/4.0/).
Share and Cite
Min, T.L.; Allen, J.W.; Velazquez Vega, J.E.; Neill, S.G.; Weinberg, B.D. MRI Imaging Characteristics of Glioblastoma with Concurrent Gain of Chromosomes 19 and 20. Tomography 2021, 7, 228-237. https://doi.org/10.3390/tomography7020021
Min TL, Allen JW, Velazquez Vega JE, Neill SG, Weinberg BD. MRI Imaging Characteristics of Glioblastoma with Concurrent Gain of Chromosomes 19 and 20. Tomography. 2021; 7(2):228-237. https://doi.org/10.3390/tomography7020021
Chicago/Turabian StyleMin, Taejin L., Jason W. Allen, Jose E. Velazquez Vega, Stewart G. Neill, and Brent D. Weinberg. 2021. "MRI Imaging Characteristics of Glioblastoma with Concurrent Gain of Chromosomes 19 and 20" Tomography 7, no. 2: 228-237. https://doi.org/10.3390/tomography7020021
APA StyleMin, T. L., Allen, J. W., Velazquez Vega, J. E., Neill, S. G., & Weinberg, B. D. (2021). MRI Imaging Characteristics of Glioblastoma with Concurrent Gain of Chromosomes 19 and 20. Tomography, 7(2), 228-237. https://doi.org/10.3390/tomography7020021







